Introduction
Gliomas are the most common primary brain tumors in
adults (1). The most malignant form,
glioblastoma (GBM), is resistant to current standard treatment, and
the current 5-year overall survival of grade IV GBM patients using
radiotherapy with concomitant TMZ treatment is only 9.8% (2). Data now suggest that GBMs are driven and
maintained by a rare population of stem-like transformed cells that
undergo self-renewal known as Glioblastoma stem-like cells (GSCs)
(3). In addition to their stem-like
capacity to proliferate, GSCs contribute to the malignancy of GBMs
by their relative resistance to radiotherapy (4) and chemotherapy (5), in contrast with their differentiated,
transformed form. Moreover, when implanted in immunodeficient mice,
GSCs form a highly invasive and phenotypically heterogeneous brain
tumor (6).
It is increasingly apparent that crosstalk between
cancer cells and cells of the neoplastic stroma is involved in the
acquired capability for invasive growth and metastasis (7). It has been generally assumed that GSCs
should also play a major role in determining GBM invasion into
normal brain tissue. Some studies have reported that normal brain
cells influence GBM invasion behavior (6,8). Rath
et al reported that under co-culture conditions, astrocytes
significantly enhance the invasion capacity of GSCs, but not of
non-GSCs (6). Therefore,
understanding the molecular profile of surrounding glioma cells
co-cultured GSCs could help us explore the underlying regulators
that control the GSCs invasion in tumor microenvironment (TME).
To identify the regulators of GSCs invasion in TME,
we carried out an integrative analysis to identify genes that are
important for GSC invasion and are specifically upregulated in
astroglia/microglia co-cultured GSCs. Among the genes identified,
serpin peptidase inhibitor clade A member 3 (SERPINA3) was
found to be abnormally overexpressed in astroglia/microglia
co-cultured GSCs. SERPINA3 is a 55–66 kDa secreted serine protease
inhibitor that proteolytically inhibits the activity of several
serine proteases including chymotrypsin and cathepsin G (9,10).
SERPINA3 is an acute phase protein and its gene expression is
stimulated by the presence of cytokines (10). SERPINA3 is involved in a wide range of
physiological activities such as inflammatory response (11), complement activation (12), regulation of lipid metabolic process
(13), apoptosis, and wound healing
(10). Recently, upregulated SERPINA3
expression has been reported in multiple cancer types (14–16), and
high SERPINA3 expression has been demonstrated to positively
correlate with poor prognosis in patients with colon (14), breast (15), lung (17,18), and
gastric cancers (19).
To further investigate the role of SERPINA3 in
glioma pathogenesis and prognosis, we used tissue microarray (TMA)
containing 80 lesions (including 3 normal brain, 3 pilocytic
astrocytomas, 11 diffuse astrocytomas, 1 oligodendrocytes
astrocytoma, 23 anaplastic astrocytomas, and 39 glioblastomas
multiforme samples) and immunohistochemistry (IHC) analysis to
evaluate the expression of SERPINA3 and its relation to
clinicopathologic factors and patient survival. Our data indicated
that upregulation of SERPINA3 was significantly associated with
glioma progression and worse patient survival. RNAi-mediated
silencing of SERPINA3 expression in cultured glioma cell lines
resulted in significant cell matrix invasion. These results suggest
that SERPINA3 may play an important role in glioma progression.
Materials and methods
Cell culture
Human GBM cell line U251MG was obtained from the
RIKEN Cell Bank (Tsukuba Science City, Japan). U251MG was cultured
in Dulbecco's modified Eagle's medium (DMEM; Life Technologies,
Rockville, Maryland, USA) supplemented with 10% (v/v) fetal bovine
serum.
GEO data sets analysis
Gene expression data (GSE63037, GSE37120, GSE52127
profiling data) were downloaded as raw signals from Gene Expression
Omnibus (http://www.ncbi.nlm.nih.gov/geo), interpreted,
normalized and log2 scaled using the online analysis tool GCBI
website (https://www.gcbi.com.cn). Exploring of
differentially expressed gene sets between GSCs and
astrocyte/microglia co-cultured GSCs profiles in GSE63037, GSE37120
and GSE52127 was also performed via the GCBI online tool.
TMA analysis
The glioma TMA containing 77 glioma clinic samples
of different grades with survival time were purchased from Shanghai
Outdo Biotech Company (Shanghai, China). Shanghai Outdo Biotech
Company is a daughter company of Shanghai Biochip Co., Ltd., which
is also the National Engineering Center for Biochip Design and
Engineering in Shanghai. The TMA consisted of 3 normal brain, 3
pilocytic astrocytomas, 11 diffuse astrocytomas, 1 oligodendrocytes
astrocytoma, 23 anaplastic astrocytomas, and 39 glioblastomas
multiforme samples. The tissue samples on the TMAs that we used in
this study were collected from Tai Zhou hospital of Zhejiang
province, China. All the patients had given informed consent and
the collection of tissue samples for research was approved by the
Ethics Committee of Tai Zhou hospital in January 26, 2010.
IHC staining of TMA
The IHC assay using SERPINA3 was performed as
described (20). Briefly, the slides
were first deparaffinized, followed by blocking with 30% normal
donkey serum for 10 min. Then the sections were incubated overnight
with the primary SERPINA3 rabbit monoclonal antibody (dilution
1:500, Rabbit monoclonal to SERPINA3, clone ab180492; Abcam Corp.,
Cambridge, UK) at 4°C. The slides were incubated with biotinylated
secondary antibody for 30 min and then the slides were incubated
for 30 min with streptavidine-peroxidase. Staining development was
performed with 3–3′-diaminobenzidine. Negative controls were
carried out by replacement of the primary antibody with
substituting phosphate buffer saline. The specimens were analyzed
under a light microscope (Nikon, Tokyo, Japan) by pathologists.
Quantification of SERPINA3 staining
intensity and statistic analysis
A previously described (21) 4-point scoring system was used to
determine intensity of SERPINA3 staining. Scoring was performed by
three independent scorers, including a pathologist, without access
to clinicopathological information of the sections. Discrepancies
among the scorers were resolved by obtaining a consensus score,
whereby the group evaluated the sections simultaneously using
scanned microscope images. Section staining was evaluated using the
12-point Remmele scale (22).
Briefly, staining intensity was scored as 0 (negative), 1 (weak), 2
(moderate) and 3 (strong). For each sample, five high-power fields
(magnificattion, ×200) were randomly selected. Cytoplasmic and
membranaceous staining intensity and percentage of positive tumor
cells were assessed. The percentage of SERPINA3 expressing cells
was scored into four categories: 1 (0–25%), 2 (26–50%), 3 (51–75%)
and 4 (76–100%). In the cases with a discrepancy between duplicated
cores, the higher score from the two tissue cores was taken as the
final score. The level of staining was evaluated by immunoreactive
score (IRS), which is calculated by multiplying the scores of
staining intensity and the score of the percentage of positive
cells. Based on the IRS, SERPINA3 staining pattern was defined as
weak (IRS: 0–4) and strong (IRS: 6–12).
Transwell invasion assay
Transwell invasion assay was performed using the
24-well cell culture inserts pre-coated with a growth
factor-reduced Matrigel layer to mimic basement membranes (8 µm
pore; BD Bioscience, San Jose, California, USA). A total of 500 µl
DMEM with 10% fetal bovine serum was added to the lower chamber.
U251MG cells were trypsinized 24 h post siRNA transfection and
plated in the upper chamber and allowed to invade for 24 h at 37°C.
The target short interfering RNA (siRNA) sequences of SERPINA3 are:
5′-AAGGACCATTGTGCGTTTCAA-3′. After the allotted time, the lower
side of the Transwell insert was carefully washed with cold PBS and
non-invading cells remaining on the top chamber were removed with a
cotton tip applicator, and then the membranes were fixed with
methanol and stained with crystal violet. The number of invading
cells was determined by averaging cell counts from nine randomly
selected fields (magnification, ×100).
Statistical analysis
Statistical analysis was performed with GraphPad
Prism 5 software (GraphPad Software, San Diego, CA). Differences in
SERPINA3 staining in the various stages of glioma were evaluated
using chi-squared (χ2) analysis. Kaplan-Meier survival
time analysis was used to estimate the survival time distributions,
and the log-rank test was used to assess the statistical
significance between different groups.
Results
Overexpression of SERPINA3 in
astrocyte/microglia co-cultured GSCs
To investigate the mechanisms through which TME
enhances GSC invasion, we carried out an integrative analysis to
identify genes that are important for GSC invasion and specifically
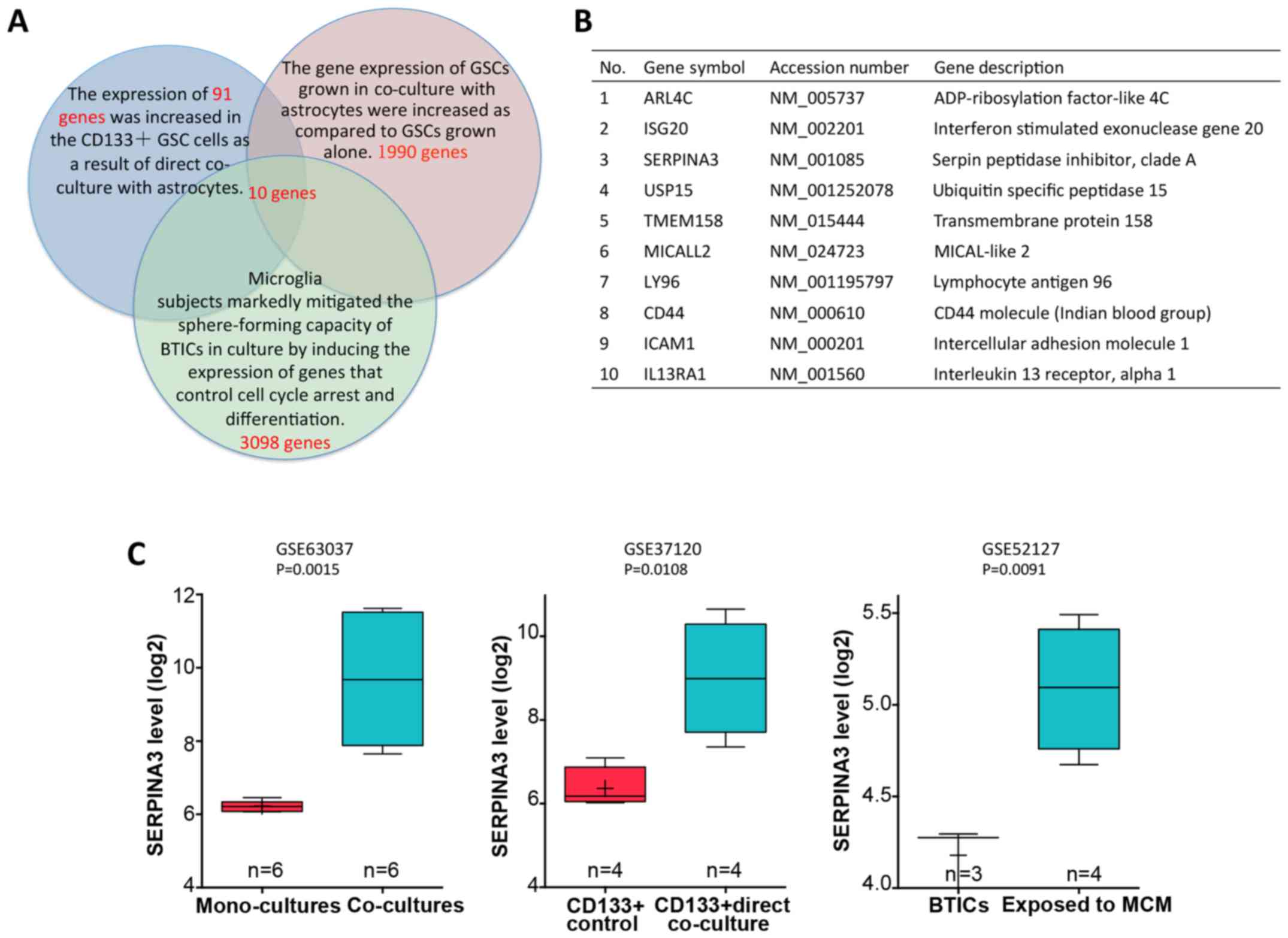
upregulated in astrocyte co-cultured GSCs. Herein, we identified a
cluster of 91 genes that were highly expressed in
CD133+GSC cells as a result of direct co-culture with
astrocytes in dataset GSE37120. To achieve this, we analyzed the
overlapping genes with the calculated 1990 genes that were
significantly upregulated in GSCs grown in co-culture with
astrocytes in GSE63037 and 3098 genes that were highly expressed in
brain tumor-initiating cells after exposure to microglia
conditioned medium (MCM) in GSE52127. We identified 10 genes as
candidate targets for GSC invasion (Fig.
1B). Among the genes identified, SERPINA3 was selected for
further study due to its reported role in regulating melanoma
migration and invasion and its capacity as a prognostic factor in
many cancers (14,17,23). To
the best of our knowledge, the role of SERPINA3 on GSC invasion has
not been well studied. Normalized expression of SERPINA3 in GSCs
vs. astrocyte co-cultured GSCs or BTICs vs. BTICs exposed to MCM
are shown in Fig. 1C.
Increase of SERPINA3 expression
correlates with glioma progression
To further investigate the role of SERPINA3 in
glioma pathogenesis and prognosis, we used TMA containing 80
melanocytic lesions (including 3 normal brain, 3 pilocytic
astrocytomas, 11 diffuse astrocytomas, 1 oligodendrocytes
astrocytoma, 23 anaplastic astrocytomas, and 39 glioblastomas
multiforme samples) and IHC to evaluate the expression of SERPINA3
and its relation to clinicopathological factors and patient
survival. The demographics and clinicopathological characteristics
of glioma patients are shown in Table
I. There were 77 glioma patients with age ranging from 21 to 75
years (median age is 53 years). According to the American Joint
Committee on Cancer (AJCC) staging system, 3 and 14 cases were
Stage I and II, respectively, while 24 and 36 cases were Stage III
and IV, respectively (Table I).
 | Table I.SERPINA3 expression in brain tumor
tissue microarray. |
Table I.
SERPINA3 expression in brain tumor
tissue microarray.
|
| SERPINA3
expression |
|
|---|
|
|
|
|
|---|
| Variable | Low no. (%) | High no. (%) | P-valuea |
|---|
| Age (years) |
|
|
|
|
<60 | 33 (77) | 10 (23) | 0.0002b |
|
≥60 | 8 (31) | 18 (69) |
|
| Sex |
|
|
|
|
Male | 22 (63) | 13 (37) | 0.5553 |
|
Female | 19 (56) | 15 (44) |
|
| Grade |
|
|
|
| Low (I
+ II) | 14 (82) | 3 (18) | 0.0412b |
| High
(III + IV) | 33 (55) | 27 (45) |
|
| Pathology |
|
|
|
|
Pilocytic astrocytoma | 3 (100) | 0 (0) | 0.0030b |
| Diffuse
astrocytoma | 9 (82) | 2 (18) |
|
|
Oligo-astrocytoma | 0 (0) | 1 (100) |
|
|
Anaplastic astrocytoma | 17 (74) | 6 (26) |
|
|
Glioblastoma | 18 (46) | 21 (54) |
|
| Normal brain
tissue | 3 (100) | 0 (0) |
|
To investigate the expression level of SERPINA3 in
the biopsies of pigmented lesions, we performed IHC staining of
normal brain, pilocytic astrocytomas, diffuse astrocytomas,
oligodendrocytes astrocytoma, anaplastic astrocytomas, and
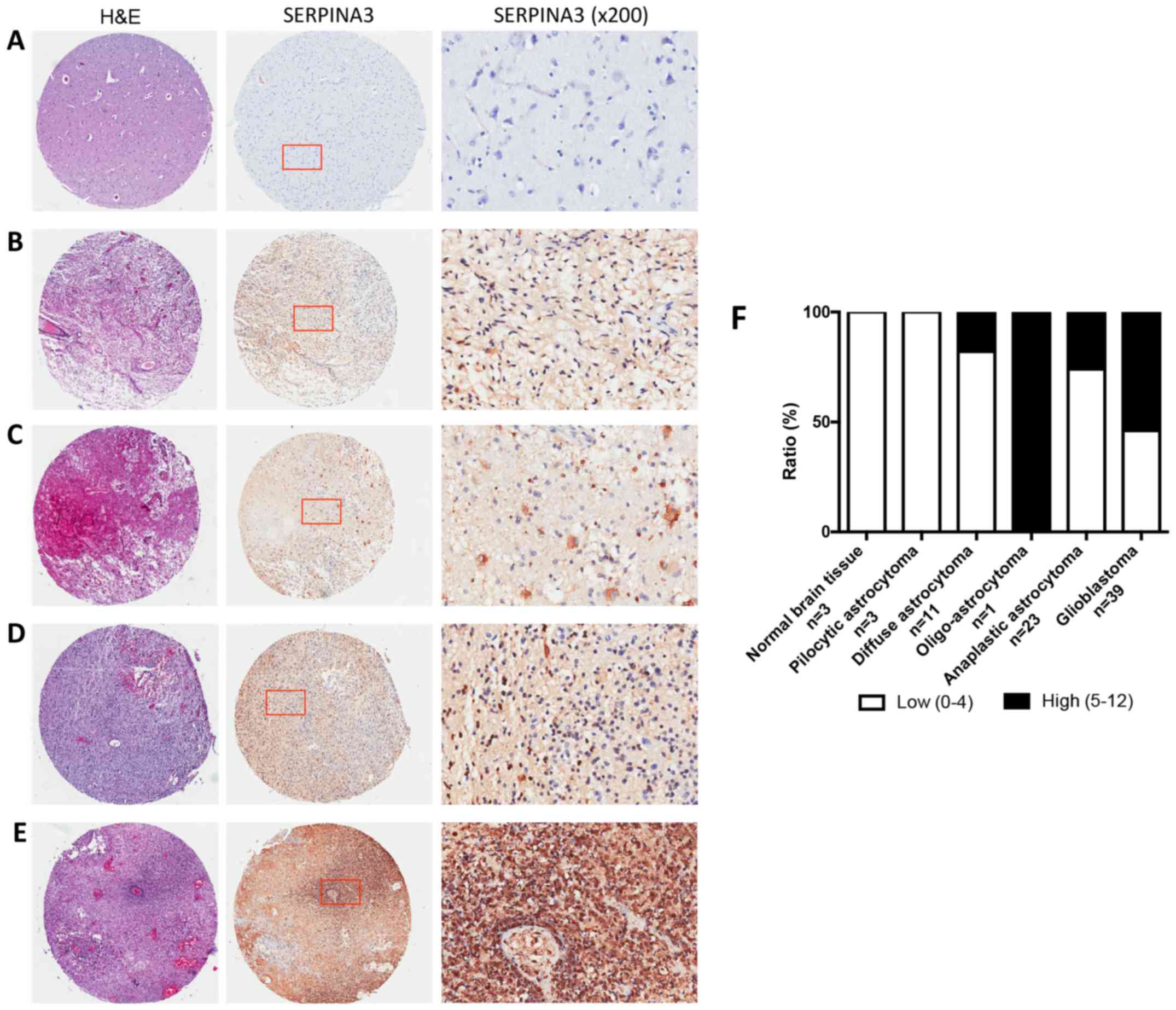
glioblastomas multiforme samples on TMA slides (Fig. 2A-E). The staining was predominantly in
the cytoplasm, therefore, only cytoplasmic staining was evaluated.
IHC results demonstrated that 0% of normal brain tissue and 0% of
the pilocytic astrocytomas vs. 26% of anaplastic astrocytomas and
54% of the glioblastomas multiforme samples showed high SERPINA3
immunostaining. Strikingly, we found marked increase of SERPINA3
expression in glioblastomas multiforme compared with normal brain
and pilocytic astrocytomas (P=0.0484, χ2 test). A
further increase was observed in glioblastomas multiforme compared
with anaplastic astrocytomas and diffuse astrocytomas (P=0.0332 and
0.0361, respectively, χ2 test) (Fig. 2F).
In the 77 glioma cases, we found that high SERPINA3
expression ratio was significantly increased from 18% in early AJCC
Stage (I + II) to 45% in Stage (III + IV) (P=0.0412, χ2
test). Interestingly, we also observed stronger staining of
SERPINA3 in older patients compared to younger subjects (P=0.0002,
χ2 test). This reason for disparity in SERPINA3
expression levels based on age is largely unknown and worth further
investigation. We did not detect significant correlations between
SERPINA3 expression and sex (Table
I).
SERPINA3 expression in glioma biopsies
is correlated with poor patient survival over 5 years
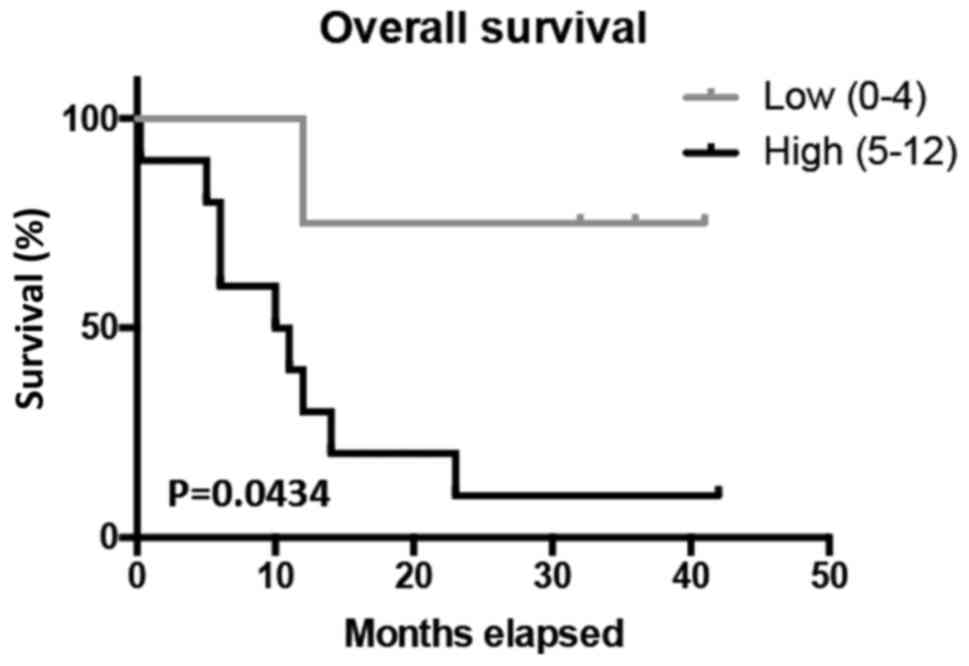
In order to determine the prognostic value of
SERPINA3, Kaplan-Meier survival tests were conducted for the 5-year
overall survival of GBM patients. The analysis revealed that
patients with high SERPINA3 expression had significantly lower
5-year overall survival (P<0.0434) (Fig. 3). Patients with higher SERPINA3
expression had a survival rate of approximately 10%, while patients
with lower SERPINA3 expression have a survival rate of
approximately 75%. Data are presented in this study using all GBM
cases; however, future studies will strive to clarify the
correlation between SERPINA3 expression and survival at all
stages.
SERPINA3 promotes glioma invasion in
extracellular matrix (ECM)
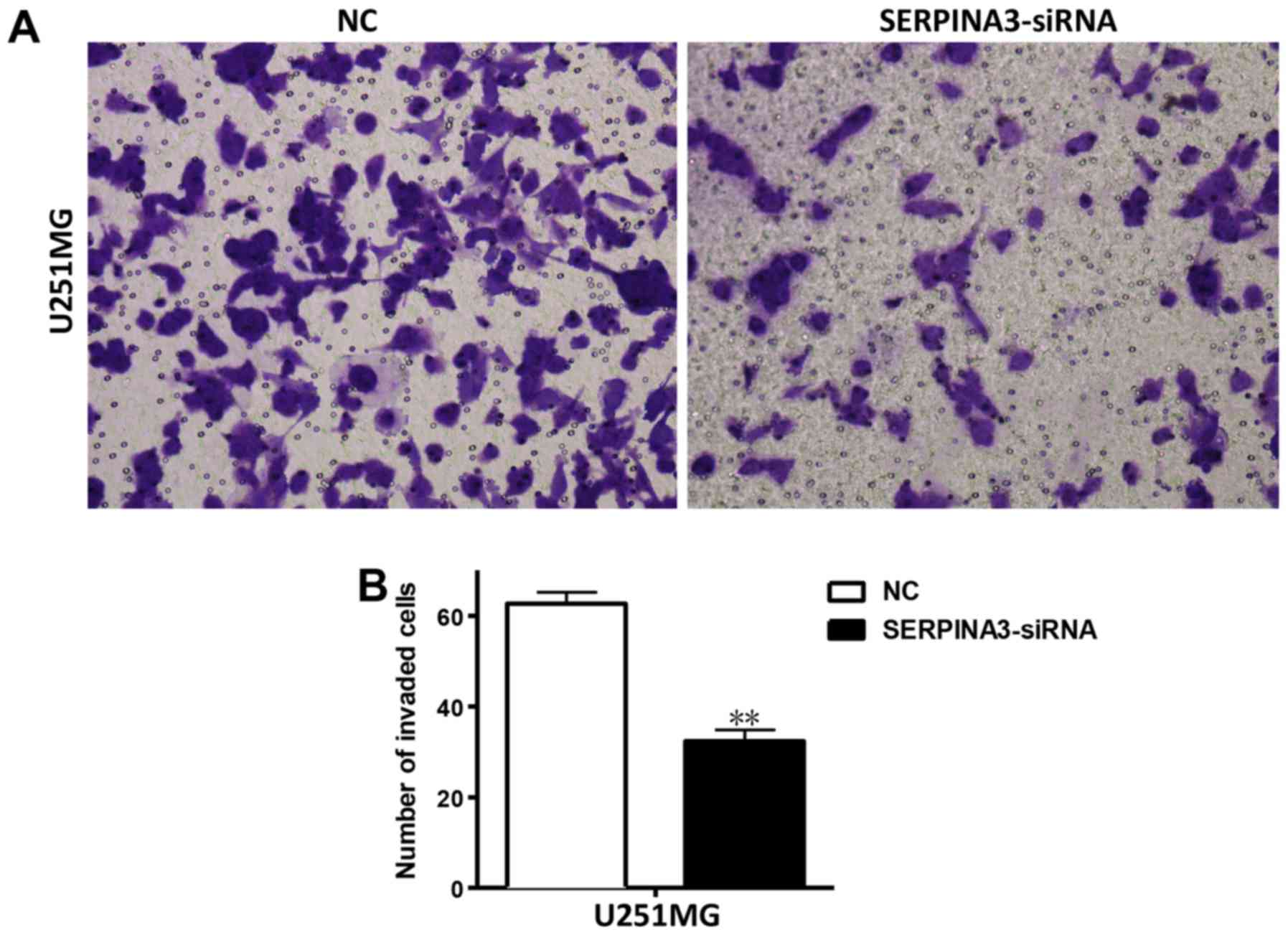
An in vitro Matrigel invasion assay was
conducted to examine the effect of SERPINA3-knockdown on cell
invasion. Matrigel is a semi-solid protein mixture that closely
mimics the ECM. As shown in Fig. 4,
the siRNA-mediated inhibition of SERPINA3 significantly reduced the
invasion of U251MG cells (Fig.
4).
Discussion
The potential contribution of GSCs to the invasive
phenotype of GBM has not been clearly defined. Invasion is a
complex process involving interactions among normal cells, tumor
cells, and the ECMs (6,24). During invasion, GBM cells interact
with a variety of surrounding glioma cells. In recent years, many
studies have been reported that surrounding glioma cells are
associated with brain cancer progression (6,25,26). Among such surrounding glioma cells,
astrocytes are the most frequent non-neuronal cell type comprising
approximately 50% of the human brain volume, and have been shown to
play a major role in the maintenance and remodeling of the brain
ECM (27). Besides astrocytes, other
surrounding glioma cells in situ are microglia, which are
innate immune cells intrinsic to the CNS. Microglia comprise a
substantial portion of the tumor mass, with some estimates being as
high as 1 in every 3 cells (26,28).
In an attempt to better define the processes and
molecules mediating GBM cell invasion within the TME, we carried
out an integrative analysis to identify genes that are important
for GSC invasion and specifically upregulated in
astroglia/microglia co-cultured GSCs. Three data sets were used
(GSE63037, GSE37120 and GSE52127) for this purpose. Gene expression
profiles were generated from GSC in mono-culture vs. 48 h after
co-culture with astrocytes for dataset GSE63037. In the data set
GSE37120, GSCs or their differentiated progeny were co-cultured for
48 h with normal human astrocytes, and the impact on
invasion-associated genes was examined. There were 6 groups
examined, including GBM CD133+ indirect co-cultured, GBM
CD133+ direct co-cultured, GBM CD133+
control, GBM CD133− indirect co-cultured, GBM
CD133− direct co-cultured, and GBM CD133−
control group. We carried out an integrative analysis between GBM
CD133+ direct co-cultured and GBM CD133+
control group, and found that 91 genes were highly expressed in
CD133+GSC cells as a result of direct co-culture with
astrocytes. In the data set GSE52127, brain tumor initiating cells
(BTICs) were subjected to microarray to determine the genes
involved in BTICs growth and differentiation when exposed to
microglia-conditioned medium (MCM) for 6 h. All the 3 data sets
used the platform of Affymetrix Human Genome U133A 2.0 Array. A
total of 10 overlapping genes were significantly upregulated within
these 3 datasets, we identified SERPINA3 as a candidate target for
enhancing the invasion potential of GSCs.
SERPINA3 is a serpin peptidase inhibitor, and has
been reported to be overexpressed in many tumor types (17–19,23),
indicating a potential role in tumor progression. Proteolytic
degradation of the ECM is considered an essential step in the
invasion and metastasis of malignant cells to distant tissues
(29,30), and proteases, such as matrix
metalloproteinases (MMPs), expressed by neoplastic and/or stromal
cells are therefore considered key players in this process. As a
consequence, protease inhibitors are intuitively expected to have
an anti-malignant role (31,32). SERPINA3 acts as an inhibitor of
several serine proteases including pancreatic chymotrypsin,
leukocyte cathepsin G, mast cell chymases, human glandular
kallikrein 2, kallikrein 3 (prostrate specific antigen), pancreatic
cationic elastase, and an uncharacterized lung serum protease
(10). The strongest association is
found with cathepsin G and is thus thought to be the major target
of SERPINA3 (33).
In this study, we examined the expression pattern of
SERPINA3 at the various stages of glioma progression, including
normal brain, pilocytic astrocytomas, diffuse astrocytomas,
oligodendrocytes astrocytoma, anaplastic astrocytomas, and
glioblastomas multiforme. SERPINA3 expression levels appeared to
strongly correlate with glioma invasion. IHC results demonstrated
that 0% of normal brain tissue and 0% of the pilocytic astrocytomas
vs. 26% of anaplastic astrocytomas and 54% of the glioblastomas
multiforme samples showed high SERPINA3 expression. Furthermore, a
significant increase of SERPINA3 expression in glioblastomas
multiforme compared with normal brain and pilocytic astrocytomas
(P=0.0484, χ2 test), and between anaplastic astrocytomas
and diffuse astrocytomas (P=0.0332 and 0.0361, respectively,
χ2 test), was observed. In the 77 glioma cases, we found
that high SERPINA3 expression ratio significantly increased from
18% in early stage GBM to 45% in late stage disease (P=0.0412,
χ2 test). Interestingly, we also observed that SERPINA3
expression was positively correlated with age (P=0.0002,
χ2 test). These results indicate that SERPINA3 could
play a critical role in glioma initiation and progression process.
Kaplan-Meier survival analysis confirmed that patients with high
SERPINA3 expression had lower survival rates, indicating possible
clinical prognostic value of SERPINA3 in GBM patients.
Serpins have multiple complex roles in tumor
biology. Although SERPINA3 has been found to be differentially
expressed in multiple tumors, its functional impact on tumor
progression remains largely unknown. To further address this
question, we performed in vitro analysis on cultured GBM
cell lines with siRNA-mediated downregulation of SERPINA3
expression. Not surprisingly, the ability of GBM cells to invade
through Matrigel was severely impaired upon downregulation of
SERPINA3 expression. Matrigel resembles the complex extracellular
environment found in many tissues and is a validated ex vivo
model of tissue matrix. It has previously been suggested that
SERPINA3 might promote GBM invasion through remodeling the
extracellular tissue matrix. Therefore, molecules targeting
SERPINA3 may have therapeutic implications for GBM in the near
future, although the precise mechanism of how SERPINA3, the serine
protease inhibitor, affects the structure of ECM remains to be
elucidated.
Acknowledgements
The authors would like to thank Dr Wang Shuhuai for
pathological evaluation of SERPINA3 staining in glioma TMA
(Department of Pathology, Cancer Hospital of Harbin Medical
University, Harbin, 150040, China). The present study was supported
by the Foundation of Health and Family Planning Commission of
Heilongjiang Province (2016–075).
References
|
1
|
Meyer MA: Malignant gliomas in adults. N
Engl J Med. 359:18502008. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Chen D, Song M, Mohamad O and Yu SP:
Inhibition of Na+/K+-ATPase induces hybrid cell death and enhanced
sensitivity to chemotherapy in human glioblastoma cells. BMC
Cancer. 14:7162014. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Binda E, Visioli A, Giani F, Trivieri N,
Palumbo O, Restelli S, Dezi F, Mazza T, Fusilli C, Legnani F, et
al: Wnt5a drives an invasive phenotype in human glioblastoma
stem-like cells. Cancer Res. 77:996–1007. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Bao S, Wu Q, McLendon RE, Hao Y, Shi Q,
Hjelmeland AB, Dewhirst MW, Bigner DD and Rich JN: Glioma stem
cells promote radioresistance by preferential activation of the DNA
damage response. Nature. 444:756–760. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Eramo A, Ricci-Vitiani L, Zeuner A,
Pallini R, Lotti F, Sette G, Pilozzi E, Larocca LM, Peschle C and
De Maria R: Chemotherapy resistance of glioblastoma stem cells.
Cell Death Differ. 13:1238–1241. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Rath BH, Fair JM, Jamal M, Camphausen K
and Tofilon PJ: Astrocytes enhance the invasion potential of
glioblastoma stem-like cells. PLoS One. 8:e547522013. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Hanahan D and Weinberg RA: Hallmarks of
cancer: The next generation. Cell. 144:646–674. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Rath BH, Wahba A, Camphausen K and Tofilon
PJ: Coculture with astrocytes reduces the radiosensitivity of
glioblastoma stem-like cells and identifies additional targets for
radiosensitization. Cancer Med. 4:1705–1716. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Kalsheker NA: Alpha 1-antichymotrypsin.
Int J Biochem Cell Biol. 28:961–964. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Baker C, Belbin O, Kalsheker N and Morgan
K: SERPINA3 (aka alpha-1-antichymotrypsin). Front Biosci.
12:2821–2835. 2007. View
Article : Google Scholar : PubMed/NCBI
|
|
11
|
Sun YX, Wright HT and Janciauskiene S:
Glioma cell activation by Alzheimer's peptide Abeta1-42,
alpha1-antichymotrypsin, and their mixture. Cell Mol Life Sci.
59:1734–1743. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Bodmer JL and Schnebli HP: Plasma
proteinase inhibitors. Schweiz Med Wochenschr. 114:1359–1363.
1984.PubMed/NCBI
|
|
13
|
Sun YX, Wright HT and Janciauskiene S:
Alpha1-antichymotrypsin/Alzheimer's peptide Abeta(1–42) complex
perturbs lipid metabolism and activates transcription factors
PPARgamma and NFkappaB in human neuroblastoma (Kelly) cells. J
Neurosci Res. 67:511–522. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Karashima S, Kataoka H, Itoh H, Maruyama R
and Koono M: Prognostic significance of alpha-1-antitrypsin in
early stage of colorectal carcinomas. Int J Cancer. 45:244–250.
1990. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Hurlimann J and van Melle G: Prognostic
value of serum proteins synthesized by breast carcinoma cells. Am J
Clin Pathol. 95:835–843. 1991. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Lein M, Stephan C, Jung K, Schnorr D and
Loening SA: Molecular forms of prostate-specific antigen and human
kallikrein 2 as possible indicators in prostatic carcinoma
diagnosis. Urologe A. 39:313–323. 2000.(In German). View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Higashiyama M, Doi O, Yokouchi H, Kodama
K, Nakamori S and Tateishi R: Alpha-1-antichymotrypsin expression
in lung adenocarcinoma and its possible association with tumor
progression. Cancer. 76:1368–1376. 1995. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Zelvyte I, Wallmark A, Piitulainen E,
Westin U and Janciauskiene S: Increased plasma levels of serine
proteinase inhibitors in lung cancer patients. Anticancer Res.
24:241–247. 2004.PubMed/NCBI
|
|
19
|
Tahara E, Ito H, Taniyama K, Yokozaki H
and Hata J: Alpha 1-antitrypsin, alpha 1-antichymotrypsin, and
alpha 2-macroglobulin in human gastric carcinomas: A retrospective
immunohistochemical study. Hum Pathol. 15:957–964. 1984. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Zhou SM, Cheng L, Guo SJ, Wang Y,
Czajkowsky DM, Gao H, Hu XF and Tao SC: Lectin RCA-I specifically
binds to metastasis-associated cell surface glycans in
triple-negative breast cancer. Breast Cancer Res. 17:362015.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Sjoestroem C, Khosravi S, Cheng Y, Safaee
Ardekani G, Martinka M and Li G: DLC1 expression is reduced in
human cutaneous melanoma and correlates with patient survival. Mod
Pathol. 27:1203–1211. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Remmele W and Stegner HE: Recommendation
for uniform definition of an immunoreactive score (IRS) for
immunohistochemical estrogen receptor detection (ER-ICA) in breast
cancer tissue. Pathologe. 8:138–140. 1987.(In German). PubMed/NCBI
|
|
23
|
Abd Hamid UM, Royle L, Saldova R,
Radcliffe CM, Harvey DJ, Storr SJ, Pardo M, Antrobus R, Chapman CJ,
Zitzmann N, et al: A strategy to reveal potential glycan markers
from serum glycoproteins associated with breast cancer progression.
Glycobiology. 18:1105–1118. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Friedl P and Alexander S: Cancer invasion
and the microenvironment: Plasticity and reciprocity. Cell.
147:992–1009. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
The tumor microenvironment mediates GBM
resistance to CSF1R blockade. Cancer Discov. 6:6902016. View Article : Google Scholar
|
|
26
|
Sarkar S, Döring A, Zemp FJ, Silva C, Lun
X, Wang X, Kelly J, Hader W, Hamilton M, Mercier P, et al:
Therapeutic activation of macrophages and microglia to suppress
brain tumor-initiating cells. Nat Neurosci. 17:46–55. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Sofroniew MV and Vinters HV: Astrocytes:
Biology and pathology. Acta Neuropathol. 119:7–35. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Charles NA, Holland EC, Gilbertson R,
Glass R and Kettenmann H: The brain tumor microenvironment. Glia.
60:502–514. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Danø K, Andreasen PA, Grøndahl-Hansen J,
Kristensen P, Nielsen LS and Skriver L: Plasminogen activators,
tissue degradation, and cancer. Adv Cancer Res. 44:139–266. 1985.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Mignatti P and Rifkin DB: Biology and
biochemistry of proteinases in tumor invasion. Physiol Rev.
73:161–195. 1993.PubMed/NCBI
|
|
31
|
Dodwell DJ: Proteinase inhibitors in
malignancy: Therapeutic promise or another white elephant? J R Soc
Med. 86:573–576. 1993.PubMed/NCBI
|
|
32
|
Liotta LA and Kohn EC: The
microenvironment of the tumour-host interface. Nature. 411:375–379.
2001. View
Article : Google Scholar : PubMed/NCBI
|
|
33
|
Beatty K, Bieth J and Travis J: Kinetics
of association of serine proteinases with native and oxidized
alpha-1-proteinase inhibitor and alpha-1-antichymotrypsin. J Biol
Chem. 255:3931–3934. 1980.PubMed/NCBI
|