Introduction
Oral squamous cell carcinoma (OSCC) is a highly
variable disease with multiple heterogeneous genetic and epigenetic
changes that accounts for ~90% of malignant oral lesions (1). Despite the improvement in OSCC
treatments, the overall survival of patients has not improved
significantly during the past 20 years, with 5-year survival rates
of 45–60% (2,3). Local recurrence and progression to
metastatic disease are the primary causes of treatment failures,
while the cellular and molecular mechanisms underlying OSCC cell
biological behaviors remain to be elucidated. Therefore, a
comprehensive investigation of the factors and molecular events
that contribute to the local recurrence and invasion of OSCC are
necessary for the development of novel strategies for diagnosing
symptoms and treatment.
Calpains belong to a family of intracellular
Ca2+-dependent cysteine proteases and are widely
expressed with ubiquitous and tissue specific isoforms in higher
organisms (4). The calpain family is
classified according to their localization or to the presence or
absence of EF-hands, structures that allow calcium binding
(5). Calpain1 and Calpain-2, which
were named on the basis of the concentration of calcium ions
required for their activity in vitro, are commonly described
and ubiquitously expressed in the majority of tissues (5). The catalytic subunits differing between
µ-calpain and m-calpain are formed by calpain1 (encoded by CAPN1)
and calpain-2 (encoded by CAPN2), respectively (5–7). The
calpain system is involved in numerous cellular functions,
including cytoskeletal remodeling, cellular signaling and apoptosis
(8). Previously, cumulative evidence
has demonstrated that the expression of calpains is associated with
more aggressive tumor behavior in a number of tumor types,
including breast (9,10), ovarian (11), gastro-esophageal (12) and endometrial cancer (13). However, to the best of our knowledge,
the role of calpain isoforms has not been investigated in OSCC.
The aims of the present study were to investigate
the pattern and the prognostic impact of calpain1 expression in
OSCC. Furthermore, the functional basis of calpain1 as a potential
molecular marker was discussed when assessing the effect of
knockdown on OSCC cell migration, invasion, cell cycle stage and
apoptosis.
Materials and methods
Cell lines and cell culture
HSC-3 cells were purchased from the Cell Bank of
Japanese Collection of Research Bioresource (Shinjuku, Japan).
CAL27 cells were purchased from American Type Culture Collection
(Manassas, USA). CAL33 and HSC6 cells and the NOK-SI normal
keratinocyte cell line were provided by Professor J. Silvio Gutkind
(National Institute of Dental and Craniofacial Research, NIH,
Bethesda, MD, USA) subsequent to authentication by polymerase chain
reaction (PCR) amplification of short tandem repeats to ensure cell
identity. The OSCC cells were cultured in Dulbecco's modified
Eagle's medium (DMEM) containing 4.5 mM glucose (cat. no. 11995;
Gibco; Thermo Fisher Scientific, Inc., Waltham, MA, USA) and
supplemented with penicillin (5 U/ml), streptomycin (5 µg/ml) and
10% heat-inactivated fetal bovine serum (FBS; cat. no. 10099-141;
Gibco; Thermo Fisher Scientific, Inc.). NOK-SI cells were grown in
keratinocyte-SFMKSFM (cat. no. 17005-042; Gibco; Thermo Fisher
Scientific, Inc.).
Patients
The present study protocol was approved by the
Institutional Review Boards of the Hospital of Stomatology, Sun
Yat-Sen University (Guangzhou, China), and written informed consent
was obtained from all patients in the current study. The study
cohort consisted of 125 postoperative patients treated at the
Hospital of Stomatology (Sun Yat-Sen University) between January
2008 and January 2010, who had complete clinical and histological
data available. All patients were followed-up with interviews and
physical examinations every three months during the first year
following surgery, every six months in the following four years,
and once a year until the study endpoint. A biopsy or positron
emission tomography-computed tomography scan was conducted when
required, based on any questionable findings (symptoms associated
with recurrence). The survival time of each patient was calculated
from the day of surgery until the time of mortality from any cause,
or end of the follow-up.
Antibodies and western blot
analysis
For western blot analysis, four OSCC cell lines and
one normal oral keratinocyte cell line were used, along with a
total of seven pairs of patient OSCC and corresponding adjacent
normal tissues. OSCC specimens were immersed and stored in liquid
nitrogen immediately subsequent to surgical resection until
examination. The radioimmunoprecipitation assay (RIPA) lysis and
extraction buffer (cat. no. 89900; Thermo Fisher Scientific, Inc.)
were used for the extraction of proteins. All subsequent
manipulations were conducted on ice. The tissue was ground in
liquid nitrogen and cells were incubated in RIPA lysis and
extraction buffer (25 mM Tris×HCl pH 7.6, 150 mM NaCl, 1% NP-40, 1%
sodium deoxycholate, 0.1% SDS) mix for 30 min at 4°C and
centrifuged at 13,000 × g at 4°C for 20 min, and the
resultant supernatant was used for following tests.
The protein concentration of each cell sample was
determined with the bicinchoninic acid protein assay reagent
(Thermo Fisher Scientific, Inc.). Samples were denatured in sodium
dodecyl sulfate sample buffer (cat. no. 9173; Takara Biotechnology
Co., Ltd., Dalian, China) and loaded onto 10% polyacrylamide gels.
Subsequent to electrophoresis, the proteins were transferred onto
polyvinylidene fluoride membranes (EMD Millipore, Billerica, MA,
USA) and immunoblotted with rabbit anti-human monoclonal
anti-calpain 1 (H-65) antibody (cat. no. sc-13990; dilution,
1:1,000; Santa Cruz Biotechnology, Inc., Dallas, TX, USA), in
universal antibody dilution buffer (cat. no. U3635; Sigma-Aldrich;
Merck Millipore, Darmstadt, Germany) at 4°C overnight. Incubation
with biotin-conjugated goat anti-rabbit immunoglobulin G secondary
antibody (cat. no. BA1003; 0.1 µg/ml; Wuhan Boster Biological
Technology, Ltd.) was conducted, and the signals were visualized
with an enhanced chemiluminescence kit (Luminata Western HRP
Substrate; Merck Millipore). The expression of protein was
determined by western blotting using anti-GAPDH antibodies (cat.
no. 2118; dilution, 1:1,000; Cell Signaling Technology, Inc.,
Danvers, MA, USA) as loading controls. All kits were completed
according to the manufacturer's protocols. The result was analyzed
using ImageJ software version 1.5 (National Institutes of Health,
Bethesda, MD, USA) for the comparison of relative target protein
expression.
RNA isolation and quantitative PCR
(qPCR)
Total RNA from all cell lines was isolated with
TRIzol® reagent (Takara Biotechnology Co., Ltd.). In
brief, 1 ml TRIzol® reagent was added to each tube, and
the tubes were incubated for 5–10 min on ice to ensure complete
dissociation of the nucleoprotein complexes. A total of 0.2 ml
chloroform was then added to each RNA specimen and this was
agitated for 30 sec, followed by incubation at room temperature for
15 min. Following centrifugation at 13,000 × g for 15 min at
4°C, each RNA specimen was divided into three layers and the
aqueous phase containing RNA (upper layer) was removed and
transferred into a fresh RNase free 1.5 ml Eppendorf tube.
Isopropanol (0.5 ml) was added and the tubes were agitated for 30
sec, followed by incubation at room temperature for 10 min. The
Eppendorf tubes were then centrifuged at 13,000 × g for 10
min at 4°C to pellet the precipitated RNA. Taking care not to
disturb the RNA pellet, the supernatant was removed and the pellet
was subsequently washed by the addition of 1 ml 75% ethanol and
vortexed.
Following centrifugation at 13,000 × g for 5
min at 4°C, supernatant was removed (this wash step was repeated).
The RNA pellet was allowed to air-dry for 5–10 min and then
resuspended in 15 µl diethylpyrocarbonate-treated water. The RNA
isolated from cell line specimens was pooled, and the quantity and
quality of extracted RNA was assessed by reading absorbance at 260,
280 and 230 nm using a NanoDrop ND-1000 (NanoDrop Technologies;
Thermo Fisher Scientific, Inc., Wilmington, DE, USA). The
first-strand cDNA was synthesized from 1 µg total RNA using Oligo
(T) primer and the GoScript™ Reverse Transcription
system (Roche Applied Science, Penzberg, Germany), according to the
manufacturer's protocol. qPCR was performed
(LightCycler® 480; Roche Applied Science) with the
LightCycler® FastStart DNA Master SYBR-Green I kit (cat.
no. 03003230001; Roche Diagnostics, Basel, Switzerland), according
to the manufacturer's protocol. The primer sequences for CAPN1 were
as follows: Forward, 5′-AACTTCCTCATCACCAAC-3′ and reverse,
5′-CTCATCCTCCTCATCCTC-3′. The cycling conditions were 95°C for 5
min, followed by 45 cycles of 95°C for 10 sec, 60°C for 20 sec and
72°C for 30 sec. The cDNA content was determined using the
2−∆∆Cq method (14). The
results are presented as relative expression normalized to that of
the internal control, GAPDH (forward, 5′-GGTGGTCTCCTCTGACTTCAAC-3′
and reverse, 5′-TCTCTCTTCCTCTTGTGTTCTTG-3′). All primer sequences
were designed and synthesized by Invitrogen (Thermo Fisher
Scientific, Inc.). All experiments were performed in
triplicate.
Tissue microarray and
immunohistochemistry
The tissue microarray (TMA) was prepared by placing
duplicate 1.5-mm tissue cores of the tumor, which was identified by
a pathologist, into a single-recipient paraffin block. Sections of
the TMA (4 µm) were mounted on poly-L-lysine coated slides.
Sections were deparaffinized, rehydrated and treated using Dako
REAL peroxidase blocking solution (cat. no. S202386; Dako; Agilent
Technologies, Inc., Santa Clara, CA, USA). Heat-based antigen
retrieval was completed prior to incubation of the tissue sections
with a rabbit anti-human monoclonal anti-calpain1 (H-65) antibody
(cat. no. sc-13990; dilution, 1:50; Santa Cruz Biotechnology, Inc.)
at 4°C overnight. Bound antibodies were visualized using an
EnVision+ kit (Dako North America, Inc.; Agilent Technologies,
Inc.) according to the manufacturer's protocol.
Subsequent to the development of the expected stain
intensity, the sections were lightly counterstained with
hematoxylin. Sections treated without primary antibodies were used
as negative controls. Each block of TMA staining results were
confirmed by each tissue slide obtained from the same patient.
Immunostained cells were evaluated over 8 visual fields at a
magnification of ×400 under a light microscope (Olympus
Corporation, Tokyo, Japan). For the statistics of the prognostic
value in the OSCC cohort, two independent observers performed
microscopy analysis. The staining was scored using a 3-point scale
on the basis of the product of the staining intensity (weak, 1;
moderate, 2; strong, 3).
RNA interference
The calpain1 small interfering RNA (siRNA) sequences
were 5′-CCACGGAACUGCUGUCAAAdTdT-3′ and
5′-dTdTGGUGCCUUGACGACAGUUU-3′. The scrambled control siRNA
sequences and all siRNAs were synthesized by Guangzhou RiboBio Co.,
Ltd. (Guangzhou, China). The HSC3 and CAL27 OSCC tongue cancer cell
lines were used for RNA interference. Cells were plated
(2×105/well) onto 6-well plates and allowed to grow for
24 h, until they reached 70% confluency. Cells were then
transfected with 100 nmol/l of calpain1 siRNA or negative control
using Lipofectamine® 2000 (Invitrogen; Thermo Fisher
Scientific Inc.), according to the manufacturer's protocol.
Following 6 h of incubation, the medium was replaced with
serum-enriched medium and the cells were cultured for an additional
48 h. Subsequently, the transfected cells were collected and
processed for qPCR, western blot analysis, proliferation, cell
cycle stage, apoptosis, migration and invasion assays.
Cell invasion assay
Chambers (cat. no. 353096; BD Biosciences, Franklin
Lakes, NJ, USA) were uniformly coated with 60 µl Matrigel (BD
Biosciences) diluted with DMEM to ~20–30% and incubated at 37°C for
2–4 h. OSCC cells (1×105/well) in serum-free DMEM (0.1
ml) were placed into the upper chamber. The 10% FBS-containing DMEM
(used as a chemoattractant, 0.5 ml) was placed in the lower
chamber. Serum-free DMEM (0.5 ml) was used for the control. Cells
were allowed to invade for 24 h at 37°C in a 5% CO2
atmosphere. Subsequent to the incubation, non-invading cells were
removed from the top of the wells with a cotton swab, and the upper
chamber was washed twice with PBS. Cells that had transferred to
the inverse surface of the membrane were fixed by 4%
paraformaldehyde at room temperature and stained for 10 min with
0.2% crystal violet.
Cell migration assay
Cells (1.5×105/well) in serum-free DMEM
(0.1 ml) were plated in the top chamber, whereas the bottom
chambers were filled with 500 µl DMEM supplemented with 10% FBS.
Subsequent to incubation at 37°C in a 5% CO2 atmosphere, the cells
were fixed and stained. All experiments were conducted in
triplicate; cells were counted under a microscope at a
magnification of ×200, and cell numbers were counted in ≤5
fields/well.
MTT assay
The calpain1 knockdown transfectants were seeded
into 96-well plates at a density of 5×103/well.
Subsequent to culturing for 24, 48, 72 or 96 h, cell growth studies
were conducted using an MTT assay (cat. no. M5655; Sigma-Aldrich;
Merck Millipore). MTT (0.01 ml, 5 mg/ml) was added to each well and
incubated at 37°C for 3–4 h. The medium was then removed and the
cells were lysed using 0.1 ml isopropanol with 0.04 N HCl. The
absorbance of the converted dye was determined by a microplate
reader (Model Elx808; BioTek Instruments, Inc. Winooski, VT, USA)
at a wavelength of 570 nm, with background subtraction at 630–690
nm, and the optical density at 570 nm of the control (CAL27 or HSC3
cells treated without supplement) was considered to be 100%.
Cell cycle analysis
CAL27 and HSC3 cell lines transfected with CAPN1
siRNA were collected as aforementioned, and then resuspended in 70%
ethanol overnight at 4°C. Subsequently, cells were stained with 500
µl PI/RNase Staining Buffer (cat. no. 556463; BD Pharmingen, San
Diego, CA, USA), according to the manufacturer's protocol. Finally,
the analysis was conducted using flow cytometry (Beckman Coulter,
Inc., Brea, CA, USA) and FlowJo software version 10.0 (Tree Star,
Inc., Ashland, OR, USA).
Apoptosis analysis
Apoptosis was detected by flow cytometry 48 h after
OSCC cells were transfected with calpain1 siRNA, using Annexin
V-fluorescein isothiocyanate (FITC) Apoptosis Detection kit (BD
Biosciences; cat. no. 556547), according to the manufacturer's
protocol. Cells were harvested, washed twice in PBS, and
resuspended in Annexin V-FITC binding buffer, following the
addition of 5 µl Annexin V-FITC and 5 µl propidium iodide (PI), and
mixed gently. Cells were then stained at room temperature in the
dark for 15 min. Finally, 400 µl buffer was added to each tube and
the cell samples were analyzed by flow cytometry. The experiment
was conducted in triplicate.
Statistical analysis
Survival analysis was determined by the Kaplan-Meier
method, and the differences were compared on the basis of the
category of the variables using the log-rank test. To determine the
hazard ratio (HR), the univariate cox proportional-hazards model
was used. The multivariate cox proportional-hazards model was then
used to assess the association of overall survival (OS) rates with
the suggested variables (P<0.20) identified by the univariate
analysis. The variables in the final model were adjusted with
respect to age (as a continuous variable) and gender. Statistical
analyses were performed using SAS software, version 9.3 (SAS
Institute Inc., Cary, NC, USA). Unless stated otherwise, two-sided
values of P<0.05 were considered to indicate a statistically
significant difference.
Results
Patient cohort
The present cohort included 24 patients with
clinical stage I, 39 patients with stage II, 37 patients with stage
III and 25 patients with stage IV cancer. A total of 88 patients
were male and 37 were female. Pretreatment clinically positive
nodes were present in 41 patients. Tumor differentiation was
classified using the World Health Organization criteria (15) (88 patients, well differentiated; 32
patients, moderately differentiated; 5 patients, poorly
differentiated). Adjuvant postoperative radiation was added for
patients with histologically positive regional lymph nodes.
Resected specimens were used to create a tissue microarray and
slides for immunohistological analysis. Clinical and histological
variables assessed included T classification, nodal status, primary
tumor differentiation and clinical stage. Patient factors analyzed
included age, gender, smoking status (57 patients, never smokers;
55 patients, past smokers; 13 patients, current smokers) and
alcohol use (57 patients, never drinkers; 57 patients, past
drinkers; 11 patients, current drinkers). The demographics of the
patient cohort are presented in Table
I. All patients were regularly followed up. The last assessment
of vital status of all patients was in May 2014; therefore, the
median follow-up time for these patients was 3.58 years.
 | Table I.Common characteristics of the study
population (n=125). |
Table I.
Common characteristics of the study
population (n=125).
| Variables | Number, n (%) |
|---|
| Age, years |
|
| ≥60 | 67 (53.60) |
|
<60 | 58 (46.40) |
| Gender |
|
|
Female | 37 (29.60) |
| Male | 88 (70.40) |
| Smoking |
|
|
Never | 57 (45.60) |
| Past | 55(44.00) |
|
Current | 13(10.40) |
| Drinking |
|
|
Never | 57 (45.60) |
| Past | 57 (45.60) |
|
Current | 11 (8.80) |
| T-stage |
|
| 1 | 34 (27.20) |
| 2 | 52 (41.60) |
| 3 | 22 (17.60) |
| 4 | 17 (13.60) |
| Lymphatic
metastasis |
|
| No | 84 (67.20) |
|
Yes | 41 (32.80) |
|
Differentiation |
|
|
Well | 88 (70.40) |
|
Moderate | 32 (25.60) |
|
Poor | 5 (4.0) |
| Clinical stage |
|
| 1 | 24 (19.20) |
| 2 | 39 (31.20) |
| 3 | 37 (29.60) |
| 4 | 25 (20.00) |
| Surgical
methoda |
|
| 1 | 53 (42.40) |
| 2 | 65 (52.00) |
| 3 | 7 (5.60) |
| Radiotherapy
(Yes) | 9 (7.20) |
| Preoperative
chemotherapy (Yes) | 8 (6.40) |
| Postoperative
chemotherapy (Yes) | 62 (49.60) |
| Calpain1 staining
intensity |
|
|
Low | 17 (13.60) |
|
Median | 35 (28.00) |
|
High | 73 (58.40) |
Expression pattern of calpain1 in OSCC
cell lines and tissues
To identify differing protein expression of calpain1
in OSCC, four OSCC cell lines and seven fresh tumor tissues
together with the paired NCMTs were examined by western blot
analysis. The present study found that calpain1 was highly
expressed in HSC3, HSC6, CAL33 and CAL27 OSCC cell lines, in
comparison with normal oral keratinocytes (Fig. 1A; HSC3, P=0.029; CAL27, P=0.0023;
CAL33, P=0.0002; HSC6, P=0.0007). In addition, calpain1 was highly
expressed in the tumor areas of 4 NCMT/tumor pairs (Fig. 1B; P=0.0008). Increased calpain1
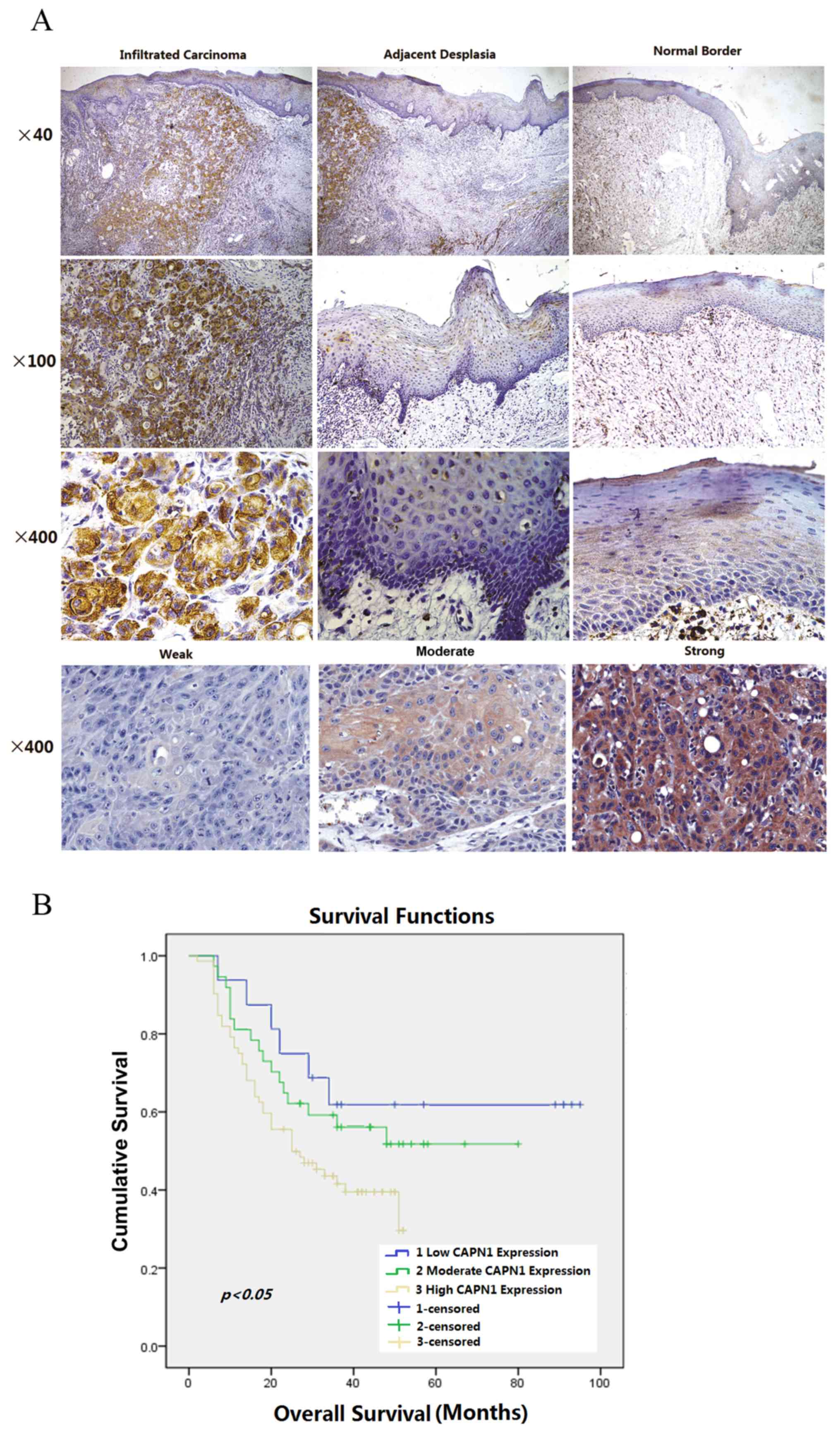
expression in tumor tissues is also presented in Fig. 2A, with representative images of normal
border epithelium, precancerous epithelium and infiltrative
carcinoma on the same tissue slide. Expression of calpain1 was
located in the cytoplasm, with a certain degree of heterogeneity
within samples. Typical staining patterns, which vary between weak,
moderate and strong, are presented in Fig. 2A.
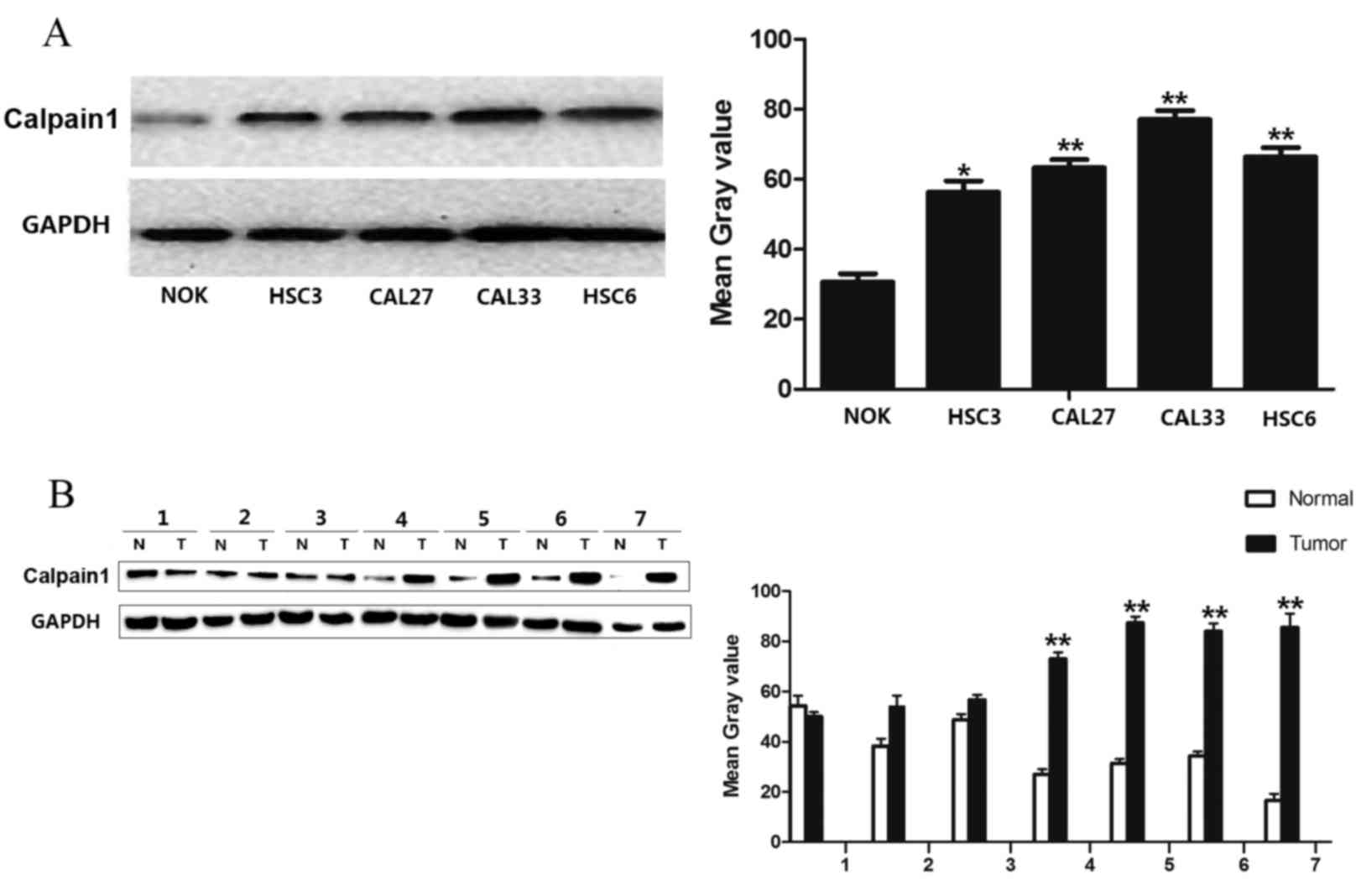 | Figure 1.Expression of calpain1 in OSCC cell
lines and tissues by western blot analysis. (A) Expression of
calpain1 in 4 oral cancer cell lines (HSC3, CAL27, CAL33 and HSC6)
and in a normal oral keratinocyte (NOK-SI) cell line. The graph
represents the mean grey value of the average data of five samples
from 3 independent experiments (*P<0.05, Student's
t-test). HSC3, CAL33, CAL27 and HSC6 cell lines exhibited
higher protein expression levels, compared with NOK. (B) Expression
of calpain1 in 7 paired NCMT (N)/OSCC (T) samples. The graph
represents the mean grey value of the average data of 7 paired
samples (**P<0.01, Student's t-test). Calpain1 was highly
expressed in the tumor areas of 4 NCMT/tumor pairs. OSCC, oral
squamous cell carcinoma; NCMT, noncancerous matched tissues; N,
normal; T, tumor. |
Association between calpain1
expression and survival analysis
Immunohistochemical analysis was completed to assess
the expression of calpain1 in 125 OSCC tissue blocks. Overall, 73
of the 125 tumor samples (58.4%) exhibited high expression of
calpain1 (immunoreactivity intensity scale, 3), whereas 17 samples
(13.6%) had low expression (immunoreactivity intensity scale, 1).
The other 35 samples (28%) exhibited moderate calpain1 expression
(immunoreactivity intensity scale, 2; Table I).
Kaplan-Meier analysis and the log-rank test were
used to analyze the effect of the calpain1 expression on OSCC
survival. The expression of calpain1 was assessed for association
with a number of clinicopathological variables. In the univariate
analysis, patient age (β=−0.022; 95% HR, 0.959–0.999; P=0.037),
cell differentiation (β=−0.561; 95% HR, 0.345–0.945; P=0.029) and
calpain1 expression (β=0.414; 95% HR, 1.039–2.204; P=0.031) were
significantly associated with overall survival. Radiotherapy and
chemotherapy (pre- and post-surgery) were not associated with
overall survival. Furthermore, multivariate analysis revealed that
calpain1 expression was an independent predictor for the overall
survival of OSCC patients (β=0.455; 95% HR, 1.069–2.322; P=0.022;
Table II). With respect to overall
survival, marked separation in curves between patients with low,
median and high calpain1 expression level was observed in the
present cohort (Fig. 2B).
 | Table II.Univariate and multivariate analyses
of overall survival among patients with oral squamous cell
carcinoma. |
Table II.
Univariate and multivariate analyses
of overall survival among patients with oral squamous cell
carcinoma.
|
| Univariate
analyses | Multivariate
analyses |
|---|
|
|
|
|
|---|
| Variable | β | 95% HR | P-value | β | 95% HR | P-value |
|---|
| Age | −0.022 | 0.959–0.959 | 0.037a | −0.337 | 0.387–1.387 | 0.28 |
| Gender | 0.26 | 0.772–2.772 | 0.326 | 0.137 | 0.646–2.646 | 0.64 |
| Smoking
history | −0.384 | 0.417–1.417 | 0.681 |
|
|
|
| Drinking
history | −0.084 | 0.562–1.562 | 0.738 |
|
|
|
| Cell
differentiation | −0.561 | 0.345–0.345 | 0.029a | −1.807 | 0.023–1.023 | 0.073 |
| Tumor size | 0.119 | 0.878–1.878 | 0.348 |
|
|
|
| Lymphatic
metastasis | 0.258 | 0.808–2.808 | 0.283 |
|
|
|
| Clinical TNM
stage | 0.217 | 0.971–1.971 | 0.085 |
|
|
|
| Radiotherapy | −1.74 | 0.024–1.024 | 0.085 |
|
|
|
| Chemotherapy before
surgery | −0.682 | 0.123–2.123 | 0.506 |
|
|
|
| Chemotherapy after
surgery | −0.234 | 0.477–1.477 | 0.366 |
|
|
|
| Calpain1 | 0.414 | 1.039–2.039 | 0.031a | 0.455 | 1.069–2.069 | 0.022a |
Effect of calpain1 gene silence on the
migration and invasion of OSCC cells
To determine the effect of calpain1 on the invasion
and migration potential of tumor cells, the present study
transfected HSC3 and CAL27 cells with calpain1 siRNA and conducted
Transwell migration and invasion assays. As observed in Fig. 3A and B, the protein and mRNA
expression of calpain1 in cancer cells was downregulated subsequent
to transfection with calpain1 siRNA, as compared with the negative
control (P<0.0001). The induced downregulation of calpain1
expression resulted in apparent CAL27 and HSC3 cell decrease in
number of invading and migrating cells, compared with the siRNA
negative controls at 24 h following seeding (P=0.00057 for
migration; P=0.0001 for invasion; Fig. 3C
and D). The results indicated that knockdown of calpain1
expression was able to suppress the mobility of CAL27 and HSC3
cells in vitro.
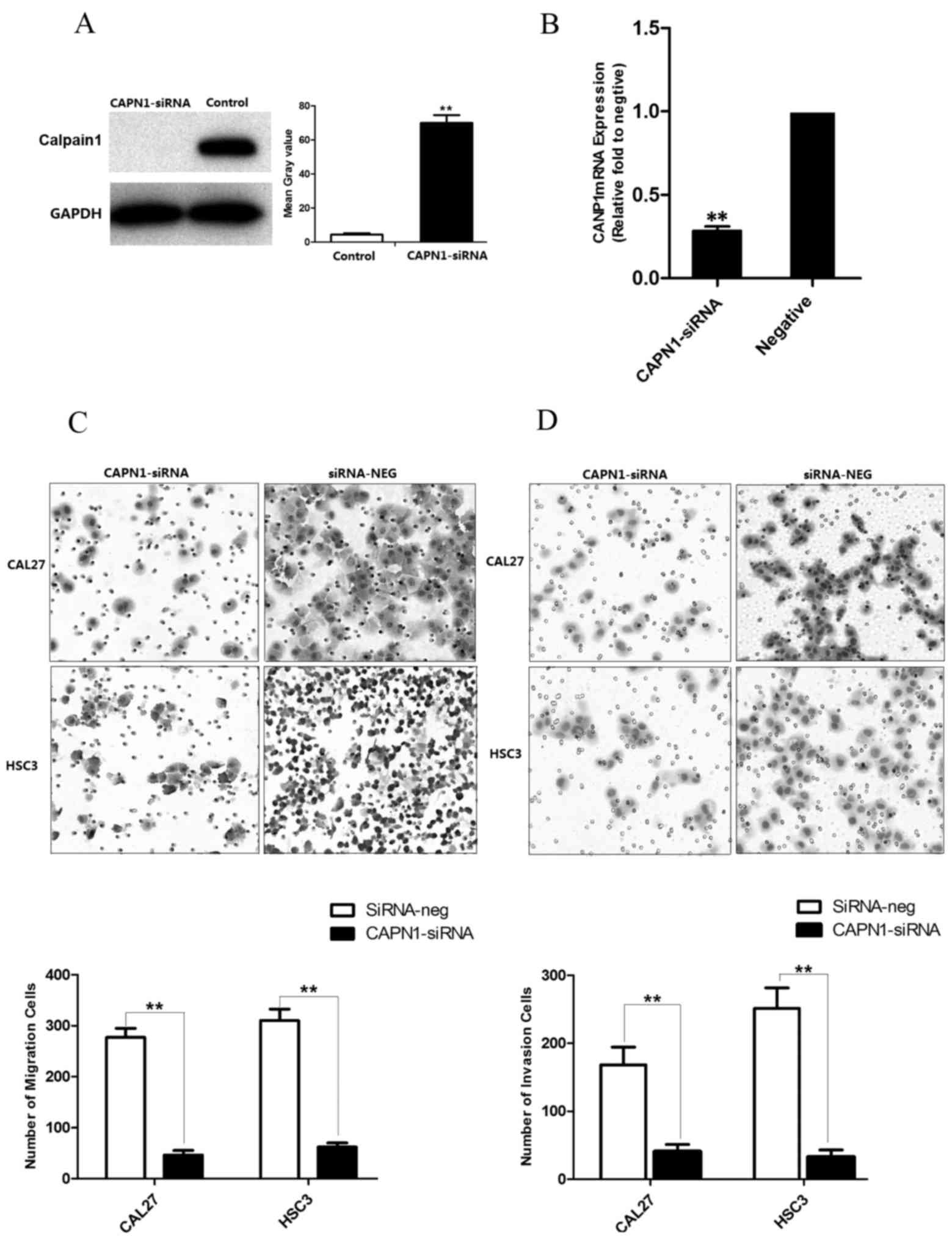 | Figure 3.Calpain1 silencing decreased the
migration and invasion capacity of OSCC cells. (A) Effective
interference of calpain1 expression in the two OSCC cell lines was
demonstrated by western blot analysis (**P<0.01, compared with
the negative control group). (B) Effective interference of calpain1
expression in OSCC cell lines was demonstrated by qPCR
(**P<0.01, compared with the negative control group). (C) Cell
migration was decreased by silencing calpain1 in OSCC cell lines
(CAL27, HSC3), as revealed by Transwell migration assay at 24 h.
Cells that had migrated to the bottom of the chamber were counted
in 5 fields at ×20 magnification. Representative images and
quantitative data are presented. Data are expressed as the mean ±
standard deviation (**P<0.01, compared with the negative control
group). (D) Cell invasion was decreased by silencing calpain1 in
OSCC cell lines, as revealed by invasion assay at 24 h (right
panels). Cells that had invaded to the bottom of the chamber were
counted in five fields ×20 magnification. Representative images and
quantitative data are presented. Data are expressed as the mean ±
standard deviation (**P<0.01, compared with the negative control
group). OSCC, oral squamous cell carcinoma; CAPN1, calpain1; siRNA,
small interfering RNA; neg, negative. |
Effect of calpain1 knockdown on the
cell proliferation
To assess the effect of siRNA-mediated calpain1
knockdown on the proliferation of OSCC cells, the cell viability of
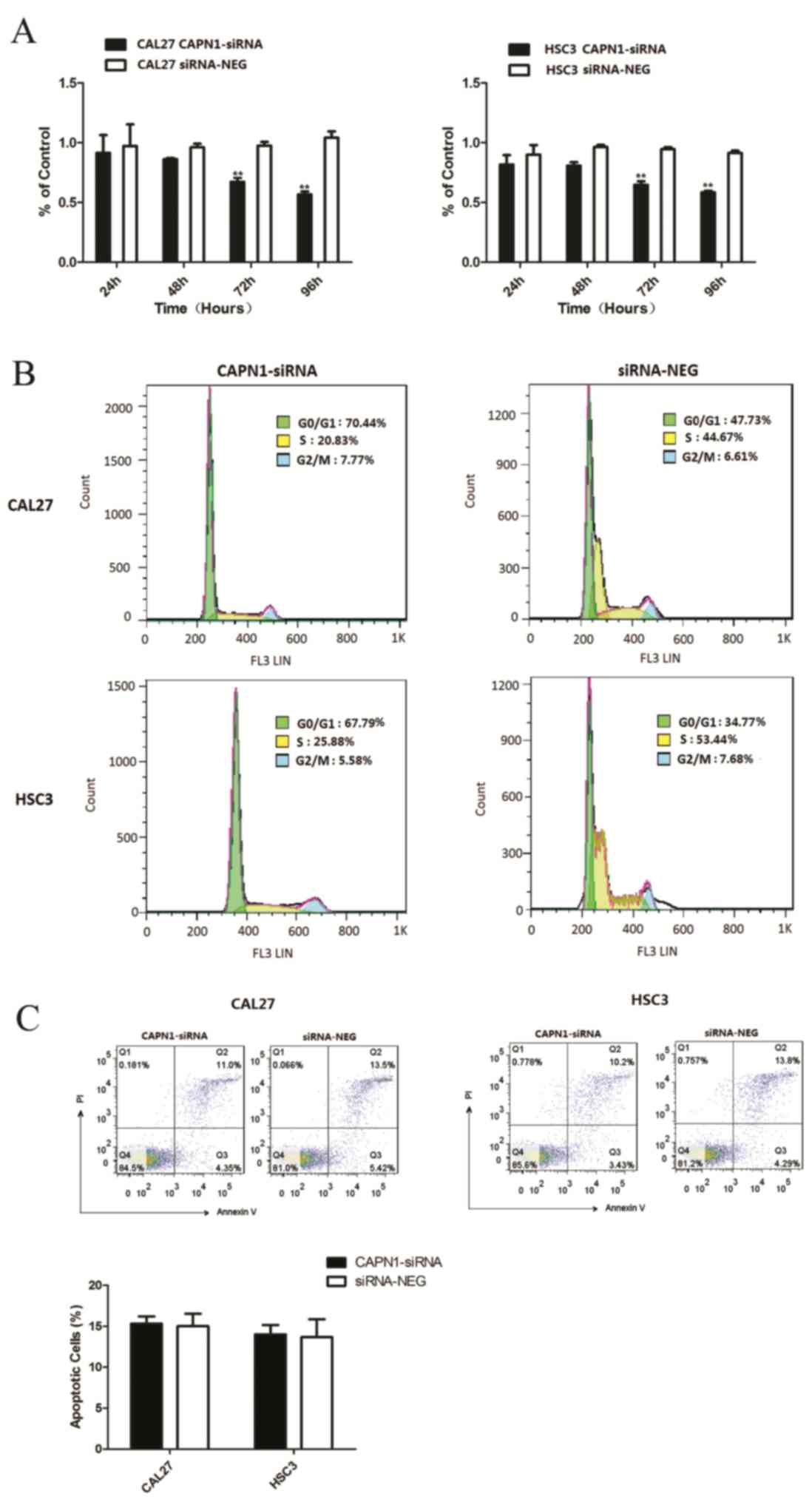
two cell lines was determined by MTT assay. In Fig. 4A, relative cell growth rate was
significantly reduced in the calpain1 siRNA group following
transfection, at 72 and 96 h, by comparison with that of the
siRNA-negative groups (72 h, P=0.0027 for CAL27, P=0.009 for HSC3;
96 h, P=0.0012 for CAL27, P=0.0001 for HSC3). The flow cytometry
analysis revealed that cells transfected with caplain-1 siRNA were
blocked in the G0/G1 phase (47.73–70.44% for CAL27, P=0.0028;
34.77–67.79% for HSC3, P=0.0014; Fig.
4B; Table III). The analysis
also revealed a concomitant decrease of cells in the S phase in the
knockdown cell line (44.67–20.83% for CAL27, P=0.006; 53.44–25.88%
for HSC3, P=0.0046). These findings suggest that the downregulation
of calpain1 resulted in cell cycle arrest and therefore inhibited
the proliferation of OSCC cells. The results indicated that
calpain1 may promote OSCC cell cycle progression.
 | Table III.The percentages of cell in various
phases of the cell cycle. |
Table III.
The percentages of cell in various
phases of the cell cycle.
|
| Cell-cycle phase
(%) |
|---|
|
|
|
|---|
| Groups |
G0/G1 | S |
G2/M |
|---|
| CAL27 |
|
|
|
|
Negative control | 43.21±4.40 | 46.12±2.36 | 9.70±2.71 |
|
CAPN1-siRNA |
71.33±1.02a |
20.41±1.42a | 7.08±0.78 |
| HSC3 |
|
|
|
|
Negative control | 33.50±1.55 | 54.24±1.92 | 9.56±2.05 |
|
CAPN1-siRNA |
69.42±1.63a |
23.99±2.14a | 5.77±0.43 |
Effect of calpain1 knockdown on the
apoptosis of OSCC cells
In order to determine whether CAPN1 knockdown may
induce apoptosis in OSCC cell lines, Annexin V/PI staining was
examined by flow cytometry analysis. As presented in Fig. 4C, the proportion of Annexin V and PI
positive cells was 12±0.16% in calpain1 siRNA transfected cells and
14±0.51% in siRNA-negative cells. The two populations of Annexin
V-positive cells and of Annexin V plus PI-positive cells in 2
groups exhibited no significant difference (P=0.8593 for CAL27;
P=0.8993 for HSC3; Fig. 4C). These
data suggest that calpain1 may have no significant effect on
apoptosis in OSCC cell lines.
Discussion
The role of calpains in cancer pathology has been
widely discussed (8). Among the
different members of the calpain family, calpain1 and −2 are most
widely implicated in the development and progression of cancer
(16). However, little is understood
about the biological significance of calpains in OSCC. The present
study demonstrated the expression of calpain1 in OSCC cell lines
and tissues. Additionally, the high expression of calpain1 was
significantly associated with the poor prognosis of patients with
OSCC. Accordingly, obstruction of calpain1 by specific siRNAs
hindered the progression of OSCC cells by affecting migration,
invasion and proliferation, but not apoptosis. To the best of our
knowledge, these results are the first observations to demonstrate
that calpain1 may potentially be important in OSCC progression, and
may be associated with poor patient prognosis.
In the present study, it was observed that calpain1
was overexpressed in OSCC cell lines and paired OSCC tissues. Its
role as a significant predictor of poor outcome was indicated in
the study. The current findings are consistent with the results of
previous studies that have demonstrated the association between
elevated calpain1 levels and poor survival rates in breast
(10) and ovarian cancer (11). As the calpain isoforms possess
tumor-type specific patterns, the opposite observations have been
made in gastro-esophageal adenocarcinoma (12). In the present study, the majority of
the common used patient variables, including epidemiological and
clinicopathological factors, were included in the regression model,
and only calpain1 expression remained a significant predictor for
poor outcome (P<0.05).
To study the effects of calpain1 in the development
of oral cancer cells, the present study selected two oral cancer
cell lines with transient calpain1 knockdown, and investigated how
this gene may affect cell invasion and migration. It was revealed
that, in addition to morphological alterations, deletion of
calpain1 may be associated with a decrease in the migratory and
invasive ability of OSCC cells. Certain studies have demonstrated
that calpain1 is able to modulate the expression and secretion of
MMPs (17,18), which are proteinases that are able to
degrade numerous extracellular matrix components, making the
invasion of neoplastic cells possible. Therefore, targeting the
activity of calpain with specific inhibitors may be a feasible way
to suppress the dissemination of tumor cells. However, treatments
targeting calpain activity exhibit no effects on invasiveness in
the case of amoeboid migration and invasion (19), calpain inhibition is even able to
promote HT1080 cell invasion that is sensitive to a Rho-associated
protein kinase inhibitor, which suppresses amoeboid invasion
(19). Therefore, further analysis of
the biological functions of calpain family members within complex
cellular or in vivo systems is important.
The calpains were notably shown to regulate the cell
cycle, particularly the transition between the G1 and S
phases through underlying mechanisms that include several crucial
regulators, including cyclin D1 and cyclin E, which are substrates
of calpain1 and calpain2 (20,21). In
cancer cells, calpains have been demonstrated to proteolyze two
inhibitors of cyclin-dependent kinases (Cdk), p21Cip1
and p27Kip1, leading to the activation of the complexes
cyclinD-Cdk4 and cyclinE-Cdk2 (22,23). By
regulating these proteins, calpains are therefore strongly involved
in the progression of the cell cycle and in cellular proliferation.
The present results also demonstrated that calpain1 knockdown
resulted in a significantly reduced growth rate in two OSCC cell
lines and triggered a marked accumulation of cells in
G0/G1 phase and a concomitant decrease of
cells in S phase. These findings suggest that the depletion of
calpain1 may account for the slow growth of the calpain1
transfected cell line.
To additionally understand the effect of calpain1
activity on OSCC cell apoptosis, the current study conducted an
apoptosis assay. The results demonstrated that calpain1 knockdown
did not induce marked cell apoptosis, compared with that in the
negative control group (P>0.05). The functions of calpain in the
perturbed cell apoptosis of cancer remain to be established.
Certain studies indicate that calpain is able to cleave wild-type
p53, regulating protein stability to prevent p53-dependent
apoptosis (24,25). However, in other studies, calpain
activity is able to promote apoptosis in neurodegenerative
disorders (26), and calpain1 was
demonstrated to suppress the differentiation of neural stem cells
(27). The present findings suggest
that calpain1 knockdown may have no significant effect on apoptosis
in OSCC cell lines. At present, there is no clear explanation for
these inconsistent results; the detailed underlying molecular
mechanisms require investigation and further studies are necessary
in a number of models.
In conclusion, the present study indicates that
calpain1 may provide important prognostic information in OSCC and
contributes to the malignant progression of OSCC. As part of future
studies, multi-center, non-selected patients with OSCC must be
investigated to test if determining calpain1 expression may be of
clinical benefit. Additional studies are warranted to enable a good
understanding of the function and mechanism of calpain1 in
vivo, which may provide a potential therapeutic target for
OSCC.
Acknowledgements
This study was supported by grants from The National
Natural Science Foundations of China (grant nos. 81272954 and
81272524).
References
|
1
|
Wu RQ, Zhao XF, Wang ZY, Zhou M and Chen
QM: Novel molecular events in oral carcinogenesis via integrative
approaches. J Dent Res. 90:561–572. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Scully C and Bagan JV: Recent advances in
oral oncology 2008; squamous cell carcinoma imaging, treatment,
prognostication and treatment outcomes. Oral Oncol. 45:e25–e30.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Rusthoven K, Ballonoff A, Raben D and Chen
C: Poor prognosis in patients with stage I and II oral tongue
squamous cell carcinoma. Cancer. 112:345–351. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Perrin BJ and Huttenlocher A: Calpain. Int
J Biochem Cell Biol. 34:722–725. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Goll DE, Thompson VF, Li H, Wei W and Cong
J: The calpain system. Physiol Rev. 83:731–801. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Ono Y and Sorimachi H: Calpains: An
elaborate proteolytic system. Biochim Biophys Acta. 1824:224–236.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Croall DE and Ersfeld K: The calpains:
Modular designs and functional diversity. Genome Biol. 8:2182007.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Storr SJ, Carragher NO, Frame MC, Parr T
and Martin SG: The calpain system and cancer. Nat Rev Cancer.
11:364–374. 2011. View
Article : Google Scholar : PubMed/NCBI
|
|
9
|
Storr SJ, Woolston CM, Barros FF, Green
AR, Shehata M, Chan SY, Ellis IO and Martin SG: Calpain-1
expression is associated with relapse-free survival in breast
cancer patients treated with trastuzumab following adjuvant
chemotherapy. Int J Cancer. 129:1773–1780. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Storr SJ, Mohammed RA, Woolston CM, Green
AR, Parr T, Spiteri I, Caldas C, Ball GR, Ellis IO and Martin SG:
Calpastatin is associated with lymphovascular invasion in breast
cancer. Breast. 20:413–418. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Storr SJ, Safuan S, Woolston CM,
Abdel-Fatah T, Deen S, Chan SY and Martin SG: Calpain-2 expression
is associated with response to platinum based chemotherapy,
progression-free and overall survival in ovarian cancer. J Cell Mol
Med. 16:2422–2428. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Storr SJ, Pu X, Davis J, Lobo D,
Reece-Smith AM, Parsons SL, Madhusudan S and Martin SG: Expression
of the calpain system is associated with poor clinical outcome in
gastro-oesophageal adenocarcinomas. J Gastroenterol. 48:1213–1221.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Salehin D, Fromberg I, Haugk C, Dohmen B,
Georg T, Bohle RM, Bauerschlag D, Maass N and Friedrich M:
Immunhistochemical analysis for expression of calpain1, calpain 2
and calpastatin in endometrial cancer. Anticancer Res.
30:2837–2843. 2010.PubMed/NCBI
|
|
14
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Hamilton SR and Aaltonen LA: World Health
Organization Classification of Tumours: Pathology and and Genetics
of Tumours of the Digestive System. IARC Press; Lyon, France:
2000
|
|
16
|
Leloup L and Wells A: Calpains as
potential anti-cancer targets. Expert Opin Ther Targets.
15:309–323. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Popp O, Heidinger M, Ruiz-Heinrich L, Ries
C, Jochum M and Gil-Parrado S: The calpastatin-derived calpain
inhibitor CP1B reduces mRNA expression of matrix
metalloproteinase-2 and −9 and invasion by leukemic THP-1 cells.
Biol Chem. 384:951–958. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Fan DG, Dai JY, Tang J, Wu MM, Sun SG,
Jiang JL and Fan QY: Silencing of calpain expression reduces the
metastatic potential of human osteosarcoma cells. Cell Biol Int.
33:1263–1267. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Carragher NO, Walker SM, Carragher LA
Scott, Harris F, Sawyer TK, Brunton VG, Ozanne BW and Frame MC:
Calpain 2 and Src dependence distinguishes mesenchymal and amoeboid
modes of tumour cell invasion: A link to integrin function.
Oncogene. 25:5726–5740. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Choi YH, Lee SJ, Nguyen P, Jang JS, Lee J,
Wu ML, Takano E, Maki M, Henkart PA and Trepel JB: Regulation of
cyclin D1 by calpain protease. J Biol Chem. 272:28479–28484. 1997.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Wang XD, Rosales JL, Magliocco A,
Gnanakumar R and Lee KY: Cyclin E in breast tumors is cleaved into
its low molecular weight forms by calpain. Oncogene. 22:769–774.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Chen Z, Knutson E, Kurosky A and Albrecht
T: Degradation of p21cip1 in cells productively infected with human
cytomegalovirus. J Virol. 75:3613–3625. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Delmas C, Aragou N, Poussard S, Cottin P,
Darbon JM and Manenti S: MAP kinase-dependent degradation of
p27Kip1 by calpains in choroidal melanoma cells. Requirement of
p27Kip1 nuclear export. J Biol Chem. 278:12443–12451. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Atencio IA, Ramachandra M, Shabram P and
Demers GW: Calpain inhibitor 1 activates p53-dependent apoptosis in
tumor cell lines. Cell Growth Differ. 11:247–253. 2000.PubMed/NCBI
|
|
25
|
Del Bello B, Moretti D, Gamberucci A and
Maellaro E: Cross-talk between calpain and caspase-3/−7 in
cisplatin-induced apoptosis of melanoma cells: A major role of
calpain inhibition in cell death protection and p53 status.
Oncogene. 26:2717–2726. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Raynaud F and Marcilhac A: Implication of
calpain in neuronal apoptosis. A possible regulation of Alzheimer's
disease. FEBS J. 273:3437–3443. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Santos DM, Xavier JM, Morgado AL, Solá S
and Rodrigues CM: Distinct regulatory functions of calpain 1 and 2
during neural stem cell self-renewal and differentiation. PLoS One.
7:e334682012. View Article : Google Scholar : PubMed/NCBI
|