Introduction
Myocardial ischemia, a consequence of the blockage
of arteries, is believed to be a common and major cause of high
mortality in patients with heart disease worldwide (1). Reperfusion of the ischemic area is an
effective way to save patients, but it can potentially lead to
additional heart damage termed myocardial ischemia/reperfusion
(I/R) injury (2). Myocardial I/R
injury usually contributes to the exacerbation of heart disease by
causing cardiac arrhythmias, myocardial infarction, and even sudden
death (3). To date, the application
of ischemic postconditioning is believed to be a promising
cardioprotective intervention that can reduce myocardial I/R injury
(4,5). One commonly used in vitro model
for the investigation of the mechanisms of I/R injury is the
hypoxia/reoxygenation (H/R) injury model (6-8).
Propofol is a widely adopted intravenous anesthetic.
It functions as a novel ischemic postconditioning agent by
exhibiting a protective capability against myocardial I/R injury
(9-12).
It is reported that propofol possesses antioxidant capacity
(13) and free radical scavenging
activity (14) and thus ameliorates
myocardial I/R injury (15).
Therefore, the characterization of the molecular mechanism of
propofol in exerting cardioprotection is of great significance and
could provide novel applications for the treatment of myocardial
I/R injury. In the present study, H9C2 cells (embryonic rat
heart-derived) were cultured and exposed to H/R injury to mimic the
I/R challenge and gain insights into the mechanism of propofol.
The relationship between microRNAs (miRNAs/miRs) and
myocardial I/R injury is reported in emerging available data.
Previous studies revealed that the concentrations of
miR-145(16), miR-327(17), and miR-29(18) were enriched in myocardial I/R
injury, and suppression of these miRNAs improved the effect on the
cardiomyocytes with I/R injury. Apart from these miRNAs, miR-449a
has also been well established as a factor that aggravates
hypoxia-induced damage of cardiomyocytes (19). In line with this, the detrimental
effect of miR-449a on H9C2 cells subjected to H/R injury was also
demonstrated in another study (20). Based on these previous reports, the
present study attempted to gain a deeper insight into the role of
miR-449a in myocardial I/R injury through its relationship with
propofol; to the best of our knowledge, its involvement remains
unclear.
Nuclear receptor subfamily 4 group A member 2
(NR4A2) is often studied in Parkinson's disease due to its roles in
neuroprotection and its symptomatic improvement (21). The NR4A family consists of three
members: NR4A1, NR4A2, and NR4A3. They are expressed in
cardiomyocytes (22) and quickly
react in response to ischemic stroke (23). A previous study reported that NR4A2
was enhanced in cardiomyocytes subjected to ischemia and exerted a
suppressive effect on ischemia-induced apoptosis, which was
regulated by miR-212-3p (24). The
present study focused on the role of NR4A2 in H9C2 cells exposed to
H/R challenge in the presence of propofol and following miR-449a
expression manipulation. The current results may provide novel
insights into the cardioprotective mechanism of propofol against
I/R injury.
Materials and methods
Bioinformatics analysis
The GSE4386 dataset (25) (from the GEO DataSets; https://www.ncbi.nlm.nih.gov/gds) is a previously
published mRNA microarray analysis involving atrial samples with or
without propofol. The upregulated differentially expressed genes
(DEGs) in atrial samples with propofol were determined by limma
3.26.8 (Bioconductor) using P<0.05 and log [fold change (FC)]≥2.
The STRING (https://string-db.org/) database was
then used to construct the interaction network for the top 20
upregulated DEGs to identify the key genes. Finally, the Tarbase
(http://carolina.imis.athena-innovation.gr/diana_tools/web/index.php?r=tarbasev8/index)
and miRDB (http://mirdb.org/) databases were used
to predict the potential upstream miRNAs of the key gene.
Cell culture and H/R challenge
The H9C2 (embryonic rat heart-derived cell) cell
line was purchased from the American Type Culture Collection and
cultured in DMEM medium (Sigma-Aldrich; Merck KGaA) with 10% fetal
bovine serum (Gibco; Thermo Fisher Scientific, Inc.) and 1%
penicillin/streptomycin (Invitrogen; Thermo Fisher Scientific,
Inc.) in an incubator with 5% CO2 at 37˚C. To simulate
I/R injury, H/R injury was induced by placing H9C2 cells in hypoxic
conditions (1% O2, 95% N2, and 5%
CO2) for 12 h. Then, reoxygenation was achieved by
transferring the cells to normal culture conditions for 6 h. Cells
in the control group were cultured normally. Propofol with
concentrations of 0, 25, 50, and 100 µmol/l was added into the
fresh medium before H/R challenge, and cell viability was detected
by the CCK-8 assay. The dose of 50 µmol/l propofol was selected for
subsequent experimental treatments.
Cell transfection
miR-449a inhibitor (cat. no. miR20001541-1-5),
miR-449a mimic (cat. no. miR10001541-1-5), corresponding negative
controls (mimic NC, cat. no. miR1N0000003-1-5 and inhibitor-NC,
cat. no. miR2N0000003-1-5), NR4A2-targeting small interfering (si-)
RNA (cat. no. siB14428110628-1-5), and non-targeting control siRNA
(si-NC; cat. no. siN0000001-1-5) (all from Guangzhou RiboBio Co.,
Ltd.) were transfected into the H9C2 cells using the Lipofectamine™
2000 reagent (Thermo Fisher Scientific, Inc.) at 37˚C according to
the manufacturer's protocol. Transfections were conducted when
cells were at ~70% confluency. The transfection concentrations were
3 pmol for a 96-well plate, 30 pmol for a 12-well plate and 75 pmol
for a 6-well plate. Subsequent experiments were performed at 48 h
following transfection.
Reverse transcription-quantitative PCR
(RT-qPCR)
Total RNA was extracted using the TRIzol reagent
(Invitrogen; Thermo Fisher Scientific, Inc.). To analyze miR-449a
expression levels, reverse transcription was conducted using the
All-in-One miRNA First-Strand cDNA Synthesis kit (GeneCopoeia,
Inc.) and qPCR was performed with the All-in-One miRNA qRT-PCR
Detection kit 2.0 (GeneCopoeia, Inc.). The thermocycling conditions
were: 95˚C initial denaturation for 10 min, followed by 40 cycles
of 95˚C denaturation for 10 sec, 60˚C annealing for 20 sec, and
72˚C extension for 10 sec. U6 served as the internal control for
miR-449a. To analyze NR4A2 mRNA expression levels, FastStart
Universal SYBR Green Master (ROX) kit (Roche Diagnostics) was used.
A total of 1 µl of the enzyme mix was used for 25 µl of each RT-PCR
reaction. The reverse transcription step was performed at 16˚C for
30 min without the RT-PCR enzyme mix, then at 42˚C for 40 min with
the enzyme mix, followed by 30 cycles of amplification at 95˚C for
20 sec and 60˚C for 1 min. GAPDH served as the internal control for
mRNA detection. qPCR was performed in an ABI 7300 thermocycler
(Applied Biosystems; Thermo Fisher Scientific, Inc.) and the
2−ΔΔCq method (26) was
used for quantification of the results. The primer sequences are
listed in Table I.
 | Table ISequences of primers used in reverse
transcription-quantitative PCR. |
Table I
Sequences of primers used in reverse
transcription-quantitative PCR.
| Gene | Primer | Sequence
(5'-3') |
|---|
| miR-449a | Forward |
GCAGTGTATTGTTAGCTG |
| | Reverse |
GAACATGTCTGCGTATCTC |
| U6 | Forward |
CTCGCTTCGGCAGCACA |
| | Reverse |
AACGCTTCACGAATTTGCGT |
| NR4A2 | Forward |
AAACTGCCCAGTGGACAAGCGT |
| | Reverse |
GCTCTTCGGTTTCGAGGGCAAA |
| GAPDH | Forward |
CACCATTGGCAATGAGCGGTTC |
| | Reverse |
AGGTCTTTGCGGATGTCCACGT |
Cell counting kit-8 (CCK-8) assay
Cell viability was assessed by the CCK-8 assay
(MedChemExpres). H9C2 cells were plated in 96-well plates at
2x103 cells/well. The detection time points were set at
1, 6, 12, 24, and 48 h post-transfection. For detection, cells were
treated with 100 µl of the CCK-8 solution for 4 h, and optical
density was measured using a microplate reader (BioTek Instruments,
Inc.) at the wavelength of 450 nm.
BrdU assay
The proliferation of H9C2 cells was detected by the
Cell Proliferation Assay kit (Cell Signaling Technology, Inc.).
Briefly, 10 µM of BrdU solution was added into each well
(containing 2x103 H9C2 cells/well) in a 96-well plate
and incubated for 2 h. Next, the absorbance at 450 nm was
determined using a microplate reader (BioTek Instruments,
Inc.).
EdU assay
H9C2 cells were seeded in 96-well plates at a
density of 5x103 cells/well for 48 h and cell
proliferation was evaluated using the Cell-Light EdU Apollo567 kit
(Guangzhou RiboBio Co., Ltd.). Briefly, 50 µmol/l of EdU was added
into the cell culture medium and incubated for 2 h. The cells were
then treated with 4% paraformaldehyde for 30 min and 0.5% Triton
X-100 for 20 min. After washing with PBS, the cells were stained at
room temperature for 30 min with the anti-EdU working solution and
incubated with 100 µl of the Hoechst 33342 stain (5 µg/ml).
Finally, a fluorescence microscope (Olympus Corporation) was used
to capture images of the cells, and the percentage of EdU-positive
cells was calculated from 5 optical fields/well.
Caspase-3 activity assay
Cell apoptosis was evaluated using the Caspase-3
Activity Assay kit (Cell Signaling Technology, Inc.). H9C2 cells
(2x103 cells/well) were cultured in 96-well plates.
Briefly, the collected cells were lysed with the lysis buffer.
Then, 0.2 mM of the caspase-3 substrate Ac-DEVD-pNA was added into
the cell supernatant obtained from high-speed centrifugation and
incubated for 2 h at 37˚C. Finally, caspase-3 activity was
determined at 405 nm using a microplate reader (BioTek Instruments,
Inc.).
Dual-luciferase reporter assay
The NR4A2 wild-type (wt) sequence containing the
predicted miR-449a binding sites, and a NR4A2 mutant (mut) sequence
containing the altered miR-449a binding sites, were amplified by
PCR and subcloned into the pGL3 luciferase reporter vectors
(Promega Corporation). The mutant miR-449a binding sites were
produced by replacing GUGACGG with CCCUUAA. Next, the constructed
NR4A2 wt or NR4A2 mut reporters were co-transfected into the H9C2
cells together with the miR-449a mimic or the negative control
using the Lipofectamine 2000 reagent (Thermo Fisher Scientific,
Inc.). After 48 h of the co-transfection, the relative luciferase
activity was determined using the dual-luciferase reporter assay
(Promega Corporation). Firefly luciferase activity was normalized
to Renilla luciferase activity.
Western blotting
Cells were lyzed by using an ice-cold RIPA buffer
(Thermo Fisher Scientific, Inc.) containing the protease inhibitor
phenylmethylsulfonyl fluoride. The concentrations of total proteins
were analyzed using a bicinchoninic acid protein assay kit (Thermo
Fisher Scientific, Inc.). An equal amount of protein (20 µg/lane)
was analyzed by 12% SDS-PAGE for 1-h separation at 100 V. After
separation, the proteins were transferred onto polyvinylidene
difluoride membranes and then blocked in 5% skimmed milk for 2 h at
room temperature. After blocking, the membranes were incubated
overnight at 4˚C with primary antibodies against NR4A2 (cat. no.
bs-0178R; 1:2,000; Bioss), Bax (cat. no. bs-20377R; 1:1,000;
Bioss), Bcl-2 (cat. no. bs-20351R; 1:2,000; Bioss), cleaved
caspase-3 (cat. no. ab32042; 1:500; Abcam), total caspase-3 (cat.
no. ab32351; 1:5,000; Abcam), and GAPDH (cat. no. ab9485; 1:2,000;
Bioss). After washing, the membranes were treated with horseradish
peroxidase-conjugated goat anti-rabbit IgG secondary antibody (cat.
no. ab6721; 1:1,0000; Bioss) for 1 h at 37˚C. The protein bands
were detected in an Odyssey CLx Infrared Imaging System (LI-COR
Biosciences). The images were analyzed using ImageJ 1.8.0 (National
Institutes of Health).
Statistical analysis
Data from three independent experiments were
represented as the mean ± SD. All statistical tests were performed
using GraphPad Prism version 8.0 (GraphPad Software, Inc.). One-way
ANOVA followed by Dunnett's multiple comparisons was used to
analyze the differences among multiple experimental groups.
Unpaired student's t-test was used to analyze the differences
between two experimental groups. P<0.05 was considered to
indicate a statistically significant difference.
Results
Identification of the genes of
interest in the present study
First, the top 20 upregulated DEGs from the GSE4386
dataset were acquired with the criteria of adjusted P-value
<0.05 and log[FC]≥2 (Table II).
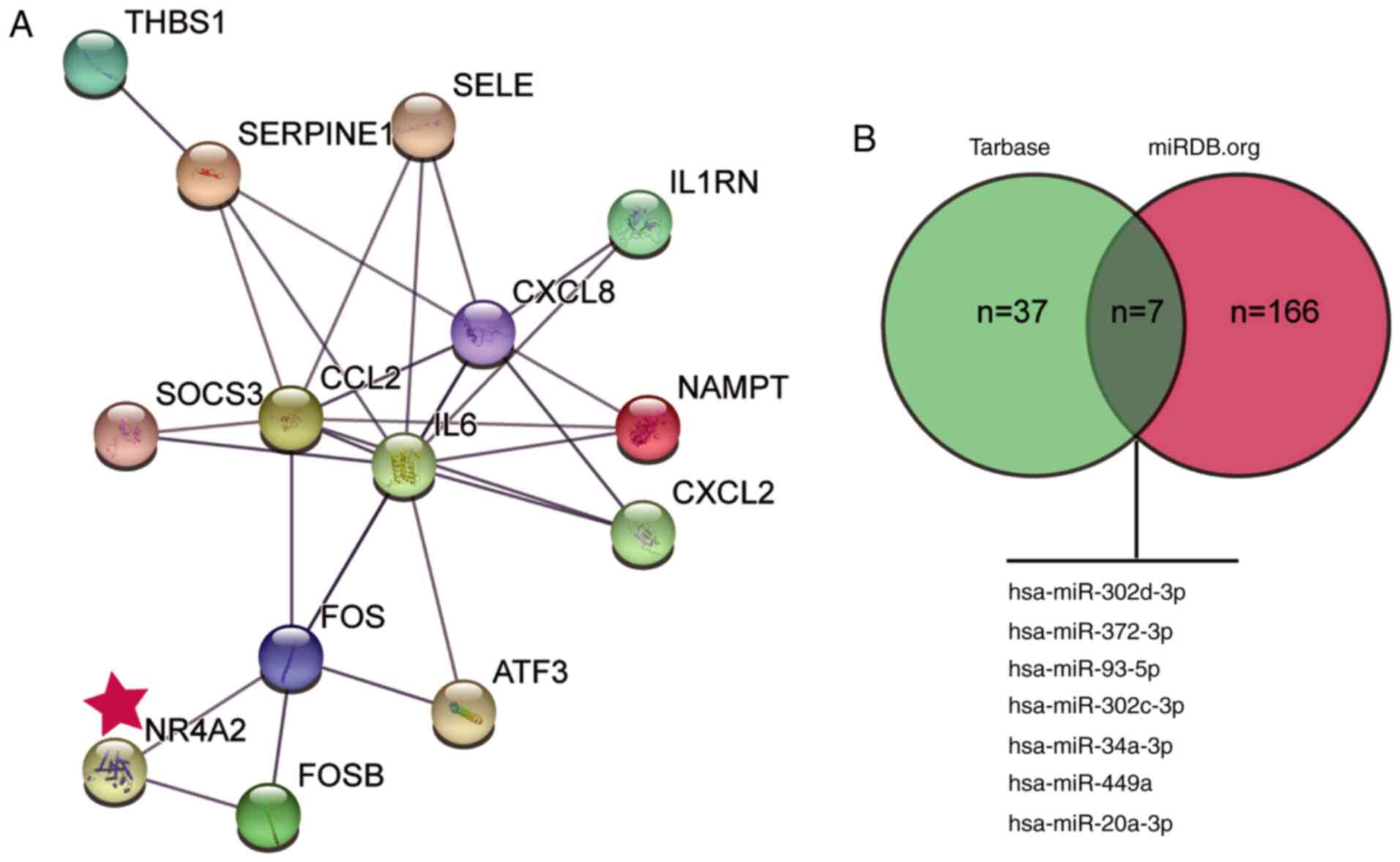
These 20 genes were then used as input for STRING analysis to
discover their interaction network (Fig. 1A). Notably, among the top 14 genes,
RT-qPCR result showed that NR4A2 mRNA expression levels were found
to be the lowest in H9C2 cells treated with H/R compared with
control cells (Table SI). NR4A2
has been studied previously in myocardial infarction injury
(24). Propofol is considered to be
a cardioprotective reagent (9,10).
However, the involvement of NR4A2 in cardioprotection in the
presence of propofol has not been studied yet. To explore the
potential roles of miRNAs in the effects of NR4A2, the potential
miRNAs upstream of NR4A2 were predicted using the Tarbase and miRDB
databases; seven common miRNAs were identified (Fig. 1B). Among them, miR-449a was reported
to enhance the damage of cardiomyocytes (19,20).
However, the involvement of miR-449a in damaging the cardiomyocytes
in the presence of propofol is not clear. Therefore, the present
study aimed to explore the potential interaction between miR-449a
and NR4A2 on cardiomyocyte injury in the presence of propofol.
 | Table IITop 20 differentially expressed genes
from analysis of the GSE4386 data series. |
Table II
Top 20 differentially expressed genes
from analysis of the GSE4386 data series.
| Probe ID | Adjusted
P-value | P-value | Log fold
change | Gene symbol | Gene full name |
|---|
| 205207_at |
2.25x10-15 |
1.24x10-19 | 9.036459 | IL6 | Interleukin 6 |
| 202768_at |
3.99x10-14 |
2.93x10-18 | 8.554949 | FOSB | FosB
proto-oncogene, AP-1 transcription factor subunit |
| 202859_x_at |
2.83x10-12 |
5.19x10-16 | 7.785047 | CXCL8 | C-X-C motif
chemokine ligand 8 |
| 207978_s_at |
3.46x10-21 |
6.34x10-26 | 7.310244 | NR4A3 | Nuclear receptor
subfamily 4 group A member 3 |
| 209189_at |
5.55x10-07 |
1.30x10-09 | 7.065234 | FOS | Fos proto-oncogene,
AP-1 transcription factor subunit |
| 206211_at |
1.48x10-11 |
4.61x10-15 | 7.042325 | SELE | Selectin E |
| 206115_at |
8.23x10-09 |
9.80x10-12 | 6.660009 | EGR3 | Early growth
response 3 |
| 209774_x_at |
5.44x10-11 |
2.69x10-14 | 6.547871 | CXCL2 | C-X-C motif
chemokine ligand 2 |
| 236523_at |
3.40x10-06 |
9.90x10-09 | 6.349735 | LOC285556 | Uncharacterized
LOC285556 |
| 206932_at |
3.68x10-13 |
4.04x10-17 | 6.310222 | CH25H | Cholesterol
25-hydroxylase |
| 207526_s_at |
6.09x10-08 |
1.07x10-10 | 6.109762 | IL1RL1 | Interleukin 1
receptor like 1 |
| 227697_at |
3.71x10-13 |
4.76x10-17 | 6.019778 | SOCS3 | Suppressor of
cytokine signaling 3 |
| 243296_at |
8.50x10-12 |
2.02x10-15 | 5.966462 | NAMPT | Nicotinamide
phosphoribosyltransferase |
| 202628_s_at |
4.18x10-08 |
6.66x10-11 | 5.96427 | SERPINE1 | Serpin family E
member 1 |
| 202672_s_at |
1.25x10-11 |
3.56x10-15 | 5.878868 | ATF3 | Activating
transcription factor 3 |
| 212657_s_at |
7.31x10-09 |
8.17x10-12 | 5.761876 | IL1RN | Interleukin 1
receptor antagonist |
| 201109_s_at |
3.32x10-11 |
1.28x10-14 | 5.700034 | THBS1 | Thrombospondin
1 |
| 1569003_at |
1.19x10-10 |
6.31x10-14 | 5.691674 | VMP1 | Vacuole membrane
protein 1 |
| 216598_s_at |
7.95x10-09 |
9.31x10-12 | 5.687414 | CCL2 | C-C motif chemokine
ligand 2 |
| 216248_s_at |
3.72x10-09 |
3.14x10-12 | 5.600101 | NR4A2 | Nuclear receptor
subfamily 4 group A member 2 |
Propofol treatment exhibits a
protective effect on the survival of the H9C2 cells
The CCK-8 assay was performed in the presence of
different concentrations of propofol (0, 25, 50 and 100 µmol/l) to
detect changes in the cell viability of H/R-induced H9C2 cells. The
dose of 50 µmol/l propofol resulted in the highest cell viability
and protection of H/R-induced H9C2 cells (Fig. S1). Therefore, 50 µmol/l of propofol
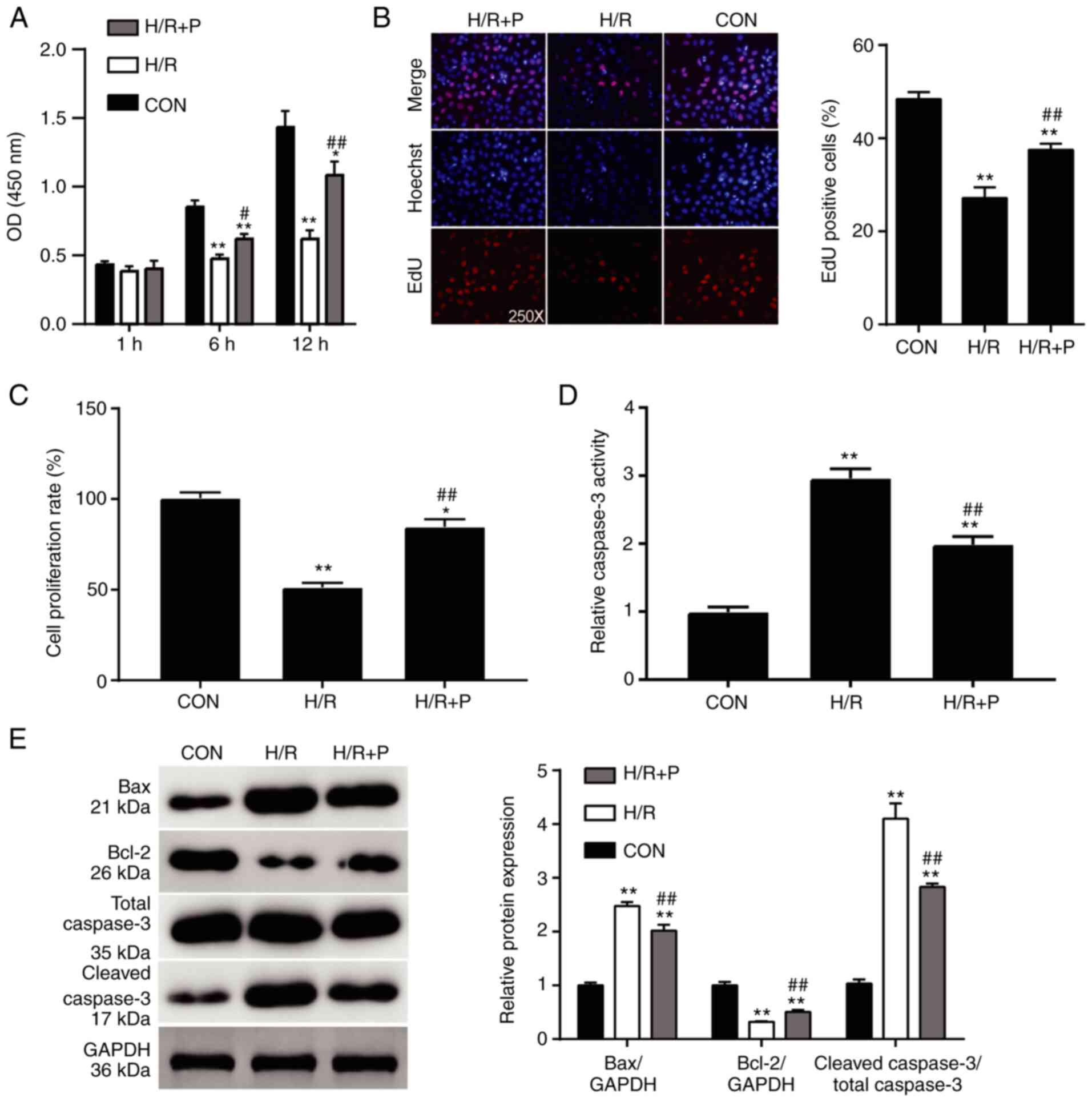
was used for subsequent experiments. The viability of the H9C2
cells subjected to H/R treatment was dramatically reduced compared
to the cells cultured normally (Fig.
2A), confirming the successful generation of the in
vitro I/R injury model. When propofol was introduced, the cell
viability of the H/R-induced cells was significantly improved
(Fig. 2A). Furthermore, the
proliferation rates of the H9C2 cells were also evaluated using an
EdU assay. Compared with the control group, the rate of
EdU-positive cells in the H/R alone group was decreased by 50%,
while the rate of EdU-positive cells in the propofol-treated groups
was reduced by 20% (Fig. 2B). The
proliferation of the H9C2 cells detected by a BrdU assay exhibited
a similar trend; the proliferation of the cells subjected to H/R
alone was suppressed by 50% compared with the control group, while
propofol compromised this reduction by 30% compared with the H/R
group (Fig. 2C). Cell apoptosis of
the H9C2 cells was measured by the caspase-3 activity assay.
Caspase-3 activity of the cells in the H/R group was almost thrice
as much as that of the control group; this increase was attenuated
by ~50% compared with that of the H/R group when propofol was
introduced (Fig. 2D). Finally, the
levels of the apoptosis-associated proteins Bax, Bcl-2, and cleaved
caspase-3 were detected by western blot analysis. Compared with the
control group, the protein expression levels of Bax and cleaved
caspase-3 were increased in the H/R group, while Bcl-2 protein
expression was decreased (Fig. 2E).
Propofol treatment resulted in lower increases in the Bax and
cleaved caspase-3 concentrations and a lower reduction in Bcl-2
(Fig. 2E). Together, these results
demonstrated that propofol exerted a protective effect on the
survival and proliferation of H9C2 cells.
Propofol attenuates H/R injury through
the inhibition of miR-449a
As shown in Fig. 3A,
the expression levels of miR-449a in the H/R group were ~2.7 times
higher compared with the control group. The expression was
relatively lower in the H/R+P group compared with that of the H/R
group (1.7-fold compared with the control group; Fig. 3A). To explore the function of
miR-449a, a specific inhibitor was used to downregulate its
expression. First, successful transfection efficiency of the
miR-449a inhibitor was confirmed in H9C2 cells under normoxic
conditions (Fig. S2A). Next, the
miR-449a inhibitor was used in the in vitro I/R model.
Compared with H/R+P group, the expression of miR-449a in
H/R+P+miR-449a inhibitor group was decreased by 50%, indicating
successful downregulation of miR-449a (Fig. 3B).
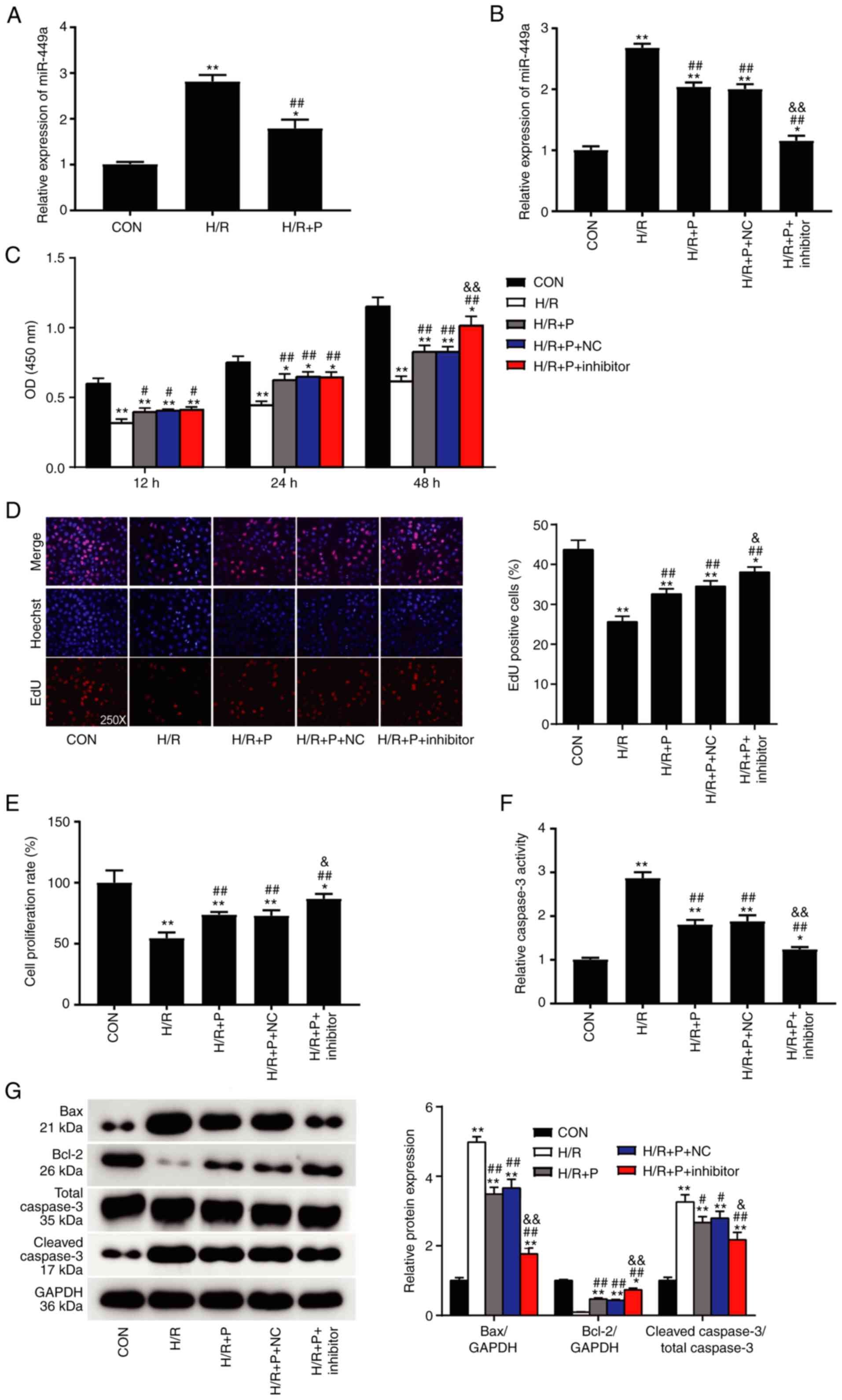 | Figure 3Propofol attenuates H/R injury
through inhibition of miR-449a. (A) miR-449a expression levels in
H9C2 cells were detected by RT-qPCR. (B) The transfection
efficiency of miR-449a inhibitor was confirmed by RT-qPCR. (C) The
effect of miR-449a inhibition on cell viability of H9C2 cells
subjected to H/R injury was measured by CCK-8 assay. (D) Cell
proliferation was measured by EdU assay and by (E) BrdU assay. (F)
Cell apoptosis was measured by caspase-3 activity assay. (G)
Western blot analysis of Bax, Bcl-2 and cleaved caspase-3 protein
expression levels Data are presented as mean ± SD (n=3).
*P<0.05, **P<0.001 compared with
control. #P<0.05, ##P<0.001 vs. H/R.
&P<0.05, &&P<0.001 vs.
H/R+P+NC. H/R, hypoxia/reoxygenation; miR, microRNA; RT-qPCR,
reverse transcription-quantitative PCR; P, propofol; Con, control;
NC, negative control; OD, optical density. |
Functional assays were then performed to assess the
role of miR-449a. The CCK-8 assay depicted that the miR-449a
inhibitor contributed to improved cell viability of the H9C2 cells
treated with H/R and propofol (Fig.
3C). The EdU assay demonstrated that the miR-449a inhibitor
increased the ratios of EdU-positive cells by 1.2-fold compared
with those in the H/R+P group (Fig.
3D). The BrdU assay showed that, despite the severe damage
caused by H/R exposure, the miR-449a inhibitor was able to promote
cell proliferation of the H9C2 cells by 25% compared with that
observed in the H/R+P group (Fig.
3E). The caspase-3 activity assay revealed that cell apoptosis
was repressed in the miR-449a inhibitor group by 30% compared with
that in the H/R+P group (Fig. 3F).
Western blot analysis showed that the protein expression levels of
Bax and cleaved caspase-3 decreased while the protein expression of
Bcl-2 increased in the miR-449a inhibitor group compared with the
H/R+P group (Fig. 3G). Therefore,
the present results demonstrated that propofol protected H9C2 cells
from H/R injury through the inhibition of miR-449a.
NR4A2 is a target gene of miR-449a in
H9C2 cells
miR449a was predicted to target the NR4A2 gene
(Fig. 1). To understand the
mechanism underlying the action of miR-449a in H9C2 cell damage
caused by H/R, the functional association between miR449a and NR4A2
was further investigated. The mRNA expression levels of NR4A2 were
measured in the H/R group and the H/R+P group by RT-qPCR. The
results demonstrated that NR4A2 mRNA expression was significantly
higher in the H/R+P group compared with that in the H/R group
(Fig. 4A). Western blot analysis
further confirmed that the protein expression levels of NR4A2 in
the H/R+P group was higher compared with that in the H/R group
(Fig. 4B). The predicted binding
sites of miR-449a on NR4A2 are depicted in Fig. 4C. As shown in Fig. 4D, luciferase activity from a wt
NR4A2 reporter plasmid was repressed when co-transfected with
miR-449a mimics compared with mimics NC, while no effect was
observed on the luciferase activity of a mutant reporter plasmid.
These results confirmed that NR4A2 was a direct target of
miR-449a.
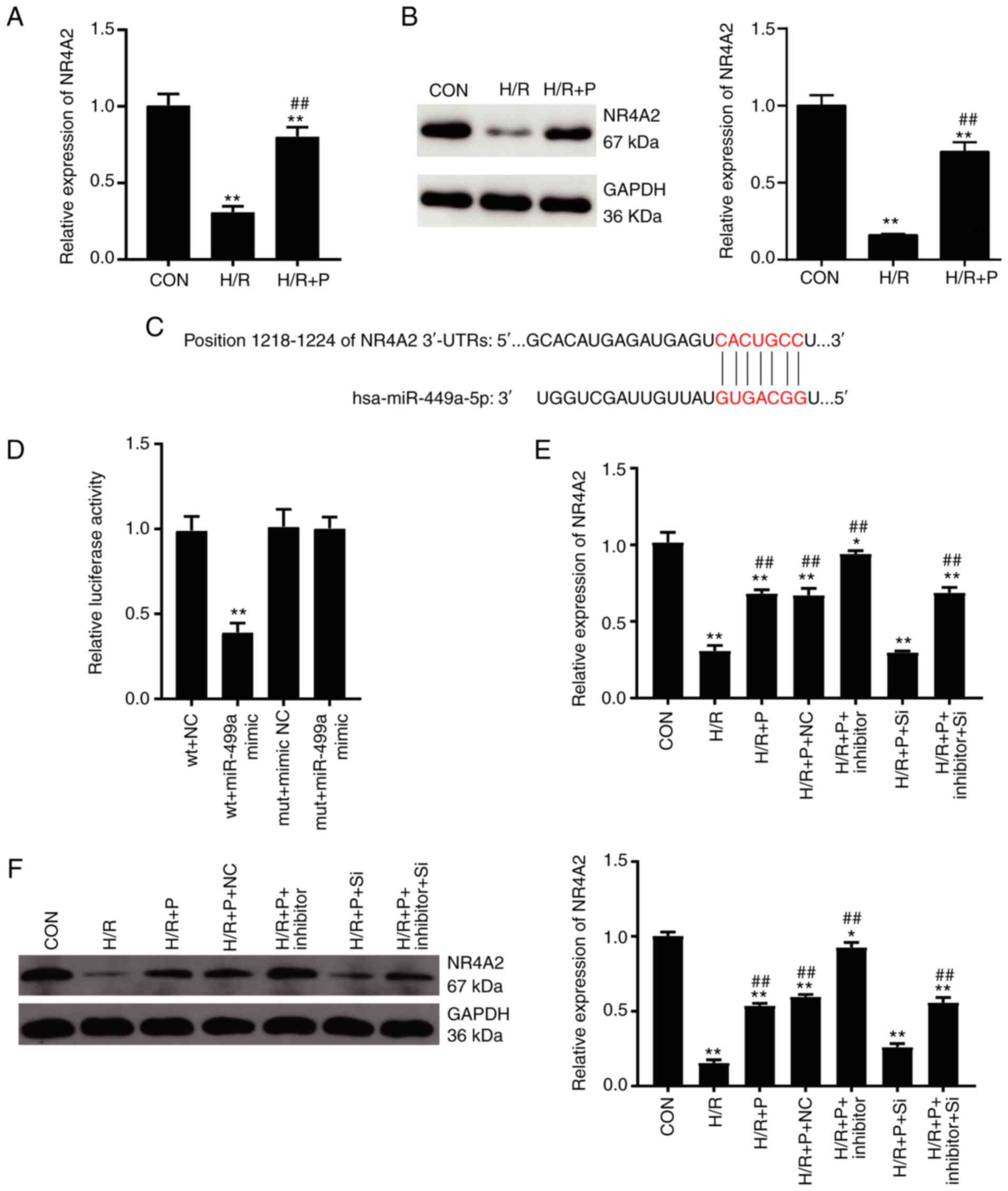 | Figure 4NR4A2 is a target gene of miR-449a.
(A) NR4A2 mRNA expression levels in H9C2 cells were detected by
RT-qPCR. (B) NR4A2 protein expression levels in H9C2 cells were
detected by western blotting. (C) Schematic of the predicted
binding sites between miR-449a and the NR4A2 3'-UTR. (D)
Dual-luciferase reporter assay was performed to confirm the target
relationship between miR-449a and NR4A2. (E) NR4A2 mRNA expression
levels were detected by RT-qPCR following siRNA transfection. (F)
NR4A2 protein expression levels were detected by western blotting
following siRNA transfection. Data are presented as mean ± SD
(n=3). *P<0.05, **P<0.001 compared with
control. ##P<0.001 vs. H/R. NR4A2, nuclear receptor
subfamily 4 group A member 2; miR, microRNA; RT-qPCR, reverse
transcription-quantitative PCR; UTR, untranslated region; siRNA,
small interfering RNA; Con, control; H/R, hypoxia/reoxygenation; P,
propofol; wt, wild-type; mut, mutant; NC, negative control; si-NC,
negative control siRNA; si, NR4A2-targeting siRNA. |
Next, NR4A2 was silenced by transfection with a
specific siRNA. First, the knockdown efficiency was confirmed in
H9C2 cells under normoxic conditions (Fig. S2B and C). In addition, NR4A2 siRNA transfection
could significantly reduce the NR4A2 expression at the mRNA
(Fig. 4E) as well as protein levels
(Fig. 4F) under the H/R and
propofol treatment conditions. Collectively, these results revealed
that miR-449a targeted NR4A2 in the propofol protection against H/R
injury in H9C2 cells.
NR4A2 exerts a protective effect on
H9C2 cells mediated by miR-449a
Given that miR-449a can target NR4A2, further
experiments were designed to explore if NR4A2 had a positive impact
on the viability of H9C2 cells after exposure to H/R injury. The
CCK-8 assay revealed that NR4A2 silencing dramatically repressed
the cell viability of the H9C2 cells with H/R injury and propofol,
while this repression could be restored by the miR-449a inhibitor
(Fig. 5A). In the EdU assay, NR4A2
silencing reduced the ratio of EdU-positive cells in the H/R injury
and propofol-treated group, while the miR-449a inhibitor reversed
this reduction (Fig. 5B). The BrdU
assay indicated that NR4A2 silencing hindered cell proliferation of
the H9C2 cells by 20% compared with that of the H/R+P group, which
was 40% lower than that of the control group (Fig. 5C). However, this repression could be
restored by the miR-449a inhibitor (Fig. 5B). The Caspase-3 activity assay
demonstrated that NR4A2 silencing induced apoptosis by 3-fold of
the H9C2 cells in the H/R group compared with that in the control.
However, this promotion could be reversed by the miR-449a inhibitor
(Fig. 5D). Western blot analysis
showed that the protein expression levels of Bax and cleaved
caspase-3 in the H9C2 cells following NR4A2 silencing were
increased compared with those in the H/R+P group, while the
expression levels of Bcl-2 were decreased (Fig. 5E). Of note, the miR-449a inhibitor
reversed these effects (Fig. 5E).
Taken together, the current results indicated that NR4A2 exerted a
protective effect on H9C2 cells, and this effect was regulated by
miR-449a.
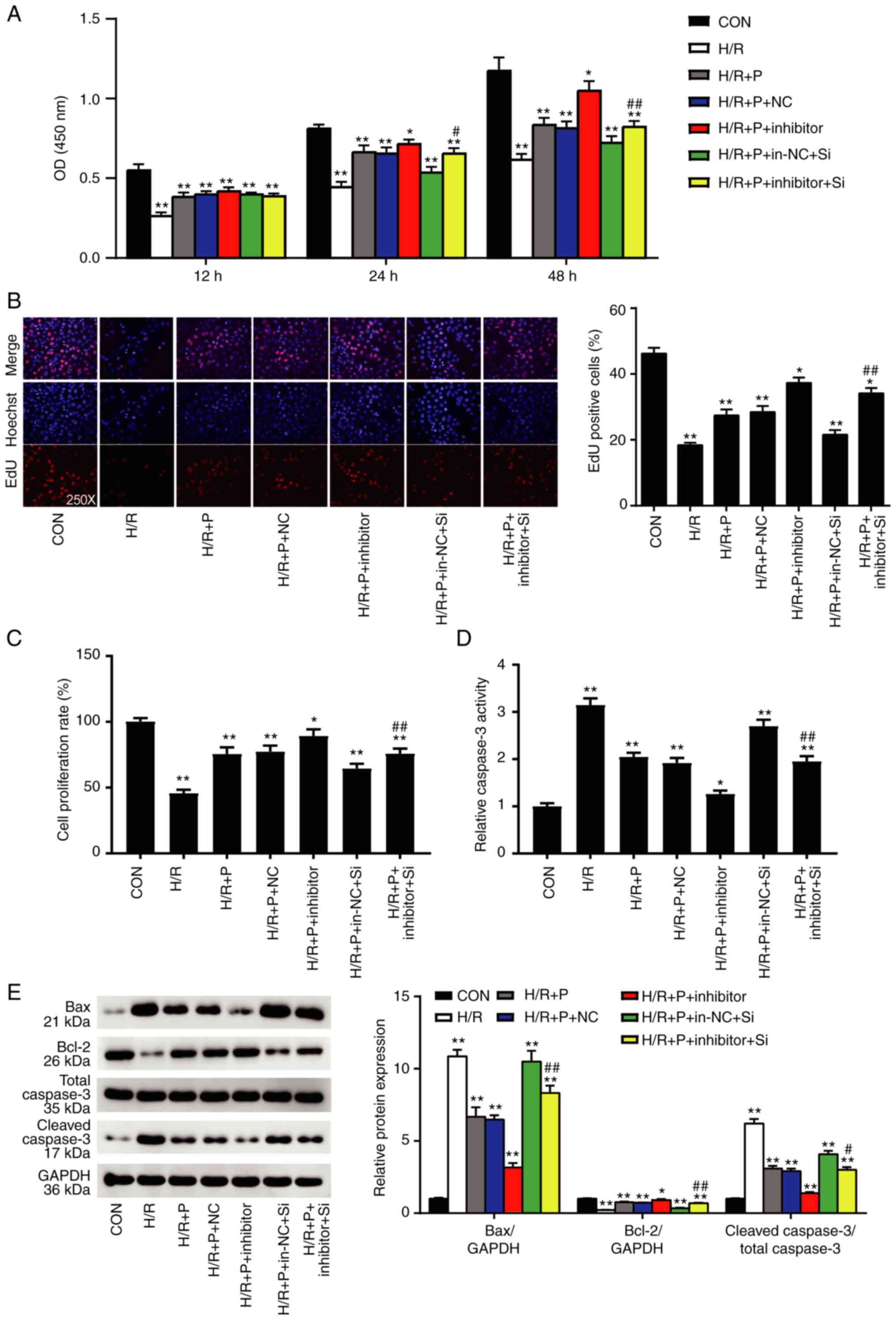 | Figure 5NR4A2 exerts a protective effect on
H9C2 cells mediated by miR-449a. H9C2 cells were subjected to H/R
injury in the presence or absence of propofol, and co-transfected
with miR-449a inhibitor and/or NR4A2-targeting siRNA or their
respective controls. (A) Cell viability was measured by CCK-8
assay. (B) Cell proliferation was measured by EdU assay and by (C)
BrdU assay. (D) Apoptosis was measured by caspase-3 activity assay.
(E) Western blot analysis of Bax, Bcl-2 and cleaved caspase-3
protein expression levels. Data are presented as mean ± SD (n=3).
*P<0.05, **P<0.001 compared with
control. #P<0.05, ##P<0.001 vs.
H/R+P+in-NC+Si. NR4A2, nuclear receptor subfamily 4 group A member
2; miR, microRNA; siRNA, small interfering RNA; Con, control; H/R,
hypoxia/reoxygenation; P, propofol; NC, negative control; in-NC,
inhibitor negative control. si-NC, negative control siRNA; si,
NR4A2-targeting siRNA. |
Discussion
In the present study, H/R-induced H9C2 cells were
used to mimic I/R injury in vitro. It was observed that
propofol treatment significantly alleviated H/R injury in
cardiomyocytes, by improving cell viability and proliferation while
repressing cell apoptosis of the H9C2 cells. Additionally, the
present results suggested that the cardioprotective effect of
propofol was closely related to the regulation of miR-449a and its
target gene NR4A2. Specifically, miR-449a inhibition promoted cell
viability and proliferation and repressed cell apoptosis of the
H9C2 cells, thus enhancing the cardioprotective effect of propofol.
By contrast, silencing of NR4A2, the target gene of miR-449a,
hindered the alleviation effect of propofol on H/R injury.
According to previous studies, it is established
that propofol protects cells against oxidative stress from hydrogen
peroxide (27,28) and oxygen-glucose deprivation
(29). The present results further
confirmed the cell-protective role of propofol. Propofol-dependent
mechanisms have been reported in several studies concerning
myocardial I/R injury. A previous study reported that propofol
exerted cardioprotection against myocardial I/R injury through the
miR-451/HMGB1 axis in a H/R injury model of H9C2 cardiomyocytes
(10). Another study reported that
propofol exerted a protective effect on the H/R-challenged
cardiomyocytes by activation of Akt phosphorylation (30). The present study confirmed that
propofol was significantly beneficial to the survival and activity
of cardiomyocytes subjected to H/R injury. The benefits of propofol
could be enhanced by miR-449a inhibition and hindered by NR4A2
silencing.
miR-449a has been widely studied in various cancer
models and acts as a tumor suppressor. For example, the
tumor-suppressive role of miR-449a has been demonstrated in breast
cancer via targeting PLAG1 like zinc finger 2(31). In addition, miR-449a was
demonstrated to be sponged by circGFRA1, thus hindering the
inhibition effect of miR-449a on ovarian cancer progression
(32). The role of miR-449a in
cancer has been studied extensively. However, the contribution of
miR-449a in protecting the heart from myocardial I/R injury or H/R
injury needs further research. Two studies determined that the
downregulation of miR-449a helped to protect cardiomyocytes from
H/R injury (19,20). The current study also described that
the inhibition of miR-449a enhanced the cardioprotective effect in
the H9C2 cells against the stress from H/R injury. It was observed
that miR-449a inhibition could not only promote cell viability and
proliferation but also inhibit cell apoptosis in H9C2 cells.
To further elucidate the role of miR-449a in
cardiomyocyte protection from H/R injury, NR4A2 was predicted and
confirmed as a target gene for miR-449a. A previous study
determined that there was a close relationship between the
haplotypes of the NR4A2 gene and cardiovascular disease (33). The role of NR4A2 in cardiomyocyte
apoptosis has been illustrated previously (24), showing that NR4A2 knockdown
aggravated the cardiomyocyte apoptosis in response to I/R injury.
The present study clarified the function of NR4A2 in cardiomyocyte
survival from the aspect of cell viability, proliferation, and
apoptosis. NR4A2 was able to impede cell apoptosis of
cardiomyocytes following H/R injury. This apoptosis-suppressive
effect of NR4A2 was consistent with previous literature (24). In addition to the suppression of
cell apoptosis, the present results determined a promotional effect
of NR4A2 on cell viability and proliferation. Collectively, NR4A2
protected H9C2 cells from H/R injury, which was consistent with the
role of propofol but opposite to that of miR-449a.
The present study explored the mechanism of propofol
in H/R-mediated cell injury, but certain limitations remain. First,
the present study was performed in vitro, and there is a
lack of in vivo experiments to further confirm the effects
of propofol and miR-449a/NR4A2 on myocardial I/R injury. In
addition, the effects of propofol and miR-449a/NR4A2 on myocardial
I/R injury need to be further explored in clinical samples. Further
studies are underway to investigate the effects of propofol and
miR-449a/NR4A2 on myocardial I/R injury in vivo.
In summary, the present results elucidated how
propofol, miR-449a, and its target gene NR4A2 interact in
protecting cardiomyocytes subjected to H/R injury. The observations
can serve as a novel clue in designing therapeutic strategies for
cardiovascular diseases against myocardial I/R injury.
Supplementary Material
Effect of different concentrations of
propofol on cell viability of H9C2 cells subjected to
hypoxia/reoxygenation injury. Cell viability was measured by CCK-8
assay. Data are presented as mean ± SD (n=3).
*P<0.05, **P<0.001 compared with
control untreated cells. OD, optical density.
Transfection efficiency of miR-449a
inhibitor and NR4A2 siRNA in untreated cells. (A) miR-499a
expression levels were evaluated by reverse
transcription-quantitative PCR in untreated parental H9C2 cells
(control), and in cells transfected with NC or inhibitor. (B) NR4A2
mRNA and (C) protein expression levels were evaluated in untreated
parental H9C2 cells (control), and in cells transfected with
NR4A2-targeting siRNA or si-NC. Data are presented as mean ± SD
(n=3). **P<0.001 compared with control untreated
cells. miR, miRNA; NR4A2, nuclear receptor subfamily 4 group A
member 2; siRNA, small interfering RNA; NC, negative control; Con,
control; si-NC, negative control siRNA; si, NR4A2-targeting
siRNA.
mRNA expression levels in H9C2 cells
treated with H/R injury.
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
All data generated or analyzed during this study are
included in this published article.
Authors' contributions
QQ performed the experiments and data analysis. YX
conceived and designed the study. QQ wrote the paper. YX reviewed
and edited the manuscript. All authors read and approved the
manuscript. QQ and YX confirm the authenticity of all the raw
data.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Nadatani Y, Watanabe T, Shimada S, Otani
K, Tanigawa T and Fujiwara Y: Microbiome and intestinal
ischemia/reperfusion injury. J Clin Biochem Nutr. 63:26–32.
2018.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Binder A, Ali A, Chawla R, Aziz HA, Abbate
A and Jovin IS: Myocardial protection from ischemia-reperfusion
injury post coronary revascularization. Expert Rev Cardiovasc Ther.
13:1045–1057. 2015.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Ferdinandy P, Schulz R and Baxter GF:
Interaction of cardiovascular risk factors with myocardial
ischemia/reperfusion injury, preconditioning, and postconditioning.
Pharmacol Rev. 59:418–458. 2007.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Zhao ZQ and Vinten-Johansen J:
Postconditioning: Reduction of reperfusion-induced injury.
Cardiovasc Res. 70:200–211. 2006.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Qiao SG, Sun Y, Sun B, Wang A, Qiu J, Hong
L, An JZ, Wang C and Zhang HL: Sevoflurane postconditioning
protects against myocardial ischemia/reperfusion injury by
restoring autophagic flux via an NO-dependent mechanism. Acta
Pharmacol Sin. 40:35–45. 2019.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Li Z, Zhang Y, Ding N, Zhao Y, Ye Z, Shen
L, Yi H and Zhu Y: Inhibition of lncRNA XIST improves myocardial
I/R injury by targeting miR-133a through inhibition of autophagy
and regulation of SOCS2. Mol Ther Nucleic Acids. 18:764–773.
2019.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Yao L, Chen H, Wu Q and Xie K:
Hydrogen-rich saline alleviates inflammation and apoptosis in
myocardial I/R injury via PINK-mediated autophagy. Int J Mol Med.
44:1048–1062. 2019.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Li W, Li Y, Chu Y, Wu W, Yu Q, Zhu X and
Wang Q: PLCE1 promotes myocardial ischemia-reperfusion injury in
H/R H9c2 cells and I/R rats by promoting inflammation. Biosci Rep.
39(BSR20181613)2019.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Lotz C, Stumpner J and Smul TM:
Sevoflurane as opposed to propofol anesthesia preserves
mitochondrial function and alleviates myocardial
ischemia/reperfusion injury. Biomed Pharmacother.
129(110417)2020.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Li YM, Sun JG, Hu LH, Ma XC, Zhou G and
Huang XZ: Propofol-mediated cardioprotection dependent of
microRNA-451/HMGB1 against myocardial ischemia-reperfusion injury.
J Cell Physiol. 234:23289–23301. 2019.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Li H, Zhang X, Tan J, Sun L, Xu LH, Jiang
YG, Lou JS, Shi XY and Mi WD: Propofol postconditioning protects
H9c2 cells from hypoxia/reoxygenation injury by inducing autophagy
via the SAPK/JNK pathway. Mol Med Rep. 17:4573–4580.
2018.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Zhao D, Li Q, Huang Q, Li X, Yin M, Wang Z
and Hong J: Cardioprotective effect of propofol against oxygen
glucose deprivation and reperfusion injury in H9c2 cells. Oxid Med
Cell Longev. 2015(184938)2015.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Vasileiou I, Xanthos T, Koudouna E, Perrea
D, Klonaris C, Katsargyris A and Papadimitriou L: Propofol: A
review of its non-anaesthetic effects. Eur J Pharmacol. 605:1–8.
2009.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Green TR, Bennett SR and Nelson VM:
Specificity and properties of propofol as an antioxidant free
radical scavenger. Toxicol Appl Pharmacol. 129:163–169.
1994.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Hanouz JL, Yvon A, Flais F, Rouet R,
Ducouret P, Bricard H and Gérard JL: Propofol decreases
reperfusion-induced arrhythmias in a model of ‘border zone’ between
normal and ischemic-reperfused guinea pig myocardium. Anesth Analg.
97:1230–1238. 2003.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Hu S, Cao S, Tong Z and Liu J: FGF21
protects myocardial ischemia-reperfusion injury through reduction
of miR-145-mediated autophagy. Am J Transl Res. 10:3677–3688.
2018.PubMed/NCBI
|
|
17
|
Yang Y, Yang J, Liu XW, Ding JW, Li S, Guo
X, Yang CJ, Fan ZX, Wang HB, Li Q, et al: Down-regulation of
miR-327 alleviates ischemia/reperfusion-induced myocardial damage
by targeting RP105. Cell Physiol Biochem. 49:1049–1063.
2018.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Ye Y, Hu Z, Lin Y, Zhang C and Perez-Polo
JR: Downregulation of microRNA-29 by antisense inhibitors and a
PPAR-gamma agonist protects against myocardial
ischaemia-reperfusion injury. Cardiovasc Res. 87:535–544.
2010.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Cheng J, Wu Q, Lv R, Huang L, Xu B, Wang
X, Chen A and He F: MicroRNA-449a inhibition protects H9C2 cells
against hypoxia/reoxygenation-induced injury by targeting the
Notch-1 signaling pathway. Cell Physiol Biochem. 46:2587–2600.
2018.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Zhang X, Dong H, Liu Y, Han J, Tang S and
Si J: Tetramethylpyrazine partially relieves hypoxia-caused damage
of cardiomyocytes H9c2 by downregulation of miR-449a. J Cell
Physiol, Feb 15, 2019 (Epub ahead of print).
|
|
21
|
Spathis AD, Asvos X, Ziavra D, Karampelas
T, Topouzis S, Cournia Z, Qing X, Alexakos P, Smits LM, Dalla C, et
al: Nurr1:RXRα heterodimer activation as monotherapy for
Parkinson's disease. Proc Natl Acad Sci USA. 114:3999–4004.
2017.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Medzikovic L, Schumacher CA, Verkerk AO,
van Deel ED, Wolswinkel R, van der Made I, Bleeker N, Cakici D, van
den Hoogenhof MM, Meggouh F, et al: Orphan nuclear receptor Nur77
affects cardiomyocyte calcium homeostasis and adverse cardiac
remodelling. Sci Rep. 5(15404)2015.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Xiao G, Sun T, Songming C and Cao Y: NR4A1
enhances neural survival following oxygen and glucose deprivation:
An in vitro study. J Neurol Sci. 330:78–84. 2013.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Liu H, Liu P, Shi X, Yin D and Zhao J:
NR4A2 protects cardiomyocytes against myocardial infarction injury
by promoting autophagy. Cell Death Discov. 4(27)2018.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Lucchinetti E, Hofer C, Bestmann L,
Hersberger M, Feng J, Zhu M, Furrer L, Schaub MC, Tavakoli R,
Genoni M, et al: Gene regulatory control of myocardial energy
metabolism predicts postoperative cardiac function in patients
undergoing off-pump coronary artery bypass graft surgery:
Inhalational versus intravenous anesthetics. Anesthesiology.
106:444–457. 2007.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Ming N, Na HST, He JL, Meng QT and Xia ZY:
Propofol alleviates oxidative stress via upregulating
lncRNA-TUG1/Brg1 pathway in hypoxia/reoxygenation hepatic cells. J
Biochem. 166:415–421. 2019.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Kim EJ, Choi IS, Yoon JY, Park BS, Yoon JU
and Kim CH: Effects of propofol-induced autophagy against oxidative
stress in human osteoblasts. J Dent Anesth Pain Med. 16:39–47.
2016.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Wang Z, Yang P and Qi Y: Role of
microRNA-134 in the neuroprotective effects of propofol against
oxygen-glucose deprivation and related mechanisms. Int J Clin Exp
Med. 8:20617–20623. 2015.PubMed/NCBI
|
|
30
|
Ma K, Qiu J, Zhou M, Yang Y and Ye X:
Cox-2 Negatively affects the protective role of propofol against
hypoxia/reoxygenation induced cardiomyocytes apoptosis through
suppressing Akt signaling. Biomed Res Int.
2019(7587451)2019.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Xu B, Zhang X, Wang S and Shi B: MiR-449a
suppresses cell migration and invasion by targeting PLAGL2 in
breast cancer. Pathol Res Pract. 214:790–795. 2018.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Liu J, Yu F, Wang S, Zhao X, Jiang F, Xie
J and Deng M: circGFRA1 promotes ovarian cancer progression by
sponging miR-449a. J Cancer. 10:3908–3913. 2019.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Kardys I, van Tiel CM, de Vries CJ,
Pannekoek H, Uitterlinden AG, Hofman A, Witteman JC and de Maat MP:
Haplotypes of the NR4A2/NURR1 gene and cardiovascular disease: The
Rotterdam Study. Hum Mutat. 30:417–423. 2009.PubMed/NCBI View Article : Google Scholar
|