|
1
|
Bosetti C, Bertuccio P, Chatenoud L, Negri
E, La Vecchia C and Levi F: Trends in mortality from urologic
cancers in Europe, 1970–2008. Eur Urol. 60:1–15. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Chung BH: The role of radical
prostatectomy in high-risk prostate cancer. Prostate Int. 1:95–101.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2016. CA Cancer J Clin. 66:7–30. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Dell'Oglio P, Karnes RJ, Joniau S, Spahn
M, Gontero P, Tosco L, Fossati N, Kneitz B, Chlosta P and Graefen
M: Very long-term survival patterns of young patients treated with
radical prostatectomy for high-risk prostate cancer. Urol Oncol.
34:234. e13–9. 2016.
|
|
5
|
Saad F and Miller K: Treatment options in
castration-resistant prostate cancer: Current therapies and
emerging docetaxel-based regimens. Urol Oncol. 32:70–79. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Thomsen FB, Brasso K, Klotz LH, Røder MA,
Berg KD and Iversen P: Active surveillance for clinically localized
prostate cancer-a systematic review. J Surg Oncol. 109:830–835.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Cole G, McCaffrey J, Ali AA and McCarthy
HO: DNA vaccination for prostate cancer: Key concepts and
considerations. Cancer Nanotechnol. 6:22015. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Cross D and Burmester JK: Gene therapy for
cancer treatment: Past, present and future. Clin Med Res. 4:218–27.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
van Asperen CJ, Brohet RM,
Meijers-Heijboer EJ, Hoogerbrugge N, Verhoef S, Vasen HF, Ausems
MG, Menko FH, Garcia Gomez EB and Klijn JG: Cancer risks in BRCA2
families: Estimates for sites other than breast and ovary. J Med
Genet. 42:711–719. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Leongamornlert D, Mahmud N, Tymrakiewicz
M, Saunders E, Dadaev T, Castro E, Goh C, Govindasami K, Guy M and
O'Brien L: Germline BRCA1 mutations increase prostate cancer risk.
Br J Cancer. 106:1697–1701. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Khandrika L, Kumar B, Koul S, Maroni P and
Koul HK: Oxidative stress in prostate cancer. Cancer Lett.
282:125–136. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Campaner S and Amati B: Two sides of the
Myc-induced DNA damage response: From tumor suppression to tumor
maintenance. Cell Div. 7:62012. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Toren P and Zoubeidi A: Targeting the
PI3K/Akt pathway in prostate cancer: Challenges and opportunities
(review). Int J Oncol. 45:1793–1801. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Ngalame NN, Tokar EJ, Person RJ, Xu Y and
Waalkes MP: Aberrant microRNA expression likely controls RAS
oncogene activation during malignant transformation of human
prostate epithelial and stem cells by arsenic. Toxicol Sci.
138:268–277. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Varambally S, Dhanasekaran SM, Zhou M,
Barrette TR, Kumar-Sinha C, Sanda MG, Ghosh D, Pienta KJ, Sewalt RG
and Otte AP: The polycomb group protein EZH2 is involved in
progression of prostate cancer. Nature. 419:624–629. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Conesa-Zamora P, García-Solano J,
García-García F, Mdel Turpin C, Trujillo-Santos J, Torres-Moreno D,
Oviedo-Ramírez I, Carbonell-Muñoz R, Muñoz-Delgado E and
Rodriguez-Braun E: Expression profiling shows differential
molecular pathways and provides potential new diagnostic biomarkers
for colorectal serrated adenocarcinoma. Int J Cancer. 132:297–307.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Wu C, Zhu J and Zhang X: Network-based
differential gene expression analysis suggests cell cycle related
genes regulated by E2F1 underlie the molecular difference between
smoker and non-smoker lung adenocarcinoma. BMC Bioinformatics.
14:3652013. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Li Y, Vongsangnak W, Chen L and Shen B:
Integrative analysis reveals disease-associated genes and
biomarkers for prostate cancer progression. BMC Med Genomics. 7
(Suppl 1):S32014. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Aryee MJ, Liu W, Engelmann JC, Nuhn P,
Gurel M, Haffner MC, Esopi D, Irizarry RA, Getzenberg RH and Nelson
WG: DNA methylation alterations exhibit intraindividual stability
and interindividual heterogeneity in prostate cancer metastases.
Sci Transl Med. 5:169ra102013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Lapointe J, Li C, Higgins JP, van de Rijn
M, Bair E, Montgomery K, Ferrari M, Egevad L, Rayford W and
Bergerheim U: Gene expression profiling identifies clinically
relevant subtypes of prostate cancer. Proc Natl Acad Sci USA.
101:811–816. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Johnson WE, Li C and Rabinovic A:
Adjusting batch effects in microarray expression data using
empirical Bayes methods. Biostatistics. 8:118–127. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Leek JT and Storey JD: A general framework
for multiple testing dependence. Proc Natl Acad Sci USA.
105:18718–18723. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW,
Shi W and Smyth GK: limma powers differential expression analyses
for RNA-sequencing and microarray studies. Nucleic Acids Res.
43:e472015. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Pounds S and Morris SW: Estimating the
occurrence of false positives and false negatives in microarray
studies by approximating and partitioning the empirical
distribution of P-values. Bioinformatics. 19:1236–1242. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Shoop E, Casaes P, Onsongo G, Lesnett L,
Petursdottir EO, Donkor EK, Tkach D and Cosimini M: Data
exploration tools for the gene ontology database. Bioinformatics.
20:3442–3454. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Kanehisa M, Goto S, Kawashima S, Okuno Y
and Hattori M: The KEGG resource for deciphering the genome.
Nucleic Acids Res. 32:D277–280. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
da Huang W, Sherman BT and Lempicki RA:
Systematic and integrative analysis of large gene lists using DAVID
bioinformatics resources. Nature Protoc. 4:44–57. 2009. View Article : Google Scholar
|
|
28
|
da Huang W, Sherman BT and Lempicki RA:
Bioinformatics enrichment tools: Paths toward the comprehensive
functional analysis of large gene lists. Nucleic Acids Res.
37:1–13. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
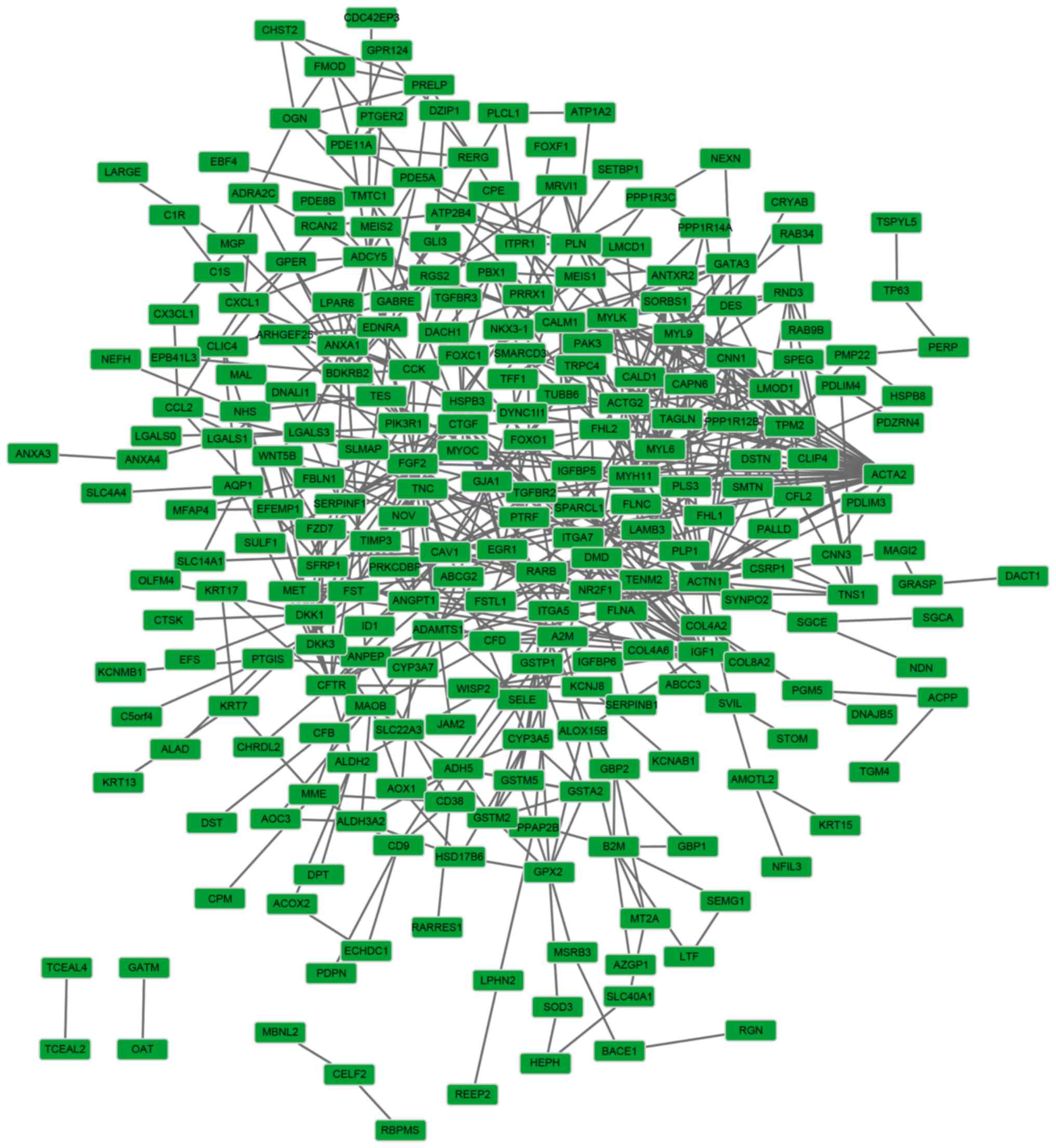
Franceschini A, Szklarczyk D, Frankild S,
Kuhn M, Simonovic M, Roth A, Lin J, Minguez P, Bork P, von Mering C
and Jensen LJ: STRING v9.1: Protein-protein interaction networks,
with increased coverage and integration. Nucleic Acids Res.
41:D808–D815. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Kohl M, Wiese S and Warscheid B:
Cytoscape: Software for visualization and analysis of biological
networks. Methods Mol Biol. 696:291–303. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Albert R, Albert I and Nakarado GL:
Structural vulnerability of the North American power grid. Physical
Review E. 69:0251032004. View Article : Google Scholar
|
|
32
|
Bader GD and Hogue CW: An automated method
for finding molecular complexes in large protein interaction
networks. BMC Bioinformatics. 4:22003. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Matys V, Kel-Margoulis OV, Fricke E,
Liebich I, Land S, Barre-Dirrie A, Reuter I, Chekmenev D, Krull M
and Hornischer K: TRANSFAC and its module TRANSCompel:
Transcriptional gene regulation in eukaryotes. Nucleic Acids Res.
34:D108–110. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Chen K and Rajewsky N: The evolution of
gene regulation by transcription factors and microRNAs. Nat Rev
Genet. 8:93–103. 2007. View
Article : Google Scholar : PubMed/NCBI
|
|
35
|
Seong J, Wang N and Wang Y:
Mechanotransduction at focal adhesions: From physiology to cancer
development. J Cell Mol Med. 17:597–604. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Fu LJ and Wang B: Investigation of the hub
genes and related mechanism in ovarian cancer via bioinformatics
analysis. J Ovarian Res. 6:922013. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Huang SQ, Liao QJ, Wang XW, Xin DQ, Chen
SX, Wu QJ and Ye G: RNAi-mediated knockdown of pituitary
tumor-transforming gene-1 (PTTG1) suppresses the proliferation and
invasive potential of PC3 human prostate cancer cells. Braz J Med
Biol Res. 45:995–1001. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Summers MK, Pan B, Mukhyala K and Jackson
PK: The unique N terminus of the UbcH10 E2 enzyme controls the
threshold for APC activation and enhances checkpoint regulation of
the APC. Mol Cell. 31:544–556. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Chen Z, Zhang C, Wu D, Chen H, Rorick A,
Zhang X and Wang Q: Phospho-MED1-enhanced UBE2C locus looping
drives castration-resistant prostate cancer growth. EMBO J.
30:2405–2419. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Huang S, Lin J, Guo N, Zhang M, Yun X, Liu
S, Zhou J, He E and Skog S: Elevated serum thymidine kinase 1
predicts risk of pre/early cancerous progression. Asian Pac J
Cancer Prev. 12:497–505. 2011.PubMed/NCBI
|
|
41
|
Slansky JE and Farnham PJ: Introduction to
the E2F family: Protein structure and gene regulation. Curr Top
Microbiol Immunol. 208:1–30. 1996.PubMed/NCBI
|
|
42
|
Bracken AP, Ciro M, Cocito A and Helin K:
E2F target genes: Unraveling the biology. Trends Biochem Sci.
29:409–417. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Liu Y, Gong Z, Sun L and Li X: FOXM1 and
androgen receptor co-regulate CDC6 gene transcription and DNA
replication in prostate cancer cells. Biochim Biophys Acta.
1839:297–305. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Mason JM, Logan HL, Budke B, Wu M,
Pawlowski M, Weichselbaum RR, Kozikowski AP, Bishop DK and Connell
PP: The RAD51-stimulatory compound RS-1 can exploit the RAD51
overexpression that exists in cancer cells and tumors. Cancer Res.
74:3546–3555. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Reimer D, Sadr S, Wiedemair A, Concin N,
Hofstetter G, Marth C and Zeimet AG: Heterogeneous cross-talk of
E2F family members is crucially involved in growth modulatory
effects of interferon-gamma and EGF. Cancer Biol Ther. 5:771–776.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Liu C, Rangnekar VM, Adamson E and Mercola
D: Suppression of growth and transformation and induction of
apoptosis by EGR-1. Cancer Gene Ther. 5:3–28. 1998.PubMed/NCBI
|
|
47
|
Zenzmaier C, Sampson N, Plas E and Berger
P: Dickkopf-related protein 3 promotes pathogenic stromal
remodeling in benign prostatic hyperplasia and prostate cancer.
Prostate. 73:1441–1452. 2013. View Article : Google Scholar : PubMed/NCBI
|