|
1
|
Tettelin H, Nelson KE, Paulsen IT, Eisen
JA, Read TD, Peterson S, Heidelberg J, DeBoy RT, Haft DH, Dodson
RJ, et al: Complete genome sequence of a virulent isolate of
Streptococcus pneumoniae. Science. 293:498–506. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Alvarez EF, Olarte KE and Ramesh MS:
Purpura fulminans secondary to Streptococcus pneumoniae
meningitis. Case Rep Infect Dis. 2012:5085032014.
|
|
3
|
Denapaite D, Brückner R, Nuhn M, Reichmann
P, Henrich B, Maurer P, Schähle Y, Selbmann P, Zimmermann W and
Wambutt R: The genome of Streptococcus mitis B6-what is a
commensal? PloS One. 5:e94262010. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Kilian M, Poulsen K, Blomqvist T,
Håvarstein LS, Bek-Thomsen M, Tettelin H and Sørensen UB: Evolution
of Streptococcus pneumoniae and its close commensal
relatives. PloS One. 3:e26832008. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Hakenbeck R, Madhour A, Denapaite D and
Brückner R: Versatility of choline metabolism and choline-binding
proteins in Streptococcus pneumoniae and commensal
streptococci. FEMS Microbiol Rev. 33:572–586. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Kohno K, Nagafuji K, Tsukamoto H, Horiuchi
T, Takase K, Aoki K, Henzan H, Kamezaki K, Takenaka K, Miyamoto T,
et al: Infectious complications in patients receiving autologous
CD34-selected hematopoietic stem cell transplantation for severe
autoimmune diseases. Transpl Infect Dis. 11:318–323. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Han XY, Kamana M and Rolston KV: Viridans
streptococci isolated by culture from blood of cancer patients:
Clinical and microbiologic analysis of 50 cases. J Clin Microbiol.
44:160–165. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Mitchell J: Streptococcus mitis:
Walking the line between commensalism and pathogenesis. Mol Oral
Microbiol. 26:89–98. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Shelburne SA, Sahasrabhojane P, Saldana M,
Yao H, Su X, Horstmann N, Thompson E and Flores AR:
Streptococcus mitis strains causing severe clinical disease
in cancer patients. Emerg Infect Dis. 20:762–771. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Matsui N, Ito M, Kuramae H, Inukai T,
Sakai A and Okugawa M: Infective endocarditis caused by
multidrug-resistant Streptococcus mitis in a combined
immunocompromised patient: An autopsy case report. J Infect
Chemother. 19:321–325. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Bochud PY, Eggiman P, Calandra T, Van
Melle G, Saghafi L and Francioli P: Bacteremia due to viridans
Streptococcus in neutropenic patients with cancer: Clinical
spectrum and risk factors. Clin Infect Dis. 18:25–31. 1994.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Husain E, Whitehead S, Castell A, Thomas
EE and Speert DP: Viridans streptococci bacteremia in children with
malignancy: Relevance of species identification and penicillin
susceptibility. Pediatr Infect Dis J. 24:563–566. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Delany I, Rappuoli R and Seib KL:
Vaccines, reverse vaccinology, and bacterial pathogenesis. Cold
Spring Harb Perspect Med. 3:a0124762013. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Chen VL, Avci FY and Kasper DL: A maternal
vaccine against group B Streptococcus: Past, present, and
future. Vaccine. 31 Suppl 4:D13–D19. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Talukdar S, Zutshi S, Prashanth KS, Saikia
KK and Kumar P: Identification of potential vaccine candidates
against Streptococcus pneumoniae by reverse vaccinology
approach. Appl Biochem Biotechnol. 172:3026–3041. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Xiang Z and He Y: Genome-wide prediction
of vaccine targets for human herpes simplex viruses using Vaxign
reverse vaccinology. BMC Bioinformatics. 14 Suppl 4:S22013.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Caro-Gomez E, Gazi M, Goez Y and Valbuena
G: Discovery of novel cross-protective Rickettsia prowazekii
T-cell antigens using a combined reverse vaccinology and in vivo
screening approach. Vaccine. 32:4968–4976. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Maritz-Olivier C, Van Zyl W and Stutzer C:
A systematic, functional genomics, and reverse vaccinology approach
to the identification of vaccine candidates in the cattle tick,
Rhipicephalus. Ticks Tick Borne Dis. 3:179–187. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Melsted P and Pritchard JK: Efficient
counting of k-mers in DNA sequences using a bloom filter. BMC
Bioinformatics. 12:3332011. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Li R, Zhu H, Ruan J, Qian W, Fang X, Shi
Z, Li Y, Li S, Shan G and Kristiansen K: De novo assembly of human
genomes with massively parallel short read sequencing. Genome Res.
20:265–272. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Luo R, Liu B, Xie Y, Li Z, Huang W, Yuan
J, He G, Chen Y, Pan Q, Liu Y, et al: SOAPdenovo2: An empirically
improved memory-efficient short-read de novo assembler.
Gigascience. 1:182012. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Craig JW, Chang FY, Kim JH, Obiajulu SC
and Brady SF: Expanding small-molecule functional metagenomics
through parallel screening of broad-host-range cosmid environmental
DNA libraries in diverse proteobacteria. Appl Environ Microbiol.
76:1633–1641. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
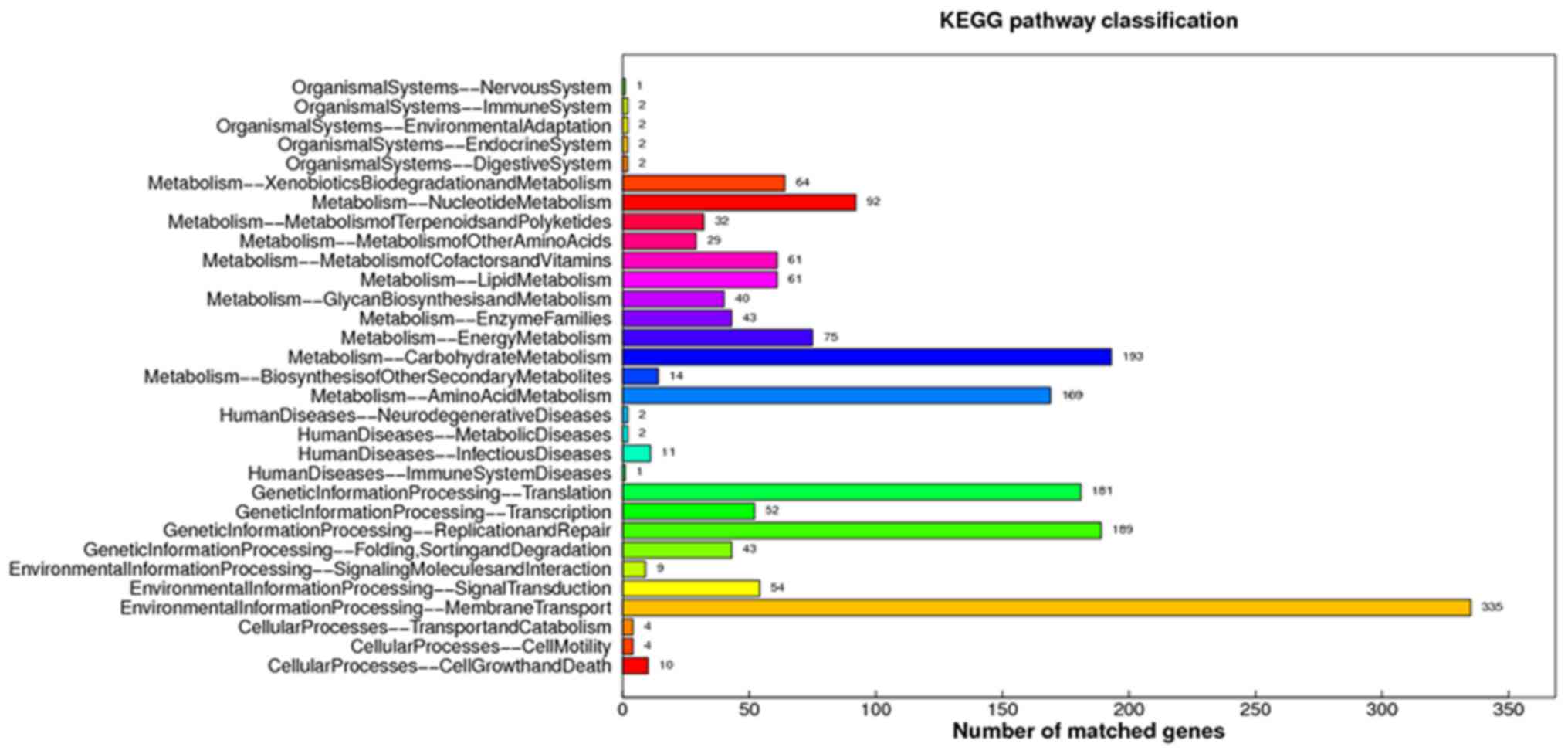
Kanehisa M, Goto S, Kawashima S, Okuno Y
and Hattori M: The KEGG resource for deciphering the genome.
Nucleic Acids Res. 32:(Database Issue). D277–D780. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
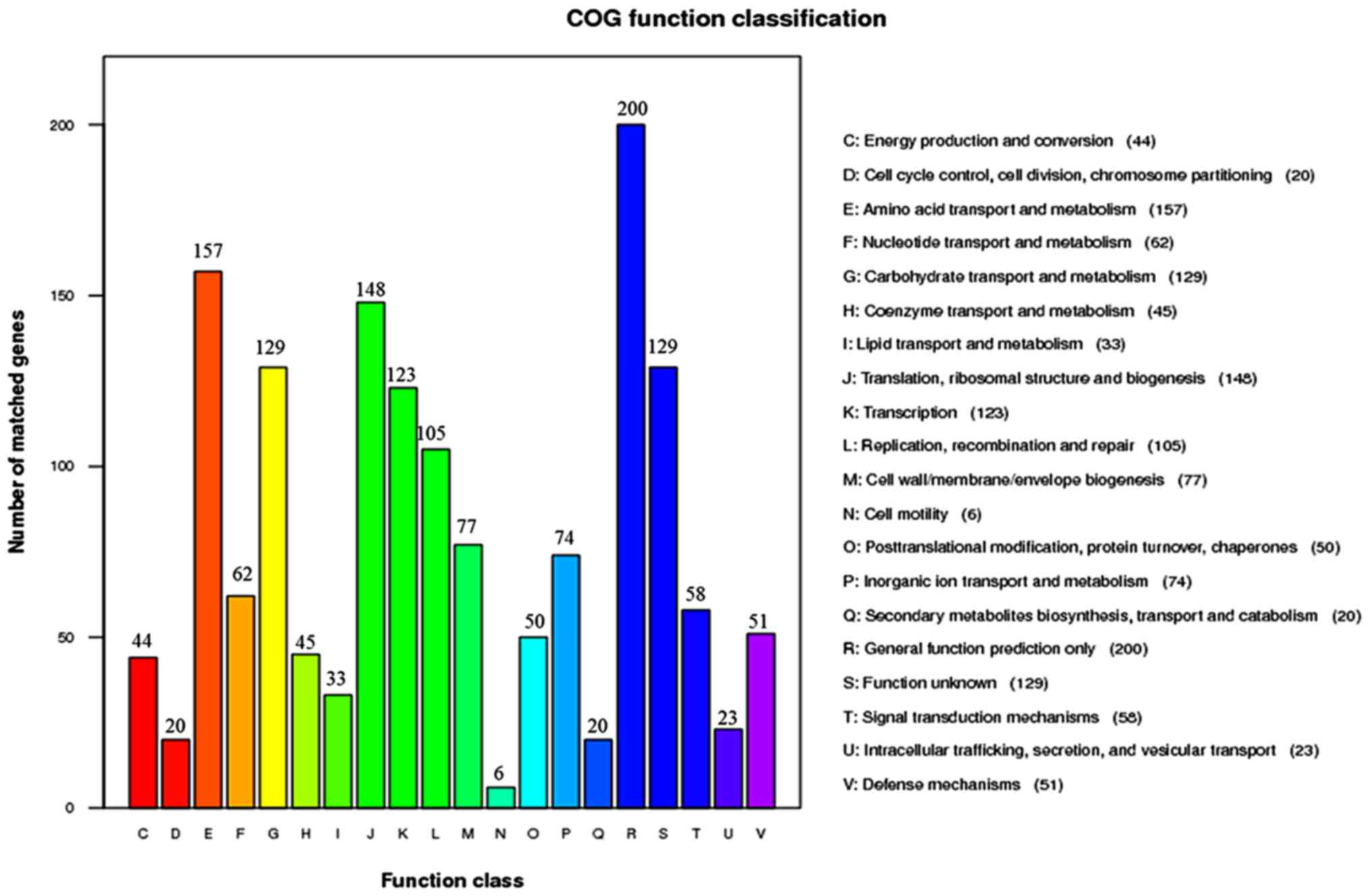
Tatusov RL, Galperin MY, Natale DA and
Koonin EV: The COG database: A tool for genome-scale analysis of
protein functions and evolution. Nucleic Acids Res. 28:33–36. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Boeckmann B, Bairoch A, Apweiler R,
Blatter MC, Estreicher A, Gasteiger E, Martin MJ, Michoud K,
O'Donovan C, Phan I, et al: The SWISS-PROT protein knowledgebase
and its supplement TrEMBL in 2003. Nucleic Acids Res. 31:365–370.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Pruitt KD, Tatusova T and Maglott DR: NCBI
reference sequence (RefSeq): A curated non-redundant sequence
database of genomes, transcripts and proteins. Nucleic Acids Res.
33:(Database Issue). D501–D5D4. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Consortium GO: The Gene Ontology (GO)
database and informatics resource. Nucleic Acids Res. 32:(Database
Issue). D258–D261. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Zdobnov EM and Apweiler R: InterProScan-an
integration platform for the signature-recognition methods in
InterPro. Bioinformatics. 17:847–848. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Sette A and Rappuoli R: Reverse
vaccinology: Developing vaccines in the era of genomics. Immunity.
33:530–541. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Maione D, Margarit I, Rinaudo CD,
Masignani V, Mora M, Scarselli M, Tettelin H, Brettoni C, Iacobini
ET, Rosini R, et al: Identification of a universal Group B
Streptococcus vaccine by multiple genome screen. Science.
309:148–150. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Chou KC and Shen HB: Cell-PLoc: A package
of Web servers for predicting subcellular localization of proteins
in various organisms. Nat Protoc. 3:153–162. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Lobo I: Basic local alignment search tool
(BLAST). Nat Educ. 1:22008.
|
|
33
|
Bateman A, Coin L, Durbin R, Finn RD,
Hollich V, Griffiths-Jones S, Khanna A, Marshall M, Moxon S,
Sonnhammer EL, et al: The Pfam protein families database. Nucleic
Acids Res. 32:(Database Issue). D138–D141. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Underwood A, Mulder A, Gharbia S and Green
J: Virulence Searcher: A tool for searching raw genome sequences
from bacterial genomes for putative virulence factors. Clin
Microbiol Infect. 11:770–772. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Zhang R and Lin Y: DEG 5.0, a database of
essential genes in both prokaryotes and eukaryotes. Nucleic Acids
Res. 37:D455–D458. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Larkin MA, Blackshields G, Brown N, Chenna
R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A and
Lopez R: Clustal W and Clustal X version 2.0. Bioinformatics.
23:2947–2948. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Guindon S, Lethiec F, Duroux P and Gascuel
O: PHYML online-a web server for fast maximum likelihood-based
phylogenetic inference. Nucleic Acids Res. 33:(Web Server Issue).
W557–W559. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Balkundi DR, Murray DL, Patterson MJ, Gera
R, Scott-Emuakpor A and Kulkarni R: Penicillin-resistant
Streptococcus mitis as a cause of septicemia with meningitis
in febrile neutropenic children. J Pediatr Hematol Oncol. 19:82–85.
1997. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Yoshpe-Besancon I, Gripon JC and
Ribadeau-Dumas B: Xaa-Pro-dipeptidyl-aminopeptidase from
Lactococcus lactis catalyses kinetically controlled
synthesis of peptide bonds involving proline. Biotechnol Appl
Biochem. 20:131–140. 1994.PubMed/NCBI
|
|
40
|
Kowluru A: Identification and
characterization of a novel protein histidine kinase in the islet β
cell: Evidence for its regulation by mastoparan, an activator of
G-proteins and insulin secretion. Biochem Pharmacol. 63:2091–2100.
2002. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Torosantucci A, Chiani P, De Bernardis F,
Cassone A, Calera JA and Calderone R: Deletion of the two-component
histidine kinase gene (CHK1) of Candida albicans contributes to
enhanced growth inhibition and killing by human neutrophils in
vitro. Infect Immun. 70:985–987. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Brenot A, King KY, Janowiak B, Griffith O
and Caparon MG: Contribution of glutathione peroxidase to the
virulence of Streptococcus pyogenes. Infect Immun.
72:408–413. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Fischer W and Haas R: The RecA protein of
Helicobacter pylori requires a posttranslational
modification for full activity. J Bacteriol. 186:777–784. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Kurokawa K, Lee H, Roh KB, Asanuma M, Kim
YS, Nakayama H, Shiratsuchi A, Choi Y, Takeuchi O, Kang HJ, et al:
The triacylated ATP binding cluster transporter substrate-binding
lipoprotein of Staphylococcus aureus functions as a native
ligand for Toll-like receptor 2. J Biol Chem. 284:8406–8411. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Kolenbrander PE, Andersen RN, Baker RA and
Jenkinson HF: The adhesion-associated sca operon in
Streptococcus gordonii encodes an inducible high-affinity
ABC transporter for Mn2+ uptake. J Bacteriol.
180:290–295. 1998.PubMed/NCBI
|
|
46
|
Griffin JE, Gawronski JD, DeJesus MA,
Ioerger TR, Akerley BJ and Sassetti CM: High-resolution phenotypic
profiling defines genes essential for mycobacterial growth and
cholesterol catabolism. PLoS Pathog. 7:e10022512011. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Akerley BJ, Rubin EJ, Novick VL, Amaya K,
Judson N and Mekalanos JJ: A genome-scale analysis for
identification of genes required for growth or survival of
Haemophilus influenzae. Proc Natl Acad Sci USA. 99:966–971. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Sassetti CM, Boyd DH and Rubin EJ:
Comprehensive identification of conditionally essential genes in
mycobacteria. Proc Natl Acad Sci USA. 98:12712–12779. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Weigel LM, Clewell DB, Gill SR, Clark NC,
McDougal LK, Flannagan SE, Kolonay JF, Shetty J, Killgore GE and
Tenover FC: Genetic analysis of a high-level vancomycin-resistant
isolate of Staphylococcus aureus. Science. 302:1569–1571.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Ramaswamy S and Musser JM: Molecular
genetic basis of antimicrobial agent resistance in Mycobacterium
tuberculosis: 1998 update. Tuber Lung Dis. 79:3–29. 1998.
View Article : Google Scholar : PubMed/NCBI
|