Introduction
Invasion and metastasis are major characteristics of
malignant tumors and are the main causes of high mortality in
ovarian cancer. During tumor growth, tumor cells are often exposed
to nutrient deficiency due to insufficient angiogenesis or drug
interventions; however, according to several experimental and
clinical research findings, environment deficiency in nutrients
does not reduce tumor growth completely. On the contrary, tumor
cells invade the matrix and blood vessels in order to survive and
subsequently metastasize to other organs and tissues where
nutrition is higher, causing high mortality (1). Starvation and nutrient deficiency have
also been reported to induce autophagy in tumor cells through the
decomposition of their organelles and other organic substrates.
This provides raw materials in the form of amino acids and other
necessary small molecules that maintain cell survival (2). These findings suggest that autophagy
and invasion constitute a self-protective mechanism of tumor cells
under nutrient deficient conditions that acts as a bottleneck,
impeding anti-angiogenesis drug effectiveness (3-5).
Therefore, an exploration of the intrinsic link between autophagy
and invasion could reveal the tumor cell metastasis mechanism and
provide a novel theoretical basis to specifically target metastasis
and reverse tumor cell resistance in nutrient deficiency
conditions.
Invadopodia are actin-rich protrusions that localize
proteolytic activity to areas of the cell in contact with the
extracellular matrix (ECM). The ability to form invadopodia is
closely associated with the invasive and metastatic potential of a
tumor. A wide variety of actin-interacting and scaffolding proteins
are involved in invadopodia formation, including cortactin,
tyrosine kinase substrate with five Src homology SH3 domains,
fascin, neural Wiskott-Aldrich syndrome protein (N-WASP), and
actin-related protein complex 2/3 (Arp2/3 complex) (6,7). In
particular, the actin-bundling protein, cortactin, appears to play
an integral role in invadopodia formation (6-9).
Src kinase activity and cortactin phosphorylation are absolute
requirements for invadopodia formation, through the phosphorylation
of serine residues 405 and 418 within the PR domain of cortactin by
extracellular signal-regulated kinase (ERK1/2) or p21-activated
kinase 1, which according to previous studies enhances cortactin
SH3 domain binding to N-WASP (10).
As a result, it has been suggested that S405/S418 phosphorylation
plays a critical role in the regulation of the cellular actin
polymerization, which is necessary for the promotion of invadopodia
function (11,12).
The selective autophagy receptor and signaling
adaptor, sequestosome-1 (SQSTM1/p62, hereafter referred to as p62)
is a multidomain protein. The presence of an LC3-interacting region
enables p62 to bind to the autophagy protein microtubule-associated
protein 1A/1B light chain 3B (LC3), and the ubiquitin-associated
domain binds to ubiquitin to mediate the selective degradation of
ubiquitinated cargo by autophagy. p62 contains a protein-protein
interaction module (PB1) that facilitates its oligomerization and
is rich in protein-interacting sequences, that function as a
signaling hub through its interaction with key components of
signaling mechanisms including the Ras/Raf/mitogen-activated
protein kinase pathway and nuclear factor κB pathway (13-17).
Tumors often exist in low nutrition environments
during their growth or treatment; however, may overcome this by
invading other organs and subsequently causing high mortality.
Materials and methods
Reagents and antibodies
ViaFect Transfection Reagent was purchased from
Promega Corporation. 3-(4, 5-Dimethylthiazol-2-yl)-2,
5-diphenyltetrazolium bromide (MTT), Hoechst 33342, Gelatin from
pig skin, and fetal bovine serum (FBS) were purchased from
Sigma-Aldrich; Merck KGaA. The nuclear and cytoplasmic protein
extraction kit was purchased from Beyotime Institute of
Biotechnology. The ERK1/2 inhibitor, SCH772984, chloroquine (CQ)
and 3-methyladenine (3-MA) were purchased from MedChemExpress. The
following antibodies were used: Anti-p62 (cat. no. ab91526; Abcam),
anti-cortactin (cat. no. ab269977; Abcam); anti-β-actin (cat. no.
MA5-15452; ProteinTech Group, Inc.); anti-ERK1/2 (cat. no.
sc-514302; Santa Cruz Biotechnology, Inc.) and anti-LC3 (cat. no.
sc-398822; Santa Cruz Biotechnology, Inc.).
Cell lines and cell culture
Human ovarian carcinoma cells (SKOV3 cells; cat. no.
TCHu185) were purchased from the The Cell Bank of Type Culture
Collection of the Chinese Academy of Sciences and Peking Union
Medical College. SKOV3 cells were cultured in Roswell Park Memorial
Institute (RPMI)-1640 culture medium (Gibco; Thermo Fisher
Scientific, Inc.) supplemented with 10% fetal bovine serum
(Invitrogen; Thermo Fisher Scientific, Inc.) at 37˚C in 5%
CO2.
Autophagy evaluation
Autophagy promoted by amino acid starvation was
examined, with the use of 70% confluent cells, washed 3 times with
modified Earle's balanced salt solution (EBSS; 0.265 g/l
CaCl2.H2O, 0.09767 g/l MgSO4, 0.4 g/l KCl,
6.8 g/l NaCl, 0.122 g/l NaH2PO4, 1.0 g/l
D-Glucose, 0.011 g/l Phenol Red.Na, 2.2 g/l NaHCO3;
Sigma-Aldrich; Merck KGaA) and then incubated with EBSS at 37˚C in
5% CO2. Autophagy evaluation was performed in accordance
with previously published studies (18-21).
Transwell invasion assay
Cells were serum-starved for 24 h, harvested and
resuspended in medium containing 1% bovine serum albumin or EBSS.
For the invasion assay, the top chambers were coated with Matrigel
(cat. no. 354234; Corning, Inc.). Cells (5x104) were
added to the top chambers of 24-well Transwell plates (Costar;
Corning, Inc.). Medium containing 10% fetal bovine serum was added
to the bottom chambers. Cells were incubated at 37˚C for 0-24 h.
The cells were fixed in 0.1% glutaraldehyde-PBS for 20 min and
stained with 0.2% crystal violet (Sigma-Aldrich; Merck KGaA) for 1
h at room temperature. Non-motile cells on the top of each filter
were then removed using a cotton swab. The number of invasive cells
was counted using an Olympus IX-71 inverted microscope (Olympus
Corporation).
Invadopodia formation assay
Cells were plated on gelatin matrix in RPMI-1640
medium or EBSS. At the indicated time points (0, 2, 4, 8, 16 and 24
h), the cells were washed, fixed and permeabilized with 4%
paraformaldehyde (cat. no. 158127; Sigma-Aldrich; Merck KGaA) and
0.2% Triton-X (cat. no. A93443; Sigma-Aldrich; Merck KGa) for 20
min at 37˚C. Additionally, the cells were stained to view punctuate
structures where F-actin and cortactin colocalized. Cells were
incubated for 37˚C, 1 h with anti-cortactin (1:100; cat. no.
ab269977; Abcam) antibody and TRITC-phalloidin (1:1,000; cat. no.
41-6559-05; Invitrogen; Thermo Fisher Scientific, Inc.). Images
were acquired by an Olympus FV1000 confocal laser microscope
(Olympus Corporation) and measured using ImageJ software 1.8.0
(National Institutes of Health).
Cell transfection
p62-siRNA and non-target siRNA (Scramble) were
obtained from Shanghai GeneChem Co., Ltd. The p62-siRNA (si-p62)
sequence was GAC-ATC-TTC-CGAATC-TAC-A and the non-target siRNA
(Scramble) was TTC-TCC-GAA-CGT-GTC-ACG-T. The pcDNA3.1-p62 and
empty pcDNA3.1 vector (NC) were constructed by Sangon Biotech Co.,
Ltd. Cell transfection was performed as previously described
(22). A total of 25x104
cells were plated in 6-well plates. After 36 h culture, cells were
transfected with siRNA (2 µg/well) or plasmids (2.5 µg/well) using
Lipofectamine™ 2000 Transfection Reagent (Invitrogen; Thermo Fisher
Scientific, Inc.11668019) according to the manufacturer's
instructions. Cells were incubated for 1-3 days at 37˚C and then
used in subsequent experiments.
Cell viability assays
A total of 8x103 cells/well were seeded
in 96-well plates. Cells were cultured for 24 h, and then the cells
were treated with EBSS or various concentrations of CQ (0, 10, 25,
50 or 100 µM)/3-MA (0, 1, 2, 4, 8 or 10 mM). MTT assay was used to
evaluate cell viability. MTT (5 mg/ml) was added to the cells and
DMSO was used to dissolve the formazan. The absorbance was measured
at 570 nm using a Vmax Microplate Reader (Molecular Devices,
LLC).
Western blot analysis
Protein expression was examined by with western blot
analysis, as previously described (22). Protein was extracted using M-PER™
Mammalian Protein Extraction Reagent (cat. no. 78505; Thermo Fisher
Scientific), the protein concentration was detected by
bicinchoninic acid method. Protein was loaded (40 µg per lane) into
10% gel and proteins were transferred to PVDF membrane. Membrane
was blocked in 1% BSA at room temperature for 1h, and incubated
with the following primary antibodies overnight at 4˚C: Anti-p62
(1:1,000; cat. no. ab91526; Abcam), anti-cortactin (1:1,000; cat.
no. ab269977; Abcam), anti-p-cortactin (1:1,000; cat. no. ab47768;
Abcam), anti-β-actin (1:1,000; cat. no. MA5-15452; ProteinTech
Group, Inc.), anti-ERK1/2 (1:1,000; cat. no. sc-514302; Santa Cruz
Biotechnology, Inc.), anti-p-ERK1/2 (1:500; cat. no. sc-136521;
Santa Cruz Biotechnology, Inc.) and anti-LC3 (1:1,000; cat. no.
sc-398822; Santa Cruz Biotechnology, Inc.) using antibody diluents
(cat. no. U3635; Sigma-Aldrich; Merck KGaA). Goat anti-mouse
horseradish peroxidase-conjugated secondary antibodies were
incubated (1:200; cat. no. SA00001; ProteinTech Group, Inc.)
according to the manufacturer's instructions. Then, immunodetection
was performed using ECL reagent (Thermo Fisher Scientific, Inc.)
and visualized using a Syngene Bio Imaging System (Syngene Europe),
and densitometry was performed using ImageJ software v1.8.0
(National Institutes of Health).
Immunofluorescence staining
SKOV3 cells were cultured on coverslips at a density
of 5x104 cells/well in 500 µl of complete medium.
Following treatment, the SKOV3 cells were washed with cold PBS
three times and fixed in 4% (w/v) paraformaldehyde/PBS for 20 min
and then washed with cold PBS three times. The fixed cells were
subsequently digested by protein enzyme K for 1 min and washed with
PBS twice. The cells were then incubated with 0.1% (v/v) Triton
X-100 for 6-10 min, washed once with PBS, and then blocked for 30
min in 5% (v/v) non-immune animal serum/PBS. The cells were
incubated with the anti-cortactin primary antibody (1:100; cat. no.
ab269977; Abcam) overnight and washed three times with PBS. They
were then incubated with goat anti-mouse IgG secondary antibody
conjugated to Alexa Fluor® Plus 488 (1:400; cat. no.
A32723; Thermo Fisher Scientific) for 30 min in the dark. Plates
were washed three times in PBS, treated with Hoechst
33342/H2O (1 µg/ml) for 2 min, and then washed three
times with PBS. The cells were examined using an Olympus FV1000
confocal laser microscope (Olympus Corporation; magnification,
x400).
Statistical analysis
Statistical analysis was performed using the
Statistical Package for the Social Sciences, version 22 (IBM
Corp.). All data are presented as the mean ± SD. The unpaired
Student's t-test was applied, in order to perform comparisons
between two groups. One-way ANOVA followed by Tukey's post hoc test
was used to analyze datasets with more than two groups. Two-way
ANOVA followed by Bonferroni's post hoc test was performed to
evaluate the effects of concentration and EBSS. P<0.05 was
considered to indicate a statistically significant difference.
Results
Autophagy inhibition reduces the
invasion of human ovarian cancer cells
The ability of cancer cells to metastasize is
dependent on their capability to invade surrounding tissues and
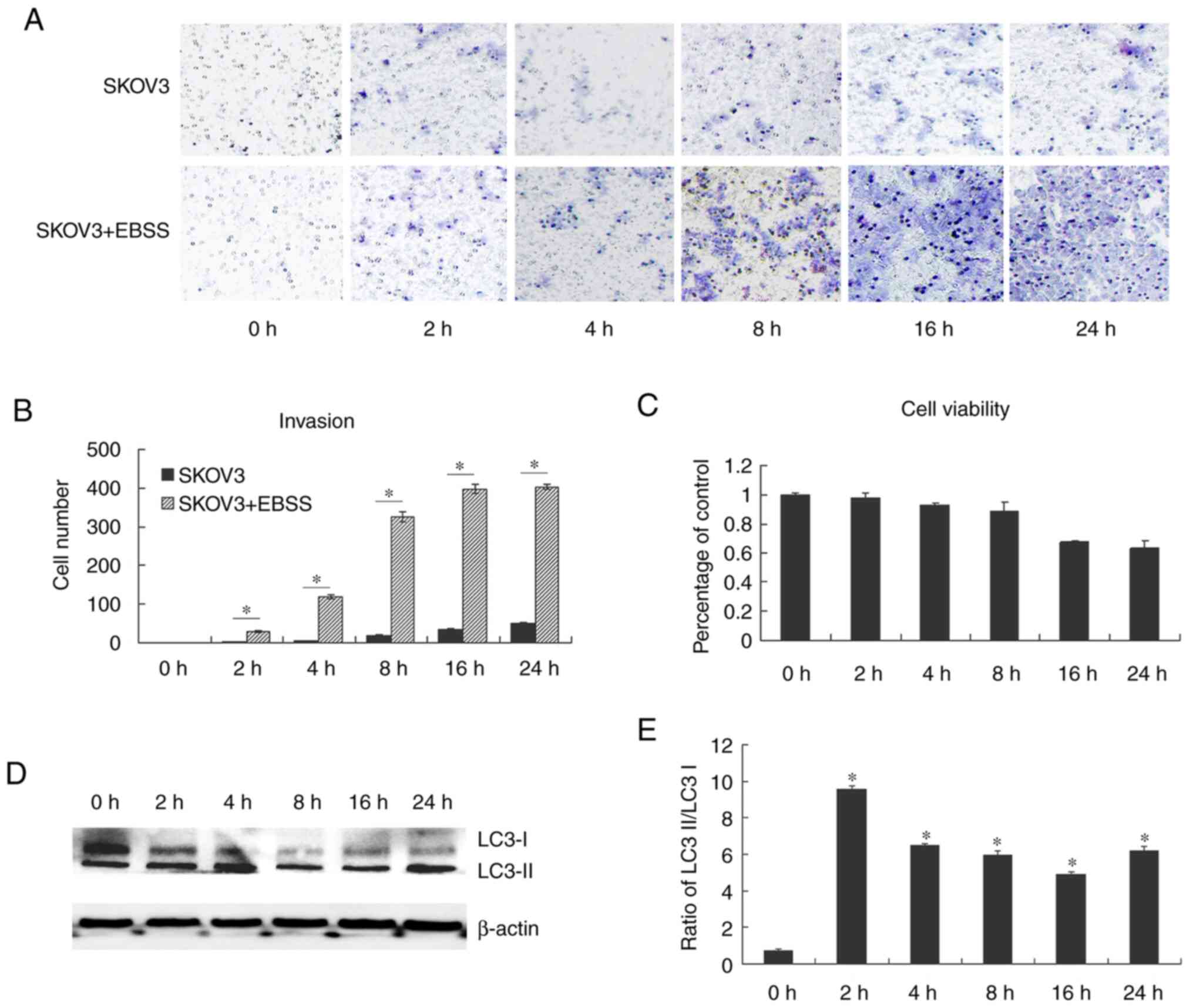
migrate from the primary tumor site. In the present study, it was
demonstrated that EBSS increased the invasion of the human ovarian
cancer cell line, SKOV3. EBSS is a balanced salt solution and does
not contain amino acids or other active substances, thus providing
a nutrient deficient environment. SKOV3 cells cultured in EBSS
exhibited an increased invasion compared with the control cells
(Fig. 1A and B). Fig. 1C
further confirmed that the increased invasion was not due to cell
viability increase. As EBSS has been previously reported to induce
autophagy (18-21),
in the present study, western blot analysis performed to examine
autophagy in SKOV3 cells treated with EBSS through the detection of
LC3 protein expression. LC3II protein levels were significantly
increased following the treatment of SKOV3 cells with EBSS for 2 h,
peaked after 8 h of treatment and then decreased (Fig. 1D and E). Subsequently, in order to examine the
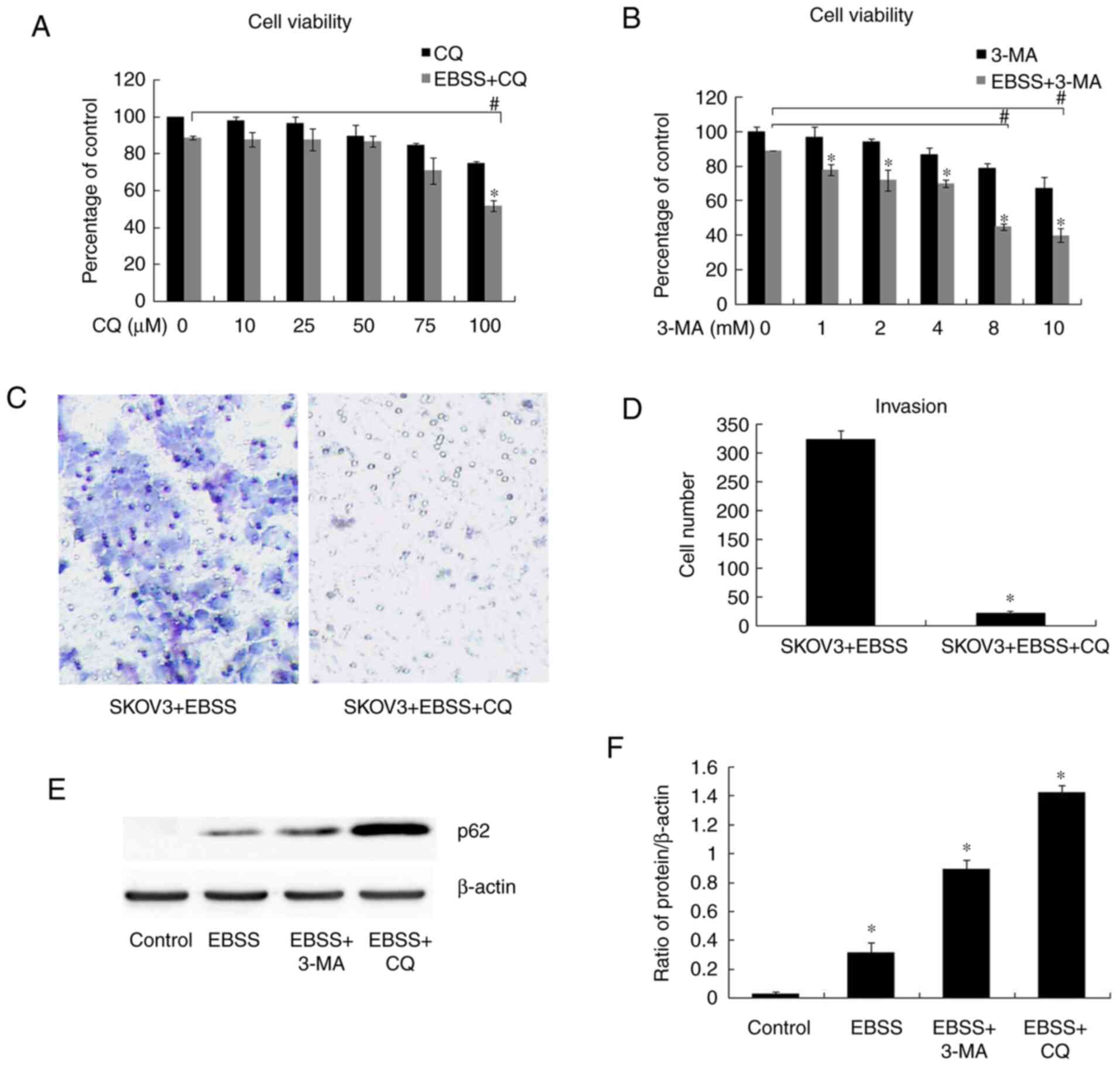
effects of autophagy in the regulation of cell migration and
invasion, the early autophagy inhibitor, 3-MA, and the late
autophagy inhibitor, CQ, were used in order to block autophagy. The
results of cell viability assay revealed that there was no
significant difference in cell viability between EBSS and EBSS + CQ
(50 µM)/EBSS and EBSS + 3-MA (4 mM) (Fig. 2A and B); the appropriate concentration of CQ and
3-MA (50 µM and 4 mM, respectively) was then selected to perform
the following assays. According to the results of western blot
analysis, autophagy was blocked by both 3-MA (4 mM) and CQ (50 µM),
as evidenced by the increased expression of p62 (Fig. 2E and F). Transwell assay for cell invasion
detection revealed that EBSS-induced invasion was inhibited when
autophagy was blocked by CQ (50 µM) (Fig. 2C and D), while there was no significant
difference in cell viability between EBSS and EBSS + CQ (50 µM)
treatment (Fig. 2A). The
aforementioned results indicated that autophagy inhibition reduced
human ovarian cancer cell invasiveness.
Autophagy stimulates invadopodia
formation
In an attempt to fully understand the association
between autophagy and invasion, the formation of invadopodia, which
are critical structures associated with tumor invasion, was
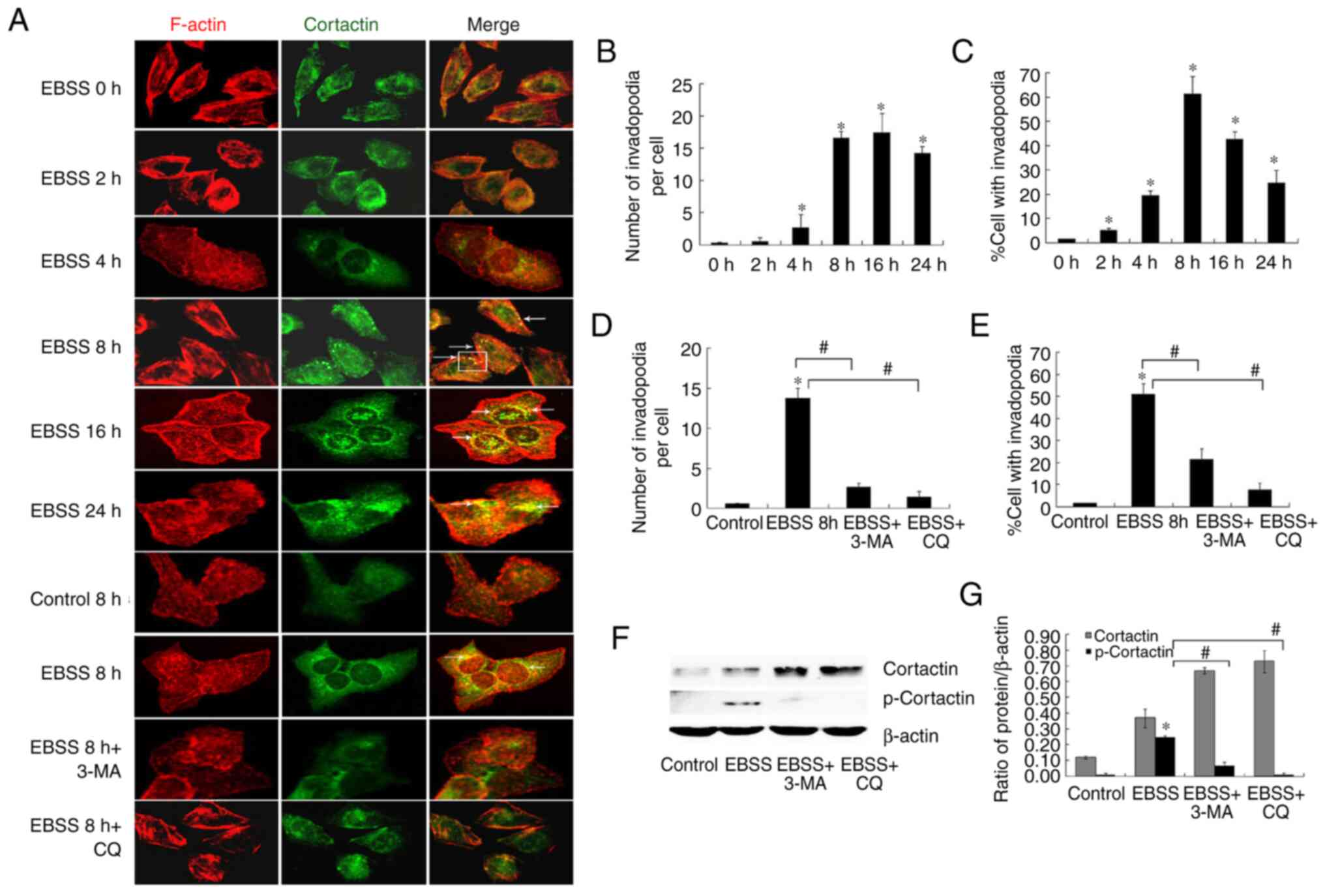
examined. Invadopodia formation is a dynamic process, characterized
by the co-localization of cortactin and F-actin in punctate or
rosette structures (6,7). The co-localization of F-actin and
cortactin as compared with control cells was observed (Fig. 3A). The application of EBSS treatment
for 8 h stimulated invadopodia formation more robustly than a
shorter 2-h treatment. The use of an autophagy inhibitor reduced
the number of cells with active invadopodia (Fig. 3A, D
and E). Moreover, treatment of the
cells with 3-MA or CQ blocked EBSS-induced invadopodia formation,
indicating the autophagy implication in the regulation of this
process.
Invadopodia formation is dependent on the activity
of cortactin, and cortactin phosphorylation is also associated with
the increased activity and maturation of invadopodia (8,9). In
the present study, it was observed that the stimulation of SKOV3
cells with EBSS increased cortactin phosphorylation. By contrast,
although the protein level of cortactin increased when autophagy
was blocked, cortactin phosphorylation was inhibited (Fig. 3F and G).
p62 is closely associated with
invadopodia formation in human ovarian cancer SKOV3 cells
p62 undergoes dynamic changes that accompany
different autophagy conditions and interacts with key components of
various signaling mechanisms. As a consequence, the present study
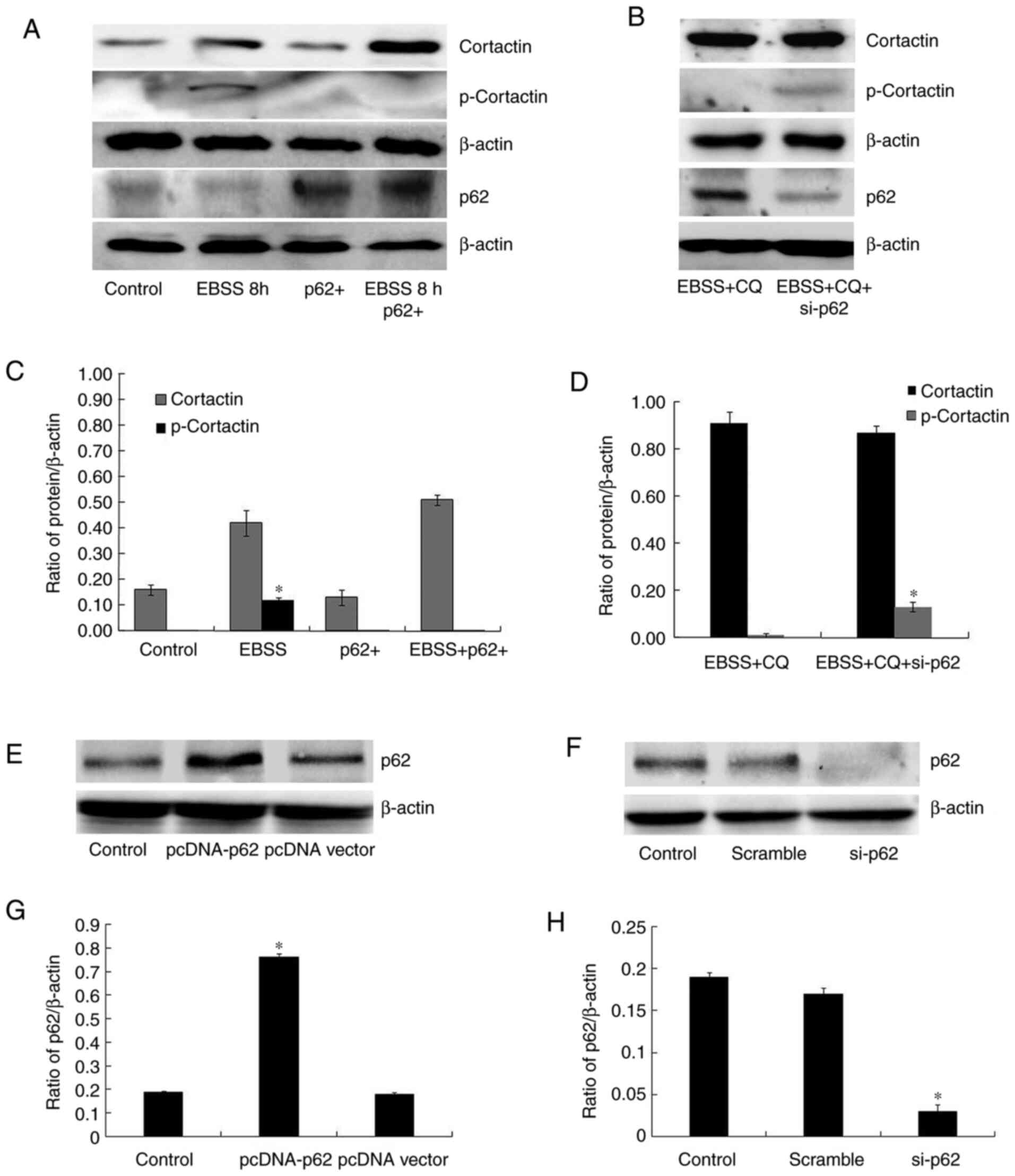
then explored the role of p62 in invadopodia formation. As shown in
Fig. 2E, p62 degradation was
observed along with autophagy following EBSS treatment for 8 h;
conversely, p62 accumulated when autophagy was blocked. It was also
demonstrated that cortactin phosphorylation was reduced in
p62-overexpressing cells, following 8 h of EBSS treatment (Fig. 4A and C), while the depletion of p62 in
autophagy-blocked cells partially recovered cortactin
phosphorylation (Fig. 4B and
D). Fig. 4E-H showed that the transfections
were successful. Although the presence or absence of EBSS resulted
in unequal cortactin protein levels, these results revealed that
the increased expression of cortactin was the stress response of
cells to the starvation environment, while p62 regulated cortactin
phosphorylation. Moreover, it was observed that exogenous p62
inhibited autophagy-induced invadopodia formation, while p62
depletion in autophagy-blocked cells partially recovered the
ability of invadopodia formation (Fig.
5A-C). Taken together, these results suggest that the p62
quantity is key to invadopodia formation regulation in
autophagy.
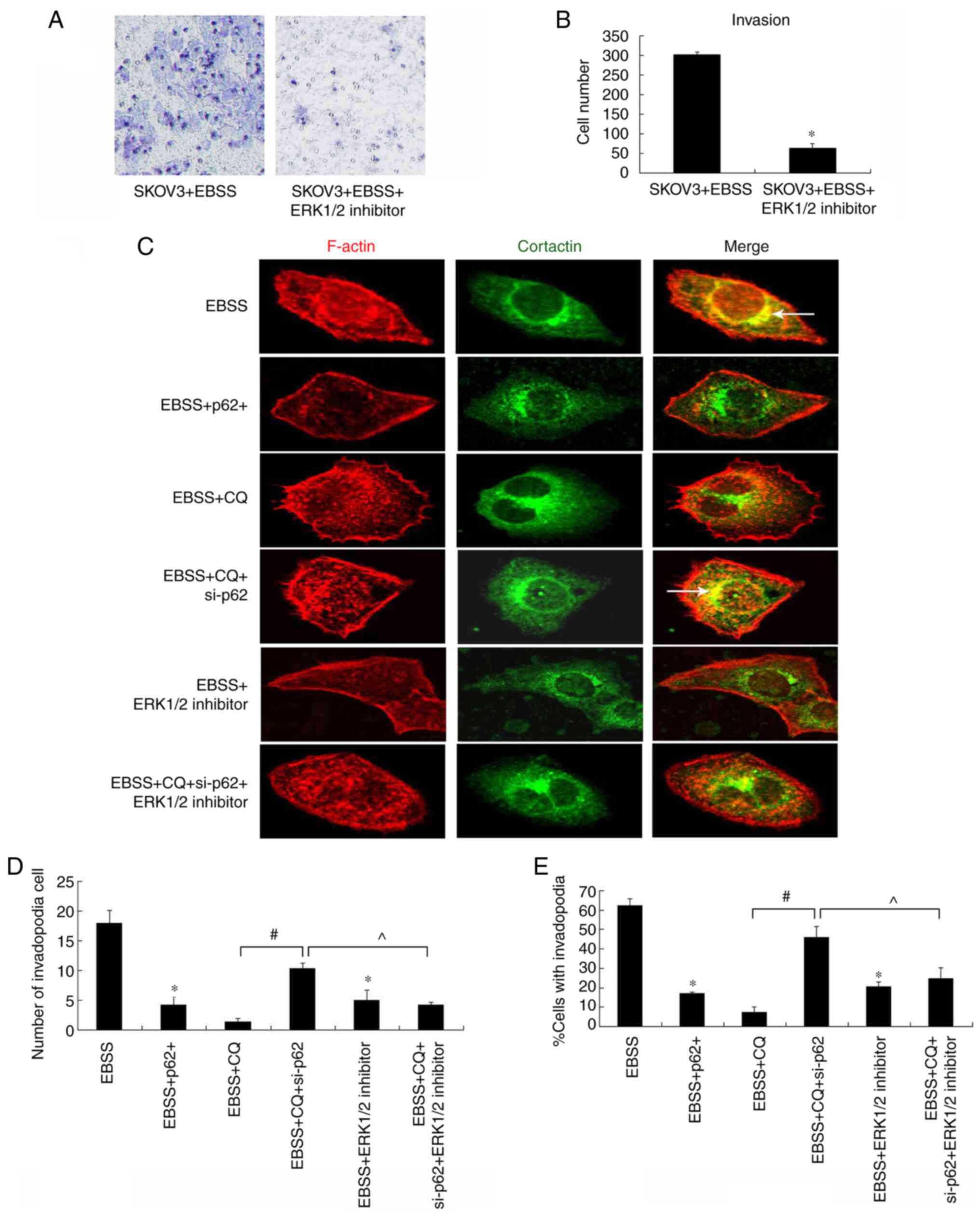
p62 regulates ERK1/2 to function in
autophagy-induced invadopodia formation
Previous studies have demonstrated that ERK1/2
promotes cortactin phosphorylation (9,10), and
that the p62 PB1 domain interacts with ERK1/2, in order to inhibit
its activity (23). To further
investigate the mechanisms of p62 in the regulation of invadopodia
formation, the effect of an ERK1/2 inhibitor on cell invasion and
invadopodia formation was examined. As shown in Fig. 5, the ERK1/2 inhibitor reduced
autophagy-induced cell invasion (Fig.
5A and B) and invadopodia
formation (Fig. 5C-E), indicating
that ERK1/2 is essential for this process. This reduction in
invadopodia formation was recovered by p62 depletion in
autophagy-blocked cells (Fig.
5C).
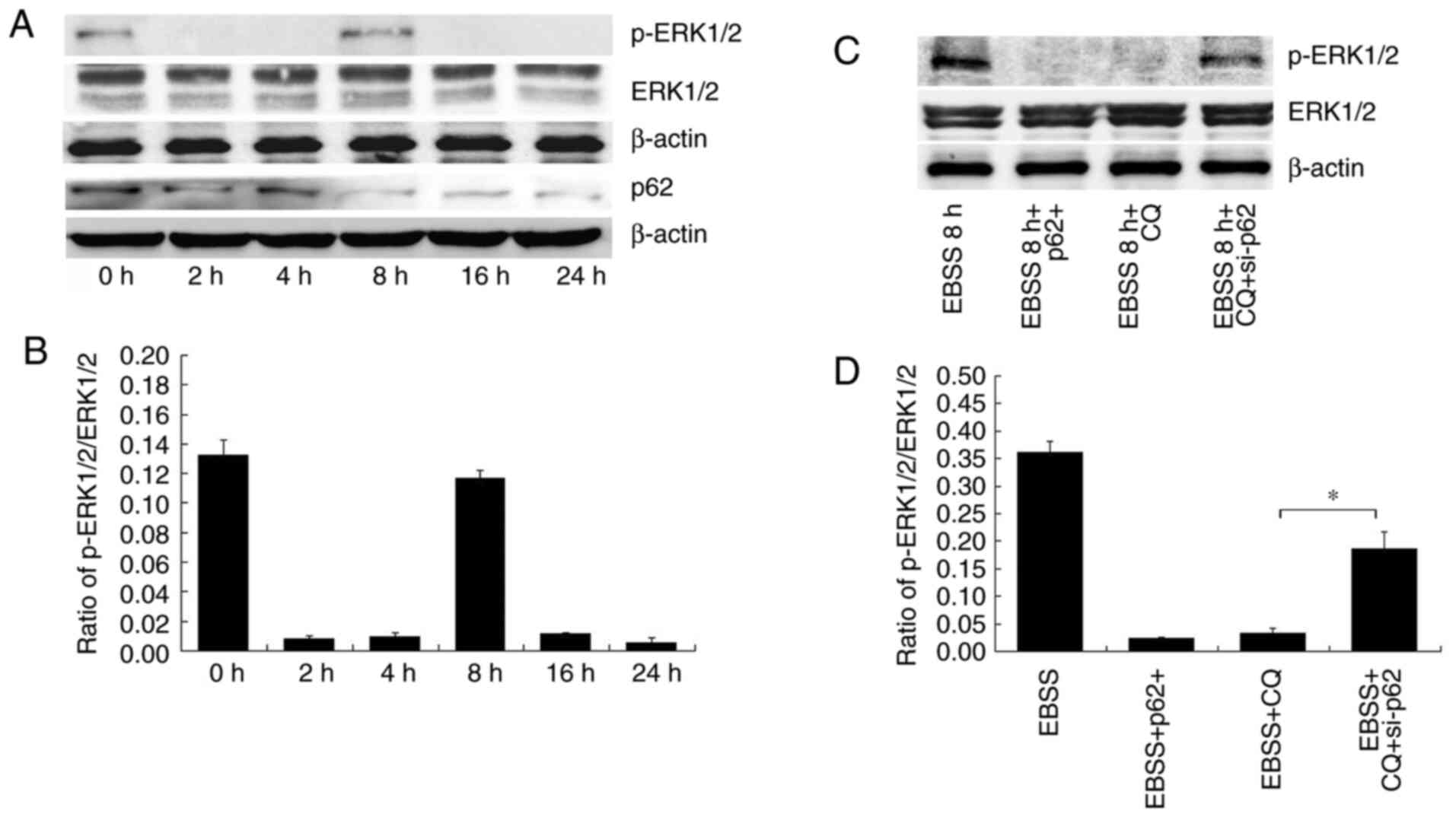
To better understand the mechanisms through which
autophagy regulates invadopodia formation, the expression and
phosphorylation of ERK1/2 in SKOV3 cells subjected to different
treatments were detected. ERK1/2 phosphorylation was shown to vary
at different times of EBSS treatment (Fig. 6A and B).
EBSS treatment initially reduced the phosphorylation
of ERK1/2, an effect which was reversed in due time. p-ERK1/2
expression was increased after 8 h of treatment with EBSS (Fig. 6A and B); this finding is consistent with the
results shown in Fig. 3, according
to which the invadopodia formation detection commenced at 8 h.
Subsequently, the p-ERK1/2 levels were observed at different p62
expression levels. In EBSS 8 h + p62+ cells (cells
overexpressing p62 and treated with EBSS), p-ERK1/2 levels
decreased. In the EBSS 8 h + CQ-treated cells, p-ERK1/2 levels
decreased. In the EBSS 8 h + CQ + si-p62 cells, p-ERK1/2 expression
levels increased. (Fig. 6C and
D). According to the aforementioned
results, ERK1/2 phosphorylation was regulated by p62 in
autophagy-induced invadopodia formation.
Discussion
The present study aimed to elucidate the mechanisms
through which autophagy regulates invadopodia formation,
implicating p62 and ERK1/2 in this process. Although there is some
controversy about the interactions between autophagy and
metastasis, to the best of our knowledge, this is the first
demonstration of the effect of autophagy on invadopodia.
ECM degradation is considered a key step in
promoting tumor invasion and metastasis. Extensive studies have
largely focused on secreted matrix metalloproteinases as key
proteases in tumor invasion (24-26).
More recently, invadopodia have been shown to be critical
structures associated with tumor invasion (27) that restrict protease activity to
areas of the cell in direct contact with the ECM, thus precisely
controlling cell invasion in vivo. In the present study, it
was demonstrated that autophagy is both necessary and sufficient to
promote invadopodia formation. Furthermore, the existence of
important regulatory mechanisms between autophagy and invasion has
been elucidated.
The mechanisms through which invadopodia formation
is regulated at the molecular level are poorly understood.
Cortactin has been reported as a pre-requisite for invadopodia
stabilization and maturation. Furthermore, in previous studies,
cortactin phosphorylation has been shown to result in actin
assembly for the direct formation of actin-rich invadopodia puncta
(8,28). ERK1/2, Src and certain other kinases
may be implicated in cortactin phosphorylation and the promotion of
invadopodia formation (10,11,29).
The formation of invadopodia includes structural maturation and
functional maturation. In the present study, the structural
maturation of invadopodia was depicted by invadopodia formation
assay. In addition, the functional maturation of invadopodia was
illustrated by Transwell invasion assay indirectly. However, the
lack of fluorescent gelatin-degradation assay is a limitation of
the present study. In the present study, it was also found that p62
may be a negative regulator of invadopodia formation in human
ovarian cancer SKOV3 cells. In a nutritionally deficient
environment, autophagy directly induces the activation of ERK1/2
through p62, thus promoting cortactin phosphorylation and
invadopodia formation. The authors aim to further investigate the
mechanism through which autophagy acts on invadopodia in future
research.
p62 was initially isolated, and shown to be mainly
interacting with atypical protein kinase C. It functions as a
selective autophagy receptor that recognizes and shuttles
ubiquitinated proteins to the autophagosome for subsequent
degradation in autophagy (10).
Previous studies have revealed that p62 acts as a multidomain
signaling hub through its ability to recruit, organize, and
oligomerize important signaling molecules in cytosolic speckles, in
order to control cell survival and apoptosis; moreover, the
elimination of p62 by autophagy has been reported to suppress
tumorigenesis (13,15,17,30,31).
The present study provides evidence of the role of p62 as an
intermediary between autophagy and invasion. The fluctuation of p62
protein levels is dynamic during autophagy. It is described as the
result of the interaction between two aspects: On the one hand, the
p62 mRNA expression levels increased when cells were treated with
EBSS, on the other hand, p62 degrades as autophagy progresses
further (16,17,22,23,30-32).
In addition, ERK1/2 is a crucial kinase which participates in many
cell activities, such as cell division, cell adhesion and
apoptosis. . The variation in p-ERK1/2 levels may be attributed to
their different location, combination and aggregation state in
cells (10,33-35).
As a result, the regulatory mechanism is a complex phenomenon.
Additionally, according to other previous studies, p62 expression
exerts an inhibitory effect on ERK1/2 activity (13,23,36-38).
Little is known about the functional association
between autophagy and invadopodia formation. Autophagy and invasion
are different cell activities, and following EBSS treatment, their
occurrences are not synchronous. Autophagy initially occurs,
digesting part of the intracellular structure in order to ensure
survival. Afterwards, invasion leads to the evasion of malignant
cells from the adverse environment. This was a speculated
self-protective mechanism of tumor cells in the present study,
which has been initially proved through the experiments. Autophagy
was detected 2 h after EBSS treatment, and invadopodia formation
was detected 8 h later. LC3 II marker level variance was used to
evaluate autophagy, and a minimal increase in autophagy was
detected at 24 h, the authors aim to investigate this further in
future studies, in order to elucidate whether there are other
mechanisms involved. In addition, under various stimuli and at
different time points, ERK1/2 participates in a number of cell
activities. As a result, p62 expression did not decrease up to a
certain level earlier than 8 h, and ERK1/2 was not activated, a
conclusion consistent with the invadopodia formation detection
after the application of 8-h EBSS treatment.
Under conditions of nutritional deficiency, tumor
cells enhance autophagy to provide nutrition for survival (20,21,39),
thus invading other tissues and organs to escape the
nutrient-deficient environment. The present study demonstrated that
autophagy may not only help cell survival, but may also increase
tumor cell invasiveness under nutrient-deficient conditions. The
interaction between autophagy and invasion may be a self-protective
mechanism for tumor cells in a harmful environment. The underlying
mechanisms of the autophagy and invasion interaction are being
actively explored by the authors with the aim of gaining new
insight regarding the prevention of tumor metastasis.
Acknowledgements
Not applicable.
Funding
Funding: The present study was supported by National Natural
Science Foundation of China (grant nos. 81772794, 81672948,
81472419), Jilin Provincial Industrial Innovation Project (grant
no. 2018C052-7) and the Fundamental Research Funds for the Central
Universities, JLU.
Availability of data and materials
All data generated or analyzed during this study are
included in this published article.
Authors' contributions
ZZ and JZ performed the experiments. YL, XY and HS
collected and analyzed the data, MX and JS designed the study. All
authors contributed to writing and reviewing the manuscript. ZZ and
JZ confirm the authenticity of all the raw data. All authors have
read and approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
De Bock K, Mazzone M and Carmeliet P:
Antiangiogenic therapy, hypoxia, and metastasis: Risky liaisons, or
not? Nat Rev Clin Oncol. 8:393–404. 2011.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Fulda S and Kögel D: Cell death by
autophagy: Emerging molecular mechanisms and implications for
cancer therapy. Oncogene. 34:5105–5113. 2015.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Hou J, Han Z, Zhao N and Wei L: Autophagy
and tumour metastasis. Adv Exp Med Biol. 1207:315–338.
2020.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Perez-Montoyo H: Therapeutic potential of
autophagy modulation in cholangiocarcinoma. Cells.
9(614)2020.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Garcia J, Hurwitz HI, Sandler AB, Miles D,
Coleman RL, Deurloo R and Chinot OL: Bevacizumab (Avastin®) in
cancer treatment: A review of 15 years of clinical experience and
future outlook. Cancer Treat Rev. 86(102017)2020.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Murphy DA and Courtneidge SA: The ‘ins’
and ‘outs’ of podosomes and invadopodia: characteristics, formation
and function. Nat Rev Mol Cell Biol. 12:413–426. 2011.PubMed/NCBI View
Article : Google Scholar
|
|
7
|
Leong HS, Robertson AE, Stoletov K, Leith
SJ, Chin CA, Chien AE, Hague MN, Ablack A, Carmine-Simmen K,
McPherson VA, et al: Invadopodia are required for cancer cell
extravasation and are a therapeutic target for metastasis. Cell
Rep. 8:1558–1570. 2014.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Clark ES, Whigham AS, Yarbrough WG and
Weaver AM: . Cortactin is an essential regulator of matrix
metalloproteinase secretion and extracellular matrix degradation in
invadopodia. Cancer Res. 67:4227–4235. 2007.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Goertzen CG, Dragan M, Turley E, Babwah AV
and Bhattacharya M: KISS1R signaling promotes invadopodia formation
in human breast cancer cell via β-arrestin2/ERK. Cell Signal.
28:165–176. 2016.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Kelley LC, Hayes KE, Ammer AG, Martin KH
and Weed SA: Revisiting the ERK/Src cortactin switch. Commun Integr
Biol. 4:205–207. 2011.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Samuelson DR and Konkel ME: Serine
phosphorylation of cortactin is required for maximal host cell
invasion by Campylobacter jejuni. Cell Commun Signal.
11(82)2013.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Navratil AM, Dozier MG, Whitesell JD, Clay
CM and Roberson MS: Role of cortactin in dynamic actin remodeling
events in gonadotrope cells. Endocrinology. 155:548–557.
2014.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Moscat J and Diaz-Meco MT: p62: a
versatile multitasker takes on cancer. Trends Biochem Sci.
37:230–236. 2012.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Anand PK, Tait SW, Lamkanfi M, Amer AO,
Nunez G, Pagès G, Pouysségur J, McGargill MA, Green DR and
Kanneganti TD: TLR2 and RIP2 pathways mediate autophagy of Listeria
monocytogenes via extracellular signal-regulated kinase (ERK)
activation. J Biol Chem. 286:42981–42991. 2011.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Ciuffa R, Lamark T, Tarafder AK, Guesdon
A, Rybina S, Hagen WJ, Johansen T and Sachse C: The selective
autophagy receptor p62 forms a flexible filamentous helical
scaffold. Cell Rep. 11:748–758. 2015.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Matsumoto G, Wada K, Okuno M, Kurosawa M
and Nukina N: Serine 403 phosphorylation of p62/SQSTM1 regulates
selective autophagic clearance of ubiquitinated proteins. Mol Cell.
44:279–289. 2011.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Ishimura R, Tanaka K and Komatsu M:
Dissection of the role of p62/Sqstm1 in activation of Nrf2 during
xenophagy. FEBS Lett. 588:822–828. 2014.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Mohamed NV, Plouffe V, Rémillard-Labrosse
G, Planel E and Leclerc N: Starvation and inhibition of lysosomal
function increased tau secretion by primary cortical neurons. Sci
Rep. 4(5715)2014.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Barutcu SA, Girnius N, Vernia S and Davis
RJ: Role of the MAPK/cJun NH2-terminal kinase signaling pathway in
starvation-induced autophagy. Autophagy. 14:1586–1595.
2018.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Chang L, Chai X, Chen P, Cao J, Xie H and
Zhu J: miR-181b-5p suppresses starvation-induced cardiomyocyte
autophagy by targeting Hspa5. Int J Mol Med. 43:143–154.
2019.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Zhang Y, Ren S, Liu Y, Gao K, Liu Z and
Zhang Z: Inhibition of Starvation-Triggered Endoplasmic Reticulum
Stress, Autophagy, and Apoptosis in ARPE-19 Cells by Taurine
through Modulating the Expression of Calpain-1 and Calpain-2. Int J
Mol Sci. 18(E2146)2017.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Yan XY, Zhang Y, Zhang JJ, Zhang LC, Liu
YN, Wu Y, Xue YN, Lu SY, Su J and Sun LK: p62/SQSTM1 as an
oncotarget mediates cisplatin resistance through activating
RIP1-NF-κB pathway in human ovarian cancer cells. Cancer Sci.
108:1405–1413. 2017.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Moscat J and Diaz-Meco MT: p62 at the
crossroads of autophagy, apoptosis, and cancer. Cell.
137:1001–1004. 2009.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Roy R, Morad G, Jedinak A and Moses MA:
Metalloproteinases and their roles in human cancer. Anat Rec
(Hoboken). 303:1557–1572. 2020.PubMed/NCBI View
Article : Google Scholar
|
|
25
|
Gonzalez-Avila G, Sommer B,
García-Hernández AA and Ramos C: Matrix metalloproteinases' role in
tumor microenvironment. Adv Exp Med Biol. 1245:97–131.
2020.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Cerofolini L, Fragai M and Luchinat C:
Mechanism and inhibition of matrix metalloproteinases. Curr Med
Chem. 26:2609–2633. 2019.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Eddy RJ, Weidmann MD, Sharma VP and
Condeelis JS: Tumor cell invadopodia: Invasive protrusions that
orchestrate metastasis. Trends Cell Biol. 27:595–607.
2017.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Yin M, Ma W and An L: Cortactin in cancer
cell migration and invasion. Oncotarget. 8:88232–88243.
2017.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Martinez-Quiles N, Ho HY, Kirschner MW,
Ramesh N and Geha RS: Erk/Src phosphorylation of cortactin acts as
a switch on-switch off mechanism that controls its ability to
activate N-WASP. Mol Cell Biol. 24:5269–5280. 2004.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Islam MA, Sooro MA and Zhang P: Autophagic
regulation of p62 is critical for cancer therapy. Int J Mol Sci.
19(E1405)2018.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Moscat J, Karin M and Diaz-Meco MT: p62 in
cancer: Signaling adaptor beyond autophagy. Cell. 167:606–609.
2016.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Sánchez-Martín P, Saito T and Komatsu M:
p62/SQSTM1: ‘Jack of all trades’ in health and cancer. FEBS J.
286:8–23. 2019.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Cook SJ, Stuart K, Gilley R and Sale MJ:
Control of cell death and mitochondrial fission by ERK1/2 MAP
kinase signalling. FEBS J. 284:4177–4195. 2017.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Sun Y, Liu W-Z, Liu T, Feng X, Yang N and
Zhou H-F: Signaling pathway of MAPK/ERK in cell proliferation,
differentiation, migration, senescence and apoptosis. J Recept
Signal Transduct Res. 35:600–604. 2015.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Roskoski R Jr: Targeting ERK1/2
protein-serine/threonine kinases in human cancers. Pharmacol Res.
142:151–168. 2019.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Gao H, Zhang Y, Dong L, Qu XY, Tao LN,
Zhang YM, Zhai JH and Song YQ: Triptolide induces autophagy and
apoptosis through ERK activation in human breast cancer MCF-7
cells. Exp Ther Med. 15:3413–3419. 2018.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Hou XO, Si JM, Ren HG, Chen D, Wang HF,
Ying Z, Hu QS, Gao F and Wang GH: Parkin represses
6-hydroxydopamine-induced apoptosis via stabilizing scaffold
protein p62 in PC12 cells. Acta Pharmacol Sin. 36:1300–1307.
2015.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Li C and Siragy HM: Autophagy upregulates
(pro)renin receptor expression via reduction of P62/SQSTM1 and
activation of ERK1/2 signaling pathway in podocytes. Am J Physiol
Regul Integr Comp Physiol. 313:R58–R64. 2017.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Gatica D and Klionsky DJ: Towards
understanding mRNA-binding protein specificity: Lessons from
post-transcriptional regulation of ATG mRNA during nitrogen
starvation-induced autophagy. Curr Genet. 65:847–849.
2019.PubMed/NCBI View Article : Google Scholar
|