Introduction
Bladder cancer is the most common malignancy of the
urinary system worldwide, with >76,960 new cases and 16,390
mortalities estimated in 2016(1).
Bladder cancer is divided into non-muscle-invasive bladder cancer,
characterized by a high recurrence rate (70%), and muscle-invasive
bladder cancer, which has a <50% of 5-year overall survival
according to biological characteristics (2). Thus, it is important to identify the
molecular mechanism of cellular proliferation in bladder cancer to
facilitate the development of a novel and more effective
therapy.
As a class of non-coding RNA without the capacity of
coding proteins, long non-coding RNAs (lncRNA) are >200
nucleotides in length (3,4). Previous studies have reported that
lncRNA are involved in epigenetic, transcriptional and
post-transcriptional modulation in various biological processes,
including proliferation, differentiation, migration and apoptosis
(5,6). lncRNA small nucleolar RNA host gene 1
(SNHG1) located at 11q12.3 locus, functions as an oncogene in a
number of cancer types. For instance, Cui et al (7) revealed that high SNHG1 expression in
non-small cell lung cancer (NSCLC) was significantly correlated
with larger tumor size, advanced TNM stage, lymph node metastasis
and poor overall survival. Moreover, Hu et al (8) reported that knockdown of SNHG1
suppressed gastric cancer cell proliferation both in vitro
and in vivo. Liu et al (9) also found that SNHG1 inhibition
significantly inhibited cervical cancer cell proliferation,
migration and invasion. However, the relationship between SNHG1
expression and bladder cancer, as well as the underlying molecular
mechanisms of the oncogenic functions of SNHG1, remains unknown and
require further investigation. Furthermore, identifying the
downstream targets of SNHG1 will elucidate its critical role in
bladder cancer progression.
The PI3K/AKT signaling pathway regulates multiple
biological processes, including cell proliferation and apoptosis
(10). In most cases, inappropriate
activation of PI3K/AKT is speculated to induce tumor formation
(11). Increased SNHG1 expression
has also been reported to promote the activity of the PI3K/AKT
signaling pathway (12). However,
the molecular mechanisms underlying this phenomenon are not fully
understood.
The aim of the present study was to elucidate the
key functions of SNHG1 in the proliferation, apoptosis, migration
and invasion of bladder cancer cells in vitro, in addition
to investigating the signaling pathways that are possibly
implicated in this process.
Materials and methods
Tissues
Tumor tissues and homologous adjacent healthy
tissues were donated by 60 patients with bladder cancer, who
underwent surgery between July 2016 and January 2018 during their
hospitalization in Shaanxi Provincial People's Hospital (Xi'an,
China), and were stored at -80˚C prior to RNA isolation. The
inclusion criteria were as follows: i) Patients diagnosed by
pathological biopsies; ii) patients at stage I or II based on the
TNM staging system (13); iii)
patients with complete medical record; and iv) patients and their
families who were willing to participate. The exclusion criteria
were as follows: i) Patients who were treated within 3 months
before admission; ii) patients who had other diseases, such as
chronic diseases and metabolic diseases; and iii) patients who were
not willing to donate plasma samples. The 60 patients with bladder
cancer included 34 males and 26 females (age range, 27-67 years;
mean age, 46.4±5.1 years). Informed consent was obtained and all
experimental procedures were approved by the Human Ethics Committee
of Shaanxi Provincial People's Hospital.
Cell culture, transfection and
treatment
In total, four human bladder cancer cell lines (T24,
SW780, J82 and RT4) and normal bladder epithelial HCV-29 cells were
purchased from the American Type Culture Collection (ATCC). Cells
were incubated in DMEM supplemented with 10% FBS, 100 U/ml
penicillin and 100 µg/ml streptomycin (all from Invitrogen; Thermo
Fisher Scientific, Inc.) in a humidified atmosphere with 5%
CO2 at 37˚C.
For knockdown of SNHG1 in SW780 cells, the sequences
of short hairpin (sh)RNA targeting for SNHG1
(5'-CAGCAGTTGAGGGTTTGCTGTGTAT-3') were designed by Shanghai
GenePharma Co., Ltd. For overexpression, the full-length SNHG1 was
cloned into a pcDNA3.1 vector (Shanghai GenePharma Co., Ltd.) to
overexpress SNHG1 in RT4 cells, and the primer sequences were as
follows: SNHG1 forward, 5'-GGGGTACCGTTCTCATTTTTCTACTGCTCGTG-3' and
reverse, 5'-CGGGATCCATGTAATCAATCATTTTATTATTTTCATC-3'. The empty
vector and scrambled shRNA for SNGH1 were used as negative
controls. Cell transfections were conducted using shRNAs and
plasmids, both at a final concentration of 100 nM, using
Lipofectamine® 2000 reagent (Invitrogen; Thermo Fisher
Scientific, Inc.) following the manufacturer's instructions. Cells
were treated with the PI3K agonist 740Y-P (50 µM; Sigma-Aldrich;
Merck KGaA) or the PI3K inhibitor LY294002 (50 µM; Sigma-Aldrich;
Merck KGaA) for 24 h at 37˚C, 48 h after transfection.
Reverse transcription-quantitative PCR
(RT-qPCR)
Total RNA was extracted from tissues or T24, SW780,
J82, RT4, and HCV-29 cells using the TRIzol® Reagent
(Thermo Fisher Scientific, Inc.) and then converted into cDNA by
reverse transcription using PrimeScript™ RT Master Mix (cat. no.
RR036Q; Takara Biotechnology Co., Ltd.), according to the
manufacturer's protocol. qPCR was performed using Path-ID™
Multiplex One-Step RT-PCR Kit (cat. no. 4442136; Applied
Biosystems; Thermo Fisher Scientific, Inc.) according to
manufacturer's protocol in an ABI 7500 RT PCR system (Applied
Biosystems; Thermo Fisher Scientific, Inc.) to determine the
relative quantification of SNHG1 expression. A total of 1 µg total
RNA was reversely transcribed using oligo(dT) primer at 42˚C for 1
h and 2 µl the reverse transcription reaction mix was amplified by
PCR with denaturation at 95˚C for 2 min, followed by 50 cycles of
95˚C for 30 sec, 55˚C for 30 sec and 72˚C for 1 min. The
thermocycling conditions were as follows: Initial denaturation at
95˚C for 30 sec, followed by 40 cycles of 95˚C for 5 sec and 60˚C
for 30 sec. Relative expression was calculated with the
2-ΔΔCq method as previously
described (14), where GAPDH was
used as the internal control. The primer sequences were as follows:
SNHG1 forward, 5'-AGGCTGAAGTTACAGGTC-3' and reverse,
5'-TTGGCTCCCAGTGTCTTA-3'; and GAPDH forward,
5'-GTCAACGGATTTGGTCTGTATT-3' and reverse,
5'-AGTCTTCTGGGTGGCAGTGAT-3'.
Cell Counting Kit-8 (CCK-8) assay
Cell proliferation was assessed using a CCK-8 assay
(Beyotime Institute of Biotechnology). At 0, 24, 48, 72 and 96 h
post-transfection, cells were re-seeded into culture medium on
96-well plates at a density of 5x103 cells/well and
incubated at 37˚C overnight. CCK-8 solution (Beyotime Institute of
Biotechnology) with 10 µl/well was then added and incubated at 37˚C
for another 2 h according to the manufacturer's instruction.
Absorbance was measured with a microplate reader at a wavelength of
450 nm.
Colony formation assay
After 24 h transfection, SW780 and RT4 cells were
re-seeded onto 6-well plates at a density of 500 cells per well and
cultured for 3 days. After washing with PBS, cells were fixed with
4% paraformaldehyde solution at 37˚C for 15 min and stained with
0.1% crystal violet solutions at room temperature for 10 min. The
number of colonies containing >50 cells was analyzed manually
using light microscope (magnification, x200). The experiments were
performed in triplicate.
Apoptosis detection
For the detection of the percentage of early
apoptotic cells, SW780 and RT4 cells were seeded in 24-well plates
(2x105 cells/well) for 48 h, and stained using Annexin
V-FITC apoptosis assay (5 µl; Invitrogen; Thermo Fisher Scientific,
Inc.) at room temperature for 20 min. The stained cells were
analyzed using BD FACSCalibur™ flow cytometer (Beckman Coulter,
Inc.) and FlowJo software (version X; FlowJo LLC). Living cells
were in the lower left quadrant. The upper right quadrant
represented necrotic and late apoptotic cells, whilst the lower
right quadrant represented early apoptotic cells, which were
quantified.
Migration assay
Wound-healing assay was performed to measure the
cell migration capacity of SW780 and RT4 cells. Transfected cells
after 48 h transfection were subsequently cultured in DMEM in a
six-well culture plate at a density of 5x105 cells/well.
When the confluence reached 95%, the cells were washed and the
culture medium was replaced with serum-free DMEM. A scratch was
made through the single cell layer using a 10 µl pipette tip and
the cells were washed again with warmed PBS. After 24 h incubation
in serum-free medium, images of the migrating cells were captured
using a light microscope (magnification, x200; Nikon Corporation).
Representative images after wounding were captured with a light
microscope. The percent closure of the cells into the wound field
was measured using ImageJ software (version 1.46r; National
Institutes of Health). The rate of wound healing was calculated as:
Wound healing rate=(wound width at 0 h-wound width at 24 h)/wound
width at 0 h x100%.
Transwell assay
Cell invasion was analyzed with Transwell chambers
(8.0-µm pore size with polycarbonate membrane; BD Biosciences) that
were pre-coated with Matrigel (BD Biosciences) for 6 h at 37˚C.
Cells resuspended in serum-free medium at a density of
1x104 cells/well were added to the upper chamber, while
500 µl DMEM with 10% FBS was added to the bottom chamber. After 24
h, cells that failed to migrate were removed from the upper part of
the filters by scrubbing with a cotton swab, and the membrane was
fixed with 4% formaldehyde at room temperature for 5 min and
stained with 0.5% crystal violet at room temperature for 10 min.
The invasive cells were counted using light microscope at x200
magnification from 10 different fields of each filter. The invaded
cell rates were calculated using the following formula: Mean test
group invaded cell number/mean blank control invaded cell number
x100%.
Western blotting
Total protein from cultured cells were extracted
using RIPA buffer (Beyotime institute of Biotechnology). Protein
concentration was quantified using a bicinchoninic acid protein
assay kit (Thermo Fisher Scientific, Inc.). Equal amounts of
protein (40 µg) were electrophoresed in a 10% SDS-PAGE and then
electro-blotted onto a PVDF membrane (EMD Millipore). The membranes
were incubated with primary antibodies against GAPDH (cat. no.
5174; 1:1,000 dilution; Cell Signaling Technology, Inc.), PI3K
(cat. no. 17366; 1:1,000 dilution; Cell Signaling Technology,
Inc.), phosphorylated (p)-AKT (cat. no. 4060; 1:1,000 dilution;
Cell Signaling Technology, Inc.), AKT (cat. no. 10176-2-AP; 1:1,000
dilution; ProteinTech Group, Inc.), proliferating cell nuclear
antigen (PCNA; cat. no. 13110; 1:1,000 dilution; Cell Signaling
Technology, Inc.), Bcl-2 (cat. no. 3498; 1:1,000 dilution; Cell
Signaling Technology, Inc.), Bax (cat. no. 5023; 1:1,000 dilution;
Cell Signaling Technology, Inc.), N-Cadherin (cat. no. 22018-1-AP;
1:3,000 dilution; ProteinTech Group, Inc.) and E-Cadherin (cat. no.
20874-1-AP; 1:5,000 dilution; ProteinTech Group, Inc.) at 4˚C
overnight. The membranes were then incubated with horseradish
peroxidase-conjugated secondary antibodies (cat. no. sc-2004;
1:5,000 dilution; Santa Cruz Biotechnology) for 1 h at room
temperature and visualized using the enhanced chemiluminescence
method (Cytiva). GAPDH was used as an internal control. Image-Pro
Plus 7.0 software (Media Cybernetics, Inc.) was used to perform the
densitometric analysis.
Statistical analysis
SPSS 19.0 software (IBM Corp.) was used to perform
statistical analysis. Data are presented as the mean ± standard
deviation from ≤3 independent experiments. Data comparisons were
performed by either unpaired (or paired for tissue samples)
Student's t-test (between two groups) or one-way ANOVA followed by
Tukey's test (among three groups). P<0.05 was considered to
indicate a statistically significant difference.
Results
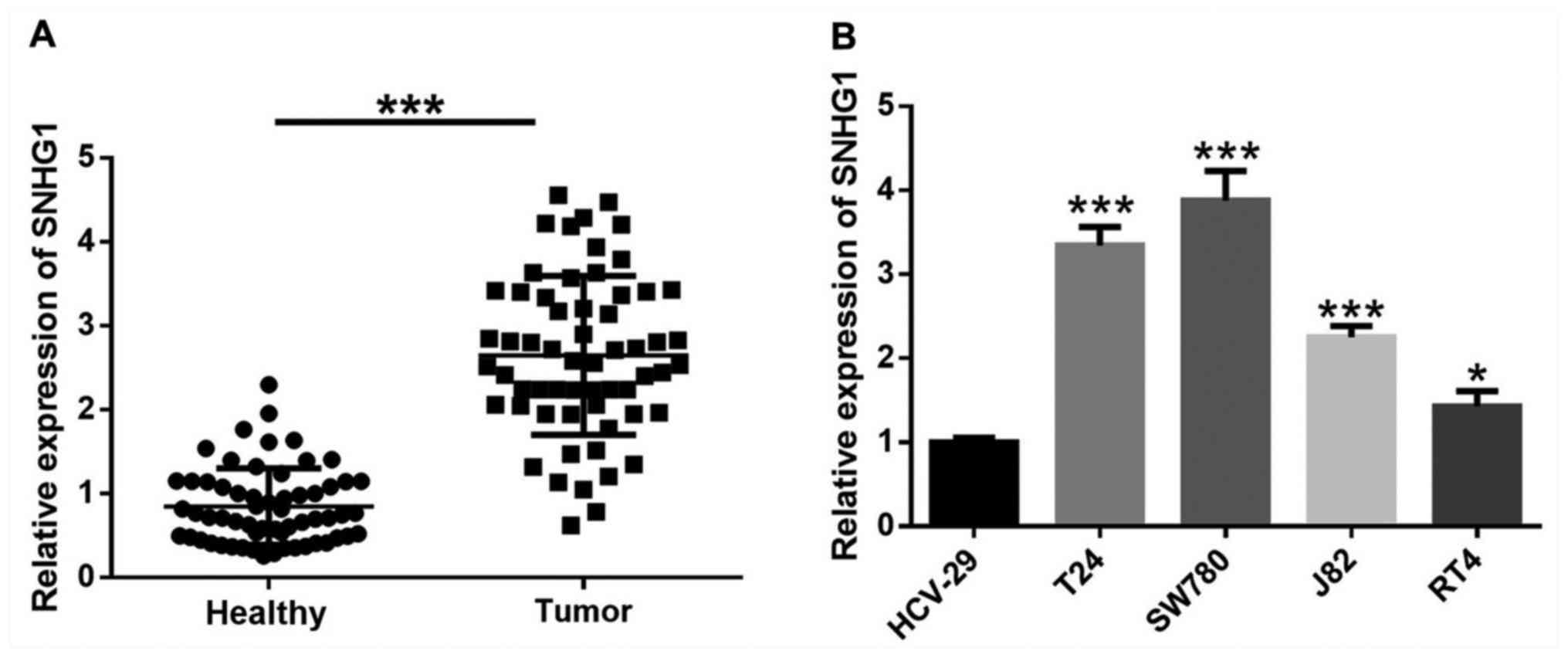
lncRNA SNHG1 expression is
significantly upregulated in bladder cancer tissues and cells
The expression of SNHG1 in bladder cancer tissues
was significantly higher compared with the adjacent healthy tissues
(Fig. 1A). Moreover, SNHG1
expression was significantly increased in four bladder cancer cell
lines, T24, SW780, J82 and RT4 compared with human immortalized
bladder epithelial HCV-29 cells (Fig.
1B).
lncRNA SNHG1 promotes bladder cancer
cell proliferation
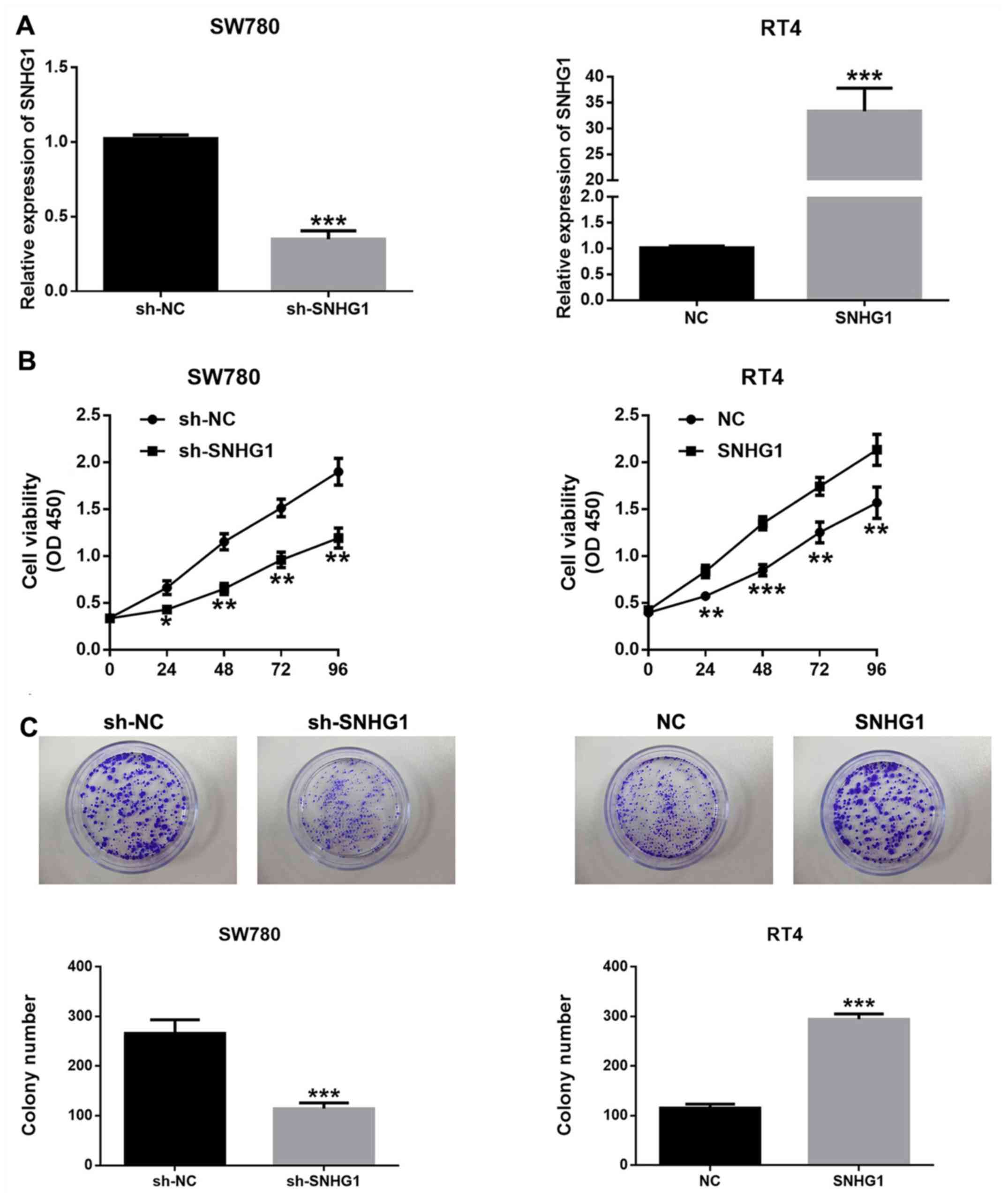
To investigate the association between SNHG1
expression and the proliferative ability of bladder cancer cells,
pcDNA-SNHG1, sh-SNHG1 and the empty plasmids were transfected into
bladder cancer cells. Transfection with sh-SNHG1 significantly
downregulated SNHG1 expression in SW780 cells, while pcDNA-SNHG1
transfection significantly promoted SNHG1 expression in RT4 cells
(Fig. 2A).
The potential biological effects of SNHG1 on the
proliferation of bladder cancer cells were detected by CCK-8 assay
and colony formation assay. It was demonstrated that SNHG1
silencing significantly inhibited the proliferation of SW780 cells,
while SNHG1 overexpression led to the opposite results (Fig. 2B and C). These findings indicated that SNHG1
facilitated the proliferation and enhanced the viability of bladder
cancer cells.
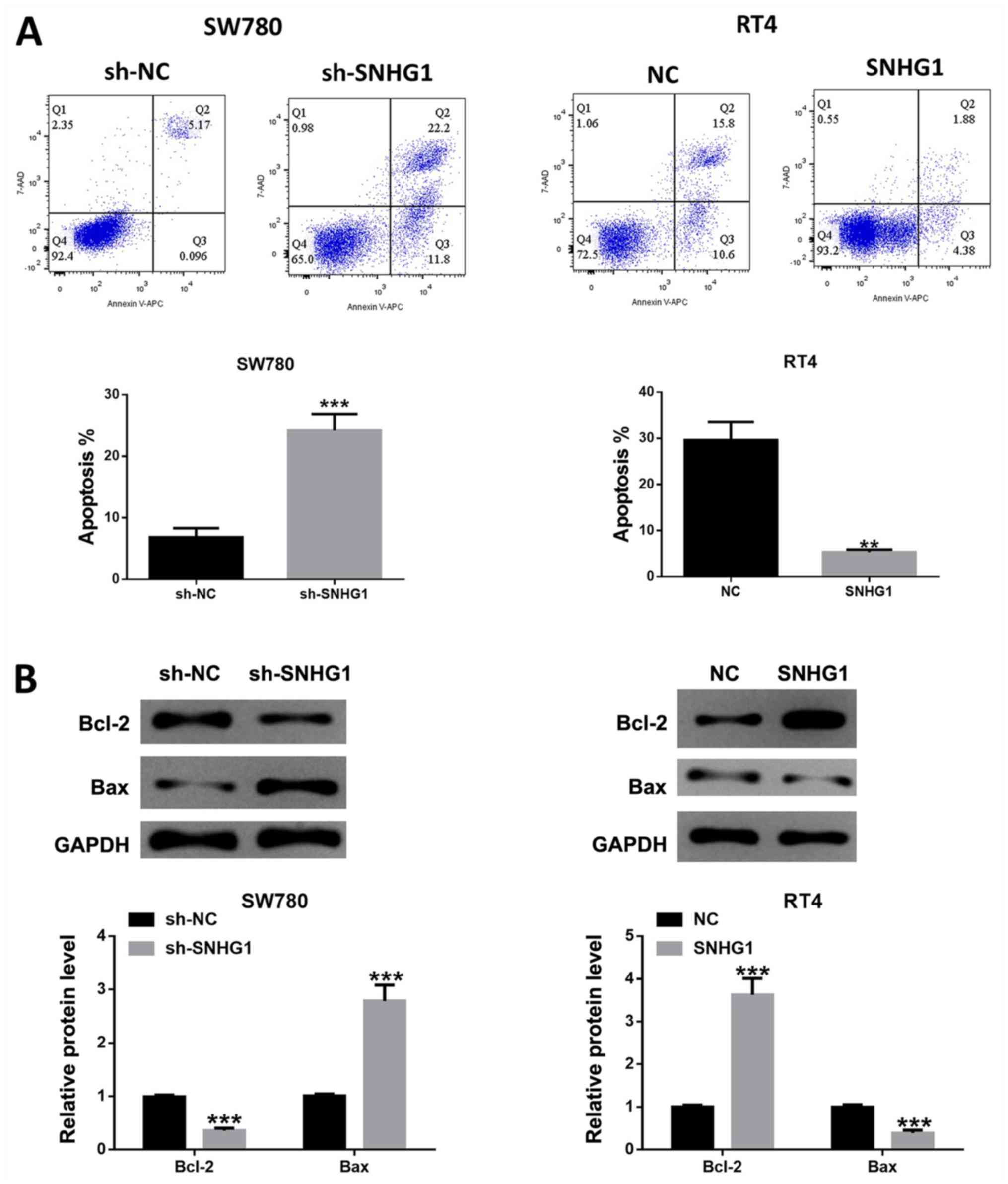
lncRNA SNHG1 inhibits bladder cancer
cell apoptosis
Cell apoptosis analysis results indicated that the
number of apoptosis cells was significantly increased in SW780
cells after SNHG1 silencing, while SNHG1 overexpression repressed
the number of apoptotic cells (Fig.
3A). Furthermore, the western blotting results demonstrated
that the ratio of /Bcl-2/Bax was significantly reduced after SNHG1
knockdown in SW780 cells, while SNHG1 overexpression in RT4 cells
led to the opposite effect (Fig.
3B), which was in line with the aforementioned flow cytometric
analysis results.
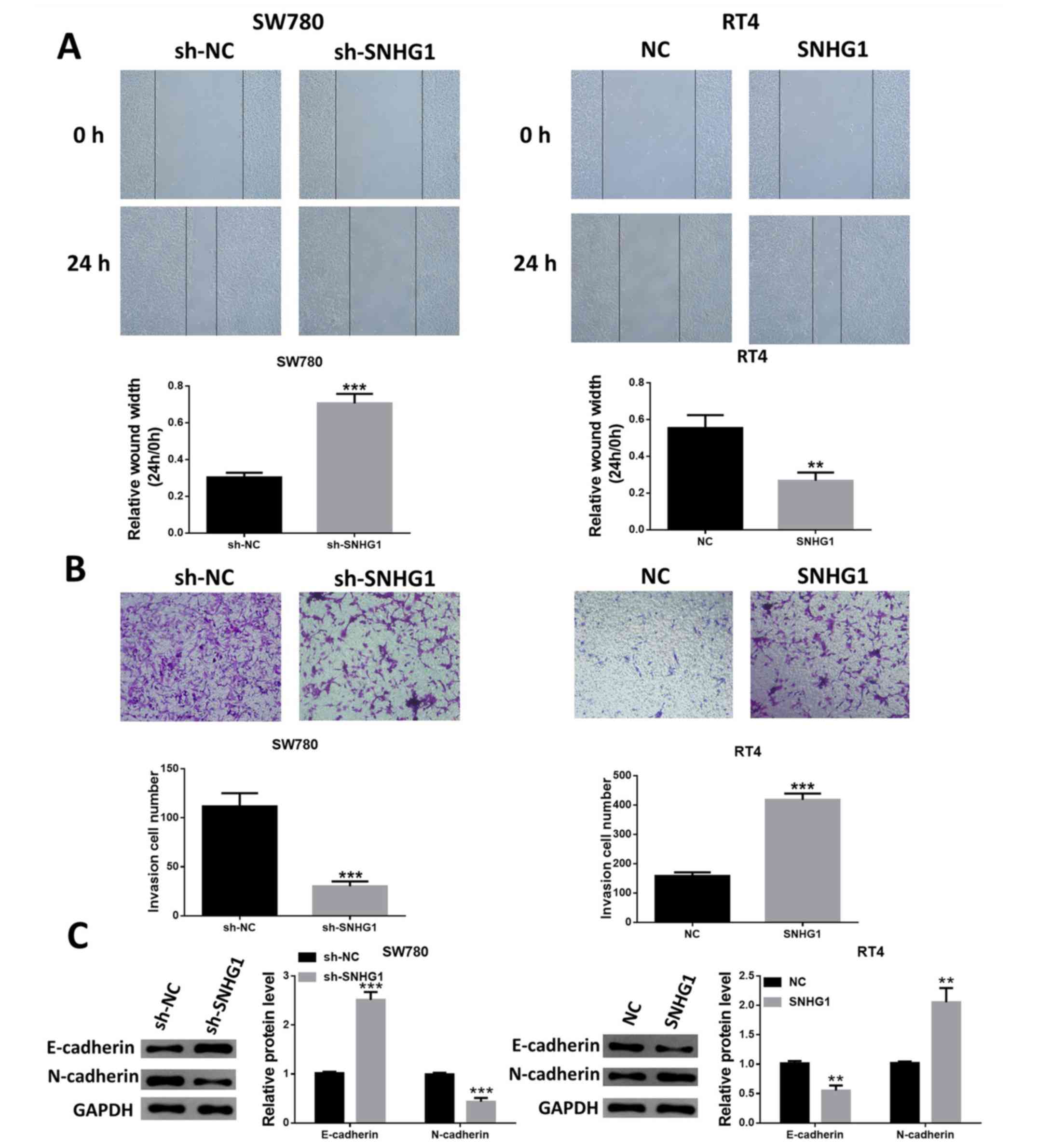
lncRNA SNHG1 increases bladder cancer
cell migration and invasion
The suppression of SNHG1 expression decreased the
migratory capacity of SW780 cells, while the migratory capacity of
RT4 cells was increased in SNHG1-overexpressing cells (Fig. 4A). In addition, the Transwell
invasion assay identified similar results, indicating that SW780
cells with SNHG1 knockdown had reduced cell invasion, while RT4
cells with SNHG1 overexpression exhibited enhanced invasive ability
(Fig. 4B).
The expression level changes of the
epithelial-mesenchymal transition markers E-cadherin and N-cadherin
were also examined. Silencing of SNHG1 upregulated E-cadherin
expression, but resulted in N-cadherin downregulation. Furthermore,
the expression of E-cadherin was significantly reduced, while the
expression of N-cadherin was upregulated following the
overexpression of SNHG1 in RT4 cells (Fig. 4C).
PI3K/AKT axis affects the
carcinogenesis of SNHG1
The addition of 740Y-P, a PI3K activator, reversed
the downregulation of PI3K and AKT phosphorylation levels and the
expression levels of PCNA, Bcl-2 and N-cadherin, which were caused
by silenced SNHG1. However, 740Y-P reduced the enhanced Bax and
E-cadherin protein expression levels induced by silenced SNHG1
(Fig. 5A).
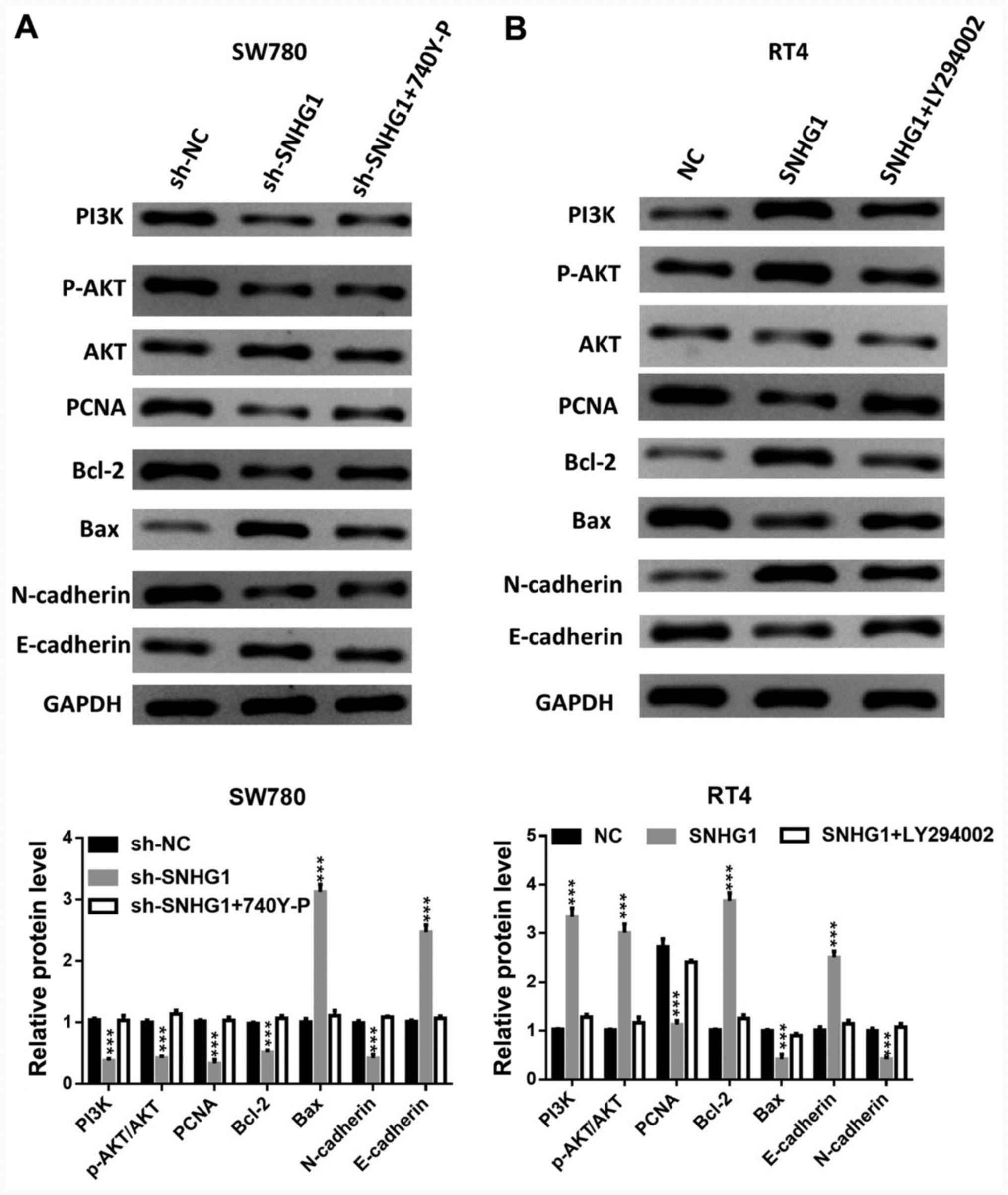 | Figure 5PI3K/AKT axis affects the
carcinogenesis of SNHG1. The phosphorylation levels of PI3K and
AKT, the protein expression levels of AKT, PCNA, Bcl-2, Bax,
N-cadherin and E-cadherin in (A) SW780 cells transfected with
sh-SNHG1 + PI3K activator 740Y-P, and (B) in RT4 cells transfected
with pcDNA-SNHG1 + PI3K inhibitor LY294002.
***P<0.001 vs. sh-NC or NC. NC, negative control; sh,
short hairpin RNA; SNHG1, small nucleolar RNA host gene 1; p-,
phosphorylated; PCNA, proliferating cell nuclear antigen. |
The application of LY294002, a PI3K inhibitor, led
to a partial abrogation of SNHG1 overexpression-induced PI3K and
AKT phosphorylation, Bcl-2 and E-cadherin upregulation, and PCNA,
Bax and N-cadherin downregulation (Fig.
5B).
Discussion
The initiation and development of bladder cancer
involves multiple molecular mechanisms, including abnormal
expression of growth factors, adhesion molecules and angiogenic
factors (15-17)
Therefore, it is important to identify novel biomarker that may be
useful for tumor prevention and therapy. In the present study,
upregulated SNHG1 expression was identified in bladder cancer
tissues and cells. Furthermore, SNHG1 knockdown significantly
suppressed bladder cancer cell proliferation, migration and
invasion, as well as promoted apoptosis. However, the
overexpression of SNHG1 led to an opposite effect via activation of
the PI3K/AKT signaling pathway.
lncRNAs are associated with tumor growth and
metastatic potentials in several types of cancer, including bladder
cancer. For example, Pei et al (18) reported that lncRNA cancer
susceptibility candidate 2 via the inhibition of the Wnt/β-catenin
signaling pathway, inhibited bladder cancer cell proliferation,
migration and invasion, but promoted apoptosis. Wang et al
(19) also showed that high
expression of lncRNA OIP5 antisense RNA 1 (OIP5-AS1) was a poor
predictor of bladder cancer prognosis, and the knockdown of
OIP5-AS1 expression decreased cell viability, as well as promoted
cell-cycle arrest and apoptosis in bladder cancer. Moreover, Gao
et al (20) revealed that
lncRNA zinc finger E-box binding homeobox 2 AS1 increased the
proliferation, migration and invasion, but reduced the apoptosis of
bladder cancer cells as competing endogenous RNA sponges microRNA
(miR)-200b to elevate the expression of fascin-1. Previous studies
have also reported that SNHG1 serves important roles in the
progression of cancer types, including NSCLC (21), glioma (9), pancreatic cancer (22), cholangiocarcinoma (23) and osteosarcoma (24).
Tumor progression is associated with the expression
or modulation of several gene products that control apoptosis and
proliferation. Apoptosis is an important negative growth regulatory
mechanism in tumors (25). In some
malignancies, the apoptotic index may reflect the degree of
carcinogenicity (26). Bcl-2 is a
potent inhibitor of apoptosis and increases proliferation (27). Furthermore, the Bcl-2/Bax ratio is
the critical determinant for the induction or inhibition of
apoptosis (28). PCNA is present in
nuclei throughout the cell cycle and is synthesized in the late
G1 and S phases (29).
Cell migration is a process that is essential during embryonic
development, throughout adult life and in some pathological
conditions (30). Cadherins,
specifically the neural cell adhesion molecule N-cadherin, play an
important role in migration (31).
In cancer, cadherins control the balance between suppression and
promotion of invasion (32). For
example, E-cadherin functions as an invasion suppressor and is
downregulated in most carcinomas, while N-cadherin, as an invasion
promoter, is frequently upregulated (33). The present results suggested that
lncRNA SNHG1 was significantly upregulated in bladder cancer
tissues and cells. Moreover, shRNA-mediated SNHG1 downregulation
impaired cell proliferation, migration and invasion, but
facilitated cell apoptosis; however, SNHG1 overexpression led to
opposite results.
The PI3K/AKT pathway has been reported to exert
important roles in regulating cell cycle and the proliferative,
antiapoptotic, metastatic and invasive abilities of cancer cells.
For example, the overexpression of lncRNA AB073614 significantly
improved the proliferation, migration and invasion of colorectal
cancer cells, and decreased the rates of apoptosis and
G1 phase cell cycle arrest by targeting the PI3K/AKT
signaling pathway (34). It has
also been revealed that miR-802 expression inhibited NSCLC tumor
growth by deactivating the PI3K/AKT/mTOR pathway by targeting
fibroblast growth factor receptor 1(35). In addition, Zhang et al
(36) demonstrated that laminin
subunit β-3 promoted pancreatic ductal adenocarcinoma (PDAC) cell
cycle progression, proliferation, invasion and migration, as well
as inhibited apoptosis by upregulating the PI3K/AKT signaling
pathway. A previous study reported also that SNHG1 acted as an
oncogenic lncRNA, and promoted tumorigenesis in PDAC via the
PI3K/AKT signaling pathway (12).
The present study investigated whether SNHG1 modulated the PI3K/AKT
pathway in bladder cancer cells. It was demonstrated that SNHG1
silencing reduced the phosphorylation levels of PI3K and AKT,
whilst SNHG1 overexpression induced the activation of the PI3K/AKT
axis. The PI3K activator 740Y-P and the inhibitor LY294002 reversed
the previous effects of SNHG1 knockdown in SW780 cells and SNHG1
overexpression in RT4 cells, respectively. Based on these findings,
it was speculated that SNHG1 may promote the tumorigenic process,
at least partly via activating the PI3K/AKT pathway in bladder
cancer.
However, several limitations should considered when
interpreting the present results. For instance, the number of
patients was limited and in vivo experiments were not
performed. Furthermore, the other molecular mechanisms that may be
involved require further investigation.
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
QD conducted the majority of the experiments, wrote
the manuscript and analyzed the data. JC designed the study and
revised the manuscript. Both authors read and approved the final
manuscript.
Ethics approval and consent to
participate
All experimental procedures were approved by the
Human Ethics Committee of Shaanxi Provincial People's Hospital
(Xi'an, China).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2016. CA Cancer J Clin. 66:7–30. 2016.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Ploussard G, Shariat SF, Dragomir A, Kluth
LA, Xylinas E, Masson-Lecomte A, Rieken M, Rink M, Matsumoto K,
Kikuchi E, et al: Conditional survival after radical cystectomy for
bladder cancer: Evidence for a patient changing risk profile over
time. Eur Urol. 66:361–370. 2014.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Batista PJ and Chang HY: Long noncoding
RNAs: Cellular address codes in development and disease. Cell.
152:1298–1307. 2013.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Ling H, Fabbri M and Calin GA: MicroRNAs
and other non-coding RNAs as targets for anticancer drug
development. Nat Rev Drug Discov. 12:847–865. 2013.PubMed/NCBI View
Article : Google Scholar
|
|
5
|
Dykes IM and Emanueli C: Transcriptional
and Post-transcriptional Gene Regulation by Long Non-coding RNA.
Genomics Proteomics Bioinformatics. 15:177–186. 2017.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Carlson HL, Quinn JJ, Yang YW, Thornburg
CK, Chang HY and Stadler HS: lncRNA-HIT functions as an epigenetic
regulator of chondrogenesis through its recruitment of p100/CBP
Complexes. PLoS Genet. 11(e1005680)2015.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Cui Y, Zhang F, Zhu C, Geng L, Tian T and
Liu H: Upregulated lncRNA SNHG1 contributes to progression of
non-small cell lung cancer through inhibition of miR-101-3p and
activation of Wnt/β-catenin signaling pathway. Oncotarget.
8:17785–17794. 2017.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Hu Y, Ma Z, He Y, Liu W, Su Y and Tang Z:
lncRNA-SNHG1 contributes to gastric cancer cell proliferation by
regulating DNMT1. Biochem Biophys Res Commun. 491:926–931.
2017.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Liu Y, Yang Y, Li L, Liu Y, Geng P, Li G
and Song H: lncRNA SNHG1 enhances cell proliferation, migration,
and invasion in cervical cancer. Biochem Cell Biol. 96:38–43.
2018.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Chen Y, Wang T, Du J, Li Y, Wang X, Zhou
Y, Yu X, Fan W, Zhu Q, Tong X and Wang Y: The Critical Role of
PTEN/PI3K/AKT signaling pathway in shikonin-induced apoptosis and
proliferation inhibition of chronic myeloid leukemia. Cell Physiol
Biochem. 47:981–993. 2018.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Singh S, Asal R and Bhagat S:
Multifunctional antioxidant nanoliposome-mediated delivery of PTEN
plasmids restore the expression of tumor suppressor protein and
induce apoptosis in prostate cancer cells. J Biomed Mater Res A.
106:3152–3164. 2018.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Zhang Y, Zhang R, Luo G and Ai K: Long
noncoding RNA SNHG1 promotes cell proliferation through PI3K/AKT
signaling pathway in pancreatic ductal adenocarcinoma. J Cancer.
9:2713–2722. 2018.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Ward JF and Margulis V: Continous
improvement of TNM staging system for bladder cancer. Cancer.
115:704–705. 2009.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Poyet C, Thomas L, Benoit TM, Delmo DA,
Luberto L, Banzola I, Günthart MS, Sais G, Eberli D, Sulser T and
Provenzano M: Implication of vascular endothelial growth factor A
and C in revealing diagnostic lymphangiogenic markers in
node-positive bladder cancer. Oncotarget. 8:21871–2183.
2017.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Annels NE, Arif M, Simpson GR, Denyer M,
Moller-Levet C, Mansfield D, Butler R, Shafren D, Au G, Knowles M,
et al: Oncolytic immunotherapy for bladder cancer using Coxsackie
A21 virus. Mol Ther Oncolytics. 9:1–12. 2018.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Gao Y, Wu K, Chen Y, Zhou J, Du C, Shi Q,
Xu S, Jia J, Tang X, Li F, et al: Beyond proliferation: KLF5
promotes angiogenesis of bladder cancer through directly regulating
VEGFA transcription. Oncotarget. 6:43791–43805. 2015.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Pei Z, Du X, Song Y, Fan L, Li F, Gao Y,
Wu R, Chen Y, Li W, Zhou H, et al: Down-regulation of lncRNA CASC2
promotes cell proliferation and metastasis of bladder cancer by
activation of the Wnt/β-catenin signaling pathway. Oncotarget.
8:18145–18153. 2017.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Wang Y, Shi F, Xia Y and Zhao H: lncRNA
OIP5-AS1 predicts poor prognosis and regulates cell proliferation
and apoptosis in bladder cancer. J Cell Biochem: Nov 28, 2018 (Epub
ahead of print). doi: 10.1002/jcb.28024.
|
|
20
|
Gao R, Zhang N, Yang J, Zhu Y, Zhang Z,
Wang J, Xu X, Li Z, Liu X, Li Z, et al: Long non-coding RNA
ZEB1-AS1 regulates miR-200b/FSCN1 signaling and enhances migration
and invasion induced by TGF-β1 in bladder cancer cells. J Exp Clin
Cancer Res. 38(111)2019.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Lu Q, Shan S, Li Y, Zhu D, Jin W and Ren
T: Long noncoding RNA SNHG1 promotes non-small cell lung cancer
progression by up-regulating MTDH via sponging miR-145-5p. FASEB J.
32:3957–3967. 2018.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Cui L, Dong Y, Wang X, Zhao X, Kong C, Liu
Y, Jiang X and Zhang X: Downregulation of long noncoding RNA SNHG1
inhibits cell proliferation, metastasis, and invasion by
suppressing the Notch-1 signaling pathway in pancreatic cancer. J
Cell Biochem. 120:6106–6112. 2019.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Li Z, Li X, Du X, Zhang H, Wu Z, Ren K and
Han X: The Interaction Between lncRNA SNHG1 and miR-140 in
regulating growth and tumorigenesis via the TLR4/NF-κB pathway in
cholangiocarcinoma. Oncol Res. 27:663–672. 2019.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Deng R, Zhang J and Chen J: lncRNA SNHG1
negatively regulates miRNA1013p to enhance the expression of ROCK1
and promote cell proliferation, migration and invasion in
osteosarcoma. Int J Mol Med. 43:1157–1166. 2019.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Pistritto G, Trisciuoglio D, Ceci C,
Garufi A and D'Orazi G: Apoptosis as anticancer mechanism: Function
and dysfunction of its modulators and targeted therapeutic
strategies. Aging (Albany NY). 8:603–619. 2016.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Gan H, Zhang Y, Zhou Q, Zheng L, Xie X,
Veeraraghavan VP and Mohan SK: Zingerone induced caspase-dependent
apoptosis in MCF-7 cells and prevents
7,12-dimethylbenz(a)anthracene-induced mammary carcinogenesis in
experimental rats. J Biochem Mol Toxicol. 33(e22387)2019.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Warren CFA, Wong-Brown MW and Bowden NA:
BCL-2 family isoforms in apoptosis and cancer. Cell Death Dis.
10(177)2019.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Mirakhor Samani S, Ezazi Bojnordi T,
Zarghampour M, Merat S and Fouladi DF: Expression of p53, Bcl-2 and
Bax in endometrial carcinoma, endometrial hyperplasia and normal
endometrium: A histopathological study. J Obstet Gynaecol.
38:999–1004. 2018.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Tjalma WA, Weyler JJ, Bogers JJ,
Pollefliet C, Baay M, Goovaerts GC, Vermorken JB, van Dam PA, van
Marck EA and Buytaert PM: The importance of biological factors
(bcl-2, bax, p53, PCNA, MI, HPV and angiogenesis) in invasive
cervical cancer. Eur J Obstet Gynecol Reprod Biol. 97:223–230.
2001.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Paluch EK, Aspalter IM and Sixt M: Focal
adhesion-independent cell migration. Annu Rev Cell Dev Biol.
32:469–490. 2016.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Cao ZQ, Wang Z and Leng P: Aberrant
N-cadherin expression in cancer. Biomed Pharmacother.
118(109320)2019.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Siret C, Terciolo C, Dobric A, Habib MC,
Germain S, Bonnier R, Lombardo D, Rigot V and André F: Interplay
between cadherins and α2β1 integrin differentially regulates
melanoma cell invasion. Br J Cancer. 113:1445–1453. 2015.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Derycke LD and Bracke ME: N-cadherin in
the spotlight of cell-cell adhesion, differentiation,
embryogenesis, invasion and signalling. Int J Dev Biol. 48:463–476.
2004.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Wang Y, Kuang H, Xue J, Liao L, Yin F and
Zhou X: lncRNA AB073614 regulates proliferation and metastasis of
colorectal cancer cells via the PI3K/AKT signaling pathway. Biomed
Pharmacother. 93:1230–1237. 2017.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Zhang J, Li J, Li S, Zhou C, Qin Y and Li
X: miR802 inhibits the aggressive behaviors of nonsmall cell lung
cancer cells by directly targeting FGFR1. Int J Oncol.
54:2211–2222. 12019.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Zhang H, Pan YZ, Cheung M, Cao M, Yu C,
Chen L, Zhan L, He ZW and Sun CY: LAMB3 mediates apoptotic,
proliferative, invasive, and metastatic behaviors in pancreatic
cancer by regulating the PI3K/Akt signaling pathway. Cell Death
Dis. 10(230)2019.PubMed/NCBI View Article : Google Scholar
|