Introduction
Acute myeloid leukemia (AML) is a highly
heterogeneous form of hematopoietic malignancies; aberrantly
differentiated myeloid cells grow rapidly in the bone marrow and
other tissues, thereby inhibiting normal hematopoiesis and immune
function, and infiltrating other organs (1,2). AML
incidence increases with age and older patients typically exhibit a
lower long-term overall survival rate (3). The formation of AML has been associated
with multiple factors, including radiation, chemical degradation,
viral infection and multiple susceptible genes (4). Although the overall prognosis of AML
has improved markedly in recent years, the pathophysiological
mechanism of AML development remains unknown. To increase the
survival rate of patients with AML, it is of paramount importance
to improve comprehension of its molecular pathogenesis and identify
prognostic markers of AML.
Long non-coding RNAs (lncRNAs) are transcripts
>200 nucleotides long that are not translated into proteins
(5). Previous studies have indicated
that various lncRNAs may be associated with specific biological
processes, including chromatin modification, epigenetic regulation,
the cell cycle, cell apoptosis and differentiation (6–8). LncRNAs
are commonly dysregulated in the pathological processes of cancer,
and may therefore serve as prognostic markers (9,10).
It has been reported that zinc finger antisense 1
(ZFAS1), a newly identified lncRNA that maps to chromosome
20q13.13, is highly expressed in the mammary gland and
downregulated in breast tumors (11). Li et al (12) and Thorenoor et al (13) reported that ZFAS1 functions as an
oncogene in hepatocellular and colorectal carcinoma progression,
and is associated with cancer cell cycle progression, metastasis
and poor prognosis. Previous studies have also demonstrated that a
variety of lncRNAs are closely associated with hematological
malignancies (14–16). However, the role of ZFAS1 in AML
remains unknown; therefore, the present study investigated its
effects in various AML cell lines. Cell proliferation and
apoptosis, which serve important roles in AML cell development and
progression, were the primary mechanisms assessed to determine the
effects of ZFAS1 on AML in the present study.
Materials and methods
Cell lines
Four human AML cell lines (HL-60, KG-1, ML-1 and
SKNO-1), a T lymphocytic leukemia cell line (Jurkat) and a
Burkitt's lymphoma cell line (Raji) were purchased from the Cell
Bank of Chinese Academy of Sciences (Shanghai, China). The Jurkat
and Raji cell lines were used as controls. The cells were all
cultured in RPMI-1640 medium (Gibco; Thermo Fisher Scientific,
Inc., Waltham, MA, USA) supplemented with 10% fetal bovine serum
(Gibco; Thermo Fisher Scientific, Inc.) at 37°C in a humidified
incubator in the presence of 5% CO2. All cell lines were
passaged for fewer than six months.
RNA extraction and reverse
transcription-quantitative polymerase chain reaction (RT-qPCR)
Total RNA was extracted from all cell lines using
TRIzol® reagent (Invitrogen; Thermo Fisher Scientific,
Inc.). 1 µg Total RNA was reverse transcribed into cDNA using the
PrimeScript™ RT Master Mix (Takara Biotechnology Co., Ltd., Dalian,
China), according to the manufacturer's protocols. RT-qPCR was
performed using the SYBR® Premix DimerEraser™ (Takara
Biotechnology Co., Ltd.) and an ABI Prism 7500 instrument (Applied
Biosystems; Thermo Fisher Scientific, Inc.). The total PCR reaction
volume was 20 µl, including SYBR Premix DimerEraser (2X) 10 µl, PCR
forward primer (10 µM) 0.6 µl, PCR reverse primer (10 µM) 0.6 µl,
ROX Reference Dye II (50X) 0.4 µl, template 2 µl and sterile water
6.4 µl. The PCR cycling conditions were as follows: 95°C for 30
sec, followed by 40 cycles at 95°C for 5 sec and 60°C for 34 sec,
and a final extension step of 95°C for 15 sec, 60°C for 1 min, 95°C
for 15 sec and 60°C for 15 sec. GAPDH was amplified as an internal
control. The specific primer pairs were as follows: ZFAS1, forward,
5′-GCGAAAGCCATCTTTGGTTA-3′ and reverse, 5′-GGGCAGGACAATAGCGTATG-3′;
and GAPDH, forward, 5′-GGACCTGACCTGCCGTCTAG-3′ and reverse,
5′-GTAGCCCAGGATGCCCTTGA-3′. Relative mRNA expression was determined
using the 2−ΔΔCq method (17). All experiments were independently
repeated at least three times.
Transient transfection of small
interfering RNA (siRNA)
The sequences of ZFAS1 siRNA were as follows:
Forward, 5′-UCCAAAAUCCAUUCUGUACCC-3′ and reverse,
5′-GUACAGAAUGGAUUUUGGAAG-3′. ZFAS1 siRNA and the negative control
siRNA were purchased from Shanghai GenePharma Co., Ltd. (Shanghai,
China). Transfection of siRNA into the AML cell lines HL-60 and
SKNO-1 was performed with Lipofectamine® 2000
(Invitrogen; Thermo Fisher Scientific, Inc.) according to the
manufacturer's instructions. Following 48 h transfection, cells
were collected and applied to the subsequent assay.
Cell proliferation assay
Following 24 h transfection with siRNA,
1×103 cells/well were seeded in 96-well plates with 100
µl culture medium in triplicate. Cell proliferation was evaluated
by water-soluble tetrazolium salt at the indicated time points
using a Cell Counting kit-8 (CCK-8, Beyotime Institute of
Biotechnology, Haimen, China). Briefly, 10 µl CCK-8 solution was
added to each well and the mixture was incubated for 3 h at 37°C,
5% CO2. Absorbance (450 nm) was detected at 24, 48, 72
and 96 h using a microplate reader (Epoch; BioTek Instruments,
Inc., Winooski, VT, USA).
Flow cytometric analysis
For flow cytometric analysis, siRNA transfected
HL-60 and SKNO-1 cells were collected following 48 h transfection.
HL-60 and SKNO-1 cells were cultured in RPMI-1640 medium (Gibco;
Thermo Fisher Scientific, Inc.) supplemented with 10% fetal bovine
serum (Gibco; Thermo Fisher Scientific, Inc.). For cell cycle
analysis, cells were fixed overnight in 70% cold ethanol, then
resuspended in 20 mg/ml propidium iodide (PI; BD Biosciences,
Franklin Lakes, NJ, USA). For apoptosis analysis, cells were
stained with Annexin V-fluorescein isothiocyanate (FITC) and PI.
Cells were then washed twice with ice-cold PBS and put into binding
buffer (5 µl Annexin V-FITC and 5 µl PI) for a 30 min incubation. A
FACSCalibur™ flow cytometer (BD Biosciences) was used for analyzing
the cell cycle or apoptosis. All experiments were independently
repeated at least three times.
Statistical analysis
All data are expressed as mean ± standard deviation.
Differences between groups were analyzed using an
independent-samples t-test. All P-values were two-sided and
P<0.05 was considered to indicate a statistically significant
difference.
Results
Expression of ZFAS1 in AML cell
lines
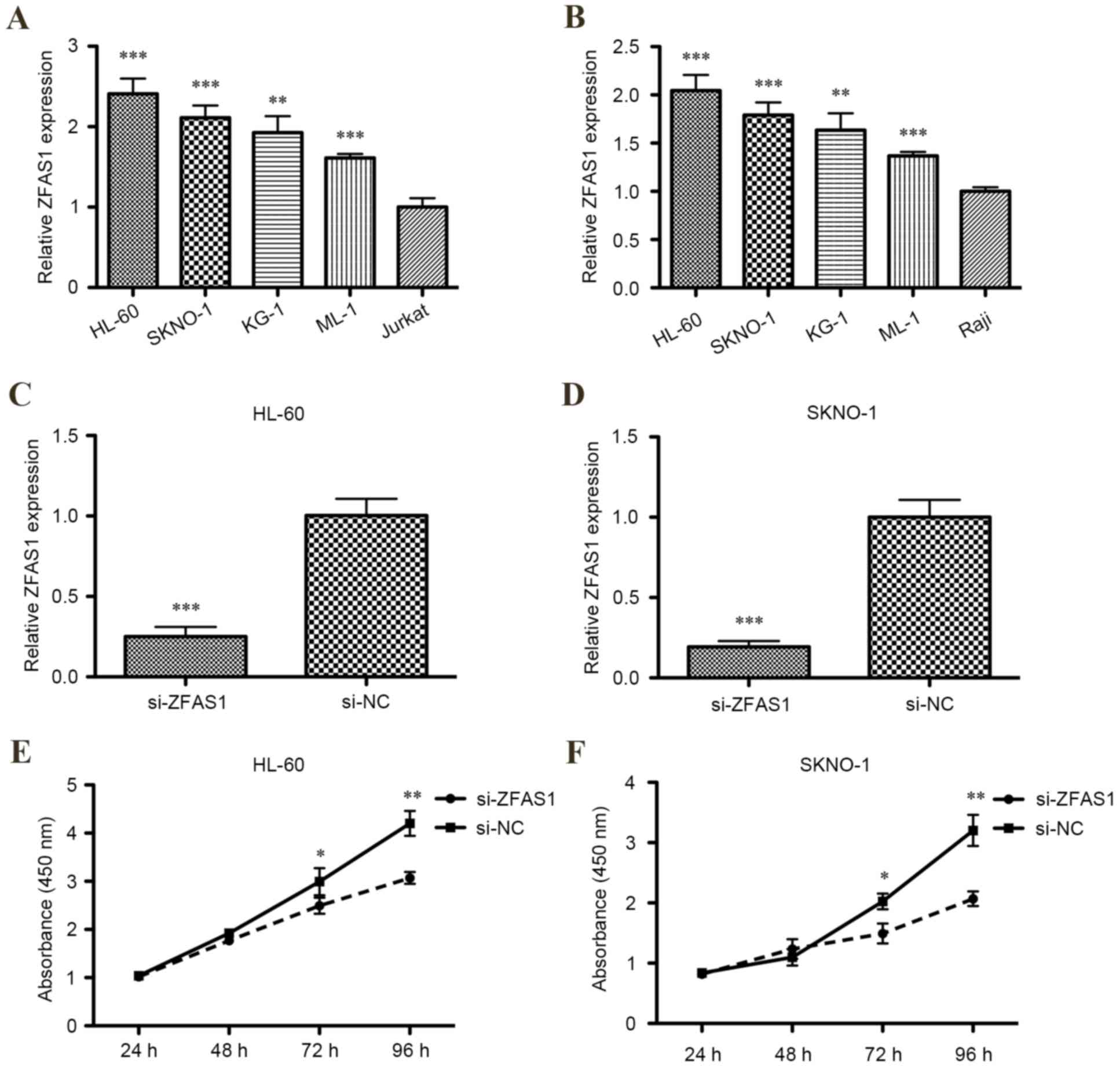
To determine whether ZFAS1 expression was
dysregulated in AML cell lines, levels of ZFAS1 mRNA expression in
HL-60, KG-1, ML-1, SKNO-1, Jurkat and Raji cell lines were measured
using RT-qPCR. Relative levels of ZFAS1 mRNA were significantly
increased in all human four AML cell lines compared with the T
lymphocytic leukemia cell line or Burkitt's lymphoma cell line
(P<0.001 for HL-60, SKNO-1 and ML-1; P<0.01 for KG-1;
Fig. 1A and B).
Efficiency of siRNA in downregulating
ZFAS1 expression in AML cells
According to the aforementioned result, two AML cell
lines (HL-60 and SKNO-1) were selected for ZFAS1 knockdown by
transfection of siRNA, as expression of ZFAS1 was markedly
increased in these cell lines compared with the KG-1 and ML-1 cell
lines. RT-qPCR demonstrated that ZFAS1 was significantly decreased
in HL-60 and SKNO-1 cells transfected with ZFAS1 siRNA compared
with the respective negative control (P<0.001; Fig. 1C and D).
Downregulating ZFAS1 expression
inhibits AML cell proliferation
The CCK-8 assay determined that the proliferation of
HL-60 and SKNO-1 cells were both significantly inhibited following
transfection with ZFAS1 siRNA after 72 h (both P<0.05; Fig. 1E) and 96 h (both P<0.01; Fig. 1F). Additionally, flow cytometric
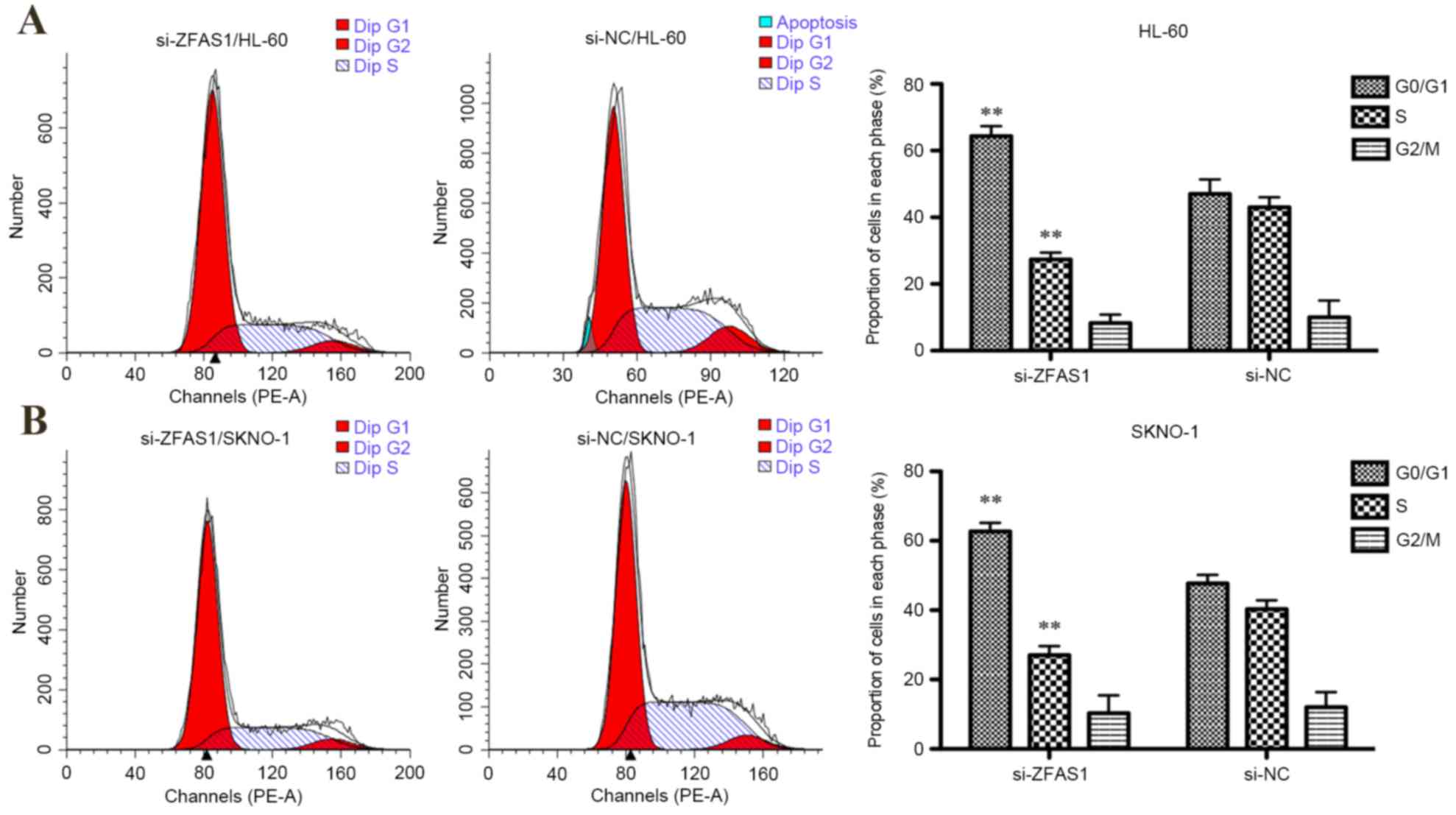
analysis of cell cycle distribution indicated that ZFAS1 knockdown
significantly increased the percentage of G0/G1 phase HL-60 (from
47–64%; P<0.01; Fig. 2A) and
SKNO-1 cells (from 48–63%; P<0.01; Fig. 2B), and decreased the percentage of
S-phase HL-60 (from 27–43%; P<0.01; Fig. 2A) and SKNO-1 cells (from 27–40%;
P<0.01; Fig. 2B). In summary,
these results indicate that ZFAS1 promotes AML cell growth in
vitro.
Downregulating ZFAS1 expression
promotes AML cell apoptosis
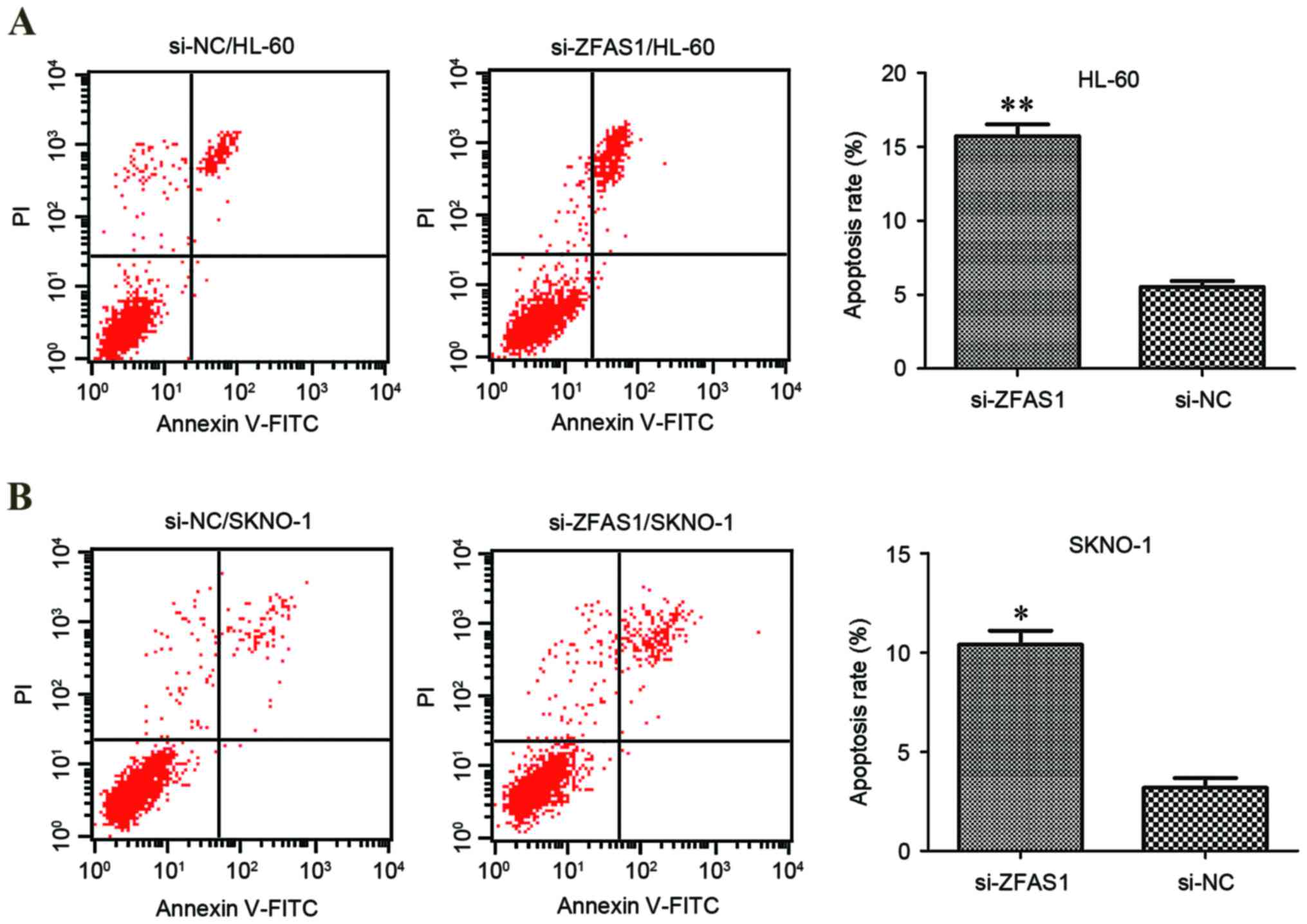
To elucidate the function of ZFAS1 in the regulation
of apoptosis in AML cells, flow cytometric analysis of HL-60 and
SKNO-1 cell lines was performed. Compared with the respective
negative control, ZFAS1 knockdown significantly increased the
percentage of apoptotic AML HL-60 (5.5±0.6% vs. 15.7±0.9%;
P<0.01; Fig. 3A) and SKNO-1 cells
(3.2±0.7% vs. 10.4±0.8%; P<0.05; Fig.
3B). These results suggest that ZFAS1 knockdown promotes AML
cell apoptosis, which may contribute to the inhibition of AML
progression.
Discussion
LncRNAs are a newly identified type of non-coding
RNA that have been reported to be dysregulated in a number of
different diseases, including carcinoma (18,19). The
importance of lncRNAs in the development of cancer may be
associated with their ability to influence cellular function
through different mechanisms, such as proliferation, apoptosis and
differentiation (20–22). In present study, CCK-8 assay and cell
cycle distribution analysis demonstrated that ZFAS1 was able to
induce AML cell proliferation. Furthermore, the apoptosis assay
indicated that ZFAS1 inhibited apoptosis in AML cells. These
results suggest that ZFAS1 knockdown suppresses AML HL-60 and
SKNO-1 cell proliferation and survival, and that ZFAS1 may act as
an oncogenic lncRNA.
It has previously been determined that ZFAS1 is
downregulated in breast cancer and that it acts as a putative tumor
suppressor (11). Previous studies
have reported that ZFAS1 is overexpressed in hepatocellular and
colorectal cancer tissues and cell lines (12,13). In
the current study, it was demonstrated that ZFAS1 has an oncogenic
role in AML cell lines. However, the underlying mechanism of ZFAS1
in the progression of AML remains unclear.
In recent years, a number of articles have reported
that lncRNAs have the ability to target and regulate microRNAs
(miRs) (23,24). It is widely acknowledged that miRNAs
regulate the expression of multiple target genes that encode
proteins, which may lead to biological alterations in function
(25,26). Li et al (12) identified that ZFAS1 functions as an
oncogene in hepatocellular carcinoma progression by binding miR-150
and abrogating its tumor-suppressive function. Furthermore, Li
et al (12) indicated that
miR-150 was able to repress hepatocellular carcinoma cell invasion
by inhibiting ZEB1 and the matrix metalloproteinases (MMPs), MMP14
and MMP16. It was also demonstrated that other lncRNAs are
associated with cell growth and apoptosis regulation by regulating
multiple target genes, including phosphoinositide-3-kinase, Akt and
extracellular signal-regulated kinases (27,28).
Although it has been demonstrated that lncRNAs influence epigenetic
gene regulation (6,7), proliferation (7), apoptosis (6,7) and
prognosis (8) in solid tumors, few
studies have identified their effects on AML. Garzon et al
(29) assessed whether a lncRNA
expression profile was associated with clinical features, molecular
abnormalities and outcome in older patients with cytogenetically
normal AML and revealed that patients with an unfavorable lncRNA
score had a shorter overall survival and disease-free survival rate
than those with a favorable lncRNA score. These results are
consistent with those from the present study.
In conclusion, the present study demonstrated that
lncRNA ZFAS1 promoted the proliferation and suppressed the
apoptotic rate of AML cells. Additionally, the results of the
present study indicated that ZFAS1 may act as an oncogenic
lncRNA.
Acknowledgements
The present study was supported by funding from the
Departments of Hematology and Medical Oncology, Third Affiliated
Hospital of Wenzhou Medical University, Ruian, China). The funding
sources had no role in the study design, in the collection,
analysis and interpretation of data, writing of the manuscript or
in the decision to submit the manuscript for publication.
References
|
1
|
Sanz GF, Sanz MA, Vallespí T, Cañizo MC,
Torrabadella M, García S, Irriguible D and San Miguel JF: Two
regression models and a scoring system for predicting survival and
planning treatment in myelodysplastic syndromes: A multivariate
analysis of prognostic factors in 370 patients. Blood. 74:395–408.
1989.PubMed/NCBI
|
|
2
|
Leith CP, Kopecky KJ, Godwin J, McConnell
T, Slovak ML, Chen IM, Head DR, Appelbaum FR and Willman CL: Acute
myeloid leukemia in the elderly: Assessment of multidrug resistance
(MDR1) and cytogenetics distinguishes biologic subgroups with
remarkably distinct responses to standard chemotherapy. A Southwest
Oncology Group study. Blood. 89:3323–3329. 1997.PubMed/NCBI
|
|
3
|
Appelbaum FR, Baer MR, Carabasi MH, Coutre
SE, Erba HP, Estey E, Glenn MJ, Kraut EH, Maslak P, Millenson M, et
al: NCCN practice guidelines for acute myelogenous leukemia.
Oncology (Williston Park). 14:53–61. 2000.PubMed/NCBI
|
|
4
|
Hope KJ, Jin L and Dick JE: Human acute
myeloid leukemia stem cells. Arch Med Res. 34:507–514. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Ding J, Lu B, Wang J, Wang J, Shi Y, Lian
Y, Zhu Y, Wang J, Fan Y, Wang Z, et al: Long non-coding RNA
Loc554202 induces apoptosis in colorectal cancer cells via the
caspase cleavage cascades. J Exp Clin Cancer Res. 34:1002015.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Mercer TR, Dinger ME and Mattick JS: Long
non-coding RNAs: Insights into functions. Nat Rev Genet.
10:155–159. 2009. View
Article : Google Scholar : PubMed/NCBI
|
|
7
|
Fachel AA, Tahira AC, Vilella-Arias SA,
Maracaja-Coutinho V, Gimba ER, Vignal GM, Campos FS, Reis EM and
Verjovski-Almeida S: Expression analysis and in silico
characterization of intronic long noncoding RNAs in renal cell
carcinoma: Emerging functional associations. Mol Cancer.
12:1402013. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Kim K, Jutooru I, Chadalapaka G, Johnson
G, Frank J, Burghardt R, Kim S and Safe S: HOTAIR is a negative
prognostic factor and exhibits pro-oncogenic activity in pancreatic
cancer. Oncogene. 32:1616–1625. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Ellinger J, Alam J, Rothenburg J, Deng M,
Schmidt D, Syring I, Miersch H, Perner S and Müller SC: The long
non-coding RNA lnc-ZNF180-2 is a prognostic biomarker in patients
with clear cell renal cell carcinoma. Am J Cancer Res. 5:2799–2807.
2015.PubMed/NCBI
|
|
10
|
Chen X, Liu L and Zhu W: Up-regulation of
long non-coding RNA CCAT2 correlates with tumor metastasis and poor
prognosis in cervical squamous cell cancer patients. Int J Clin Exp
Pathol. 8:13261–13266. 2015.PubMed/NCBI
|
|
11
|
Askarian-Amiri ME, Crawford J, French JD,
Smart CE, Smith MA, Clark MB, Ru K, Mercer TR, Thompson ER, Lakhani
SR, et al: SNORD-host RNA Zfas1 is a regulator of mammary
development and a potential marker for breast cancer. RNA.
17:878–891. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Li T, Xie J, Shen C, Cheng D, Shi Y, Wu Z,
Deng X, Chen H, Shen B, Peng C, et al: Amplification of long
noncoding RNA ZFAS1 promotes metastasis in hepatocellular
carcinoma. Cancer Res. 75:3181–3191. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Thorenoor N, Faltejskova-Vychytilova P,
Hombach S, Mlcochova J, Kretz M, Svoboda M and Slaby O: Long
non-coding RNA ZFAS1 interacts with CDK1 and is involved in
p53-dependent cell cycle control and apoptosis in colorectal
cancer. Oncotarget. 7:622–637. 2016.PubMed/NCBI
|
|
14
|
Xing CY, Hu XQ, Xie FY, Yu ZJ, Li HY, Bin
Zhou, Wu JB, Tang LY and Gao SM: Long non-coding RNA HOTAIR
modulates c-KIT expression through sponging miR-193a in acute
myeloid leukemia. FEBS Lett. 589:1981–1987. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Hirano T, Yoshikawa R, Harada H, Harada Y,
Ishida A and Yamazaki T: Long noncoding RNA CCDC26, controls
myeloid leukemia cell growth through regulation of KIT expression.
Mol Cancer. 14:902015. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Alvarez-Dominguez JR, Hu W, Gromatzky AA
and Lodish HF: Long noncoding RNAs during normal and malignant
hematopoiesis. Int J Hematol. 99:531–541. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) Method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Yang F, Huo XS, Yuan SX, Zhang L, Zhou WP,
Wang F and Sun SH: Repression of the long noncoding RNA-LET by
histone deacetylase 3 contributes to hypoxia-mediated metastasis.
Mol Cell. 49:1083–1096. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Gupta RA, Shah N, Wang KC, Kim J, Horlings
HM, Wong DJ, Tsai MC, Hung T, Argani P, Rinn JL, et al: Long
non-coding RNA HOTAIR reprograms chromatin state to promote cancer
metastasis. Nature. 464:1071–1076. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Ma MZ, Li CX, Zhang Y, Weng MZ, Zhang MD,
Qin YY, Gong W and Quan ZW: Long non-coding RNA HOTAIR, a c-Myc
activated driver of malignancy, negatively regulates miRNA-130a in
gallbladder cancer. Mol Cancer. 13:1562014. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Wu XS, Wang XA, Wu WG, Hu YP, Li ML, Ding
Q, Weng H, Shu YJ, Liu TY, Jiang L, et al: MALAT1 promotes the
proliferation and metastasis of gallbladder cancer cells by
activating the ERK/MAPK pathway. Cancer Biol Ther. 15:806–814.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Chen CL, Tseng YW, Wu JC, Chen GY, Lin KC,
Hwang SM and Hu YC: Suppression of hepatocellular carcinoma by
baculovirus-mediated expression of long non-coding RNA PTENP1 and
MicroRNA regulation. Biomaterials. 44:71–81. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Nie W, Ge HJ, Yang XQ, Sun X, Huang H, Tao
X, Chen WS and Li B: LncRNA-UCA1 exerts oncogenic functions in
non-small cell lung cancer by targeting miR-193a-3p. Cancer Lett.
371:99–106. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Kang Y, Song J, Kim D, Ahn C, Park S, Chun
CH and Jin EJ: PCGEM1 stimulates proliferation of osteoarthritic
synoviocytes by acting as a sponge for miR-770. J Orthop Res.
34:412–418. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Han F, Wu Y and Jiang W: MicroRNA-18a
decreases choroidal endothelial cell proliferation and migration by
inhibiting HIF1A expression. Med Sci Monit. 21:1642–1647. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Yan G, Li B, Xin X, Xu M, Ji G and Yu H:
MicroRNA-34a promotes hepatic stellate cell activation via
targeting ACSL1. Med Sci Monit. 21:3008–3015. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Redis RS, Sieuwerts AM, Look MP, Tudoran
O, Ivan C, Spizzo R, Zhang X, De Weerd V, Shimizu M, Ling H, et al:
CCAT2, a novel long non-coding RNA in breast cancer: Expression
study and clinical correlations. Oncotarget. 4:1748–1762. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Cai Y, He J and Zhang D: Long noncoding
RNA CCAT2 promotes breast tumor growth by regulating the Wnt
signaling pathway. Onco Targets Ther. 8:2657–2664. 2015.PubMed/NCBI
|
|
29
|
Garzon R, Volinia S, Papaioannou D,
Nicolet D, Kohlschmidt J, Yan PS, Mrózek K, Bucci D, Carroll AJ,
Baer MR, et al: Expression and prognostic impact of lncRNAs in
acute myeloid leukemia. Proc Natl Acad Sci U S A. 111:pp.
18679–18684. 2014; View Article : Google Scholar : PubMed/NCBI
|