Introduction
Atrial fibrillation (AF) is the most common type of
arrhythmia and afflicts numerous individuals worldwide. AF
increases the probability of thromboembolism, ischemic stroke,
congestive heart failure, psychological distress and death
(1). AF impairs patient quality of
life (2-4),
especially those who require open-heart surgery (5). AF occurs alone or concomitantly with
other cardiovascular diseases including valvular heart, coronary
artery disease, hypertension and congestive heart failure (6).
AF usually begins in a self-terminating paroxysmal
form (ParoAF, defined by episodes lasting <7 days). Over time,
the AF pattern often evolves to become persistent (PersAF, duration
of episodes >7 days) and non-terminating within 7 days (7). Significant differences exist between
ParoAF and PersAF in terms of clinical features, responsiveness to
antiarrhythmic drugs and ablation therapy (8). However, the underlying molecular
mechanism of the occurrence and development of AF is poorly
understood, as well as factors regulating the progression from
ParoAF to PersAF.
Gene expression changes are associated with the
progression from ParoAF to PersAF (9,10).
Studies have primarily focused on the association between microRNAs
(miRNAs/miRs) and AF. Several have revealed that miRNAs regulate AF
or other cardiovascular diseases by promoting electrical or
structural remodeling of the atrium (11-13).
Dawson et al (11) reported
that miR29 likely regulates atrial fibrotic remodeling and may
represent a biomarker and/or therapeutic target. In recent years,
long non-coding RNAs (lncRNAs), which are longer than 200 nt in
length with a nonprotein-coding function, and mRNA expression have
gained interest among researchers (14,15).
lncRNA plays a central role in many processes during heart
development and various heart diseases, including cardiac
hypertrophy, cardiac fibrosis, AF and heart failure (16-21).
lncRNA AK055347 was upregulated in AF and shown to regulate
mitochondrial energy production in myocardiocytes (18). Several mRNAs were demonstrated to be
contributed to ParoAF pathogenesis via the gonadotropin releasing
hormone receptor and p53 pathways (22). However, these molecular markers were
investigated either on patients with ParoAF or PersAF. Few studies
have been performed on the differences in the molecular mechanism
of patients with ParoAF and PersAF.
In the present study, RNA sequencing (RNA-Seq)
technology was used to identify mRNAs and lncRNAs associated with
ParoAF and PersAF and to explore the underlying disease mechanisms.
Differentially expressed mRNAs (DE mRNAs) and lncRNAs (DE lncRNAs)
in ParoAF and PersAF were identified and compared with controls.
The putative function of the DE mRNAs and lncRNAs were determined
by Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes
(KEGG) pathway analyses. A co-expression network for lncRNA-mRNA
was subsequently constructed. The results provide valuable
molecular markers associated with the occurrence and development of
ParoAF and PersAF.
Materials and methods
Patients and tissue collection
This study was permitted by the Human Ethics
Committee of the First Affiliated Hospital of Nanjing Medical
University (approval number: 2020-SRFA-340). All patients with AF
and healthy donors provided written informed consent for the use of
their tissue in this study. The informed consent of the donors was
signed by one of their lineal relative members.
Patients who underwent heart transplantation in the
Department of Cardiovascular Surgery at the First Affiliated
Hospital of Nanjing Medical University (Nanjing, China) were
recruited. Patients categorized as ParoAF and PersAF met the
following criteria: i) Over 18 years old; ii) exhibited obvious
symptoms of AF as confirmed by electrocardiogram and had a clear
medical history; iii) minimally invasive radiofrequency ablation
performed for the first time; and iv) one-lung ventilation
implemented. The exclusion criteria were: i) Below 18 years old or
over 70 years old; ii) unclear medical history data; iii) combined
with other cardiac surgery or undergoing secondary surgery; and iv)
with psychiatric illness. The donor criteria included: i) Age
<50 years old, with some marginal donors <60 years old; ii)
estimated cold ischemia time <8 h; iii) no long-term or repeated
history of cardiopulmonary resuscitation; iv) serological
examination without hepatitis B/C, acquired immunodeficiency
syndrome; v) no existing bacteremia; and vi) no malignant tumors
other than primary brain tumors. The exclusion criteria for donors
were: i) Brain death due to cardiac arrest; ii) heart contusion;
iii) intractable ventricular arrhythmia; iv) long-term or repeated
cardiopulmonary resuscitation; v) past heart disease, particularly
congenital heart malformations; vi) after actively optimizing the
before and after load, the support of a super-large dose of
inotropic drugs still needed; vii) echocardiography findings of
severe heart wall motion abnormalities and reduced sustained left
ventricular ejection fraction (after optimized afterload, positive
muscle support and other treatments, remained 40% lower); and viii)
severe left ventricular hypertrophy, with ventricular septum >13
mm and accompanied by electrocardiogram manifestations of left
ventricular hypertrophy.
Left atrial appendage (LAA) tissues collected from
the hearts of patients with AF (patients with ParoAF=3 and patients
with PersAF=3) and donors (n=3) were used for RNA-Seq analysis. All
healthy donors exhibited a sinus rhythm and were in good condition.
The LAA tissues were selected from patients with ParoAF and PersAF,
who underwent maze surgery. The tissues were quickly cut into
pieces, placed into liquid nitrogen for at least 4 h, and then
stored at -80˚C for further use.
RNA extraction
Total RNA was extracted using TRIzol reagent
(Invitrogen; Thermo Fisher Scientific, Inc.) according to the
manufacturer's instructions and quantified using a Qubit 3.0
(Invitrogen; Thermo Fisher Scientific, Inc.). RNA integrity (RIN)
was evaluated with an Agilent 2100 Bioanalyzer (Agilent
Technologies, Inc.). Samples with
OD260/280 values >1.9 and RIN
>7 were used for library construction.
Whole transcriptome library
preparation and sequencing
Ribosomal RNA (rRNA) was removed from the total RNA
samples using the Ribo-Minus kit (Thermo Fisher Scientific, Inc.).
Next, each RNA sample was quantified and used for library
preparation using the TruSeq RNA Library Preparation kit version 2
(Illumina, Inc.). All libraries were loaded into one lane on the
Illumina HiSeq X ten (Illumina, Inc.) platform, followed by 2x150
bp pair-end sequencing.
Bioinformatics analysis
FastQC (v0.11.5; http://www.bioinformatics.babraham.ac.uk/projects/fastqc/)
was used to process the raw sequencing data. The clean reads were
mapped onto the Homo sapiens (assembly GRCh38. P12) sequence
using TopHat (v2.0.12; https://ccb.jhu.edu/software/tophat/index.shtml),
followed by assembly using Cufflinks (v2.2.1; http://cufflinks.cbcb.umd.edu). Transcripts with reads
per kilobase of exon per million reads mapped (RPKM>0) were
retained for further analyses. EdgeR (http://www.bioconductor.org/packages/release/bioc/html/edgeR.html)
was utilized to identify the DE lncRNAs and mRNAs by pairwise
comparisons, using thresholds of |Log2 fold change|>2
and P<0.05.
As reported, lncRNAs usually work through
cis- and trans-elements (23). In the present study, the mRNAs
detected by cis function for the DE lncRNAs were analyzed by
GO (http://www.Geneontology.org/) and KEGG
(http://www.genome.jp/kegg/) with a
corrected P<0.05 considered as significantly enriched for terms
and pathways, respectively. Trans function targeted mRNAs
were predicted by correlations ≥0.9 or ≤-0.9.
mRNA-lncRNA co-expression network
The DE mRNAs upstream or downstream of the DE
lncRNAs were identified and their associations were visualized
using Cytoscape software (v3.6.1, https://cytoscape.org/). The DE mRNAs targeted by DE
lncRNA were also calculated, and the co-expression network was
displayed by Cytoscape software (v3.6.1).
Protein-protein interaction (PPI)
network
The association between DE mRNAs was also revealed
using a PPI network. The STRING database (https://string-db.org/cgi/input.pl) was used for an
interaction correlation study and correlations of r>0.7 were
displayed.
Reverse transcription-quantitative PCR
(RT-qPCR)
RT-qPCR was used to validate the sequencing analysis
results for the DE lncRNAs and mRNAs. Expression levels of
collagen, type I, alpha 1 (COL1A1); collagen, type I, alpha
2 (COL2A1); collagen, type VI, alpha 1 (COL6A1);
dopachrome tautomerase (DCT); phosphodiesterase 4D,
cAMP-specific (PDE4D); RP11-428C19.4; GPC-AS2;
and XLOC_110310 were measured in ParoAF_Control and
PersAF_Control samples and normalized by β-actin expression. RNA
samples were prepared as aforementioned. The primers were designed
according to the sequences provided by the RNA-Seq data and
synthesized by Genewiz Inc. (https://www.genewiz.com.cn/). The primers were
provided in Table SI. RT-PCR was
performed using HiScript® II One Step RT-PCR Kit (Vazyme
Biotech Co., Ltd.) kit according to the manufacturer's instruction.
The RT-qPCR reaction was performed using the AceQ qPCR SYBR Green
Master Mix (Vazyme Biotech Co., Ltd.) with a 20 µl PCR reaction
systems, including 0.5 µl of each primer (10 µM), 2 µl of cDNA, 10
µl of AceQ qPCR SYBR Green Master Mix and 7 µl of RNase free
H2O. The mixture was put on the ABI 7500 system
real-time PCR instrument (Applied Biosystems, Thermo Fisher
Scientific, Inc.), and the amplification conditions were as
follows: Pre-denaturation at 95˚C for 30 sec; denaturation at 95˚C
for 5 sec, annealing at 60˚C for 30 sec (39 cycles); extension at
95˚C for 10 sec. All the experiments were repeated in triplicate.
The mRNA and lncRNA expression levels were calculated using the
2-ΔΔCq method (24).
Statistical analysis
All statistical analyses were performed using IBM
SPSS Statistics v22.0 (IBM Corp.) or GraphPad Prism 8 (GraphPad
Software, Inc.) software. Normal data are shown as means ± standard
error of means. Comparison among three groups was done using the
non-parametric Kruskal-Wallis H test followed by Dunn's test.
Comparisons between two groups were performed using the
non-parametric Mann-Whitney U test. P<0.05 was considered to
indicate a statistically significant difference.
Results
Clinical characteristics
There were no significant differences in age, body
mass index, left ventricular diastolic dimension (LVDD), left
ventricular systolic diameter, ventricular rate, systolic blood
pressure and diastolic blood pressure across control donors or
patients with ParoAF and PersAF (P>0.05; Table I). The CHA2DS2-VASc score (that
considers congestive heart failure/LV dysfunction, hypertension,
age ≥75, diabetes mellitus, stroke/TIA/TE, vascular disease,
age=65-74 and sex category), HASBLED score (that considers
parameters of hypertension, abnormal renal function, abnormal liver
function, previous stroke, bleeding history or predisposition,
history of labile international normalized ratio, age ≥65,
concomitant aspirin or nonsteroidal anti-inflammatory drug therapy
and substantial alcohol intake) and European Hearth Rhythm
Association showed no significant differences between the ParoAF
and PersAF groups.
 | Table IClinical characteristics of subjects
used for the sequencing. |
Table I
Clinical characteristics of subjects
used for the sequencing.
| Characteristic | Control, n=3 | ParoAF, n=3 | PersAF, n=3 | P-value |
|---|
| Age, years | 39.67±1.86 | 53.67±3.76 | 57.33±3.76 | 0.441a |
| BMI,
kg/m2 | 21.70±1.62 | 23.07±0.55 | 25.17±0.44 | 0.059a |
| LA, mm | 35.67±1.20 | 36.33±3.48 | 44.33±3.38 | 0.086a |
| LVDD, mm | 44.33±2.73 | 46.67±1.20 | 45.33±1.33 | 0.429a |
| LVSD, mm | 27.33±1.76 | 32.00±2.00 | 29.67±0.88 | 0.097a |
| Ventricular
rate | 56.00±4.16 | 91.33±8.41 | 73.67±8.09 | 0.083a |
| Systolic blood
pressure | 125.0±5.51 | 114.0±6.47 | 121.3±8.05 | 0.287a |
| Diastolic blood
pressure | 78.05±6.89 | 80.24±5.11 | 80.46±4.10 | 0.617a |
| CHA2DS2-VASc
score | – | 3.280±0.85 | 2.962±0.88 | 0.478b |
| HASBLED score | – | 2.852±0.61 | 2.846±0.74 | 0.988b |
| EHRA | – | I(1)/II(2) | I(1)/II(2) | – |
Differentially expressed lncRNAs and
mRNAs
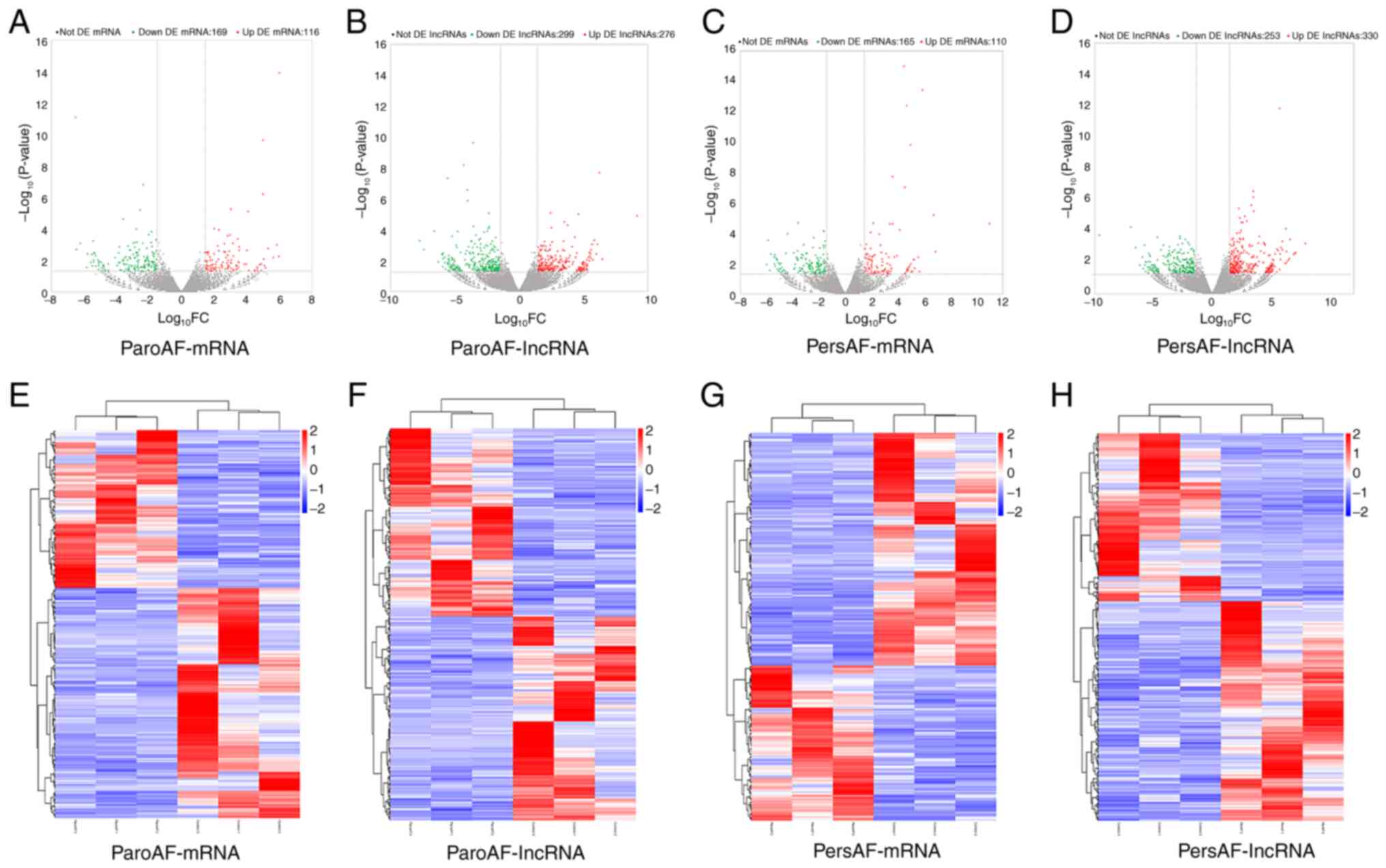
A total of 285 (116 upregulated and 169
downregulated) and 275 (110 upregulated and 165 downregulated) DE
mRNAs were identified in patients with ParoAF and PersAF compared
with the control group, respectively. A total of 575 (276
upregulated and 299 downregulated) and 583 (330 upregulated and 253
downregulated) DE lncRNAs were detected in the ParoAF_Control and
PersAF_Control samples, respectively. The volcano map of DE mRNAs
and lncRNAs in the ParoAF_Control and PersAF_Control samples are
shown in Fig. 1A-D. In addition,
the expression levels of DE mRNA and lncRNAs in the ParoAF_Control
and PersAF_Control samples are shown in Fig. 1E-H. The DE mRNAs and lncRNAs in the
ParoAF_Control sample are provided in Tables SII and SIII, respectively, whereas the DE mRNAs
and lncRNAs in the PersAF_Control samples are listed in Tables SIV and SV, respectively.
GO and KEGG pathway analysis of DE
mRNAs
There were two GO terms for ‘extracellular matrix’
and ‘platelet-derived growth factor binding’ that were
significantly enriched in the DE mRNA profile of the ParoAF_Control
sample. These two GO terms were represented by COL1A2
(upregulated), COL6A1 (upregulated), COL3A1
(upregulated), COL2A1 (downregulated) and so forth.
Moreover, COL1A2 (upregulated), COL6A1 (upregulated),
and COL3A1 (upregulated) were also significantly enriched in
the GO terms ‘collagen type I’ and ‘platelet-derived growth factor
binding’ in the PersAF_Control samples (Table II).
 | Table IIEnriched GO terms of ParoAF and
PersAF when compared to control. |
Table II
Enriched GO terms of ParoAF and
PersAF when compared to control.
| A,
ParoAF_Control |
|---|
| GO terms | Term name | GO
classification | Corrected
P-value | Input genes |
|---|
| GO:0031012 | Extracellular
matrix | CC |
3x10-2 | COL3A1,
COL2A1, THBS1, LAMC1, SPARC,
LAD1, COL6A6, COL1A2, GPC1,
THSD4, WISP1, LPL, ENTPD2,
C1QTNF9B, LAMB2, IMPG1, TIMP4,
ADAMTSL4, COL6A1, VCAN, SERPINE2,
DCN, ADAM11 |
| B,
PersAF_Control |
| GO terms | Term name | GO
classification | Corrected
P-value | Input genes |
| GO:0048407 | Platelet-derived
growth factor binding | MF |
5x10-3 | COL1A2,
COL6A1, COL3A1, COL1A1 |
| GO:0005584 | Collagen type
I | CC |
5x10-2 | COL1A2,
COL1A1 |
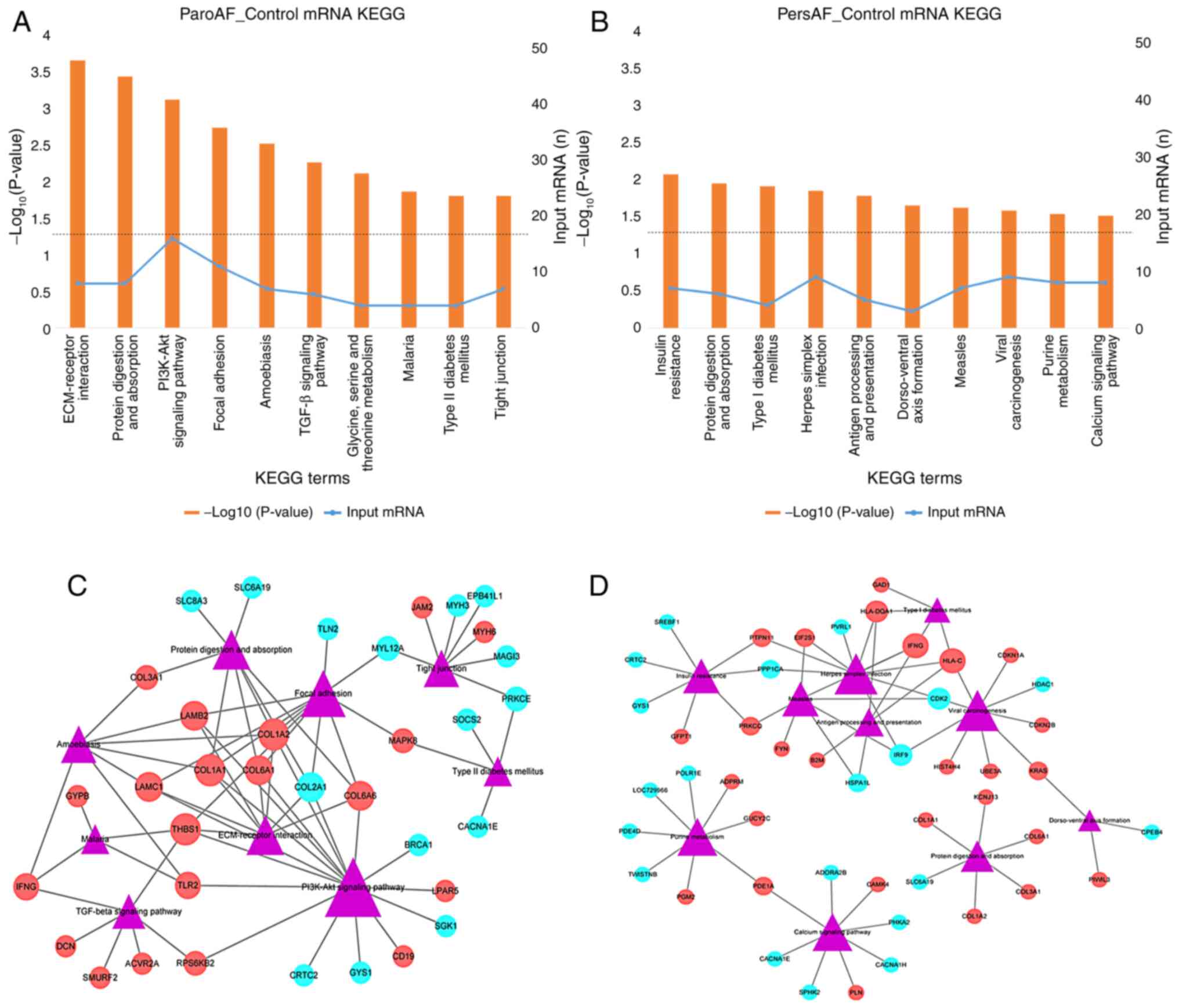
The top five enriched pathways were ‘ECM-receptor
interaction,’ ‘protein digestion and absorption,’ ‘PI3K-Akt
signaling pathway Focal adhesion Amoebiasis,’ ‘TGF-beta signaling
pathway,’ and ‘Amoebiasis’ in the ParoAF_Control samples (Fig. 2A). Upregulated DE mRNAs, such as
COL1A1, COL1A2 and laminin β2 (LAMB2), were
associated with most of the top ten pathways (Fig. 2C). Downregulated COL2A1 was
associated with most of the top ten pathways. Decorin (DCN),
SMAD specific E3 ubiquitin protein ligase 2 (SMURF2), and
interferon, γ (IFNG) are known to participate in the TGF-β
signaling pathway.
The top five enriched pathways in the PersAF_Control
were ‘insulin resistance,’ ‘protein digestion and absorption,’
‘type I diabetes mellitus,’ ‘herpes simplex infection,’ and
‘antigen processing and presentation dorso-ventral axis formation’.
They were enriched in IFNG (upregulated), major
histocompatibility complex class I C (HLA-C; upregulated),
and interferon regulatory factor 9 (IRF9; downregulated)
(Fig. 2B and D).
Function analysis of DE lncRNAs
through cis
The upstream mRNAs of the DE lncRNAs were enriched
in ‘phosphoric diester hydrolase activity,’ ‘regulation of
nucleotide metabolic process,’ and ‘cytoskeleton’. The GO terms
‘sequence-specific DNA binding RNA polymerase II transcription
factor activity,’ ‘response to dietary excess,’ and ‘positive
regulation of gene expression’ were enriched among the mRNAs
downstream of the DE lncRNAs. The top ten GO terms of the upstream
and downstream mRNAs of the DE lncRNAs are listed in Table III.
 | Table IIITop ten GO terms for cis
target genes of upstream and downstream DE lncRNAs. |
Table III
Top ten GO terms for cis
target genes of upstream and downstream DE lncRNAs.
| A, Upstream |
|---|
| GO Terms | Term name | Corrected
P-value | Input
genesa |
|---|
| GO:0008081_MF | Phosphoric diester
hydrolase activity |
5.73x10-6 | PLCXD3,
PDE7B, GPCPD1, ENPP2, PDE5A,
PDE3A, LOC101928269, PLCH1, CHRM3,
PLCD1 |
| GO:0006140_BP | Regulation of
nucleotide metabolic process |
1.6x10-4 | ITGB1,
ADRB2, MCF2L2, RIN3, ARHGAP24,
NF1, CRHR1, BPGM, RASAL2,
GRM3 |
| GO:0005856_CC | Cytoskeleton |
3.9x10-4 | GRM3,
HSPB7, ATP6V1D, BCL10, MPLKIP,
CTNNB1, PCGF5, RILPL1, NFE2L2,
FRMD4A |
| GO:0042995_CC | Cell
projection |
5x10-4 | RILPL1,
CTNNB1, SPG11, TIAM2, FAM49A,
GRM3, ATP6V1D, NF1, ARHGAP24,
UNC13B |
| GO:1900542_BP | Regulation of
purine nucleotide metabolic process |
5x10-4 | HTR1B,
FBXO8, BCAS3, ABR, MYO9A, BCAR3,
RAP1B, ARF4, NRG4, NTRK2 |
| GO:0043005_CC | Neuron
projection |
5.9x10-4 | APC,
AMFR, NTRK2, PDE5A, PRSS23,
ANK3, KCNIP1, SEMA6A, CAMK1G,
DLG2 |
| GO:1901701_BP | Cellular response
to oxygen-containing compound |
7.1x10-4 | TGFB1,
PIK3CA, CALCRL, PDE4D, RORA,
WDTC1, ATP6V1D, CTNNB1, FZD4,
CMPK2 |
| GO:0008092_MF | Cytoskeletal
protein binding |
1.05x10-3 | DAAM1,
PPARGC1A, NCK2, DST, SHROOM3,
DYNLL1, MYRIP, ACTR3C, ITGB1,
XIRP2 |
| GO:0045202_CC | Synapse |
1.41x10-3 | CLSTN1,
GRM1, CAMK2D, SYNE1, SLC30A3,
UNC13B, RIMS4, CACNA1C, ITGB1,
SPG11 |
| GO:0048731_BP | System
development |
1.63x10-3 | ARHGAP24,
ADAMTS18, MAPK10, UNC13B, ITGB1,
SPG11, SYCP2, COL18A1, PLK5,
FGF14 |
| B, Downstream |
| GO Terms | Term name | Corrected
P-value | Input
genesa |
| GO:0000981_MF | Sequence-specific
DNA binding RNA polymerase II transcription factor activity |
3.12x10-5 | TGIF1,
ZSCAN5A, RORA, NFE2L2, ERG,
GATA6, CSRNP3, RDH13, NFIB,
ZBTB16 |
| GO:0002021_BP | Response to dietary
excess |
1x10-4 | BMP8A,
TBL1XR1, ADRB2, RMI1, CCKAR,
PPARGC1A, MC4R |
| GO:0010628_BP | Positive regulation
of gene expression |
2.2x10-4 | HIF1A,
FOXD2, MED12L, NFIA, GTF2F2,
TFAP2C, HIPK2, MAPK10, FGF7,
TLR2 |
| GO:1902531_BP | Regulation of
intracellular signal transduction |
6.5x10-4 | C12orf60,
PDGFD, TAOK3, MTA3, FOXM1,
MCF2L2, EPHB1, MUL1, OTUD7A,
TBPL1 |
| GO:0045893_BP | Positive regulation
of transcription, DNA-dependent |
2.33x10-3 | ARF4,
GATA6, TET2, BCL9, TBL1XR1,
BCL10, BCAS3, NFE2L2, SMARCA2,
ZBTB17 |
| GO:1902532_BP | Negative regulation
of intracellular signal transduction |
2.42x10-3 | SORL1,
SPRED2, NF1, VGLL4, RORA,
NFE2L2, RDH13, VDAC2, CNKSR3,
TIMP3 |
| GO:0000977_MF | RNA polymerase II
regulatory region sequence-specific DNA binding |
4.8x10-3 | NHLH2,
ZNF486, ZNF812, AEBP2, E2F8,
NFIA, FOXD2, ZNF736, MTA3,
ZNF98 |
| GO:0001012_MF | RNA polymerase II
regulatory region DNA binding |
5.22x10-3 | FBXO16,
ZNF506, ZBTB16, NFIB, SMARCC2,
RORA, ZNF708, NFE2L2, TGIF1,
GATA6 |
| GO:0045944_BP | Positive regulation
of transcription from RNA polymerase II promoter |
6.37x10-3 | RDH13,
CSRNP3, CTNNB1, ADRB2, TMEM173,
TGFB3, ATXN7, SMARCA2, NFIB,
TET2 |
| GO:0001071_MF | Nucleic acid
binding transcription factor activity |
8.72x10-3 | HIF1A,
C12orf60, NFIA, FOXD2, TBX20,
EPAS1, MTA3, ZNF98, AEBP2 |
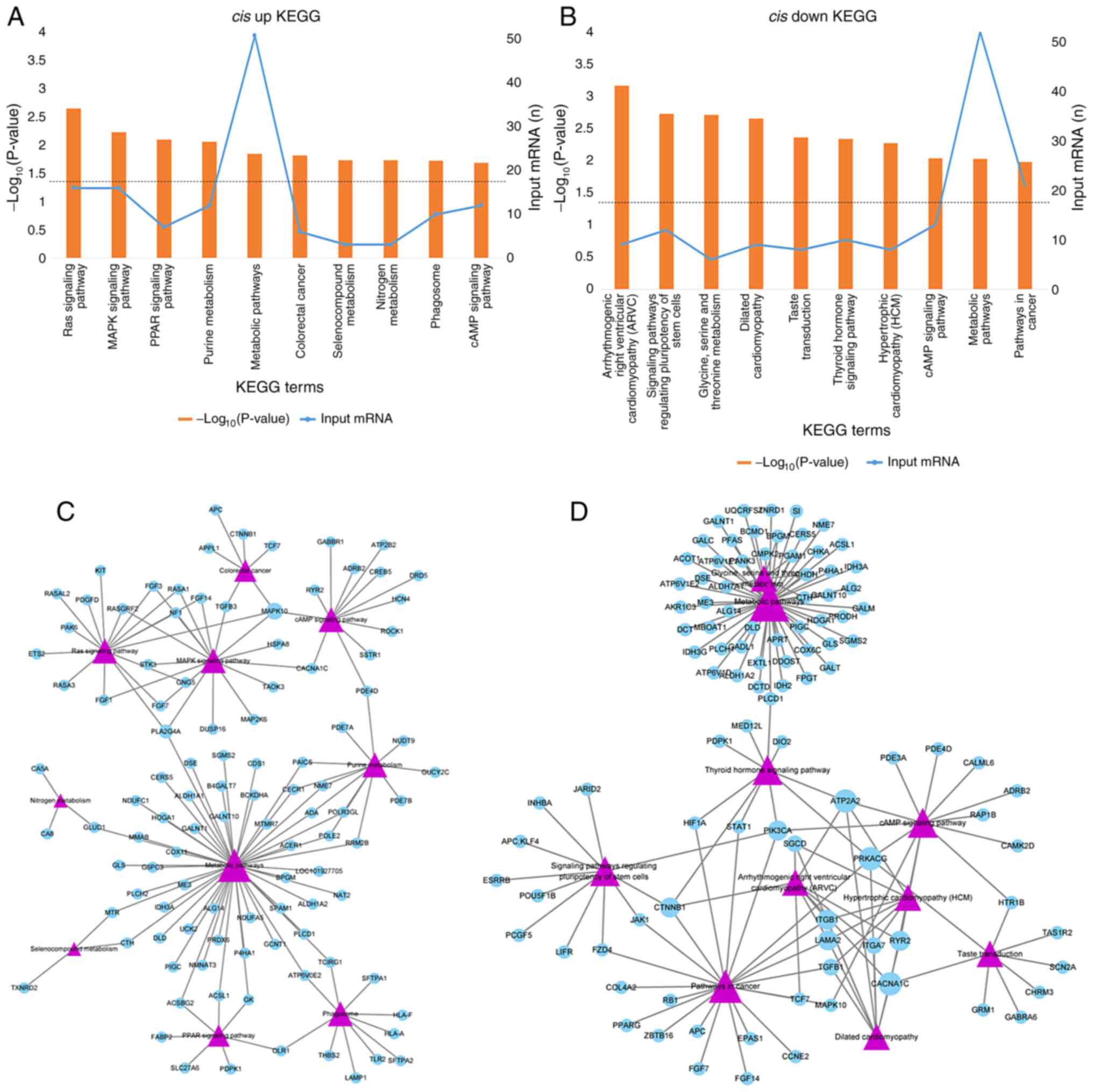
In addition, 23 and 47 KEGG pathways were enriched
in upstream and downstream mRNAs respectively. The upstream mRNAs
were mainly enriched in the ‘Ras signaling pathway,’ ‘MAPK
signaling pathway,’ and ‘PPAR signaling pathway’ (Fig. 3A), whereas the downstream mRNAs were
involved in ‘Arrhythmogenic right ventricular cardiomyopathy,’
‘Signaling pathways regulating pluripotency of stem cells,’ and
‘Glycine, serine and threonine metabolism’ (Fig. 3B). Mitogen-activated protein kinase
10 (MAPK10) and phospholipase A2, group IVA (PLA2G4A)
were involved in more pathways compared with the others for
upstream DE mRNAs (Fig. 3C). For
downstream mRNAs, calcium channel, voltage-dependent, L-type, alpha
1C subunit (CACNA1C), ATPase, Ca++ transporting,
cardiac muscle, slow twitch 2 (ATP2A2), catenin
(cadherin-associated protein), beta 1, 88 kDa (CTNNB1),
phosphatidylinositol-4,5-bisphosphate 3-kinase, catalytic subunit
alpha (PIK3CA), and protein kinase, cAMP-dependent,
catalytic, γ (PRKACG) involved in no less than four pathways
(Fig. 3D).
mRNA-lncRNA co-expression network
The DE mRNAs nearby the DE lncRNAs were identified
and are show in Fig. 4A and
B. The association between DE mRNAs
and lncRNAs in the ParoAF_Control were as follows: DCT was
downstream of GPC5-AS2, methyltransferase like 15 (METTL15)
at the downstream of BDNF-AS (Fig.
4A). The association between DE mRNAs and DE lncRNAs in
PersAF_Control, including DCT at the downstream of
GPC5-AS2, chromosome 21 open reading frame 90
(C21orf90) at the downstream of TSPEAR-AS1, and Mdm1 nuclear
protein homolog (MDM1) at the downstream of RP11-81H14.2
(Fig. 4B).
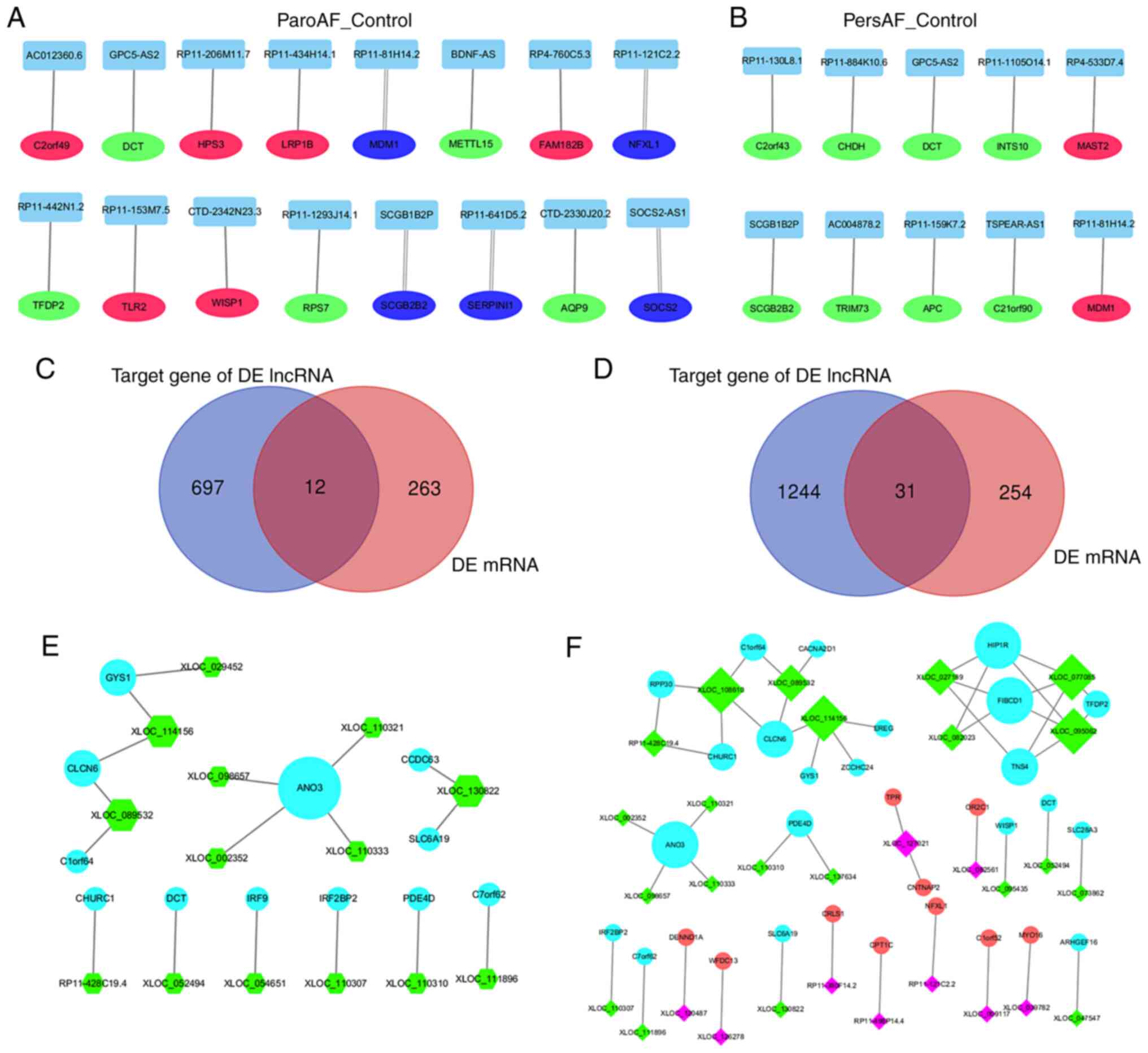 | Figure 4mRNA and lncRNA interaction
co-expression network of ParoAF_Control and PersAF_Control. (A and
B) DE lncRNAs and their upstream or downstream DE mRNAs. The red
circle node, green circle node, and blue represents the upstream DE
mRNAs, downstream DE mRNAs, and the DE mRNAs that were both
upstream and downstream, respectively. The double line indicates
that the DE mRNAs passes through the DE lncRNAs. The Venn map in (C
and D) showed the number of target genes of DE lncRNA and DE mRNAs
of ParoAF_Control and PersAF_Control, respectively. (E and F) The
co-expression network of DE lncRNA and DE mRNAs of ParoAF_Control
and PersAF_Control, respectively. The ‘diamond node’ represents DE
lncRNAs, and ‘circles node’ represent as mRNAs. The rose-red and
green colors represent upregulated and downregulated DE lncRNAs,
respectively. The red and blue represent upregulated and
downregulated DE mRNAs, respectively. The size of the node in A, B,
E and F all represent the DE lncRNAs or DE mRNAs that were
connected. DE, differentially expressed; ParoAF, paroxysmal atrial
fibrillation; PersAF, persistent atrial fibrillation; lncRNA, long
non-coding RNA. |
Next, the possible target mRNAs of the DE lncRNAs
(Trans prediction) was assessed and a total of 709 and 1,275
mRNAs were identified in ParoAF_Control and PersAF_Control,
respectively. DE mRNAs that were also targeted by DE lncRNAs were
selected, which are shown in Fig.
4C and D. There were 12 and 31
DE lncRNAs and DE mRNAs co-expressed in ParoAF and PersAF samples
compared with the control, respectively.
The selected mRNAs were then used for an mRNA-lncRNA
network analysis. The mRNA-lncRNA co-expression network of
ParoAF_Control revealed that anoctamin 3 (ANO3) exhibited a
concomitant downregulation trend with XLOC_002352,
XLOC_110321, XLOC_098657 and XLOC_110333.
Churchill domain containing 1 (CHURC1) showed downregulated
expression consistent with RP11-428C19.4 (Fig. 4E). ANO3 and CHURC1
co-expression were also detected in the PersAF_Control.
Furthermore, CHURC1 was also found to be co-expressed with
XLOC_108610. The upregulated trend was also detected, as
cardiolipin synthase 1 (CRLS1) was associated with
RP11-380f14.2, carnitine palmitoyltransferase 1C (CPT1C)
with RP11-498P14.4, and NFXL1 with RP11-121C2.2 (Fig. 4F).
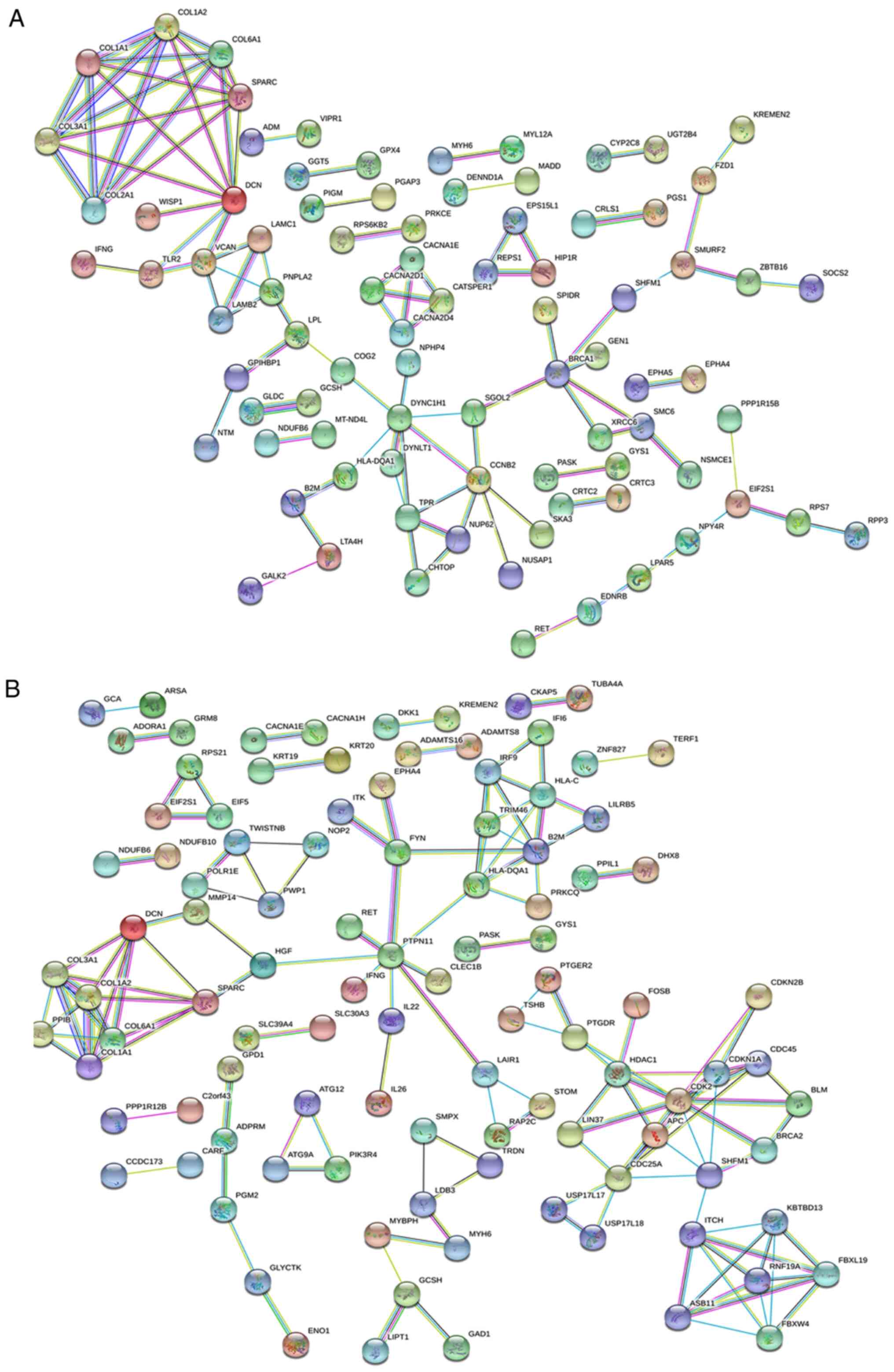
PPI network analysis
The PPI network in ParoAF_Control and PersAF_Control
is shown in Fig. 5A and B. Fig. 5A
shows that COL3A1, COL1A1, secreted protein acidic
cysteine-rich (SPARC), COL6A1, COL1A2,
DCN, and COL2A1 exhibited a strong interaction with
another. Downregulated genes, including breast cancer 1 early onset
(BRCA1) and COL2A1 are key genes that may interact
with many other DE mRNAs in the ParoAF_Control network (Fig. 5A).
In PersAF_Control, upregulated genes for protein
tyrosine phosphatase nonreceptor type 11 (PTPN11), major
histocompatibility complex class II DQ alpha 1 (HLA-DQA1),
cell division cycle 25A (CDC25A), β-2-microglobulin
(B2M), COL6A1, COL3A1, SPARC,
COL1A1 and COL1A2, and downregulated DE mRNAs,
including cyclin-dependent kinase 2 (CDK2), kelch repeat and
BTB (POZ) domain containing 13 (KBTBD13), histone
deacetylase 1 (HDAC1), split hand/foot malformation
(ectrodactyly) type 1 (SHFM1), F-box and leucine-rich repeat
protein 19 (FBXL19), and ring finger protein 19A, RBR E3
ubiquitin protein ligase (RNF19A) are essential genes that
may interact with other DE mRNAs (Fig.
5B).
Validation of differentially expressed
lncRNAs and mRNAs using RT-qPCR
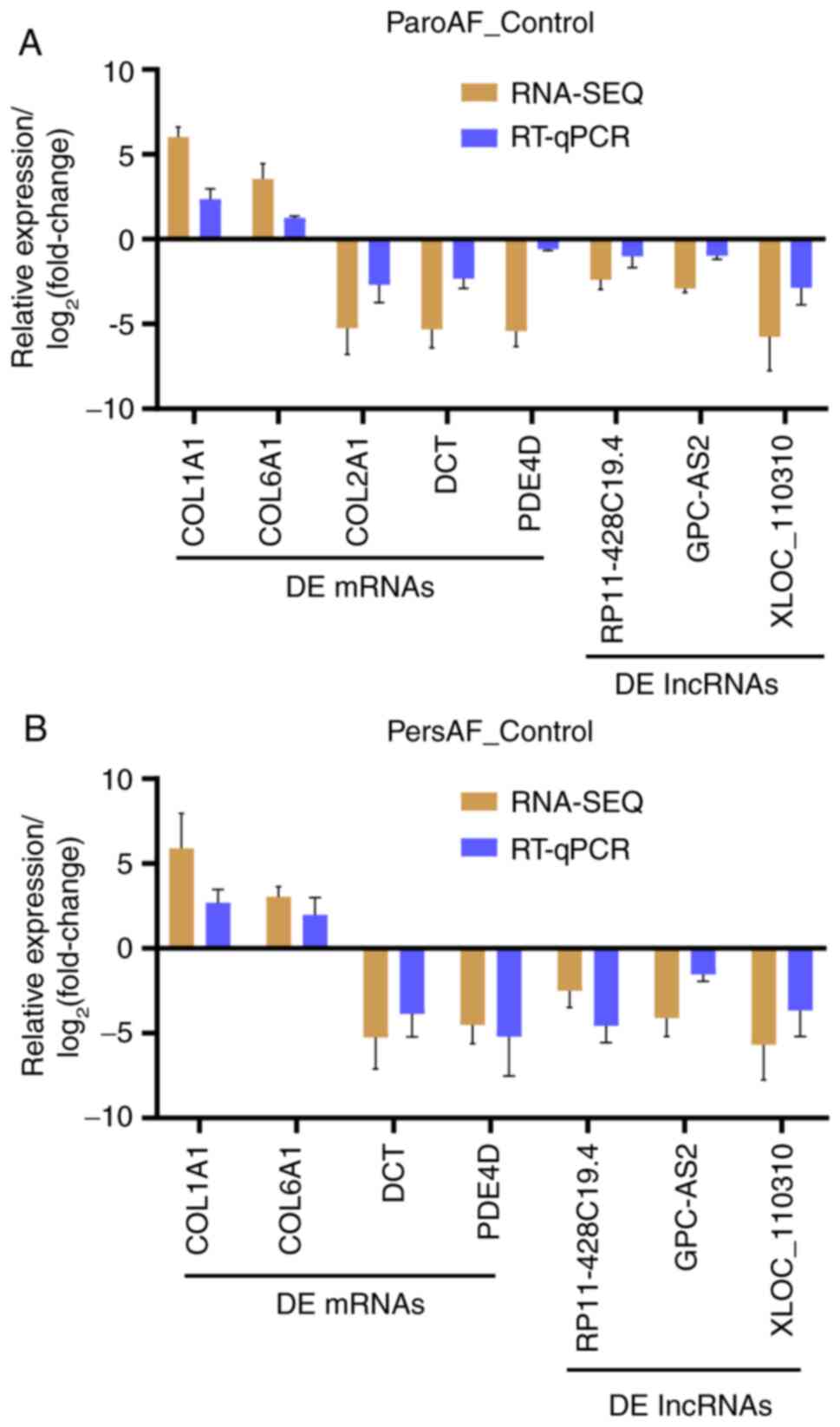
The expression of the DE mRNAs and lncRNAs measured
by RT-qPCR exhibited a similar trend in expression consistent with
that of RNA-Seq (Fig. 6). Moreover,
mRNA-lncRNA co-expression interactions of DCT with
GPC-AS2, and PDE4D with XLOC_110310 were
demonstrated.
Discussion
AF is the most prevalent heart disease worldwide,
with different subtypes causing different clinical features.
Therefore, these subtypes should be treated differently. In recent
years, the molecular mechanism of AF has been studied in a variety
of ways. However, the studies on the differences of lncRNAs and
mRNAs between ParoAF and PersAF have not been sufficient. In the
present study, ParoAF, PersAF and healthy donors samples were
sequenced, and found that DE mRNAs and lncRNAs vary considerably.
The putative function of DE mRNAs and DE lncRNA in ParoAF_Control
and PersAF_Control were also different.
COL1A2, was associated with platelet-derived
growth factor binding in the comparison of both ParoAF_Control and
PersAF_Control. Moreover, COL1A2 and COL2A1 were also
associated with most of the top ten KEGG pathways in patients with
ParoAF. In a study by Zhou et al (25), upregulated COL1A2 in patients
with AF were observed when compared with a control, suggesting that
it may participate in the occurrence and development of AF. Gambini
et al (26) demonstrated
that TGF-β1 could induce the upregulation of COL1A2 in human
cardiac mesenchymal progenitor cells from PersAF specimens. The
expression level of COL1A2 in these two studies was
consistent with that of the present results. In the study by Dawson
et al (11), it was reported
that COL1A2 was a target of miR29, which may play a role in
atrial fibrotic remodeling and is considered to be a biomarker or
therapeutic target. In the present study, COL1A2 also
exhibited a strong connection with other recombinant collagen type
I, III and IV family proteins, as determined by PPI analysis. These
results suggest an important role for COL1A2 in the
pathophysiology of both patients with ParoAF and PersAF.
The PI3K/Akt and TGF-β signaling pathways were
significantly enriched in the ParoAF_Control, whereas the calcium
signaling pathway was significantly enriched in the PersAF_Control.
Studies have demonstrated that some herbs can decrease the
incidence of AF through the PI3K/Akt signaling pathway (27,28).
Other studies have revealed that activating this pathway may
reverse atrial remodeling and inhibit the occurrence of AF
(29,30). Most collagen family genes detected
in the present study were associated with enriched pathways of both
the ParoAF_Control and PersAF_Control. Of note, COL2A1 was
only downregulated in the ParoAF_Control and is involved in
PI3K-Akt signaling pathway. However, there have been no reports
describing any associations among COL2A1, ParoAF and the
PI3K/Akt signaling pathway.
TGF-β signaling is involved in the course of atrial
structural remodeling in AF and is associated with the occurrence
and maintenance of AF (31). Fu
et al (32) showed that
overexpressed Gal-3 in AF could subsequently activate the
TGF-β1/α-SMA/Col I pathway in cardiac fibroblasts, thus
strengthening atrial fibrosis. Moreover, suppressing the
overexpression of TGF-β could prevent atrial remodeling (33-35).
TGF-β has also been demonstrated to be a promising therapeutic
target for decreasing cardiac fibrosis (36). In the present study, the TGF-β
signaling pathway was significantly enriched in the ParoAF_Control
along with SMURF2 and DCN. SMURF2, which was
upregulated in the present study, can induce proteasomal
degradation of Smad7 and activate the TGF-β signaling pathway
(37,38). DCN, a proteoglycan that can
bind to collagen fibrils in the ECM (extracellular matrix),
interacts with many growth factors, and inhibits TGF-β activity
(37). Moreover, DCN
exhibited a strong connection with other DE mRNAs in the PPI
analysis in the present study. Taken together, it was hypothesized
that the TGF-β signaling pathway and DCN may play essential
roles in AF occurrence and maintenance. Moreover, TGF-β may
represent an inhibitor for preventing AF occurrence.
In the present study, the calcium signaling pathway
was significantly enriched in the PersAF_Control. Calcium signaling
pathway was considered to play a vital role in electrical
remodeling and promoting the recurrence of AF (39). Tan et al (38) have demonstrated that the lncRNA
HOTAIR was involved in the modulation of calcium homeostasis
in human cardiomyocytes. It was also confirmed that HOTAIR
inhibited intracellular Ca2+ content by regulating
L-type calcium channels. In a study on mice with ParoAF and PersAF
(40), PersAF mice exhibited a
phenomenon of enhanced diastolic Ca2+ release, marked
conduction abnormalities and atrial enlargement. The absence of
PLN increased Ca2+ transient amplitude and a
faster Ca2+ decay rate (39). The combined results of these studies
with the present study indicate that the calcium signaling pathway
may play an essential role in the processing of ParoAF to PersAF.
Moreover, it was hypothesized that developing inhibitors to this
pathway may inhibit the ParoAF to PersAF transition. Unfortunately,
the study did not provide detailed information regarding ParoAF and
PersAF and was not conducive for the comparison analysis.
lncRNAs play essential roles in ParoAF and PersAF
through cis-elements. In the present study, DCT
interacted with GPC5-AS2, and METTL15 interacted with
BDNF-AS in the ParoAF_Control. DCT interacted with
GPC5-AS2, C21orf90 interacted with TSPEAR-AS1,
and MDM1 interacted with RP11-81H14.2 in the
PersAF_Control. These DE lncRNAs may perform their function through
nearby DE mRNAs. Levin et al (41) revealed that adult mice lacking
DCT display normal cardiac development but an increased
susceptibility to atrial arrhythmias. This suggests that lncRNA
GPC-AS2 inhibits the expression of DCT to influence
atrial arrhythmias, which warrants further study. Through our
trans analysis, mRNAs that were differentially expressed and
targeted DE lncRNAs were identified. These DE mRNAs exhibited
co-expression with DE lncRNAs. PDE4D, a biomarker of
myocardial infarction and heart failure (42), was downregulated in both
ParoAF_Control and PersAF_Control and was targeted by the DE
lncRNAs of XLOC_110310 and XLOC_137634. Further
studies are needed to understand the molecular mechanism of DE mRNA
and lncRNA interactions.
The function of DE lncRNAs in ParoAF_Control and
PersAF_Control was analyzed through cis-elements. As a
nearby gene of DE lncRNAs, CACNA1C was enriched in both the
MAPK and cAMP signaling pathways. Zhang et al (43) revealed that the MAPKs/TGF-β1/TRAF6
signaling pathways participate in atrial fibrosis in patients with
rheumatic heart disease, causing the occurrence of AF following
cardiac surgery. Inhibiting the MAPK pathways can prevent atrial
parasympathetic remodeling and the occurrence of AF (44). The induction of AF and structural
remodeling was associated with MAPK expression and the decrease in
collagenase activity (15).
CACNA1C is enriched in the MAPK signal pathway which is the
direct target gene of miR-29a-3p. Zhao et al (45) found that decreased expression of
CACNA1C caused by overexpression of microRNA-29a underlies
the pathogenesis of AF. Thus, it was assumed that miR-29a-3p may be
a potential therapeutic target in AF. These studies combined with
the present result indicate that the MAPK signaling pathway is an
important pathway that participates in AF occurrence by preventing
atrial parasympathetic remodeling.
There were some limitations in the present study.
Firstly, the original intention was to study the molecular
mechanism of all AF types including permanent AF, ParoAF and
PersAF. However, the permanent AF samples were difficult to
collect. Secondly, the sample number used in the present study was
low, as more than five would have been better. Thirdly, the present
study lacked functional verification of the mRNAs and lncRNAs,
which will be the subject of a future study.
The present study analyzed the DE mRNAs and lncRNAs
and their putative roles in ParoAF and PersAF. It was found that
PI3K/Akt and TGF-β signaling were significantly enriched in the
ParoAF_Control, and the calcium signaling pathway was significantly
enriched in the PersAF_Control. The cis and trans
analyses revealed some important interactions between DE mRNAs and
lncRNAs including GPC-AS2 with DCT, and PDE4D
with XLOC_110310 and XLOC_137634. In summary, the
present study provided molecular theoretical data for further
clinical studies involving ParoAF and PersAF.
Supplementary Material
Primer list of genes used for reverse
transcription-quantitative PCR validation.
The differentially expressed mRNAs in
the ParoAF_Control group.
The differentially expressed lncRNAs
in the ParoAF_Control group.
The differentially expressed mRNAs in
the PersAF_Control group.
The differentially expressed lncRNAs
in the PersAF_Control group.
Acknowledgements
Not applicable.
Funding
Funding: Not applicable.
Availability of data and materials
The raw data were deposited on the NCBI Sequence
Read Archive (SRA), with the SRA accession number of
PRJNA531935.
Authors' contributions
HS designed the study, collected the samples,
conducted the experiment, analyzed the data, and wrote the
manuscript. YS equally designed the study, analyzed the data,
provided the foundation and revised the manuscript. Both authors
read and approved the final manuscript.
Ethics approval and consent to
participate
This study was permitted by the Human Ethics
Committee of the Jiangsu People Hospital, (approval number,
2020-SRFA-340). All patients with AF and control patients have been
informed of writing consent to use their tissue for this study.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Marrouche NF, Brachmann J, Andresen D,
Siebels J, Boersma L, Jordaens L, Merkely B, Pokushalov E, Sanders
P, Proff J, et al: Catheter ablation for atrial fibrillation with
heart failure. N Engl J Med. 378:417–427. 2018.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Kalman JM, Sanders P, Rosso R and Calkins
H: Should we perform catheter ablation for asymptomatic atrial
fibrillation? Circulation. 136:490–499. 2017.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Zoni Berisso M, Landolina M, Ermini G,
Parretti D, Zingarini GL, Degli Esposti L, Cricelli C and Boriani
G: The cost of atrial fibrillation in Italy: A five-year analysis
of healthcare expenditure in the general population. From the
Italian survey of atrial fibrillation management (ISAF) study. Eur
Rev Med Pharmacol Sci. 21:175–183. 2017.PubMed/NCBI
|
|
4
|
Hu CY, Wang CY, Li JY, Ma J and Li ZQ:
Relationship between atrial fibrillation and heart failure. Eur Rev
Med Pharmacol Sci. 20:4593–4600. 2016.PubMed/NCBI
|
|
5
|
Yang S, Mei B, Feng K, Lin W, Chen G,
Liang M, Zhang X and Wu Z: Long-term results of surgical atrial
fibrillation radiofrequency ablation: Comparison of two methods.
Heart Lung Circ. 27:621–628. 2018.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Zhao L, Wang WYS and Yang X:
Anticoagulation in atrial fibrillation with heart failure. Heart
Fail Rev. 23:563–571. 2018.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Heijman J, Voigt N, Nattel S and Dobrev D:
Cellular and molecular electrophysiology of atrial fibrillation
initiation, maintenance, and progression. Circ Res. 114:1483–1499.
2014.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Schotten U, Dobrev D, Platonov P, Kottkamp
H and Hindricks G: Current controversies in determining the main
mechanisms of atrial fibrillation. J Intern Med. 279:428–438.
2016.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Liu H, Chen GX, Liang MY, Qin H, Rong J,
Yao JP and Wu ZK: Atrial fibrillation alters the microRNA
expression profiles of the left atria of patients with mitral
stenosis. BMC Cardiovasc Disord. 14(10)2014.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Yubing W, Yanping X, Zhiyu L, Weijie C, Li
S, Huaan D, Peilin X, Zengzhang L and Yuehui Y: Long-term outcome
of radiofrequency catheter ablation for persistent atrial
fibrillation. Medicine (Baltimore). 97(e11520)2018.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Dawson K, Wakili R, Ördög B, Clauss S,
Chen Y, Iwasaki Y, Voigt N, Qi XY, Sinner MF, Dobrev D, et al:
MicroRNA29: A mechanistic contributor and potential biomarker in
atrial fibrillation. Circulation. 127:1466–1475, 1475e1-28.
2013.PubMed/NCBI View Article : Google Scholar
|
|
12
|
McManus DD, Lin H, Tanriverdi K, Quercio
M, Yin X, Larson MG, Ellinor PT, Levy D, Freedman JE and Benjamin
EJ: Relations between circulating microRNAs and atrial
fibrillation: Data from the Framingham offspring study. Heart
Rhythm. 11:663–669. 2014.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Wang Y, Cai H, Li H, Gao Z and Song K:
Atrial overexpression of microRNA-27b attenuates angiotensin
II-induced atrial fibrosis and fibrillation by targeting ALK5. Hum
Cell. 31:251–260. 2018.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Saxena A and Carninci P: Long non-coding
RNA modifies chromatin: Epigenetic silencing by long non-coding
RNAs. Bioessays. 33:830–839. 2011.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Ruan Z, Sun X, Sheng H and Zhu L: Long
non-coding RNA expression profile in atrial fibrillation. Int J
Clin Exp Pathol. 8:8402–8410. 2015.PubMed/NCBI
|
|
16
|
Song C, Zhang J, Liu Y, Pan H, Qi HP, Cao
YG, Zhao JM, Li S, Guo J, Sun HL and Li CQ: Construction and
analysis of cardiac hypertrophy-associated lncRNA-mRNA network
based on competitive endogenous RNA reveal functional lncRNAs in
cardiac hypertrophy. Oncotarget. 7:10827–10840. 2016.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Liang H, Pan Z, Zhao X, Liu L, Sun J, Su
X, Xu C, Zhou Y, Zhao D, Xu B, et al: lncRNA PFL contributes to
cardiac fibrosis by acting as a competing endogenous RNA of let-7d.
Theranostics. 8:1180–1194. 2018.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Chen G, Guo H, Song Y, Chang H, Wang S,
Zhang M and Liu C: Long non-coding RNA AK055347 is upregulated in
patients with atrial fibrillation and regulates mitochondrial
energy production in myocardiocytes. Mol Med Rep. 14:5311–5317.
2016.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Chen L, Yan KP, Liu XC, Wang W, Li C, Li M
and Qiu CG: Valsartan regulates TGF-β/Smads and TGF-β/p38 pathways
through lncRNA CHRF to improve doxorubicin-induced heart failure.
Arch Pharm Res. 41:101–109. 2018.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Su Y, Li L, Zhao S, Yue Y and Yang S: The
long noncoding RNA expression profiles of paroxysmal atrial
fibrillation identified by microarray analysis. Gene. 642:125–134.
2018.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Yu XJ, Zou LH, Jin JH, Xiao F, Li L, Liu
N, Yang JF and Zou T: Long noncoding RNAs and novel inflammatory
genes determined by RNA sequencing in human lymphocytes are
up-regulated in permanent atrial fibrillation. Am J Transl Res.
9:2314–2336. 2017.PubMed/NCBI
|
|
22
|
Chiang DY, Zhang M, Voigt N, Alsina KM,
Jakob H, Martin JF, Dobrev D, Wehrens XHT and Li N: Identification
of microRNA-mRNA dysregulations in paroxysmal atrial fibrillation.
Int J Cardiol. 184:190–197. 2015.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Paralkar VR, Taborda CC, Huang P, Yao Y,
Kossenkov AV, Prasad R, Luan J, Davies JO, Hughes JR, Hardison RC,
et al: Unlinking an lncRNA from its associated cis element.
Mol Cell. 62:104–110. 2016.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Schmittgen TD and Livak KJ: Analyzing
real-time PCR data by the comparative C(T) method. Nat Protoc.
3:1101–1108. 2008.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Zhou DD, Jiang YY, Yuan C and Shi C: A
study on expression and distribution of collagen in rheumatic heart
disease with atrial fibrillation. Guangdong Med J, 2017.
|
|
26
|
Gambini E, Perrucci GL, Bassetti B,
Spaltro G, Campostrini G, Lionetti MC, Pilozzi A, Martinelli F,
Farruggia A, DiFrancesco D, et al: Preferential myofibroblast
differentiation of cardiac mesenchymal progenitor cells in the
presence of atrial fibrillation. Transl Res. 192:54–67.
2018.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Mao T, Zhang J, Qiao Y, Liu B and Zhang S:
Uncovering synergistic mechanism of Chinese herbal medicine in the
treatment of atrial fibrillation with obstructive sleep apnea
hypopnea syndrome by network pharmacology. Evid Based Complement
Alternat Med. 2019(8691608)2019.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Chong E, Chang SL, Hsiao YW, Singhal R,
Liu SH, Leha T, Lin WY, Hsu CP, Chen YC, Chen YJ, et al:
Resveratrol, a red wine antioxidant, reduces atrial fibrillation
susceptibility in the failing heart by PI3K/AKT/eNOS signaling
pathway activation. Heart Rhythm. 12:1046–1056. 2015.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Liu SH, Hsiao YW, Chong E, Singhal R, Fong
MC, Tsai YN, Hsu CP, Chen YC, Chen YJ, Chiou CW, et al: Rhodiola
inhibits atrial arrhythmogenesis in a heart failure model. J
Cardiovasc Electrophysiol. 27:1093–1101. 2016.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Mira YE, Muhuyati Lu WH, He PY, Liu ZQ and
Yang YC: TGF-β1 signal pathway in the regulation of inflammation in
patients with atrial fibrillation. Asian Pac J Trop Med.
6:999–1003. 2013.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Shen H, Wang J, Min J, Xi W, Gao Y, Yin L,
Yu Y, Liu K, Xiao J, Zhang YF and Wang ZN: Activation of
TGF-β1/α-SMA/Col I profibrotic pathway in fibroblasts by galectin-3
contributes to atrial fibrosis in experimental models and patients.
Cell Physiol Biochem. 47:851–863. 2018.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Fu H, Li G, Liu C, Li J, Wang X, Cheng L
and Liu T: Probucol prevents atrial remodeling by inhibiting
oxidative stress and TNF-α/NF-κB/TGF-β signal transduction pathway
in alloxan-induced diabetic rabbits. J Cardiovasc Electrophysiol.
26:211–222. 2015.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Verheule S, Sato T, Everett T IV, Engle
SK, Otten D, Rubart-Von Der Lohe M, Nakajima HO, Nakajima H, Field
LJ and Olgin JE: Increased vulnerability to atrial fibrillation in
transgenic mice with selective atrial fibrosis caused by
overexpression of TGF-beta1. Circ Res. 94:1458–1465.
2004.PubMed/NCBI View Article : Google Scholar
|
|
34
|
He X, Zhang K, Gao X, Li L, Tan H, Chen J
and Zhou Y: Rapid atrial pacing induces myocardial fibrosis by
down-regulating Smad7 via microRNA-21 in rabbit. Heart Vessels.
31:1696–1708. 2016.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Xie J, Tu T, Zhou S and Liu Q:
Transforming growth factor (TGF)-β1 signal pathway: A promising
therapeutic target for attenuating cardiac fibrosis. Int J Cardiol.
239(9)2017.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Cunnington RH, Nazari M and Dixon IM:
c-Ski, Smurf2, and Arkadia as regulators of TGF-beta signaling: New
targets for managing myofibroblast function and cardiac fibrosis.
Can J Physiol Pharmacol. 87:764–772. 2009.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Järvinen TAH and Ruoslahti E: Generation
of a multi-functional, target organ-specific, anti-fibrotic
molecule by molecular engineering of the extracellular matrix
protein, decorin. Br J Pharmacol. 176:16–25. 2019.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Tan WL, Xu M, Liu Z, Wu TY, Yang Y, Luo J,
Yang J and Luo Y: HOTAIR inhibited intracellular Ca2+
via regulation of Cav1.2 channel in human cardiomyocytes. Cell Mol
Biol (Noisy-le-grand). 61:79–83. 2015.PubMed/NCBI
|
|
39
|
Baskin KK, Makarewich CA, DeLeon SM, Ye W,
Chen B, Beetz N, Schrewe H, Bassel-Duby R and Olson EN: MED12
regulates a transcriptional network of calcium-handling genes in
the heart. JCI Insight. 2(e91920)2017.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Li N, Chiang DY, Wang S, Wang Q, Sun L,
Voigt N, Respress JL, Ather S, Skapura DG, Jordan VK, et al:
Ryanodine receptor-mediated calcium leak drives progressive
development of an atrial fibrillation substrate in a transgenic
mouse model. Circulation. 129:1276–1285. 2014.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Levin MD, Lu MM, Petrenko NB, Hawkins BJ,
Gupta TH, Lang D, Buckley PT, Jochems J, Liu F, Spurney CF, et al:
Melanocyte-like cells in the heart and pulmonary veins contribute
to atrial arrhythmia triggers. J Clin Invest. 119:3420–3436.
2009.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Swyngedouw NE and Jickling GC: RNA as a
stroke biomarker. Fut Neurol. 12:71–78. 2017.
|
|
43
|
Zhang D, Chen X, Wang Q, Wu S, Zheng Y and
Liu X: Role of the MAPKs/TGF-β1/TRAF6 signaling pathway in
postoperative atrial fibrillation. PLoS One.
12(e0173759)2017.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Liu L, Geng J, Zhao H, Yun F, Wang X, Yan
S, Ding X, Li W, Wang D, Li J, et al: Valsartan reduced atrial
fibrillation susceptibility by inhibiting atrial parasympathetic
remodeling through MAPKs/neurturin pathway. Cell Physiol Biochem.
36:2039–2050. 2015.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Zhao Y, Yuan Y and Qiu C: Underexpression
of CACNA1C caused by overexpression of microRNA-29a underlies the
pathogenesis of atrial fibrillation. Med Sci Monit. 22:2175–2181.
2016.PubMed/NCBI View Article : Google Scholar
|