Introduction
Pancreatic ductal adenocarcinoma (PDAC) is a serious
disease with the lowest five-year survival rate among all solid
cancers (<5%) (1). At present,
PDAC is the fourth leading cause of cancer-associated mortality in
the USA and its incidence has increased in past decades (2). Poor survival is probably due to early
local invasion and metastasis, which means that potentially
curative surgery is no longer an option for 85% of patients at the
point of diagnosis (3). Furthermore,
patients who have undergone surgery may experience local recurrence
or metastasis within 1 year (4).
Therefore, an improved understanding of the biological processes
underlying PDAC, in particular the aggressive nature of its
invasion, is warranted.
Receptor for activated C kinase 1 (RACK1), also
known as GNB2L1, is a 36-kilodalton cytosolic protein (5). It was first reported as an anchoring
protein with seven WD40 (Trp-Asp) repeats (6). RACK1 has been revealed to interact with
multiple signaling molecules, including protein kinase C (PKC),
period circadian clock 1 and Src (7,8). It is
regarded as a platform for various signal transduction pathways.
RACK1 is involved in cell division, invasion and migration in
cancer (5,9,10).
However, the function of RACK1 in pancreatic ductal adenocarcinoma
has not yet been investigated.
The aim of the present study was to assess the
expression of RACK1 in PDAC tumor tissues and adjacent noncancerous
tissues, and to compare RACK1 expression with clinicopathological
characteristics. The value of RACK1 as a prognostic biomarker for
patients with PDAC was then evaluated.
Materials and methods
Patients and tissue samples
Archived and formalin-fixed, paraffin-embedded tumor
tissues were obtained from 157 patients with PDAC, who were
underwent surgery between September 2003 and March 2011 in the
Department of Hepatobiliary Surgery, The Second Affiliated Hospital
of Xi'an Jiaotong University (Xi'an, China). Informed consent was
given by all patients prior to surgery. The group consisted of 76
men and 81 women with an average age of 56 years (range, 29–81
years). The clinicopathological characteristics are presented in
Table I. The diagnosis of all
patients was confirmed by pathologists. The histological
classification and tumor stages were evaluated according to the
tumor, node and metastasis classification of malignant tumors
defined by the American Joint Committee on Cancer (11). All the specimens were directly
snap-frozen in liquid nitrogen, and stored at −130°C for the
extraction of RNA and total protein. The other parts of the
specimens were fixed in buffered formalin for 48 h, embedded in
paraffin, and cut into 4 µm sections for immunohistochemical
detection.
 | Table I.Clinical characteristics of respective
discovery and verification sets populations. |
Table I.
Clinical characteristics of respective
discovery and verification sets populations.
| Characteristic | Value |
|---|
| No. of patients | 157 |
| Sex, male/female | 76/81 |
| Age, years; median,
range | 56 (35–76) |
| CA19-9 level, KU/l;
median, range | 131.5
(62.1–154.9) |
| Tumor size, mm;
median, range | 30 (29.2–35) |
| Pathologic
differentiation, well/moderate/poor | 10/31/9 |
| Clinical stage,
I/II/III/IV | 7/22/16/5 |
Immumohistochemical staining
Paraffin-embedded samples were cut into 4 µm
sections and stained with hematoxylin and eosin for tumor
confirmation. RACK1 protein expression was visualized using a
Streptavidin-Biotin Complex immumohistochemical assay kit (cat. no.
SA1027; Wuhan Boster Technology, Ltd., Wuhan, China) according to
the manufacturer's protocol. Briefly, the endogenous peroxidase
activity of sections was blocked with H2O2
methanol at room temperature for 10 min and then incubated in 5%
goat antiserum (Wuhan Boster Biological Technology, Ltd., Wuhan,
China) for 15 min at 37°C. The sections were then sequentially
incubated with a mouse anti-human RACK1 monoclonal antibody (cat.
no. sc-17754; dilution, 1:500; Santa Cruz Biotechnology, Inc.,
Dallas, TX, USA) overnight at 4°C. The sections were then incubated
with biotinylated rabbit anti-mouse immunoglobulin G (cat. no.
sc-358914; dilution, 1:2,000; Santa Cruz Biotechnology, Inc.) at
37°C for 15 min. A streptavidin-peroxidase complex was added and
then 3′,3′-diaminobenzidine-H2O2 was used for
the color reaction. Images of four representative fields were
captured. The RACK1 positive rate (brown-yellow colored cells) was
automatically measured using the Biological Image Analysis System
2000 (Kontron, Eching, Germany).
Immumohistochemical staining was evaluated
independently by two pathologists. The level of RACK1 staining was
based on the intensity of staining and the proportion of positively
stained cancer cells. The following staining scores were applied:
Intensity [0 (no staining), 1 (light yellow), 2 (yellow brown), 3
(strong brown color)]; the proportion of positive tumor cells [0
(≤5% positive tumor cells), 1 (6–25% positive tumor cells), 2
(26–50% positive tumor cells), 3 (51–75% positive tumor cells) and
4 (≥76% positive tumor cells)]. The final basis [immunoreactivity
score (IS)] for grouping was the product of the staining area score
and staining intensity as follows: 0, negative; 1–4, weak positive;
5–8, positive; 9–12, strong positive.
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
Total RNA from 8 randomly selected primary tumor and
adjacent non-tumor tissue samples were extracted using TRIzol
reagent (Invitrogen; Thermo Fisher Scientific, Inc.) according to
the manufacturer's protocol. RNA (2 µg) from each sample was used
for complementary DNA synthesis using the Primescript RT reagent
kit (Takara Biotechnology Co., Ltd., Dailan, China). To amplify the
spliced form of human RACK1, the following primer sequences were
used: forward, 5′-GATTCTGGAAATATTGACTCTT-3′ and reverse,
5′-AACTGGGCCCTTCTGGGTAGAC-3′. GAPDH was used as an internal control
and the primer sequences were as follows: forward,
5′-GCACCGTCAAGGCTGAGAAC-3′ and reverse, 5′-TGGTGAAGACGCCAGTGGA-3′.
PCR was performed on the ABI prism 7500 sequence detection system
(Applied Biosystems; Thermo Fisher Scientific, Inc., USA) and using
SYBR Green PCR Master Mix (Applied Biosystems, Thermo Fisher
Scientific, Inc., USA), the thermocycler conditions were as
follows: 50°C for 2 min and 95°C for 10 min, followed by 40 cycles
at 95°C for 15 sec and 60°C for 1 min. The relative expression
levels of RACK1 was normalized to that of GAPDH using the
comparative 2−ΔΔCq method (12).
Western blotting
The 8 tumor samples which was selected for RT-qPCR
were homogenized in ice-cold radio immunoprecipitation assay lysis
buffer at 4°C for 15 min (cat. no. 9806; Cell Signaling Technology,
Inc., Danvers, MA, USA) and centrifuged at 13,400 × g at 4°C for 20
min. Then protein concentrations were determined using a Bio-Rad
BCA assay kit (Bio-Rad Laboratories, Inc., Hercules, CA, USA).
Next, equal amounts of 5X Laemmli buffer was added and the protein
was boiled. 50 µg proteins were resolved in 10% SDS-PAGE and
transferred to polyvinylidene fluoride membranes. The membranes
were blocked for 1 h in 5% bovine milk diluted in TBST at room
temperature and incubated with the following antibodies: RACK1
(cat. no. ab72483; 1:500; Abcam, Cambridge, UK) and GAPDH (cat. no.
SAB1405848; 1:1,000; Sigma-Aldrich; Merck KGaA, Darmstadt, Germany)
at room temperature for 2 h. Following washing with TBST (0.05%
Tween20) three times, the membranes were subsequently incubated
with corresponding horseradish peroxidase-conjugated rabbit
anti-mouse immunoglobulin G (IgG) (cat. no. ab6728; dilution,
1:2,000; Abcam, Cambridge, UK) or goat anti-rabbit IgG (cat. no.
ab6721; dilution, 1:2,000; Abcam, Cambridge, UK) for 1 h at room
temperature, and then washed 3 times with TBST. The final band was
visualized using an enhanced chemiluminescence assay (Thermo Fisher
Scientific, Inc.) and detected using AlphaImager 2200
(ProteinSimple; Bio-Techne, Minneapolis, MN, USA). Statistical
analysis was conducted from 3 independent experiments.
Statistical analysis
Statistical analysis was performed using SPSS 18.0
software (SPSS, Inc., Chicago, IL, USA). Results were expressed as
the mean ± standard deviation. Pearson's χ2 test was
used to analyze the associations between RACK1 expression and
clinicopathological features in patients with PDAC. Univariate and
multivariate Cox regression analyses were performed to analyze the
survival data. The Kaplan-Meier method and log-rank test were used
to evaluate the survival of patients with PDAC.
Results
RACK1 protein is overexpressed in
patients with PDAC
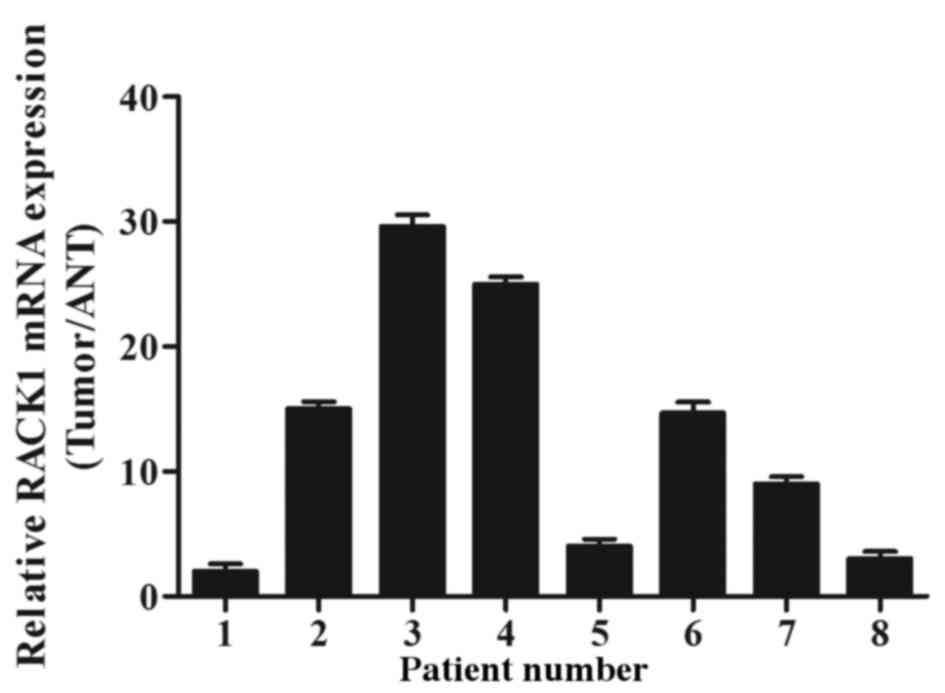
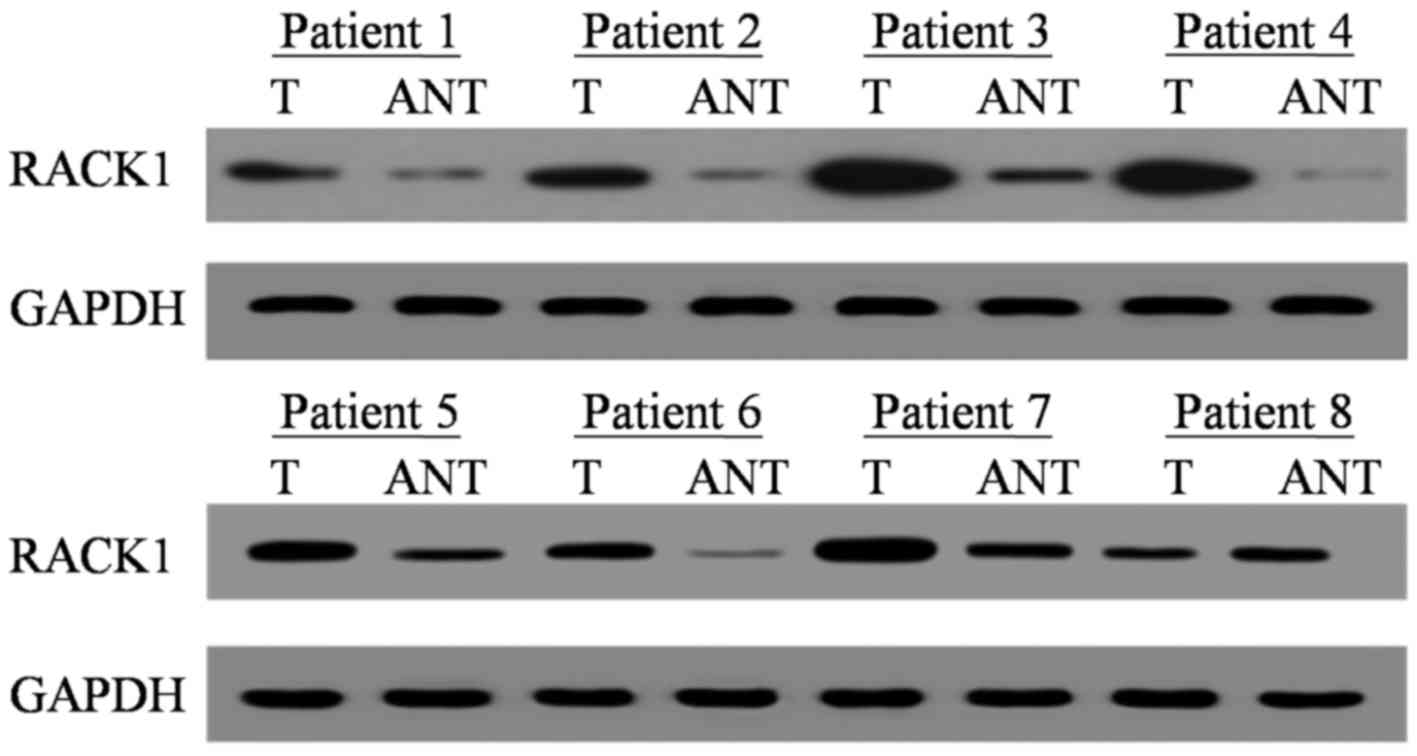
RACK1 mRNA and protein expression levels were
assessed by RT-qPCR and western blotting. Of the 8 randomly
selected cases, RACK1 mRNA and protein were overexpressed in 7 out
of 8 PDAC tumor specimens compared with in adjacent noncancerous
tissues (ANT; Figs. 1 and 2). The medians of RACK1 mRNA and protein
expression increased by 2.12 fold and 3.43 fold, respectively
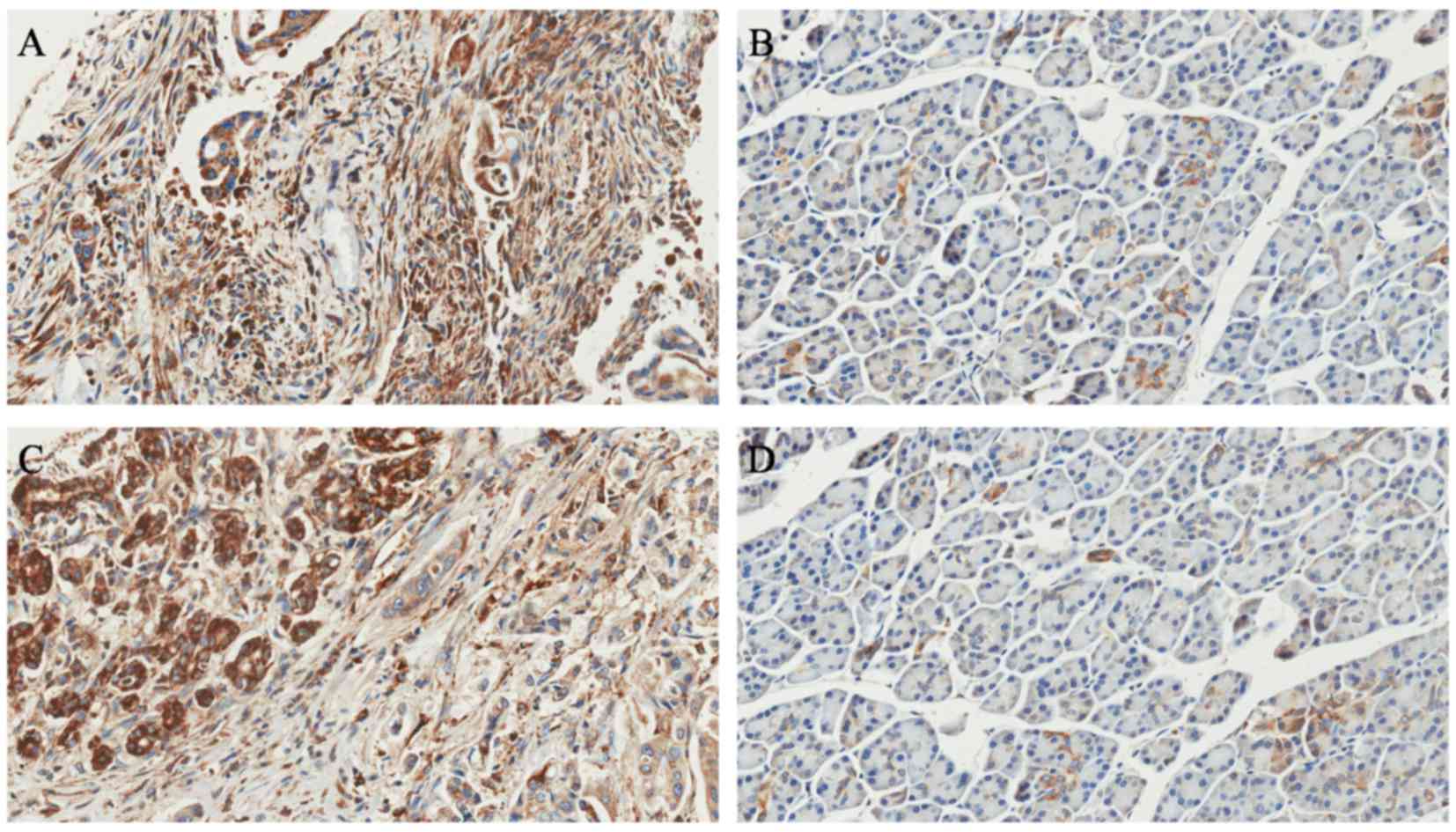
(relative ratio of tumor/ANT). In addition, immunohistochemistry
confirmed that RACK1 protein was overexpressed in the 121 PDAC
lesions compared with their matched adjacent noncancerous tissues
(Fig. 3). Furthermore, the IHC data
revealed that RACK1 was primarily localized in the cytoplasm of the
PDAC cancer cells. No positive staining was identified in the
stroma tissues of the tumor (Fig.
3).
RACK1 expression is associated with
clinical data in patients with PDAC
The immumohistochemical staining data suggested an
increase in the proportion of RACK1 in PDAC tumor tissues compared
with adjacent noncancerous tissues. A total of 121/157 (59.2%) of
the PDAC cases exhibited high expression levels of RACK1 (SI ≥6),
whereas 36/157 (40.8%) had low levels of RACK1 (IS <6). The
relationships between RACK1 expression and clinicopathological
features are summarized in Table II.
The results indicated that RACK1 expression was significantly
associated with histological differentiation (P<0.001), lymph
node invasion (P<0.001) and clinical stage (P=0.011). However,
no associations were observed between RACK1 expression and age,
sex, tumor location or tumor size.
 | Table II.Relationship between RACK1 protein
expression levels in pancreatic ductal adenocarcinoma and clinical
pathological features (n=157). |
Table II.
Relationship between RACK1 protein
expression levels in pancreatic ductal adenocarcinoma and clinical
pathological features (n=157).
|
|
| RACK1 expression |
|
|---|
|
|
|
|
|
|---|
| Clinical
characteristic | N (%) | High (n=121) | Low (n=36) | P-valuea |
|---|
| Age (years) |
|
|
|
|
| ≤60 | 45 (28.7) | 33 (73.3) | 12 (26.7) | 0.213 |
|
>60 | 112 (71.3) | 88 (78.6) | 24 (21.4) |
|
| Sex |
|
|
|
|
|
Female | 81 (51.6) | 64 (79.0) | 17 (21.0) | 0.106 |
| Male | 76 (48.4) | 57 (75.0) | 11 (25.0) |
|
| Tumor location |
|
|
|
|
| Head | 123 (78.3) | 100 (81.3) | 23 (18.7) | 0.165 |
|
Body/tail | 34 (21.7) | 21 (61.8) | 13 (38.2) |
|
| Histological
differentiation |
|
|
|
|
| Well | 22 (14) | 12 (54.5) | 10 (45.5) | <0.001 |
|
Moderate/poor | 135 (86) | 109 (80.7) | 26 (19.3) |
|
| Tumor size |
|
|
|
|
| ≤2
cm | 62 (39.5) | 43 (69.4) | 19 (30.6) | 0.123 |
| >2
cm | 95 (60.5) | 79 (83.2) | 16 (16.8) |
|
| Lymph node
invasion |
|
|
|
|
|
Absent | 32 (20.4) | 20 (62.5) | 12 (37.5) | <0.001 |
|
Present | 125 (79.6) | 101 (80.8) | 24 (19.2) |
|
| Clinical stage |
|
|
|
|
|
I+II | 87 (55.4) | 66 (75.9) | 21 (24.1) | 0.011 |
|
III+IV | 70 (44.6) | 55 (78.6) | 15 (21.4) |
|
Association between RACK1 expression
and prognosis in patients with PDAC
To determine whether RACK1 expression is a
prognostic factor for patients with PDAC, Kaplan-Meier analysis and
log-rank tests were performed using RACK1 protein expression and
clinical follow-up data in all 157 patients. A total of 147
patients died during the follow-up period, and 10 patients were
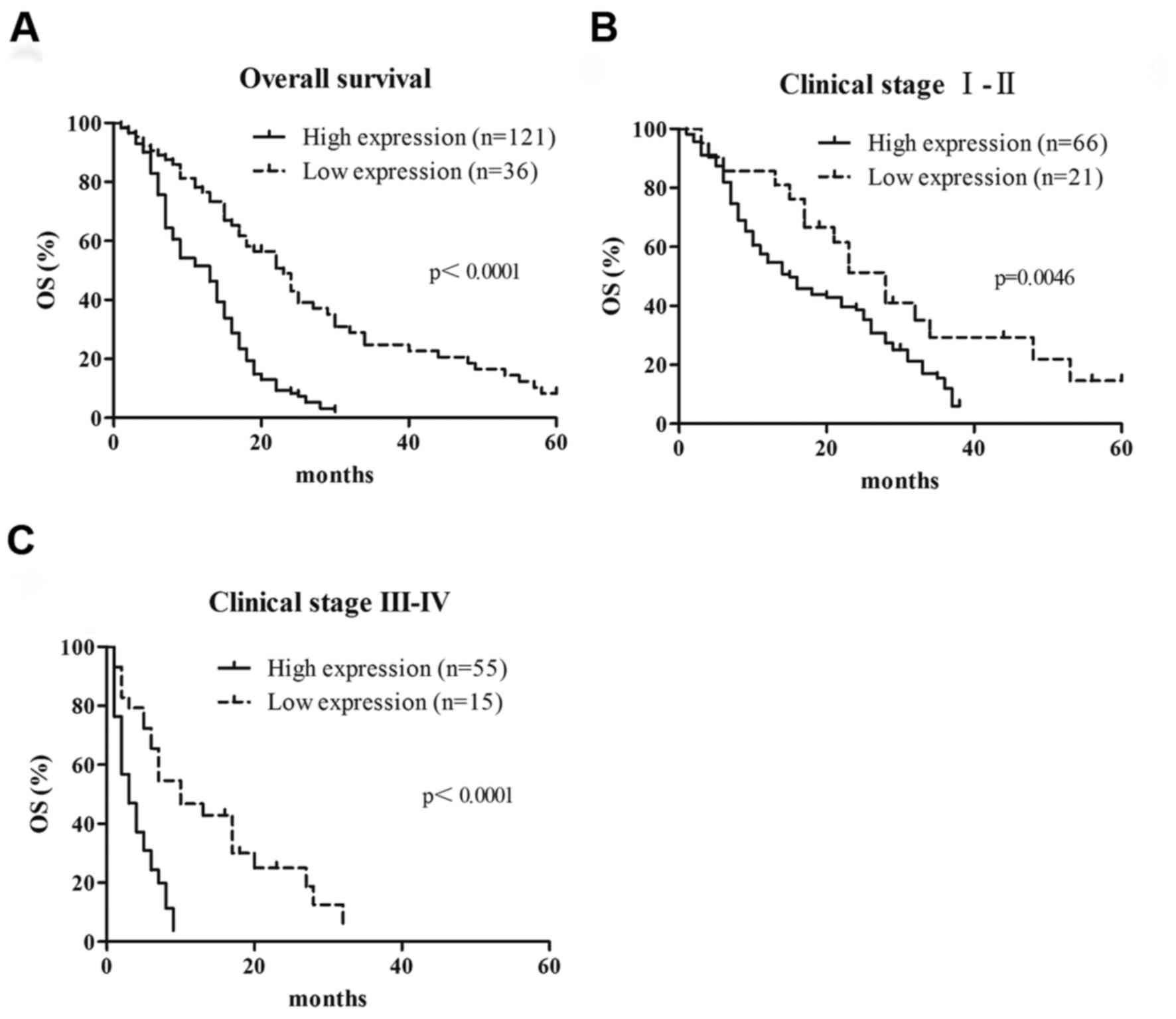
still alive. The results indicated that the median survival time in
patients with high RACK1 expression (14.3±1.6 months; n=121) was
significantly shorter compared with in patients with a low RACK1
expression (26.9±3.2 months; n=36; P<0.0001; Fig. 4A). These results revealed that high
level of RACK1 expression indicates poor prognosis for patients
with PDAC. Overall survival time was also examined in subgroups of
patients. Patients with PDAC in the high RACK1 expression group
with early (I and II) and advanced stage (III and IV) PDAC had
significantly shorter overall survival times than those in the low
RACK1 expression group (P=0.0046 and P<0.001, respectively;
Fig. 4B and C, respectively). In
addition, univariate and multivariate analyses revealed that RACK1
expression, histological differentiation, lymph node invasion and
tumor resectability were all independent prognostic factors in
patients with PDAC (Table III).
 | Table III.Univariate and multivariate analysis
of prognostic parameters for survival in patients with pancreatic
ductal adenocarcinoma. |
Table III.
Univariate and multivariate analysis
of prognostic parameters for survival in patients with pancreatic
ductal adenocarcinoma.
|
| Univariate
analysis | Multivariate
analysis |
|---|
|
|
|
|
|---|
| Parameter | RR | 95% CI | P-value | RR | 95% CI | P-value |
|---|
| Expression of
RACK1, high vs. low | 1.901 | 1.141–3.012 | 0.006 | 2.712 | 1.566–4.691 | 0.002 |
| Age, ≤60 vs.
>60 | 1.211 | 0.801–1.901 | 0.812 | 1.354 | 0.866–2.102 | 0.223 |
| Sex, male vs.
female | 0.999 | 0.678–1.499 | 0.924 | 1.112 | 0.647–1.623 | 0.857 |
| Tumor location,
head vs. body/tail | 0.821 | 0.465–1.436 | 0.554 | 0.643 | 0.379–1.242 | 0.124 |
| Histological
differentiation, well vs. moderate/poor | 1.613 | 0.876–3.512 | 0.001 | 1.721 | 0.976–2.993 | 0.005 |
| Size, ≤2 cm vs.
>2 cm | 1.966 | 1.132–3.499 | 0.011 | 1.666 | 0.899–2.902 | 0.219 |
| Lymph node
invasion, absent vs. present | 1.112 | 0.743–1.699 | 0.222 | 0.623 | 0.399–0.976 | 0.023 |
| Liver metastasis,
absent vs. present | 1.342 | 0.787–2.243 | 0.325 | 1.032 | 0.387–3.532 | 0.933 |
| Clinical stage, I
vs. II vs. III vs. IV | 1.376 | 1.154–1.599 | 0.021 | 1.498 | 0.731–2.675 | 0.221 |
| Treatment, radical
vs. palliative | 2.499 | 1.632–3.812 | 0.001 | 2.932 | 1.790–4.659 | 0.001 |
Discussion
Pancreatic ductal adenocarcinoma is known for its
poor prognosis (13). The aggressive
nature of its invasion, high incidence of early recurrence and poor
response to radiotherapy and chemotherapy all contribute to the
poor outcomes observed in patients with PDAC (14). To the best of our knowledge, the
association between RACK1 expression and prognosis in PDAC was not
reported prior to the present study, where RACK1 expression was
examined in PDAC in detail.
In the present study, RACK1 expression was revealed
to be dramatically higher in PDAC tumor tissues than in adjacent
noncancerous tissues. RACK1 expression was associated with T
classification, N classification, clinical stage and liver
metastasis, indicating that RACK1 may be a potential biomarker for
patients with metastasis. Furthermore, the results of the present
study revealed that patients with high RACK1 expression had shorter
overall survival times than those with low RACK1 expression.
Multivariate analysis revealed that RACK1 expression level was an
independent prognostic factor for patients with PDAC. These results
suggested that RACK1 was involved in PDAC progression and may be a
novel prognostic biomarker for PDAC.
Several novel prognostic factors including C-C motif
chemokine ligand 18 (15), golgi
phosphoprotein 3 (16), and UL16
binding protein 2 (17) have
previously been reported. In addition, tumor grade, lymph node
invasion and clinical stage have also been demonstrated to be
correlated with outcome in patients with PDAC (18–20).
However, the efficacy of such prognostic markers in clinical
application remains unknown. Therefore, further investigation is
warranted to discover novel prognostic markers with improved
prognostic efficacy in patients with PDAC.
RACK1, which is upregulated in several solid
malignancies, was originally reported in colorectal cancer
(21). RACK1 serves as a scaffold and
anchoring protein for PKC, and stabilizes the activated
conformation of PKC. RACK1 has been associated with several
signaling pathways, and serves a vital function in tumorigenesis
(22). RACK1 was reported to be
overexpressed in multiple cancers including melanoma (23), gastric cancer (24), hepatocellular carcinoma (25) and pulmonary adenocarcinoma (26). RACK1 is thought to promote tumor
invasion and metastasis. However, RACK1 also suppresses tumor
growth in gastric cancer, indicating that RACK1 may serve different
functions in different tumors.
Mamidipudi and Cartwright (27) reported that RACK1 inhibited colorectal
cancer cell proliferation via the regulation of Src activity.
However, Saito et al (28)
reported that RACK1 was overexpressed in colon cancer, and promoted
colon cancer cell proliferation. Thus, the function of RACK1 varies
between tumor types, and remains to be elucidated. The present
study measured RACK1 mRNA and protein expression levels in tumor
tissues and adjacent noncancerous tissues from 157 patients with
PDAC. The results revealed that RACK1 was overexpressed in tumor
tissues compared with adjacent noncancerous tissues. RACK1 was
associated with clinical stage, differentiation and lymph node
metastasis in patients with PDAC.
In conclusion, RACK1 was overexpressed in patients
with PDAC, and positively associated with the degree of malignancy
in patients with PDAC. Therefore, RACK1 may serve as a novel
biomarker and potential therapeutic target. Further investigations,
in particular those concerning the molecular mechanisms underlying
the effect of RACK1 in PDAC, are being performed.
References
|
1
|
Buekki J: Pancreatic adenocarcinoma. N
Engl J Med. 371:2139–2140. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Iacobuzio-Donahue CA and Herman JM:
Autophagy, p53, and pancreatic cancer. N Engl J Med. 370:1352–1353.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Viale A, Pettazzoni P, Lyssiotis CA, Ying
H, Sanchez N, Marchesini M, Carugo A, Green T, Seth S, Giuliani V,
et al: Oncogene ablation-resistant pancreatic cancer cells depend
on mitochondrial function. Nature. 514:628–632. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Meacham CE and Morrison SJ: Tumour
heterogeneity and cancer cell plasticity. Nature. 501:328–337.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Jin S, Mu Y, Wang X, Liu Z, Wan L, Xiong
Y, Zhang Y, Zhou L and Li L: Overexpressed RACK1 is positively
correlated with malignant degree of human colorectal carcinoma. Mol
Biol Rep. 41:3393–3399. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Hu L, Lu F, Wang Y, Liu Y, Liu D, Jiang Z,
Wan C, Zhu B, Gan L, Wang Y, et al: RACK1, a novel
hPER1-interacting protein. J Mol Neurosci. 29:55–63. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Cho IR, Kaowinn S, Moon J, Soh J, Kang HY,
Jung CR, Oh S, Song H, Koh SS and Chung YH: Oncotropic H-1
parvovirus infection degrades HIF-1alpha protein in human
pancreatic cancer cells independently of VHL and RACK1. Int J
Oncol. 46:2076–2082. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Sato T, Takahashi H, Hatakeyama S, Iguchi
A and Ariga T: The TRIM-FLMN protein TRIM45 directly interacts with
RACK1 and negatively regulates PKC-mediated signaling pathway.
Oncogene. 34:1280–1291. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Cheng D, Qian W, Wang Y, Meng M, Wei L, Li
Z, Kang L, Peng J and Xia Q: Nuclear import of transcription factor
BR-C is mediated by its interaction with RACK1. PloS One.
9:e1091112014. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Omosigho NN, Swaminathan K, Plomann M,
Mueller-Taubenberger A, Noegel AA and Riyahi TY: The Dictyostelium
discoideum RACK1 orthologue has roles in growth and development.
Cell Commun Signal. 12:372014. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Edge SB and Compton CC: The American Joint
Committee on Cancer: the 7th edition of the AJCC cancer staging
manual and the future of TNM. Ann Surg Oncol. 17:1471–1474. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Waddell N, Pajic M, Patch AM, Chang DK,
Kassahn KS, Bailey P, Johns AL, Miller D, Nones K, Quek K, et al:
Whole genomes redefine the mutational landscape of pancreatic
cancer. Nature. 518:495–501. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Junttila MR and de Sauvage FJ: Influence
of tumour micro-environment heterogeneity on therapeutic response.
Nature. 501:346–354. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Meng F, Li W, Li C, Gao Z, Guo K and Song
S: CCL18 promotes epithelial-mesenchymal transition, invasion and
migration of pancreatic cancer cells in pancreatic ductal
adenocarcinoma. Int J Oncol. 46:1109–1120. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Zhang LJ, Wang KB, Liu LS, Chen LZ, Peng
BG, Liang LJ, Li Z, Xue L, Li W and Xia JT: Overexpression of
GOLPH3 is associated with poor prognosis and clinical progression
in pancreatic ductal adenocarcinoma. BMC Cancer. 14:5712014.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Chang YT, Wu CC, Shyr YM, Chen TC, Hwang
TL, Yeh TS, Chang KP, Liu HP, Liu YL, Tsai MH, et al:
Secretome-based identification of ULBP2 as a novel serum marker for
pancreatic cancer detection. PloS One. 6:e200292011. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Oron Y: Integrin-based therapy of
pancreatic adenocarcinoma: Current status and future perspectives.
Minerva Gastroenterol Dietol. 61:71–86. 2015.PubMed/NCBI
|
|
19
|
Strobel O, Hinz U, Gluth A, Hank T,
Hackert T, Bergmann F, Werner J and Buchler MW: Pancreatic
adenocarcinoma: Number of positive nodes allows to distinguish
several N categories. Ann Surg. 261:961–969. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Bever KM, Sugar EA, Bigelow E, Sharma R,
Laheru D, Wolfgang CL, Jaffee EM, Anders RA, De Jesus-Acosta A and
Zheng L: The prognostic value of stroma in pancreatic cancer in
patients receiving adjuvant therapy. HPB (Oxford). 17:292–298.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Cui J, Chen Y, Wang HY and Wang RF:
Mechanisms and pathways of innate immune activation and regulation
in health and cancer. Hum Vaccin Immunother. 10:3270–3285. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Myklebust LM, Horvli O and Raae AJ: RACK1
(receptor for activated C-kinase 1) interactions with spectrin
repeat elements. J Mol Recognit. 28:49–58. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Campagne C, Jule S, Alleaume C, Bernex F,
Ezagal J, Chateau-Joubert S, Estrada M, Aubin-Houzelstein G,
Panthier JJ and Egidy G: Canine melanoma diagnosis: RACKI as a
potential biological marker. Vet Pathol. 50:1083–1090. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Deng YZ, Yao F, Li JJ, Mao ZF, Hu PT, Long
LY, Li G, Ji XD, Shi S, Guan DX, et al: RACK1 suppresses gastric
tumorigenesis by stabilizing the beta-Catenin destruction complex.
Gastroenterology. 142:812.e15–823.e15. 2012. View Article : Google Scholar
|
|
25
|
Ruan Y and Gu J: O-GlcNAc modification of
ribosomal RACK1 enhances hypoxia-induced epithelial-mesenchymal
transition in hepatocellular carcinoma. Glycobiology. 23:1345.
2013.
|
|
26
|
Choi YY, Lee SY, Lee WK, Jeon HS, Lee EB,
Lee HC, Choi JE, Kang HG, Lee EJ, Bae EY, et al: RACK1 is a
candidate gene associated with the prognosis of patients with early
stage non-small cell lung cancer. Oncotarget. 6:4451–4466. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Mamidipudi V and Cartwright CA: A novel
pro-apoptotic function of RACK1: Suppression of Src activity in the
intrinsic and Akt pathways. Oncogene. 28:4421–4433. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Saito A, Fujii G, Sato Y, Gotoh M,
Sakamoto M, Toda G and Hirohashi S: Detection of genes expressed in
primary colon cancers by in situ hybridisation: Overexpression of
RACK 1. Mol Pathol. 55:34–39. 2002. View Article : Google Scholar : PubMed/NCBI
|