Introduction
Renal ischemia/reperfusion (I/R) injuries are the
major causes of acute renal failure and may be involved in the
development and progression of certain forms of chronic kidney
disease (1). Ischemia is
associated with oxidative stress and apoptosis. Oxidative stress is
known to be a result of the imbalance between the production of
reactive oxygen species (ROS), antioxidants and repair processes
(2). Lipid peroxidation, mediated
by ROS, is believed to be an important cause of destruction and
damage to cell membranes during oxidative stress (3). Tissue levels of malondialdehyde (MDA)
are used as indicators of lipid peroxidation (4,5).
Glutathione (GSH) also provides major protection in oxidative
injury by participating in the cellular system of defense against
oxidative damage. Several reports have indicated that various
stimuli, such as I/R injury, cause GSH depletion (3). However, ROS also activates nuclear
factor-κB (NF-κB). NF-κB regulates the transcription of genes
involved in cellular responses to mechanical stress and is required
for tissue repair. It is recognized to have a potential role in
apoptosis and the adaptive response to stress (6,7).
Apoptosis is a form of programmed cell death, and in
this way cells that have been damaged in the organism are removed
without harming their environment (8,9). p53
is a tumor suppressor gene that plays an important role in cell
cycle control and apoptosis. If p53 is overexpressed, this may
cause certain structures to have various abnormalities (10). When tissue is exposed to I/R
injury, the DNA in tissue cells may be damaged. If DNA damage is
irreparable, p53 activates the apoptotic pathway and directs cell
death via apoptosis in order to protect the genome (11).
Nitric oxide (NO) is recognized as an important
mediator of physiological and pathological processes of renal I/R
injury (12–6). NO is produced by constitutive NO synthases (cNOSs),
including endothelial NOS (eNOS), neuronal NOS (nNOS) and inducible
NOS (iNOS). Alterations in the expression/activity of the different
NOS isoforms have been described in renal I/R (17). However, no study has been found
which simultaneously investigates the effects of quercetin on p53,
apoptosis, NF-κB and NOS gene expression in renal I/R injury.
Therefore, in the present study, we analyzed the effects of
quercetin on p53, apoptosis, eNOS and iNOS expression, as well as
NF-κB activation, in a renal I/R injury rat model.
Materials and methods
Animals
Sprague Dawley rats (n=42) weighing 250–300 g were
supplied from the Eskisehir Osmangazi University Experimental
Research Center (Turkey). Rats were housed in polycarbonate cages
in a temperature-controlled (21±1°C) and humidity-controlled
(45–55%) room, which was maintained on a 12/12 reversed light
cycle. The rats were fed with a standard rat chow (Oguzlar Yem,
Eskisehir, Turkey) and allowed to drink water ad libitum.
The present study was approved by the Eskisehir Osmangazi
University Institutional Local Animal Care and Use Committee
(67/2008). Animals were divided into three groups: group 1, control
(n=8+6); group 2, I/R (n=8+6); and group 3, I/R+quercetin (I/R+Q)
(n=8+6). The first 8 rats from each group were selected for
histological and biochemical evaluation; the remaining 6 rats were
used for real-time polymerase chain reaction (RT-PCR) analyses.
The rats were anesthetized with ketamine [50 mg/kg
intraperitoneally (i.p.)] and romphun (20 mg/kg i.p.) and placed on
a heating pad in order to maintain body temperature. Under aseptic
conditions, a midline incision was performed, and a non-traumatic
vascular clamp was applied to the left renal pedicle for 2 h and
allowed to reperfuse for 6 h. Quercetin (50 mg/kg i.p.) was
administered 1 h prior to these processes. At the end of the
reperfusion period, the left kidneys of the first 8 rats from each
group were removed. One-half of the kidney samples were used to
evaluate the biochemical parameters, such as MDA and GSH levels,
while the other half of the kidney samples were used for
immunohistochemical staining. For histological analyses, the
samples were maintained in 10% neutral formalin solution and for
biochemical analyses, the samples were kept frozen at −80°C until
analysis.
Biochemical determinations (tissue MDA
and GSH)
The kidney tissues were homogenized in 0.1 M
phosphate buffer (pH 7.4) using a homogenizer (Ultra Turrax IKA T18
basic; Wilmington, NC, USA). Homogenized samples were centrifuged
at 8,000 rpm for 10 min at 4°C, and the supernatant fractions were
analyzed for MDA and GSH. MDA and GSH levels were determined by
high-performance liquid chromatography (HPLC; Agilent 1100 Series,
Munich, Germany) with fluorescence detection using a commercial kit
(Chromsystems Diagnostics, Munich, Germany).
Histological determinations
All of the specimens were fixed in 10% neutral
formalin, dehydrated in an increasing alcohol series, cleared in
xylene and embedded in paraffin. Numerous 5-μm sections were
obtained and mounted on both poly-L-lysine-coated slides for
immunohistochemistry and classic slides for histochemistry. The
slides were stained with hematoxylin and eosin (H&E) for
histochemistry and p53, eNOS, NF-κB and the terminal
deoxynucleotidyl transferase dUTP nick end-labeling (TUNEL)
staining kit were used for immunohistochemistry. The slides were
evaluated under a light microscope (E600, Nikon, Japan).
DNA nick end-labeling of tissue
sections
For the detection of apoptosis in tissues, a
TUNEL-based apoptosis kit (FragEL DNA fragmentation kit;
Calbiochem, Darmstadt, Germany) was used. In this method, the
sections were first deparaf-finized and rehydrated. They were then
permeabilized with proteinase K and endogen peroxidase, inactivated
by 10% H2O2. For DNA labeling, Tdt Labeling
reaction mix and Tdt Enzyme mixture were used. Following
incubation, the reaction was detected with conjugate and
diaminobenzidine (DAB) solution in H2O2/urea
mixture. Methyl green (3%) was used for counter staining. The
slides were then dehydrated and mounted. The TUNEL-positive
brown-colored cells were considered to be apoptotic cells, in
agreement with the positive control supplied by the
manufacturer.
Immunohistochemistry
For the detection of p53, eNOS and NF-κB expression,
immunohistochemistry was applied. For antigen retrieval, microwave
treatment was used in 10 mM citrate buffer, pH 6.0 for all samples,
and endogenous peroxidase was inactivated by 10%
H2O2. The specimens were then reacted with
primary antibodies. The horseradish peroxidase detection system was
used as a secondary antibody and DAB was used for p53, AEC for eNOS
and NF-κB was used as a chromogen (all chemicals described above
were purchased from Lab Vision Corp., Fremont, CA, USA). Mayer’s
hematoxylin (Sigma, St. Louis, MO, USA) was used for
counterstaining. The positively stained cells were counted in image
analysis.
Image analysis
p53-, eNOS- and NF-κB-positive and apoptotic cells
in 20 different areas in each slide were counted under x20
objective magnification. For counting, UTHSCSA Image Tool for
Windows 3.0 image analysis software was used.
RNA extraction and RT-PCR
The mRNA level of iNOS in relation to the
housekeeping gene, glyceraldehyde 3-phosphate dehydrogenase
(GAPDH), was determined using RT-PCR with SYBR-Green. Total RNA was
extracted from renal tissue by the RNA stabilization reagent
(Qiagen, Valencia, CA, USA), according to the manufacturer’s
instructions, and quantified by measuring the absorbance at 260 nm
(Nanodrop1000; Thermo, Wilmington, DE, USA). Aliquots of 20 μl of
RNA from each group were applied for the production of
complementary DNA (cDNA). The newly synthesized cDNA, stored at
−20°C, was used for the mRNA assay of the iNOS isoform with RT-PCR.
cDNA (1 μl) from each group was amplified in 25 μl of reactive
mixture with 0.25X SYBR-Green Supermix (Molecular Probes,
Invitrogen, Carlsbad, CA, USA). RT-PCR was performed by monitoring
in real time the increase in the amount of SYBR-Green using
Rotor-Gene 6000 RT-PCR (Corbett Research, Sydney, Australia). The
oligonucleotide sequences of the cDNA primers were designed at Gene
Research Laboratories, UK. The forward primer for rat iNOS was
5′-CACCACCCTCCTTGTTCAAC-3′ and the reverse primer was
5′-CAATCCACAACTCGCTCCAA-3′. Sobajima et al also used GAPDH
(housekeeping gene) to normalize iNOS (target gene) data using
RT-PCR (18).
RT-PCR thermal cycling conditions were as follows: 5
min at 65°C, 60 min at 37°C for cDNA synthesis, 15 min at 95°C, 15
sec at 95°C, 1 min at 60°C for 50 cycles and 1 min at 55°C. RT-PCR
data were collected using the Rotor-Gene 6000 detection system.
Cycle threshold (CT) values were determined by automated threshold
analysis. Primer quality (lack of primer-dimer amplification) was
confirmed by melting curve analysis. Relative quantification of the
gene expression was performed using the standard curve method,
constructed with serial dilutions of control mRNA or RT-PCR
amplicons. All experiments were carried out in triplicate. iNOS
levels were standardized with GAPDH (ratio iNOS:GAPDH) to account
for loading differences. Gene expression levels (mRNA) were
reported using the median as a point estimator and the range of
values.
Statistical analysis
The statistical Package for Social Sciences (SPSS)
version for Windows 10.0 was used to evaluate the data. Results
were expressed as the means ± standard deviation (SD). Rejection of
the null hypothesis was set at p<0.05.
Biochemical and RT-PCR parameters were analyzed
using one-way ANOVA and the Tukey test for post hoc multiple
comparison. Histological parameters were analyzed using the
Mann-Whitney U test.
Results
Biochemical results
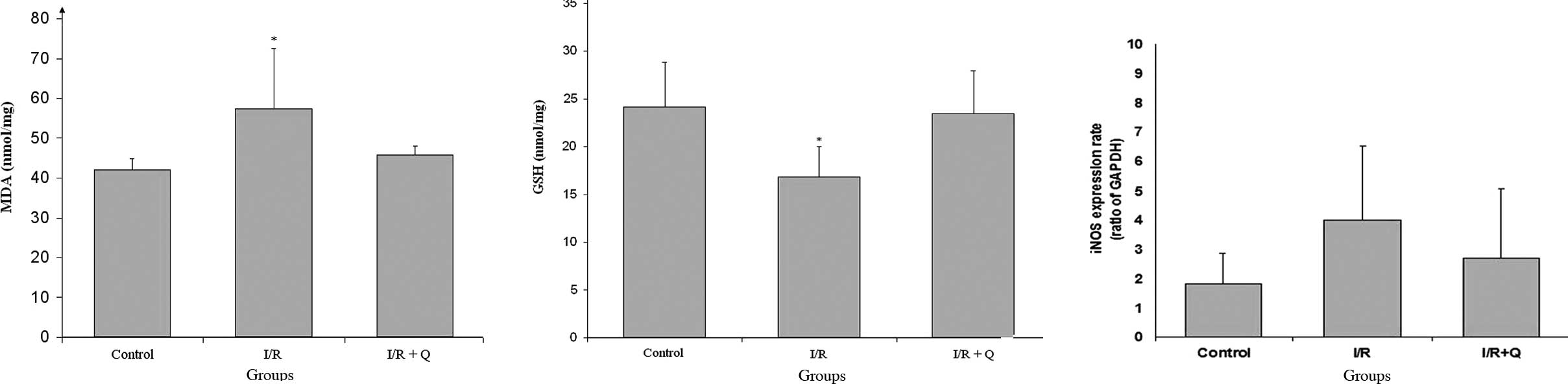
The level of MDA was significantly increased in the
I/R group when compared to the control and quercetin treatment
groups. Quercetin treatment decreased MDA levels, which was
significant when compared to the control and I/R+Q treatment groups
(p<0.05; Fig. 1A). While I/R
injury decreased GSH levels, quercetin treatment increased GSH
levels, which was significant when compared to the other groups
(p<0.05; Fig. 1B).
RT-PCR results
Expression of iNOS in renal
tissue
In iNOS, gene expression increased in the I/R group
and decreased in the I/R+Q group; however, there was no
statistically significant difference in the ratio of iNOS:GAPDH
between the I/R and I/R+Q groups (Fig.
1C).
Histological results
Microscopic determinations
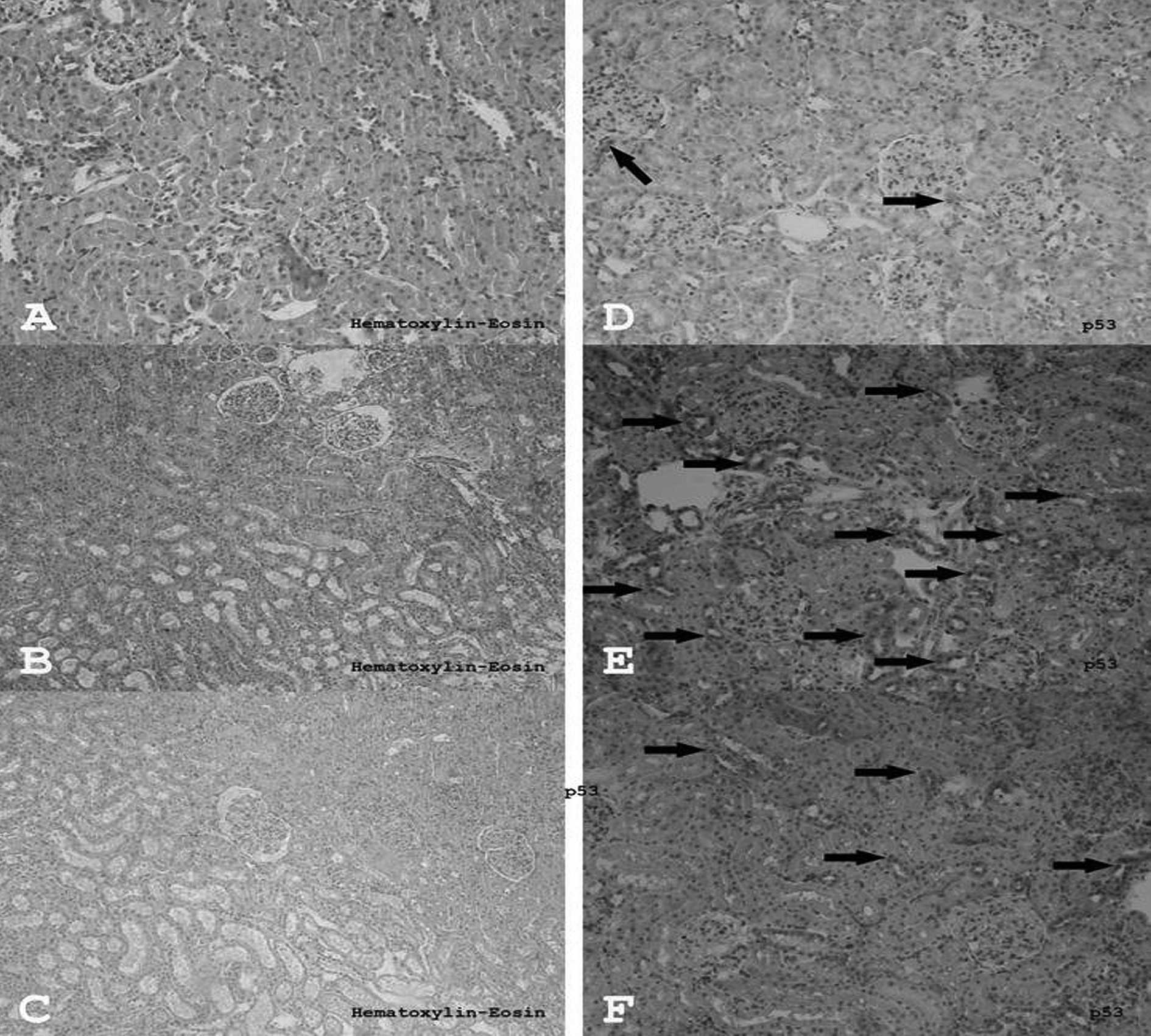
In the control group, H&E staining of all the
structures in the kidney were normal in appearance (Fig. 2A). Edema, vascularization, and
partly inflammatory fields, were observed in the I/R groups when
compared to the control and the quercetin-treated group (Fig. 2B). There were some inflammatory
cells and edema in the quercetin-treated group, however, these
fields were smaller and weaker than the inflammatory fields
observed in the I/R group (Fig.
2C). H&E-stained sections of rat kidneys are presented in
Table I.
 | Table I.H&E-stained sections of rat
kidneys. |
Table I.
H&E-stained sections of rat
kidneys.
| Histological
determinations | Control group | I/R group | I/R+Q group |
|---|
| Edema | 0 | ++ | + |
|
Vascularisation | 0 | +++ | + |
| Infiltration | 0 | ++ | + |
p53 expression
Although there were a few p53-expressing cells in
the renal tissue of the control group (Fig. 2D), more p53-expressing cells were
detected in the I/R group (Fig.
2E). Conversely, the number of p53-expressing cells in the
I/R+Q group (Fig. 2F) was
significantly decreased compared to the I/R group (p=0.001).
Apoptosis
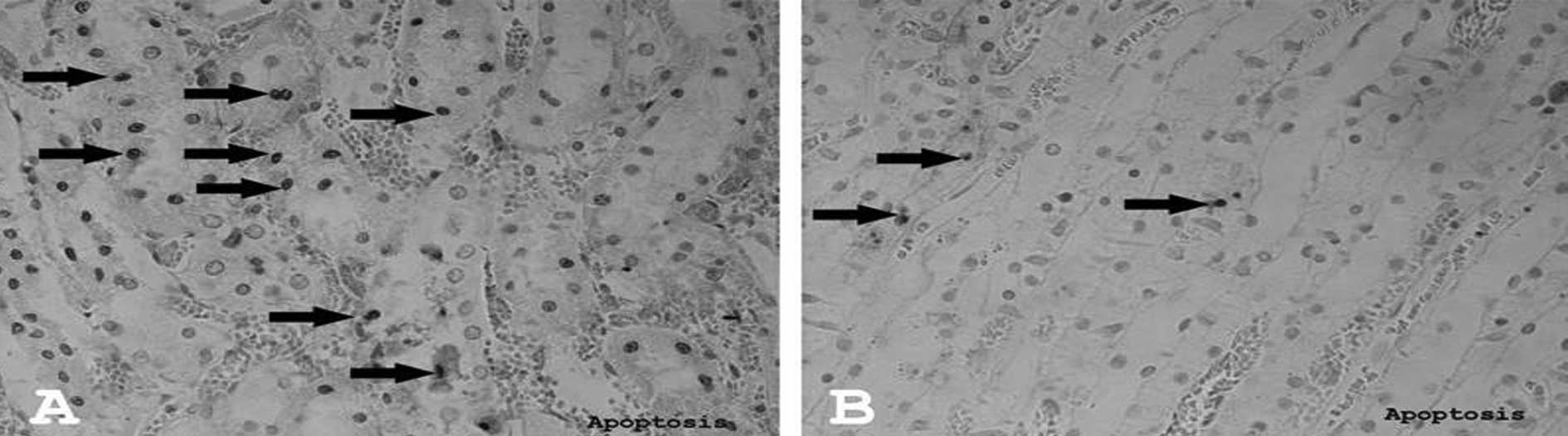
The number of apoptotic cells significantly
decreased in the I/R+Q group (Fig.
3B) compared to the I/R group (Fig. 3A) (p=0.016). There were no
apoptotic cells in the control group.
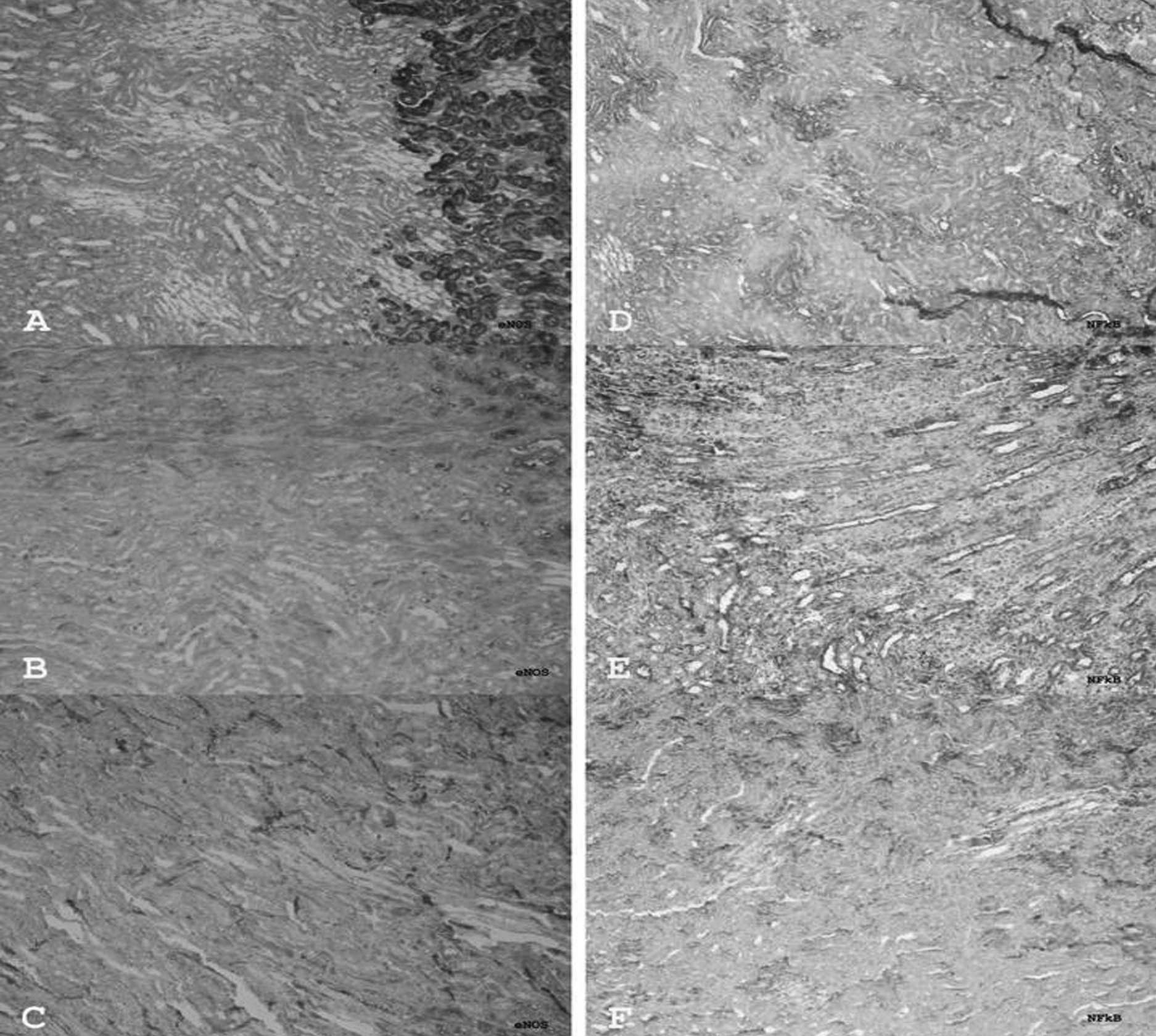
eNOS expression
In the control group, massive eNOS expression was
detected at the corticomedullar junction. This expression may be
related to a structurally dense vascular network in this area in
normal animals (Fig. 4A). However,
little eNOS expression was detected in the medullar cords and
cortex. However, in the I/R group many eNOS-expressing cells were
observed, particularly in the medullar cords (Fig. 4B). The number of these cells was
significantly decreased in the I/R+Q group (p=0.002) (Fig. 4C).
NF-κB expression
In the control group, many NF-κB-positive cells were
observed in the tissue, particularly at the corticomedullar
junction (Fig. 4D). Although this
number of cells was increased significantly in the I/R group
(p=0.000) (Fig. 4E), in the I/R+Q
group the number of these cells was clearly decreased (p=0.000)
(Fig. 4F).
Discussion
I/R induces oxidative stress in the kidneys, which
causes the enhancement of the reactive oxygen products and the
disappearance of the antioxidant defense system (8). In the present study, I/R injury was
created in the renal tissue of rats and it was found that the renal
damage created by I/R was significantly decreased by quercetin
treatment. In the present study, MDA, which indicates the degree of
lipid peroxidation, verifying the oxidative damage in tissue
(5) in the I/R-induced renal
tissue, significantly increased but then decreased with quercetin
therapy. A reduction in the GSH levels was observed in the I/R
group, whereas its level was increased by quercetin therapy. As we
have observed in our previous studies (19), a return of the increase in the
tissue MDA level in the I/R group, and a decrease in the GSH levels
back to normal with quercetin treatment, is a result of the
antioxidant effect of quercetin.
It is well known that ischemia, together with
oxidative stress and reactive oxygen products, arise as a result of
oxidative stress (2). ROS
activates NF-κB, which is a transcription factor responsible for
the production of adhesion molecules and cytokines. A number of
clinical and experimental studies have demonstrated that NF-κB, a
transcription factor, plays a central role in renal pathology
(6). Spandou et al reported
that renal I/R leads to the activation of NF-κB, which suggests
that it may be a marker of injury linked to the pathophysiology of
a variety of renal disorders, including I/R, and that its
inhibition prevents in vivo cell apoptosis associated with
renal I/R injury (20). Therefore,
inhibition of the permanent activation of NF-κB protects renal
tissue from ischemic injury. In the present study, we also detected
that NF-κB expression was increased in the renal I/R group.
However, this increase in NF-κB expression decreased with quercetin
treatment, and this demonstrates that quercetin may act like NF-κB
inhibitor.
The results of the histological examination revealed
that, upon overall tissue evaluation, significant edema,
vascularization and patchy infiltration areas formed by the
inflammatory cells were detected in the I/R group compared to the
quercetin treatment and control groups. In addition, a decreased
number of apoptotic cells was observed in the quercetin group when
compared to the I/R group. It is known that during renal I/R
injury, apoptosis is a very important contributor to kidney damage.
While evaluating experimentally created I/R, Vinas et al
demonstrated that the cells died of apoptosis or necrosis according
to the severity of the injury, due to direct cellular damage caused
by free oxygen radicals (17). In
accordance with this study, an increase in the number of apoptotic
cells subsequent to the renal I/R injury was observed in our
investigation; however, this increase was prevented by quercetin
treatment and this was due to the anti-apoptotic effect of
quercetin.
The p53 protein, which stimulates apoptosis, is a
guardian of the genome and is expressed in the presence of DNA
damage. Therefore, the expression of p53 in tissues reveals an
exposure to genotoxic stress, including radiation, chemical agents
and ischemia (21). In the present
study, we clearly detected the expression of p53 in the I/R group,
and this expression decreased in the case of quercetin
administration prior to I/R stress, which suggests that quercetin
has a protective effect on DNA damage. In addition to this decrease
in the level of apoptosis, the cell number increased in the I/R
group, suggesting that quercetin also has a protective effect on
cell injury and abnormal cell formation.
Renal I/R activates NOS and increases the expression
of NOS proteins (22,23). In spite of all the studies carried
out, the role of NO in I/R remains controversial. Several studies
have indicated that NO induces cellular cytotoxicity and tissue
injury via lipid peroxidation, as well as DNA damage and
pro-apoptotic effects in I/R injury (12,16,24).
Conversely, there are studies which demonstrate that the increased
activity of NOS is associated with reduced I/R-induced injury
(25). Alterations in the
expression of various NOS isoforms have been described in renal I/R
injury (17). Vinas et al
reported that, according to quantitative analysis, I/R increased
the renal expression of iNOS compared to the control and that no
differences were found between I/R and the various selective NOS
inhibitors (17). In the present
study, quantitative iNOS expression increased in the I/R group and
no significant difference was observed between the I/R and
quercetin treatment groups, similar to the Vinas et al
study. Immunohistochemical analysis revealed higher eNOS expression
in the I/R group; however, this was lower with quercetin
treatment.
Within the experimental studies that have been
carried out to date, we are unable to identify a study
simultaneously investigating the effects of quercetin on apoptosis,
p53, NF-κB and NOS gene expression in renal I/R injury. The results
of the present study reveal that quercetin treatment in I/R-injured
rat renal tissue reduces the injury by decreasing oxidative stress,
apoptosis and p53, NF-κB and eNOS gene expressions.
In conclusion, quercetin not only has antioxidant
and anti-apoptotic activities, but also an inhibitory effect on
eNOS and NF-κB for renal tissue protection during I/R injury in
rats. Therefore, quercetin may be used to protect renal tissue in
I/R injury, which may develop during surgical procedures.
References
|
1.
|
C ShinguH KogaS HagiwaraS MatsumotoK GotoI
YokoiT NoguchiHydrogen-rich saline solution attenuates renal
ischemia-reperfusion injuryJ
Anesth24569574201010.1007/s00540-010-0942-120480186
|
|
2.
|
MM ElahiYX KongBM MatataOxidative stress
as a mediator of cardiovascular diseaseOxid Med Cell
Longev2259269200910.4161/oxim.2.5.944120716913
|
|
3.
|
A KacmazA PolatY UserM TilkiS OzkanG
SenerOctreotide: a new approach to the management of acute
abdominal
hypertensionPeptides2413811386200310.1016/j.peptides.2003.09.00414706553
|
|
4.
|
E SahnaH ParlakpinarY TurkozA
AcetProtective effects of melatonin on myocardial
ischemia-reperfusion induced infarct size and oxidative
changesPhysiol Res54491495200515641932
|
|
5.
|
M JakesevicK AabyG BorgeB JeppssonS AhrneG
MolinAntioxidative protection of dietary bilberry, choke-berry and
Lactobacillus plantarum HEAL19 in mice subjected to
intestinal oxidative stress by ischemia-reperfusionBMC Complement
Altern Med111122011
|
|
6.
|
YM SeokJ KimMJ ParkYC BooYK
ParkWen-pi-tang-Hab-Wu-ling-san attenuates kidney fibrosis induced
by ischemia/reperfusion in micePhytother
Res2210571063200810.1002/ptr.244018570213
|
|
7.
|
EM BoyleTG CantyEN MorganW YunTH PohlmanED
VerrierTreating myocardial ischemia-reperfusion injury by targeting
endothelial cell transcriptionAnn Thorac
Surg6819491953199910.1016/S0003-4975(99)01033-410585109
|
|
8.
|
A KucukS KabadereM TosunT KokenMK KinaciB
IsikliN ErkasapProtective effects of doxycycline in
ischemia/reperfusion injury on kidneyJ Physiol
Biochem65183191200910.1007/BF0317906919886397
|
|
9.
|
DR ShultzJ WilliamJR HarringtonApoptosis:
programmed cell death at a molecular levelSemin Arthritis
Rheum32345369200310.1053/sarh.2003.5000512833244
|
|
10.
|
M TosunE TosunS KalkanMC Avundukp53
expression between 13–27 weeks old human male fetus gonadsJ Mol
Histol382712742007
|
|
11.
|
É BalintKH VousdenActivation and
activities of the p53 tumor suppressor proteinBr J
Cancer8518131823200111747320
|
|
12.
|
AF Lopez-NeblinaAJ PaezAH ToledoLH
Toledo-PereyraRole of nitric oxide in ischemia/reperfusion of the
rat kidneyCirc Shock4491951995
|
|
13.
|
MS GoligorskySV BrodskyE NoiriNitric oxide
in acute renal failure: NOS versus NOSKidney
Int61855861200210.1046/j.1523-1755.2002.00233.x11849438
|
|
14.
|
J Lopez-MartiA SolaF PiV AlfaroA MarcoG
HotterNucleotides modulate renal ischaemia-reperfusion injury by
different effects on nitric oxide and superoxideClin Exp Pharmacol
Physiol30242248200310.1046/j.1440-1681.2003.03821.x12680841
|
|
15.
|
A SolaV AlfaroJL VinasG HotterExogenous
adenosine enhances caspase-3 activity in warm renal
ischaemiaPflugers
Arch447387391200410.1007/s00424-003-1197-614605885
|
|
16.
|
LL WanJ XiaD YeJ LiuJ ChenG WangEffects of
quercetin on gene and protein expression on NOX and NOS after
myocardial ischemia and reperfusion in rabbitCardiovasc
Ther272833200910.1111/j.1755-5922.2009.00071.x19207477
|
|
17.
|
JL VinasA SolaM GensecaV AlfaroF PiG
HotterNO and NOS isoforms in the development of apoptosis in renal
ischemia/reperfusionFree Radic Biol
Med409921003200610.1016/j.freeradbiomed.2005.10.04616540395
|
|
18.
|
S SobajimaAL ShimerRC
ChadderdonQuantitative analysis of gene expression in a rabbit
model of intervertebral disc degeneration by real-time polymerase
chain reactionSpine
J51423200510.1016/j.spinee.2004.05.25115653081
|
|
19.
|
A KahramanN ErkasapM SerteserT
KokenProtective effect of quercetin on renal ischemia/reperfusion
injury in ratJ Nephrol16219224200312768068
|
|
20.
|
E SpandouI TsouchnikasG KarkavelasE
DounousiC SimeonidouO Guiba-TziampiriD TsakirisErythropoietin
attenuates renal injury in experimental acute renal failure
ischaemic/reperfusion modelNephrol Dial
Transplant21330336200610.1093/ndt/gfi17716221709
|
|
21.
|
Y ArikanM TosunS YilmazV SaykolZ
SoylemezThe comparative effects of pneumoperitoneum on apoptosis
and p53 expression in gastrointestinal organsJ Laparoendosc Adv
Surg Tech A18365713200810.1089/lap.2007.002318503368
|
|
22.
|
DM GoldbergSE HanhJG ParkesBeyond alcohol:
beverage consumption and cardiovascular mortalityClin Chim
Acta237155187199510.1016/0009-8981(95)06069-P7664473
|
|
23.
|
AI MoralesC Vicente-SanchezM JerkicEffect
of quercetin on metallothionein, nitric oxide synthases and
cyclooxygenase-2 expression on experimental chronic cadmium
nephrotoxicity in ratsToxicol Appl
Pharmacol210128135200610.1016/j.taap.2005.09.00616226777
|
|
24.
|
S Martinez-FlorezMB GutierrezS
Sanchez-CamposQuercetin prevents nitric oxide production and
nuclear factor kappa B activation in interleukin-1 beta activated
rat hepatocytesJ Nutr13513591365200515930438
|
|
25.
|
H ChenB XingX LiuB ZhanJ ZhouOzone
oxidative preconditioning protects the rat kidney from reperfusion
injury: the role of nitric oxideJ Surg
Res149287295200810.1016/j.jss.2007.12.75618262565
|