|
1
|
Zhu Z, Ding J, sky Shankowsky HA and
Tredget EE: The molecular mechanism of hypertrophic scar. J Cell
Commun Signal. 7:239–252. 2013.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Aarabi S, Longaker MT and Gurtner GC:
Hypertrophic scar formation following burns and trauma: New
approaches to treatment. PLoS Med. 4(e234)2007.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Tyack Z, Simons M, Spinks A and Wasiak J:
A systematic review of the quality of burn scar rating scales for
clinical and research use. Burns. 38:6–18. 2012.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Penn JW, Grobbelaar AO and Rolfe KJ: The
role of the TGF-β family in wound healing, burns and scarring: A
review. Int J Burns Trauma. 2:18–28. 2012.PubMed/NCBI
|
|
5
|
Niessen FB, Schalkwijk J, Vos H and Timens
W: Hypertrophic scar formation is associated with an increased
number of epidermal Langerhans cells. J Pathol. 202:121–129.
2004.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Morris DE, Wu L, Zhao LL, Bolton L, Roth
SI, Ladin DA and Mustoe TA: Acute and chronic animal models for
excessive dermal scarring: Quantitative studies. Plast Reconstr
Surg. 100:674–681. 1997.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Ding J, Hori K, Zhang R, Marcoux Y,
Honardoust D, Shankowsky HA and Tredget EE: Stromal cell-derived
factor 1 (SDF-1) and its receptor CXCR4 in the formation of
postburn hypertrophic scar (HTS). Wound Repair Regen. 19:568–578.
2011.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Yates CC, Krishna P, Whaley D, Bodnar R,
Turner T and Wells A: Lack of CXC chemokine receptor 3 signaling
leads to hypertrophic and hypercellular scarring. Am J Pathol.
176:1743–1755. 2010.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Wang J, Hori K, Ding J, Huang Y, Kwan P,
Ladak A and Tredget EE: Toll-like receptors expressed by dermal
fibroblasts contribute to hypertrophic scarring. J Cell Physiol.
226:1265–1273. 2011.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Eto H, Suga H, Aoi N, Kato H, Doi K, Kuno
S, Tabata Y and Yoshimura K: Therapeutic potential of fibroblast
growth factor-2 for hypertrophic scars: Upregulation of MMP-1 and
HGF expression. Lab Invest. 92:214–223. 2012.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Ulrich D, Noah EM, von Heimburg D and
Pallua N: TIMP-1, MMP-2, MMP-9, and PIIINP as serum markers for
skin fibrosis in patients following severe burn trauma. Plast
Reconstr Surg. 111:1423–1431. 2003.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Zhu S, Qing T, Zheng Y, Jin L and Shi L:
Advances in single-cell RNA sequencing and its applications in
cancer research. Oncotarget. 8:53763–53779. 2017.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Liu YJ, Zhang F, Liu HD and Sun X: The
application of next-generation sequencing techniques in studying
transcriptional regulation in embryonic stem cells. Yi Chuan.
39:717–725. 2017.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Sarig O and Sprecher E: The molecular
revolution in cutaneous biology: Era of next-generation sequencing.
J Invest Dermatol. 137:e79–e82. 2017.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Saulis AS, Mogford JH and Mustoe TA:
Effect of Mederma on hypertrophic scarring in the rabbit ear model.
Plast Reconstr Surg. 110:177–183. 2002.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Syed F, Ahmadi E, Iqbal SA, Singh S,
McGrouther DA and Bayat A: Fibroblasts from the growing margin of
keloid scars produce higher levels of collagen I and III compared
with intralesional and extralesional sites: Clinical implications
for lesional site-directed therapy. Br J Dermatol. 164:83–96.
2011.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
18
|
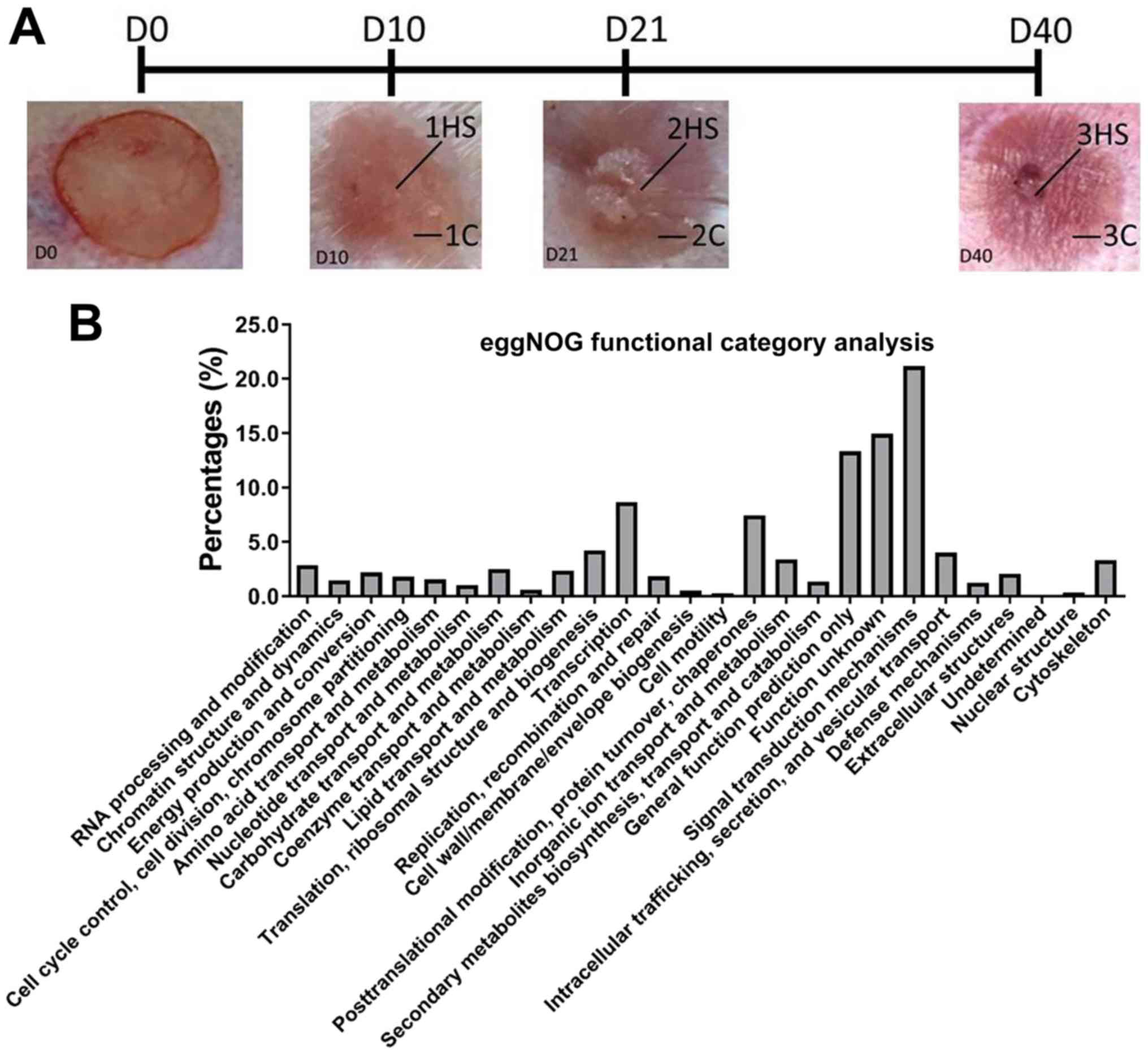
Jensen LJ, Julien P, Kuhn M, von Mering C,
Muller J, Doerks T and Bork P: eggNOG: Automated construction and
annotation of orthologous groups of genes. Nucleic Acids Res.
36:D250–D254. 2008.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Walmsley GG, Rinkevich Y and Hu MS:
Identification and targeted inhibition of a fibroblast lineage
responsible for skin scarring and cancer stroma. J Am Coll Surg.
219:S84–S85. 2014.
|
|
20
|
Scott PG, Dodd CM, Tredget EE, Ghahary A
and Rahemtulla F: Immunohistochemical localization of the
proteoglycans decorin, biglycan and versican and transforming
growth factor-beta in human post-burn hypertrophic and mature
scars. Histopathology. 26:423–431. 1995.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Aarabi S, Bhatt KA, Shi Y, Paterno J,
Chang EI, Loh SA, Holmes JW, Longaker MT, Yee H and Gurtner GC:
Mechanical load initiates hypertrophic scar formation through
decreased cellular apoptosis. FASEB J. 21:3250–3261.
2007.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Kawano Y and Kypta R: Secreted antagonists
of the Wnt signaling pathway. J Cell Sci. 116:2627–2634.
2003.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Melkonyan HS, Chang WC, Shapiro JP,
Mahadevappa M, Fitzpatrick PA, Kiefer MC, Tomei LD and Umansky SR:
SARPs: A family of secreted apoptosis-related proteins. Proc Natl
Acad Sci USA. 94:13636–13641. 1997.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Maganga R, Giles N, Adcroft K, Unni A,
Keeney D, Wood F, Fear M and Dharmarajan A: Secreted frizzled
related protein-4 (sFRP4) promotes epidermal differentiation and
apoptosis. Biochem Biophys Res Commun. 377:606–611. 2008.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Matsushima K, Suyama T, Takenaka C,
Nishishita N, Ikeda K, Ikada Y, Sawa Y, Jakt LM, Mori H and
Kawamata S: Secreted frizzled related protein 4 reduces fibrosis
scar size and ameliorates cardiac function after ischemic injury.
Tissue Eng Part A. 16:3329–3341. 2010.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Muley A, Majumder S, Kolluru GK, Parkinson
S, Viola H, Hool L, Arfuso F, Ganss R, Dharmarajan A and Chatterjee
S: Secreted frizzled-related protein 4: An angiogenesis inhibitor.
Am J Pathol. 176:1505–1516. 2010.PubMed/NCBI View Article : Google Scholar
|