Introduction
Acinetobacter baumannii, a leading nosocomial
pathogen, has been identified to induce serious infections and high
mortality rates in intensive care units (ICUs) (1,2).
Multidrug-resistant Acinetobacter baumannii (A.
baumannii) (MDRAB), with resistance to at least three classes
of antibiotics among cephalosporins, carbapenems, β-lactamases,
aminoglycosides and quinolones (3),
is only susceptible to certain agents such as tigecycline, and
polymyxins due to intrinsic and acquired resistance (4,5). In
addition, several pandrug-resistant (PDR) A. baumannii
strains with resistance to almost all available antibiotics have
been identified in previous decades (6,7).
Abuse of broad-spectrum antibiotics has been
demonstrated to be a major cause for the development of drug
resistance of A. baumannii. At present, a number of genes
responsible for drug resistance have been identified though
long-term studies on the mechanisms of bacterial resistance
(7–10). Production of β-lactamase is suggested
to be associated with the bacterial resistance to penicillin,
cephalosporins and carbapenems (9).
At present, four major categories have been available for the
β-lactamase protein-encoding genes, including narrow-spectrum
β-lactamase, extended-spectrum β-lactamase (ESBLs),
metallo-β-lactamases (MBLs) and oxacillinase (OXA)-type
carbapenemases (10). Bacterial
resistance to aminoglycoside usually results in the production of
aminoglycoside-modifying enzymes. Aminoglycoside resistance genes,
including acetyltransferase (aac), phosphotransferase
(aph) and adenylyltransferase (aad), have been
frequently identified in MDRAB (11).
For example, MDRAB has been demonstrated to acquire antimicrobial
resistance genes via class 1 integrons (Int-1), which
contain single or multiple gene cassettes (12). Carbapenemase and aminoglycoside
resistance genes were localized within Int-1 (13). In addition, 16S rRNA methylases may
confer resistance to aminoglycosides (14). The increased production of
fluoroquinolone-resistant A. baumannii was demonstrated to
be markedly associated with spontaneous mutations in the quinolone
resistance-determining regions (QR-DRs) in DNA gyrase (gyrA)
or topoisomerase IV (parC) (15).
MDRAB remains a challenge for the clinical
management of life-threatening infections, including bacteremia,
pneumonia and wound infections. At present, MDRAB is a lethal
threat to public health due to the lack of effective antimicrobial
agents available. Currently, combination therapy has been
considered to be a promising method for the management of
infection. Previous studies revealed that tigecycline and
polymyxins were active against MDRAB (4,5), but their
application is inhibited due to high toxicity and low commercial
availability in China. Consequently, the effective combinations of
clinical drugs may be an improved choice for treating MDRAB
infection. In the present study, the genotypes and encoding
resistance genes of MDRAB were determined. Based on the phenotypic
analysis, the gene structure and molecular determinants that confer
alternative MDRAB phenotypes were investigated. Furthermore, five
drugs were used to evaluate the in vitro activity of various
antibiotic combinations against MDRAB, in order to provide reliable
data to support novel clinical combination therapies.
Materials and methods
Bacterial isolates
A total of 34 consecutive and non-repetitive MDRAB
strains were identified using the MicroScan WalkAway-96 system
(Siemens AG, Munich, Germany) from the Second Xiangya Hospital
(Changsha, China) between February 2011 and May 2011. Kirby-Bauer
antibiotic susceptibility testing (K-B test) was utilized to
determine the susceptibility to several clinically significant
antimicrobial agents. MDRAB was defined as the presence of
resistance to at least three classes of antibiotics, including:
Cephalosporins, carbapenems, β-lactamases, aminoglycosides and
quinolones. In the present study, a total of 34 strains were
collected (Table I), 30 of which were
identified from sputum samples from patients with nosocomial
pneumonia, referring to criteria for radiologically-confirmed
pneumonia occurring ≥48 h after hospitalization in non-intubated
patients (16), 3 were isolated from
wound secretion and one was isolated from fluid drainage. The
samples were primarily collected from the ICU Respiratory and
Cardiothoracic Surgery departments. The interpretation of
susceptibility test and breakpoints was performed according to the
Clinical and Laboratory Standard Institute (CLSI) criteria
(17). The present study was approved
by the Ethics Committee of The Second Xiangya Hospital, Central
South University (Changsha, China). Written informed consent was
obtained from all patients.
 | Table I.Characteristics of study
isolates. |
Table I.
Characteristics of study
isolates.
|
|
|
|
|
Clinical
antimicrobial susceptibility |
|---|
|
|
|
|
|
|
|---|
| NOx | PT | Date of strain
separated | Ward | TZP | CAZ | SCF | ATM | IPM | AK | FEP | MEM | TIM | CIP | SXT | CTX | LEV | CN | SAM |
|---|
| 1AI | 51M | 3/9 | 3 | R | R | R | R | R | R | R | R | R | R | R | R | R | R | R |
| 2AI | 13M | 3/9 | 4 | R | R | R | R | R | R | R | R | R | R | R | R | R | R | R |
| 3BI | 46M | 3/23 | 1 | R | R | I | R | R | R | R | R | R | R | R | R | R | R | R |
| 4AI | 64M | 3/22 | 10 | R | R | R | R | R | R | R | R | R | R | R | R | R | R | R |
| 5AI | 34M | 3/11 | 8 | R | R | R | R | R | R | R | R | R | R | R | R | R | R | R |
| 6BI | 1M | 3/17 | 3 | R | R | I | R | R | R | R | R | R | R | R | R | R | R | R |
| 7CI | 65M | 3/12 | 1 | R | R | R | R | R | R | R | R | R | R | R | R | I | R | R |
| 8AI | 7M | 5/14 | 4 | R | R | R | R | R | R | R | R | R | R | R | R | R | R | R |
| 9DI | 88F | 3/31 | 6 | R | R | S | R | R | R | R | R | R | R | R | R | R | R | R |
|
10BI | 33F | 3/28 | 3 | R | R | I | R | R | R | R | R | R | R | R | R | R | R | R |
|
11BII | 25F | 3/30 | 3 | R | R | I | R | R | R | R | R | R | R | R | R | R | R | R |
|
12BII | 48F | 3/28 | 1 | R | R | I | R | R | R | R | R | R | R | R | R | R | R | R |
|
13BII | 30F | 3/28 | 1 | R | R | I | R | R | R | R | R | R | R | R | R | R | R | R |
|
14BII | 47F | 4/2 | 2 | R | R | I | R | R | R | R | R | R | R | R | R | R | R | R |
|
15BII | 55M | 4/21 | 3 | R | R | I | R | R | R | R | R | R | R | R | R | R | R | R |
|
16BIV | 25M | 4/21 | 1 | R | R | I | R | R | R | R | R | R | R | R | R | R | R | R |
|
17AII | 63M | 3/20 | 1 | R | R | R | R | R | R | R | R | R | R | R | R | R | R | R |
|
18BI | 72F | 3/23 | 1 | R | R | I | R | R | R | R | R | R | R | R | R | R | R | R |
|
19CI | 57M | 3/12 | 1 | R | R | R | R | R | R | R | R | R | R | R | R | I | R | R |
|
20AI | 78M | 3/18 | 1 | R | R | R | R | R | R | R | R | R | R | R | R | R | R | R |
|
21AII | 41M | 3/10 | 4 | R | R | R | R | R | R | R | R | R | R | R | R | R | R | R |
|
22EII | 80F | 3/30 | 1 | R | R | I | R | R | R | R | R | R | R | R | R | R | I | R |
|
23FI | 46M | 4/5 | 4 | R | R | I | R | R | R | I | R | R | R | R | R | R | R | R |
|
24BII | 43M | 4/3 | 9 | R | R | I | R | R | R | R | R | R | R | R | R | R | R | R |
|
25AII | 55M | 4/5 | 5 | R | R | R | R | R | R | R | R | R | R | R | R | R | R | R |
|
26AII | 55M | 4/6 | 5 | R | R | R | R | R | R | R | R | R | R | R | R | R | R | R |
|
27BII | 57F | 4/7 | 2 | R | R | I | R | R | R | R | R | R | R | R | R | R | R | R |
|
28AI | 70F | 4/7 | 2 | R | R | R | R | R | R | R | R | R | R | R | R | R | R | R |
|
29DII | 69M | 4/5 | 2 | R | R | S | R | R | R | R | R | R | R | R | R | R | R | R |
|
30GII | 87F | 2/28 | 1 | R | S | R | R | R | R | R | R | R | R | R | R | R | S | S |
|
31BIII | 56M | 2/28 | 1 | R | R | I | R | R | R | R | R | R | R | R | R | R | R | R |
|
32BI | 6F | 2/28 | 7 | R | R | I | R | R | R | R | R | R | R | R | R | R | R | R |
|
33BI | 27M | 3/4 | 1 | R | R | I | R | R | R | R | R | R | R | R | R | R | R | R |
|
34HI | 79M | 2/28 | 1 | R | S | I | R | R | R | R | R | R | R | R | R | R | R | R |
Random amplified polymorphic DNA
(RAPD) genotyping and detection of drug resistance genes by
polymerase chain reaction (PCR)
Bacterial DNA was extracted and purified by using
Tiangen UltraClean Microbial DNA Isolation kit (Tiangen Biotech
Co., Ltd., Beijing, China) according to the manufacturer's
instructions. MDRAB genotyping was conducted using RAPD (Applied
Biosystems; Thermo Fisher Scientific, Inc., Waltham, MA, USA) with
an AP2 primer (5-GTTTCGCTCC-3) designed by Primers Express
software (version 2.0; Applied Biosystems; Thermo Fisher
Scientific, Inc.) and synthesized by Sangon Biotech Co., Ltd.
(Shanghai, China), as previously described (18). The genes encoding resistance,
including β-lactamase TEM-1, AmpC,
metallo-β-lactamase IMP-5, oxacillinase (OXA)-23,
OXA-24, acetylyltransferase aac(3)-I,
aac(6′)-I, ant(3″)-I, 16S rRNA methyltransferase
armA, 16S rRNA methylase rmtA, rmtB,
phosphotransferase aph-(3), AdeB family multidrug efflux RND
transporter adeB, adeRS two-component system,
Int-1 and ParC genes, were detected by PCR. All
primers were designed based on the sequences in GenBank (19) using Primer Express software v2.0
(Applied Biosystems; Thermo Fisher Scientific, Inc., Foster City,
CA, USA) (Table II). The PCR total
reaction volume of 20 µl, containing 10 µl 2X TaqMan PCR master
mix, 1 µl forward and reverse primers (10 µmol/l), 7 µl
ddH2O and 1 µl DNA template. The amplification
conditions of the target genes were presented in Table III. Following amplification, 3 µl of
products were electrophoresed on a 1.5% agarose gel (Oxoid; Thermo
Fisher Scientific, Inc.) and visualized using ethidium bromide
(Sigma-Aldrich; Merck KGaA, Darmstadt, Germany) for 20 min at a
voltage of 150 V.
 | Table II.Primers of resistance genes. |
Table II.
Primers of resistance genes.
|
| Primer sequences
(5′-3′) |
|
|---|
|
|
|
|
|---|
| Target genes | Forward | Reverse | Size, bp |
|---|
| TEM-1 |
TTCGTGTCGCCCTTATTC |
ACGCTCGTCGTTTGGTAT | 512 |
| IMP-5 |
CTACCGCAGCAGAGTCTTTG |
AACCAGTTTTGCCTTACCAT | 587 |
| OXA-23 |
TGTCATAGTATTCGTCGTT |
TTCCCAAGCGGTAAA | 453 |
| OXA-24 |
TTTGCCGATGACCTT |
TAGCTTGCTCCACCC | 175 |
| AmpC |
CGACAGCAGGTGGAT |
GGTTAAGGTTGGCATG | 510 |
|
aac(3)-I |
ACCTACTCCCAACATCAGCC |
ATATAGATCTCACTACGCGC | 158 |
|
aac(6′)-I |
TATGAGTGGCTAAATCGA |
CCCGCTTTCTCGTAGCA | 395 |
|
ant(3″)-I |
TGATTTGCTGGTTACGGTGAC |
CGCTATGTTCTCTTGCTTTTG | 284 |
| armA |
GGGGTCTTACTATTCTG |
TTCCCTTCTCCTTTC | 503 |
| rmtA |
CCTAGCGTCCATCCTTTCCTC |
AGCGATATCCAACACACGATGG | 315 |
| rmtB |
ATGAACATCAACGATGCCCTC |
TTATCCATTCTTTTTTATCAAGTATAT | 756 |
| aph(3) |
ATACAGAGACCACCATACAGT |
GGACAATCAATAATAGCAAT | 234 |
| adeB |
GTATGAATTGATGCTGC |
CACTCGTAGCCAATACC | 1,000 |
| adeRS |
CTCAGACTCCCGTGATCATGTTG |
CGTAAGTCTTCGACTAAGTGAGA | 1,115 |
| Int-1 |
GCACCGCCAACTTTC |
CCTTGATGTTACCCGAGA | 433 |
| ParC |
CTGAACAGGCTTACTTGAA |
AAGTTATCTTGCCATTCG | 200 |
 | Table III.Polymerase chain reaction conditions
of target genes. |
Table III.
Polymerase chain reaction conditions
of target genes.
|
|
| Amplification |
|
|
|---|
|
|
|
|
|
|
|---|
| Target genes | Initialdenaturation
(°C, min) | Denaturation (°C,
sec) | Annealing (°C,
sec) | Extension (°C,
sec) | Cycles (n) | Final extension
(°C, min) |
|---|
| AP2 | 95, 6 | 95, 45 | 33, 45 | 72, 120 | 45 | 72, 10 |
| TEM-1 | 94, 5 | 94, 60 | 55, 60 | 72, 50 | 30 | 72, 7 |
| IMP | 94, 5 | 94, 60 | 55, 60 | 72, 50 | 30 | 72, 7 |
| OXA-23 | 94, 5 | 94, 30 | 48, 30 | 72, 35 | 30 | 72, 10 |
| OXA-24 | 94, 5 | 94, 30 | 48, 30 | 72, 35 | 30 | 72, 10 |
| AmpC | 94, 5 | 94, 30 | 50, 30 | 72, 50 | 30 | 72, 10 |
|
aac(3)-I | 94, 5 | 94, 30 | 55, 30 | 72, 30 | 35 | 72, 10 |
|
aac(6′)-I | 94, 5 | 94, 30 | 55, 30 | 72, 30 | 35 | 72, 10 |
|
ant(3″)-I | 94, 5 | 94, 30 | 55, 30 | 72, 30 | 35 | 72, 10 |
| rmtA | 93, 2 | 93, 20 | 55, 30 | 72, 30 | 30 | 72, 5 |
| aph (3) | 93, 2 | 93, 20 | 55, 30 | 72, 30 | 30 | 72, 5 |
| armA | 94, 5 | 94, 30 | 47, 30 | 72, 35 | 30 | 72, 10 |
| rmtB | 94, 5 | 94, 30 | 55, 30 | 72, 60 | 30 | 72, 10 |
| adeB | 95, 5 | 95, 30 | 53, 60 | 72, 60 | 30 | 72, 7 |
| adeRS | 95, 5 | 95, 30 | 53, 40 | 72, 60 | 30 | 72, 7 |
| Int-1 | 94, 5 | 94, 30 | 53, 30 | 72, 60 | 30 | 72, 10 |
| ParC | 94, 4 | 94, 30 | 53, 30 | 72, 40 | 30 | 72, 7 |
Antimicrobial agents and minimum
inhibitory concentration (MIC) assays
Antimicrobial agents used in the present study were
amikacin (AK), cefoperazone/sulbactam [SCF, SCFI 1:1
(cefoperazone:sulbactam) and SCFII 2:1 (cefoperazone:sulbactam)],
meropenem (MEM), minocycline (MINO) and ciprofloxacin (CIP). MIC
assays were performed in 96-well microtiter plates by the broth
microdilution method, according to the CLIS protocol (17). Bacteria were cultured in 10% horse
blood agar (Oxoid; Thermo Fisher Scientific, Inc.) for 20–24 h
until cells reached the exponential phase. The inoculums were
adjusted with fresh Cationic adjustment of Mueller-Hinton Broth
[Oxoid; Thermo Fisher Scientific, Inc.; CAMHB, containing
Ca2+ (10–25 mg/l) and Mg2+ (10–12.5 mg/l)] to
produce solutions with ~5×105 colony forming units
(CFUs)/ml in a final volume of 100 µl. Subsequently, the bacteria
were cultured using various concentrations of drugs: AK and SCF,
256, 128, 64, 32, 16, 8 µg/ml; MEM and MINO, 64, 32, 16, 8, 4, 2
µg/ml; CIP, 16, 8, 4, 2, 1, 0.5 µg/ml, for 18–20 h at 37°C. The
average MIC (MICG), the concentration that inhibited 50%
of growth (MIC50) or 90% of strains (MIC90)
were calculated. All tests were performed in triplicate, and growth
and sterility controls were conducted simultaneously.
Chequerboard assay
A chequerboard assay was used to determine the
potential interactions between antibiotics. In each assay, a
combination of two antibiotics randomly chosen from the total five
was used, and the range of drug concentration was identical to the
MIC assays. The drugs in the 96-well plates were diluted with CAMHB
by checkerboard method as previously described (20). The broth microdilution plates were
inoculated with each MDRAB isolate (initial concentration of
bacteria was 0.5 McFarland) for 18–24 h at 37°C, to yield
~5×105 CFU/ml in a 100 µl volume. The effect of the
combinations was analyzed through measuring the fractional
inhibitory concentration index (FIC) with the following formula:
FICA + FICB, where FICA was the
ratio of MIC of drug A in combination compared with that of drug A
used alone, and FICB was the ratio of MIC of drug B in
combination compared with that of drug B used alone. The
interaction was defined as synergy (FIC ≤0.5), addition (0.5<
FIC ≤1), indifference (1< FIC ≤2) or antagonism (FIC >2),
respectively.
Results
Antimicrobial susceptibility
The MDRAB phenotype was determined according to the
susceptibility results. In total, 34 strains were isolated, among
which 11 strains (29.41%) were PDR A. baumannii with
resistance to almost all clinically significant agents. All the
strains were resistant to piperacillin/tazobactam, imipenem,
meropenem, amikacin, cefotaxime, ticarcillin/clavulanic acid and
ciprofloxacin, while only 14 strains (41.18%) were resistant to
cefoperazone/sulbactam. The isolates were also resistant to other
tested drugs including ceftazidime (97.06%), aztreonam (97.06%),
trimethoprim/sulfamethoxazole (97.06%), cefepime (94.12%),
gentamicin (94.12%), ampicillin/sulbactam (94.12%) and levofloxacin
(94.12%). The strains were classified into 8 phenotypes (A-H) based
on the resistance to the aforementioned primary clinical drugs
(Table I).
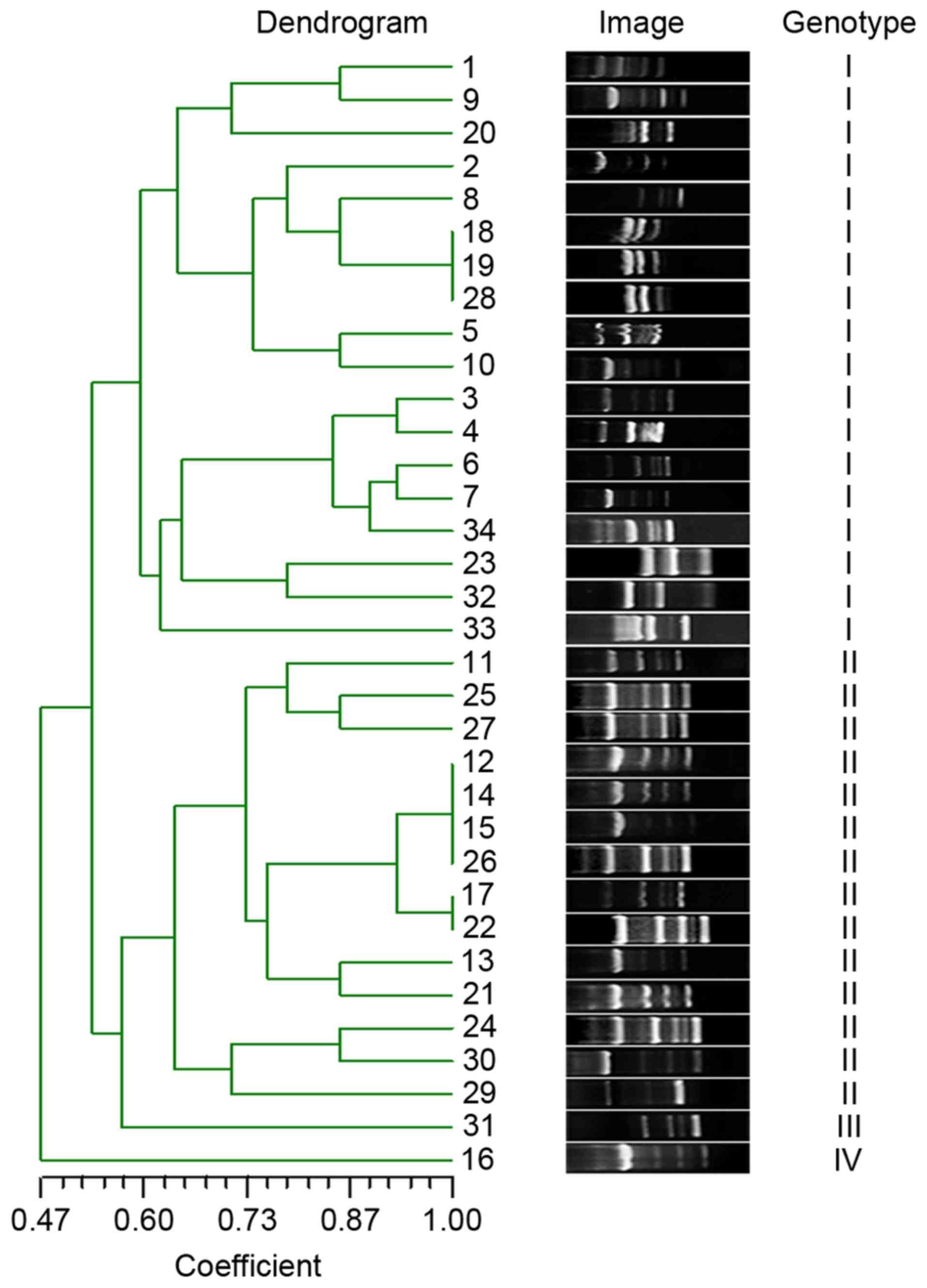
Genotypic diversity and molecular
determinants for MDR
To detect the extent of genotypic diversity of the
tested MDRAB, RAPD was performed and the fingerprint images were
analyzed by NTsys 2.10e (Exeter Software; Exeter Publishing Ltd.,
East Setauket, NY, USA) using dice similarity index for cluster
analysis and unweighted pair group average for dendrogram
construction. The isolates were clustered into four major genotypes
(I–IV) at 60% genotypic similarity threshold (Fig. 1).
Table IV summarizes
the distribution of resistance genes in the strains. TEM-1
and ParC were identified in all strains. A total of seven
genes demonstrated high positive rates, including OXA-23
(97.06%), AdeB (97.06%), AmpC (94.12%), Int-1
(94.12%), rmtA (94.12%), aph(3) (91.18%) and
armA (91.18%). The other genes, including aac(3)-I,
aac(6′-I, ant(3″)-I, rmtB, OXA-24 and
IMP-5, demonstrated positive rates of 29.41, 32.35, 76.47,
41.18, 85.29 and 64.71%, respectively.
 | Table IV.Distribution of positivity of various
resistance genes. |
Table IV.
Distribution of positivity of various
resistance genes.
| No. |
aac(3)-I |
aac(6′)-I |
ant(3”)-I | OXA-23 | OXA-24 | TEM-1 | IMP-5 | AmpC | ArmA | RmtA | RmtB | Aph(3) | AdeRS | AdeB | Int1 | ParC |
|---|
| 1 | N | N | P | P | P | P | N | P | P | P | N | P | N | P | P | P |
| 2 | N | N | P | P | P | P | P | P | N | P | P | P | N | P | P | P |
| 3 | N | N | P | P | P | P | P | P | P | P | N | P | N | P | P | P |
| 4 | N | N | P | P | P | P | P | P | P | P | N | P | N | P | P | P |
| 5 | N | N | P | P | P | P | P | P | P | P | N | P | N | P | P | P |
| 6 | N | N | P | P | P | P | P | P | P | N | N | P | N | P | P | P |
| 7 | N | N | P | P | P | P | P | P | N | P | N | P | N | P | P | P |
| 8 | N | N | P | P | P | P | P | P | P | P | P | P | N | P | P | P |
| 9 | N | N | P | P | P | P | P | P | P | P | P | P | N | P | P | P |
| 10 | N | N | P | P | P | P | N | P | P | P | N | N | N | P | P | P |
| 11 | N | N | P | P | P | P | P | P | P | P | N | P | N | P | P | P |
| 12 | N | N | P | P | P | P | P | P | P | P | N | P | N | P | P | P |
| 13 | P | N | P | P | P | P | P | P | P | P | N | P | N | P | P | P |
| 14 | N | N | P | P | P | P | P | P | P | P | P | P | N | P | P | P |
| 15 | N | N | P | P | P | P | P | P | P | P | N | P | N | P | P | P |
| 16 | N | N | P | P | P | P | P | P | P | P | P | P | N | P | P | P |
| 17 | N | N | P | P | P | P | P | P | P | P | N | P | N | P | P | P |
| 18 | N | N | P | P | P | P | P | P | P | P | N | P | N | P | P | P |
| 19 | N | N | P | P | N | P | P | P | P | P | N | P | N | P | P | P |
| 20 | N | P | N | P | P | P | P | P | P | P | P | P | N | P | P | P |
| 21 | N | N | P | P | P | P | P | P | P | P | N | P | N | P | P | P |
| 22 | N | N | P | P | P | P | P | P | P | P | P | P | N | P | P | P |
| 23 | P | P | N | P | N | P | N | P | P | P | N | P | N | P | P | P |
| 24 | P | P | N | P | N | P | N | P | P | P | N | P | N | P | P | P |
| 25 | N | N | N | N | N | P | N | P | N | P | P | P | N | P | P | P |
| 26 | P | N | N | P | P | P | N | N | P | P | P | P | N | P | N | P |
| 27 | P | P | P | P | P | P | N | P | P | P | N | P | N | P | P | P |
| 28 | P | P | N | P | P | P | N | P | P | P | P | P | N | P | N | P |
| 29 | P | P | P | P | P | P | N | P | P | P | P | P | N | P | P | P |
| 30 | N | P | N | P | N | P | N | N | P | P | P | N | N | P | P | P |
| 31 | P | P | P | P | P | P | P | P | P | P | P | P | N | P | P | P |
| 32 | P | P | P | P | P | P | P | P | P | P | P | P | N | P | P | P |
| 33 | N | P | N | P | P | P | N | P | P | P | N | P | N | P | P | P |
| 34 | P | P | P | P | P | P | N | P | P | P | N | N | N | N | P | P |
| P (%) | 29.41 | 32.35 | 76.47 | 97.06 | 85.29 | 100.00 | 64.71 | 94.12 | 91.18 | 97.06 | 41.18 | 91.18 | 0.00 | 97.06 | 94.12 | 100.00 |
MIC and the interaction of drug
combinations
The antibiotic susceptibility levels, expressed as
MIC of AK, SCF I, SCF II, MEM, MINO and CIP, were preliminarily
determined for the 34 MDRAB isolates. The distribution of
MIC50, MIC90 and MICG are
presented in Table V. The majority of
isolates were resistant to CIP (91.18%), SCF II (91.18%), amikacin
(85.29%), SCF I (82.35%) and MEM (73.53%), while less isolates
(5.88%) were resistant to MINO.
 | Table V.MIC parameters of drugs alone use or
in combination. |
Table V.
MIC parameters of drugs alone use or
in combination.
| Antibiotics
usage | MIC50
(µg/ml) | MIC90
(µg/ml) | MICG
(µg/ml) |
|---|
| AK alone | >256.00 | >256.00 | >222.24 |
| SCF I alone | 64.00 | >256.00 | >90.83 |
| SCF II alone | 128.00 | >256.00 | >130.35 |
| MEM alone | 33.60 | >64.00 | >46.39 |
| MINO alone | 3.50 | >25.00 | >5.50 |
| CIP alone | >16.00 | >16.00 | >14.13 |
| AK/SCF I |
|
|
|
| AK | 8 | 8 | 8 |
| SCF
I | 32 | 256 | 29.88 |
| AK/SCF II |
|
|
|
| AK | 8 | >256 | >24.24 |
| SCF
II | 64 | >256 | >57.41 |
| MEM/SCF I |
|
|
|
|
MEM | 2 | 32 | 2.82 |
| SCF
I | 8 | 128 | 17.18 |
| MEM/SCF II |
|
|
|
|
MEM | 2 | >64 | >6.14 |
| SCF
II | 8 | >256 | >26.12 |
| CIP/SCF I |
|
|
|
|
CIP | 0.5 | >16 | >0.96 |
| SCF
I | 32 | >256 | >48.24 |
| CIP/SCF II |
|
|
|
|
CIP | 0.5 | >16 | >1.85 |
| SCF
II | 64 | >256 | >87.76 |
| MINO/SCF I |
|
|
|
|
MINO | 2 | 2 | 2 |
| SCF
I | 8 | 8 | 8 |
| MINO/SCF II |
|
|
|
|
MINO | 2 | 4 | 3.24 |
| SCF
II | 8 | 8 | 8 |
| AK/MEM |
|
|
|
| AK | 8 | 8 | >37.18 |
|
MEM | 16 | >64 | >89.82 |
| AK/CIP |
|
|
|
| AK | >256 | >256 | >176.99 |
|
CIP | >16 | >16 | >12.25 |
| CIP/MEM |
|
|
|
|
CIP | >16 | >16 | >14.40 |
|
MEM | 16 | >64 | >24.65 |
| CIP/MINO |
|
|
|
|
CIP | 0.5 | 1 | 0.53 |
|
MINO | 2 | 4 | 2.47 |
| MEM/MINO |
|
|
|
|
MEM | 2 | 2 | 2 |
|
MINO | 2 | 2 | 2 |
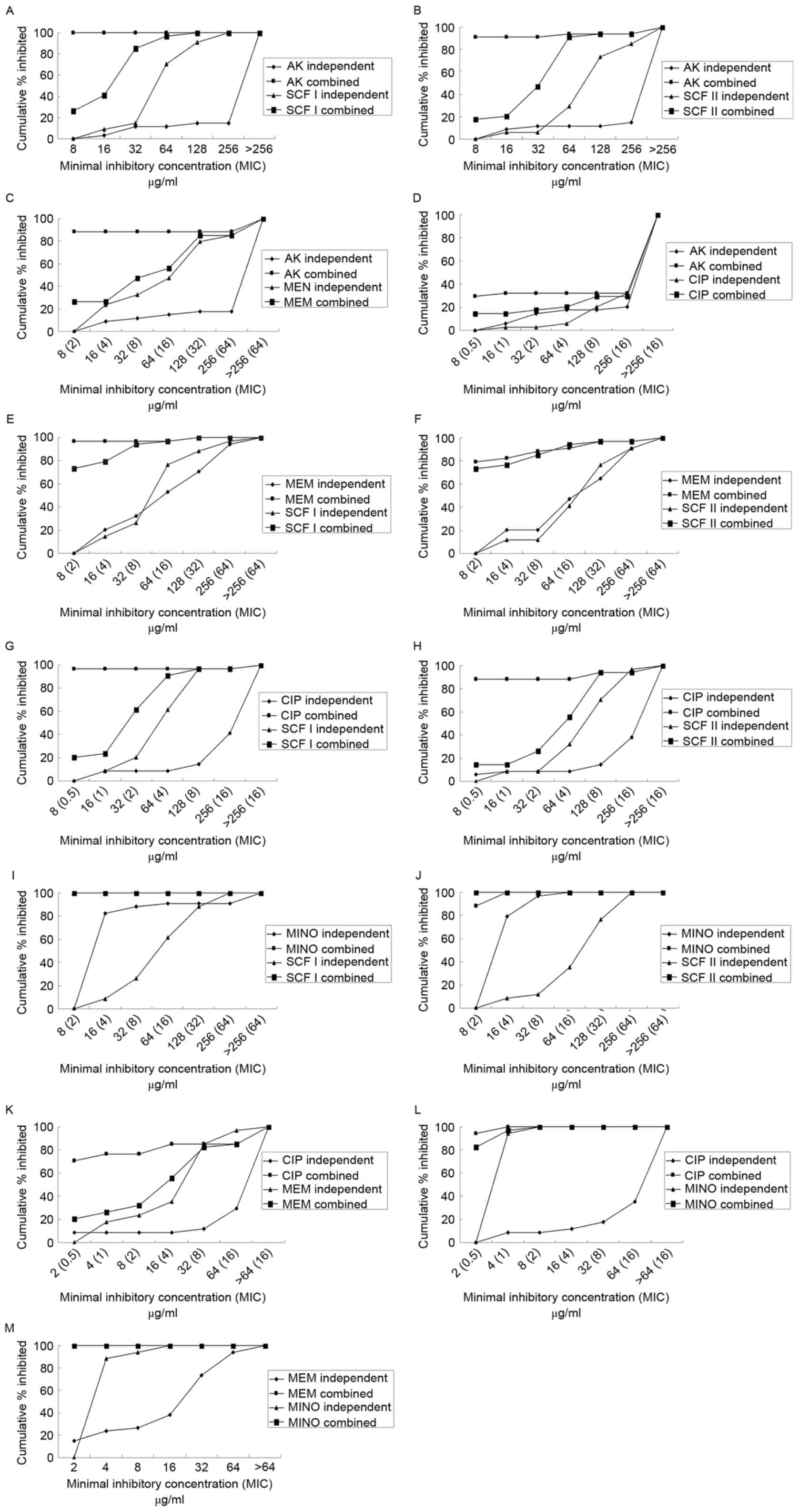
A chequerboard assay was performed with random
combinations of two drugs (Table V).
Fig. 2 demonstrates the percentage of
isolates inhibited at various MIC of antibiotics when use alone or
in combination. The majority of the drug combinations exhibited
superior inhibitory effects compared with each used alone. All the
combinations demonstrated synergism, with the exception of CIP with
MINO. MEM in combination with SCF I or SCF II primarily
demonstrated synergy, while the combination of AK+SCF I, AK+SCF II,
MINO+SCF I, MINO+SCF II, MINO+CIP, and MINO+MEM primarily exhibited
additive effects. Concurrently, the combination of AK+CIP
demonstrated evidence of antagonism (Fig.
3).
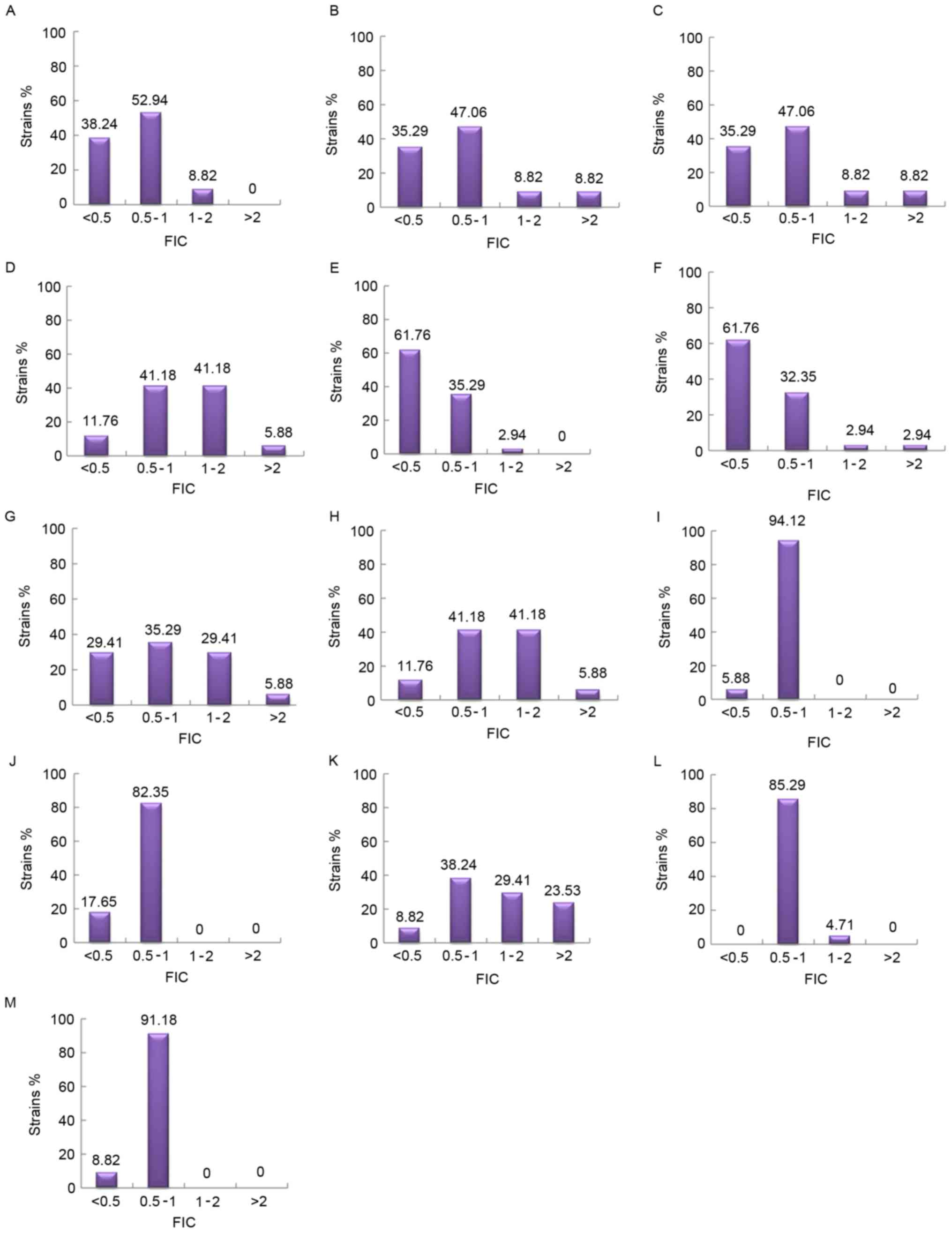 | Figure 3.The distribution of FIC of various
combinations of antimicrobial drugs. (A) AK+SCFI; (B) AK+SCFII; (C)
AK+MEM; (D) AK+CIP; (E) MEM+SCFI; (F) MEM+SCFII; (G) CIP+SCFI; (H)
CIP+SCFII; (I) MINO+SCFI; (J) MINO+SCFII; (K) CIP+MEM; (L)
CIP+MINO; and (M) MEM+MINO. Synergy, FIC ≤0.5; addition, 0.5<
FIC ≤1; indifference, 1< FIC ≤2; antagonism, FIC >2. AK,
amikacin; SCF, SCFI 1:1 and SCFII 2:1, cefoperazone/sulbactam; MEM,
meropenem; MINO, minocycline; CIP, ciprofloxacin; FIC, fractional
inhibitory concentration index. |
Discussion
Antibiotic resistance patterns observed in A.
baumannii exhibits the capacity to cause an epidemic globally.
MDRAB has been demonstrated to lead to serious hospital-acquired
infection, as limited therapeutic options are available (20). In the present study, the features of
34 MDRAB strains isolated from the Second Xiangya Hospital were
investigated, including antimicrobial susceptibility, genotypes,
screening of antibiotic resistance genes, MIC assay and antibiotic
interactions.
In the antimicrobial susceptibility study, eight
phenotypes (A-H) were classified using the K-B test. The results of
the drug susceptibility were in accordance with previous studies
(21,22) that highlighted the efficiency of
cefoperazone/sulbactam against MDRAB. However, the expression of
characteristic bacterial phenotypes may be easily affected by
various environmental factors. Therefore, only determining the
phenotype was not sufficient for the complete epidemiological
typing of various strains. Organism identification based on the
genotype is considered to be more reliable, as the genotype of each
organism is unique and invariable. In the present study, a
AP2 primer known to be efficient in RAPD genotyping was used
to amplify the genomic DNA of MDRAB strains (23). In total, four genotypes (I–IV) were
formed at a 60% similarity level. The genotypic distribution was
polyclonal, which was opposite to a previous study (24). In general, the ‘classic’ outbreaks of
MDRAB may be more frequently induced by a single clone spread among
people, whereas for the prevalence of polyclones in the Second
Xiangya Hospital, it may be associated the existence of mobile
genetic elements, for example integrons. The variety of patient
wards and isolate origin was hypothesized to be responsible for the
transmission of different subtypes. When comparing the phenotypes
with diverse fingerprinting profiles, it is noteworthy to select
isolates of the same phenotype with different genotypes, as it
manifested that the environment may affect the discrepancy between
genotype and phenotype.
To additionally investigate the resistant mechanisms
of MDRAB, the expression of the genes associated with drug
resistance was detected among the 34 isolates. Differences were
observed in the genetic characteristics of β-lactam, aminoglycoside
and quinolones resistance. Previously, the carbapenem resistance
associated with class D β-lactamase genes had been suggested to
cause serious therapeutic problems in clinical practice (25,26). The
high positive rate of OXA-23 (33/34) in the present study
was consistent with a previous study (27), while the proportion of
OXA-24-positive strains (29/34) was increased compared with
a previous study (28). The
production of OXA-23 and OXA-24 β-lactamase may be
the major cause for the selected MDRAB representing 100% resistance
to imipenem and meropenem. Other β-lactamase genes, including
TEM-1 (class A), IMP-5 (class B) and AmpC
(class C) were also identified in the present study, which may be
associated with the resistance of MDRAB to various types of
β-lactams including aztreonam, ceftazidime, cefepime, and
cefoperazone/sulbactam. At present, aminoglycoside-modifying
enzymes and 16S rRNA methylases have been suggested to be the most
important mechanism for bacterial resistance against
aminoglycosides (14). In the present
study, the co-existence of aph(3) (91.18%), ant(3″)-I
(76.47%), aac(3)-I (29.41%), aac(6′)-I (32.35%),
armA (91.18%), rmtA (94.12%) and rmtB (41.18%)
resistance genes confers the high resistance to amikacin and
gentamicin in MDRAB. Mutations in ParC were observed in all
isolates, which may assist in explaining the 100% resistance to
ciprofloxacin. The result was similar to a previous study (29). Among the isolates, the positive rates
of Int1 and adeB were 94.1 and 97.1%, respectively,
while the adeRS was completely negative. Integron and efflux
pump genes are non-specific resistance genes. Concurrently,
integrons are widely present in MDRAB, particularly Int-1,
which provides A. baumannii with a gene capture system
adapted to circumvent the challenges of multiple-antibiotic
treatment regimens (30). The results
of the present study demonstrated that Int1 serves a crucial
role in multiple drug resistance. In addition, the identification
of the co-existence of Int-1 and the majority of the
β-lactamase and aminoglycoside genes was notable, which is in
accordance with a previous study (31). This highlighted the importance of the
roles of Int-1 in the horizontal spreading of antibiotic
resistance genes, which may finally result in the polyclonal
prevalence of MDRAB in the Second Xiangya Hospital. AdeB is
a vital component of the AdeABC efflux pump, and the expression of
adeABC is regulated by the adeRS two-component regulatory
system (32). The sophisticated
feedback between them may account for the high positive ratio of
adeB and absolute absence of adeRS observed in the
present study.
The results of the present study confirmed that
several genes were associated with MDR. Abuse of antibiotic
chemotherapeutics affords major challenges in treating MDRAB
infections. At present, traditional therapy regimens are not
efficient to manage these life-threatening infections. Tigecycline
and polymyxins have been considered as the last resort for treating
MDRAB infections (4,5). However, they are still widely used
across mainland China due to the lack of any other qualified
commercial products. On this basis, the present study aimed to
identify an effective regimen to manage MDRAB infections through a
combination of antibiotics that are frequently used in clinical
practice. In the present study, 6 drugs (iAK, SCF I, SCF II, MEM,
MINO and CIP) were selected to study the in vitro activity
of drug combinations against MDRAB. The lower MIC of SCFI compared
with SCF II demonstrated a comparatively increased rate of in
vitro activity of SCF I against MDRAB, which was incompatible
with previous studies (33,34). This discrepancy may arise from the
geographical and biological evolutional differences. In addition, a
high percentage (76.47%) of MDRAB was susceptible to minocycline,
indicating the potential of this drug for the treatment of this
fatal infection (35). The
chequerboard assay indicated synergism for all the tested
combinations, particularly the combination of MEM+SCF I, and
MEM+SCF II, with the exception of CIP+MINO. Conversely, the
combinations of MINO+SCF I, MINO+SCF II, CIP+MINO and MEM+MINO
demonstrated additive effects. In summary, the combination of
cefoperazone-sulbactam or meropenem-minocycline has been indicated
to be more active compared with ciprofloxacin-amikacin, which is
similar to the recent surveillance data (16,21). The
present study also demonstrated that meropenem and
cefoperazone-sulbactam were generally active in MDRAB. In a
previous study, the combined utilization of meropenem and sulbactam
was considered a therapeutic option for A. baumannii
infection (36). However, only a
small number of studies have been focused on the study of the
efficiency of a MEM+SCF combination. The present study attempted to
investigate the in vitro activity of two types of SCF (SCF I
1:1; SCF II 2:1) combined with MEM against 34 strains of MDRAB. The
results indicated a marked synergistic interaction in the majority
tested isolates. Although no significant differences were observed
in the activity of cefoperazone-sulbactam combined with meropenem,
it revealed a novel potential option for clinical combination
therapy. Meropenem belongs to the family of β-lactam antibiotics,
while cefoperazone-sulbactam is a type of the third-generation
cephalosporin and β-lactamase inhibitor. When meropenem is combined
with cefoperazone-sulbactam, they may bind to different types of
penicillin bonding proteins, executing their bactericidal effects.
Concurrently, sulbactam may irreversibly inhibit β-lactamase
activity. This may be the most probable explanation for the
synergism observed. MINO was active against MDRAB whenever it is
used alone or combination. Although it is only a second-line
antibiotic for the majority of clinical bacterial infections, its
potential antibacterial activity against MDRAB should not be
neglected. In the present study, the combination of AK+CIP produced
an antagonism of 70.59%. Therefore, the combined use of these drugs
should be avoided in clinical practice.
In conclusion, the identification of fingerprinting
diversity highlights the issues with the polyclonal and horizontal
spread of MDRAB in the Second Xiangya Hospital. Although the
co-occurrence of numerous resistance-encoding genes presented a
completely threaten for the active therapy, the determination of
efficacious combinations among minocycline, meropenem and
cefoperazone-sulbactam, particularly MEM+SCFI and MEM+SCFII,
provides improved choices for the rational clinical combination
therapy for MDRAB infections. Additionally, MINO may be the
alternative choice to overcome the critical resistance of A.
baumannii. The present study failed to depict the
pharmacokinetics and pharmacodynamics of these drug combinations.
Future studies should focus on updating these data and proceed to
additionally identify the clinical effects of combination
therapy.
Acknowledgements
The present study was supported by the China
National Natural Scientific Foundation (grant no. 81470133), and
the Science and Technology Planning Project of Hunan Province of
China (grant no. 2015JC3035).
References
|
1
|
Visca P, Seifert H and Towner KJ:
Acinetobacter infection - an emerging threat to human health. IUBMB
Life. 63:1048–1054. 2011. View
Article : Google Scholar : PubMed/NCBI
|
|
2
|
Munoz-Price LS and Weinstein RA:
Acinetobacter infection. N Engl J Med. 358:1271–1281. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Dent LL, Marshall DR, Pratap S and Hulette
RB: Multidrug resistant Acinetobacter baumannii: A descriptive
study in a city hospital. BMC Infect Dis. 10:1962010. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Principe L, D'Arezzo S, Capone A,
Petrosillo N and Visca P: In vitro activity of tigecycline in
combination with various antimicrobials against multidrug resistant
Acinetobacter baumannii. Ann Clin Microbiol Antimicrob.
8:182009. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Lim TP, Tan TY, Lee W, Sasikala S, Tan TT,
Hsu LY and Kwa AL: In-vitro activity of polymyxin B, rifampicin,
tigecycline alone and in combination against carbapenem-resistant
Acinetobacter baumannii in Singapore. PLoS One.
6:e184852011. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Weng XB and Mi ZH: Phylogenetic analysis
on pandrug-resistant Acinetobacter baumannii strains.
Zhonghua Liu Xing Bing Xue Za Zhi. 32:948–949. 2011.(In Chinese).
PubMed/NCBI
|
|
7
|
Zhao WS, Liu GY, Mi ZH and Zhang F:
Coexistence of blaOXA-23 with armA and novel gyrA mutation in a
pandrug-resistant Acinetobacter baumannii isolate from the
blood of a patient with haematological disease in China. J Hosp
Infect. 77:278–279. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Wen Y, Ouyang Z, Yu Y, Zhou X, Pei Y,
Devreese B, Higgins PG and Zheng F: Mechanistic insight into how
multidrug resistant Acinetobacter baumannii response
regulator AdeR recognizes an intercistronic region. Nucleic Acids
Res. 45:9773–9787. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Yang H, Huang L, Barnie PA, Su Z, Mi Z,
Chen J, Aparna V, Kumar D and Xu H: Characterization and
distribution of drug resistance associated β-lactamase, membrane
porin and efflux pump genes in MDR A. Baumannii isolated from
Zhenjiang, China. Int J Clin Exp Med. 8:15393–15402.
2015.PubMed/NCBI
|
|
10
|
Jiang W, Liu H, Zhong M, Yang YC, Xiao DW
and Huang WF: Study on the resistant genes to carbapenems and
epidemiological characterization of multidrug-resistant
Acinetobacter baumannii isolates. Microb Drug Resist.
19:117–123. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Ramirez MS and Tolmasky ME: Aminoglycoside
modifying enzymes. Drug Resist Updat. 13:151–171. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Petersen A, Guardabassi L, Dalsgaard A and
Olsen JE: Class I integrons containing a dhfrl trimethoprim
resistance gene cassette in aquatic Acinetobacter spp. FEMS
Microbiol Lett. 182:73–76. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Azizi O, Shakibaie MR, Badmasti F,
Modarresi F, Ramazanzadeh R, Mansouri S and Shahcheraghi F: Class 1
integrons in non-clonal multidrug-resistant Acinetobacter
baumannii from Iran, description of the new blaIMP-55 allele in
In1243. J Med Microbiol. 65:928–936. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Tada T, Miyoshi-Akiyama T, Kato Y,
Ohmagari N, Takeshita N, Hung NV, Phuong DM, Thu TA, Binh NG, Anh
NQ, et al: Emergence of 16S rRNA methylase-producing
Acinetobacter baumannii and Pseudomonas aeruginosa
isolates in hospitals in Vietnam. BMC Infect Dis. 13:2512013.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Lee JK, Lee YS, Park YK and Kim BS:
Mutations in the gyrA and parC genes in ciprofloxacin-resistant
clinical isolates of Acinetobacter baumannii in Korea. Microbiol
Immunol. 49:647–653. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
American Thoracic Society, . Infectious
Diseases Society of America: Guidelines for the management of
adults with hospital-acquired, ventilator-associated, and
healthcare-associated pneumonia. Am J Respir Crit Care Med.
171:388–416. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Clinical and Laboratory Standards
Institute: Performance standards for antimicrobial susceptibility
testing: Twenty-second informational supplement M100-S21. CLSI.
31:64–65. 2011.
|
|
18
|
Zhang T, Wang M, Xie Y, Li X, Dong Z, Liu
Y, Wang L, Yang M, Song H, Cao H and Cao W: Active efflux pump adeB
is involved in multidrug resistance of Acinetobacter
baumannii induced by antibacterial agents. Exp Ther Med.
13:1538–1546. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Benson DA, Karsch-Mizrachi I, Lipman DJ,
Ostell J and Wheeler DL: GenBank. Nucleic Acids Res. 33:(Database
Issue). D34–D38. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Daoud Z, Mansour N and Masri K:
Synergistic combination of carbapenems and colistin against P.
aeruginosa and A. baumannii. Open J Med Microbiol.
3:253–258. 2013. View Article : Google Scholar
|
|
21
|
Huang J, Chen EZ, Qu HP, Mao EQ, Zhu ZG,
Ni YX, Han LZ and Tang YQ: Sources of multidrug-resistant
Acinetobacter baumannii and its role in respiratory tract
colonization and nosocomial pneumonia in intensive care unit
patients. Chin Med J (Engl). 126:1826–1831. 2013.PubMed/NCBI
|
|
22
|
Xu T, Xia W, Rong G, Pan S, Huang P and Gu
B: A 4-year surveillance of antimicrobial resistance patterns of
Acinetobacter baumanni in a university-affiliated hospital
in China. J Thorac Dis. 5:506–512. 2013.PubMed/NCBI
|
|
23
|
Koeleman JG, Stoof J, Biesmans DJ,
Savelkoul PH and Vandenbroucke-Grauls CM: Comparison of amplified
ribosomal DNA restriction analysis, random amplified polymorphic
DNA analysis, and amplified fragment length polymorphism
fingerprinting for identification of Acinetobacter genomic
species and typing of Acinetobacter baumannii. J Clin
Microbiol. 36:2522–2529. 1998.PubMed/NCBI
|
|
24
|
Fontana C, Favaro M, Minelli S, Bossa MC,
Testore GP, Leonardis F, Natoli S and Favalli C: Acinetobacter
baumannii in intensive care unit: A novel system to study
clonal relationship among the isolates. BMC Infect Dis. 8:792008.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Niranjan DK, Singh NP, Manchanda V, Rai S
and Kaur IR: Multiple carbapenem hydrolyzing genes in clinical
isolates of Acinetobacter baumannii. Indian J Med Microbiol.
31:237–241. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Feizabadi MM, Fathollahzadeh B,
Taherikalani M, Rasoolinejad M, Sadeghifard N, Aligholi M, Soroush
S and Mohammadi-Yegane S: Antimicrobial susceptibility patterns and
distribution of blaOXA genes among Acinetobacter spp.
Isolated from patients at Tehran hospitals. Jpn J Infect Dis.
61:274–278. 2008.PubMed/NCBI
|
|
27
|
Thapa B, Tribuddharat C, Srifuengfung S
and Dhiraputra C: High prevalence of bla(OXA)-23 in oligoclonal
carbapenem-resistant Acinetobacter baumannii from Siriraj
Hospital, Mahidol University, Bangkok, Thailand. Southeast Asian J
Trop Med Public Health. 41:625–635. 2010.PubMed/NCBI
|
|
28
|
Chen Z, Liu W, Zhang Y, Li Y, Jian Z, Deng
H, Zou M and Liu Y: Molecular epidemiology of carbapenem-resistant
Acinetobacter spp. from XiangYa Hospital, in Hunan Province,
China. J Basic Microbiol. 53:121–127. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Sung JY, Kwon KC, Cho HH and Koo SH:
Antimicrobial resistance determinants in imipenem-nonsusceptible
Acinetobacter calcoaceticus-baumannii complex isolated in
Daejeon, Korea. Korean J Lab Med. 31:265–270. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Peymani A, Farajnia S, Nahaei MR, Sohrabi
N, Abbasi L, Ansarin K and Azhari F: Prevalence of class 1 integron
among multidrug-resistant Acinetobacter baumannii in Tabriz,
northwest of Iran. Pol J Microbiol. 61:57–60. 2012.PubMed/NCBI
|
|
31
|
Farajnia S, Azhari F, Alikhani MY,
Hosseini MK, Peymani A and Sohrabi N: Prevalence of PER and VEB
type extended spectrum betalactamases among multidrug resistant
acinetobacter baumannii Isolates in North-West of Iran. Iran J
Basic Med Sci. 16:751–755. 2013.PubMed/NCBI
|
|
32
|
Yoon EJ, Courvalin P and Grillot-Courvalin
C: RND-type efflux pumps in multidrug-resistant clinical isolates
of Acinetobacter baumannii: Major role for AdeABC
overexpression and AdeRS mutations. Antimicrob Agents Chemother.
57:2989–2995. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Jones RN, Barry AL, Packer RR, Gregory WW
and Thornsberry C: In vitro antimicrobial spectrum, occurrence of
synergy, and recommendations for dilution susceptibility testing
concentrations of the cefoperazone-sulbactam combination. J Clin
Microbiol. 25:1725–1729. 1987.PubMed/NCBI
|
|
34
|
Barry AL and Jones RN: Criteria for disk
susceptibility tests and quality control guidelines for the
cefoperazone-sulbactam combination. J Clin Microbiol. 26:13–17.
1988.PubMed/NCBI
|
|
35
|
Pei G, Mao Y and Sun Y: In vitro activity
of minocycline alone and in combination with cefoperazone-sulbactam
against carbapenem-resistant Acinetobacter baumannii. Microb Drug
Resist. 18:574–577. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Deveci A, Coban AY, Acicbe O, Tanyel E,
Yaman G and Durupinar B: In vitro effects of sulbactam combinations
with different antibiotic groups against clinical Acinetobacter
baumannii isolates. J Chemother. 24:247–252. 2012. View Article : Google Scholar : PubMed/NCBI
|