Introduction
Gliomas are among the most common types of
intracranial malignancies (1).
Although the global incidence of gliomas is only 3-5.5 per 100,000
individuals, gliomas have high mortality rates (2,3).
Although the prognosis of gliomas has improved with advancements in
surgical technologies and chemotherapies, the 5-year survival rate
for advanced glioma is still <36% (4,5). The
poor prognosis of glioma is mainly due to its highly invasive
capabilities, rendering it difficult to completely resect, with
high rates of recurrence (6). In
addition, the prognosis of glioma is closely associated with
pathological grade, histological type and abnormal expression of a
number of oncogenes, including chitinase 3 like 1, MET and PTEN
(7).
Studies have indicated that abnormally expressed
genes are closely associated with the prognosis of gliomas. For
instance, several studies have reported that the malignant
progression of gliomas involved an EGFR variant, whose abnormal
expression is involved in the infiltration and proliferation of
tumor cells (4). Furthermore, TGF-β
influences the proliferation and migration of tumor cells by
affecting T-cell function (8). In
addition, certain studies have reported that in glioma patients
undergoing chemotherapy, CD133 expressed by the HOX gene is a
marker for poor prognosis (9). Hale
et al (10) revealed that
CD36 promotes the growth of cancer stem cells, thereby improving
the growth and immune resistance of gliomas, and is associated with
poor prognosis. However, these biomarkers are still controversial
and hence, novel biomarkers urgently require to be identified.
Collagen α-1 (IV) chain (COL4A1) is an important
component of the basement membrane of numerous tissues and cell
types in the human body (11). The
basement membrane is a vascular and extracellular scaffold that
supports and partially regulates cell behavior (12). A previous study reported that
mutations in COL4A1 may affect glycine residues in the protein,
which is not conducive for its structural and functional stability
(13). COL4A1 mutations may cause
several diseases, including those pertaining to the cardiovascular,
cerebrovascular, renal, gastrointestinal and circulatory systems
(14-19).
Studies have indicated that abnormal expression of COL4A1, as a
novel oncogene, is closely associated with the occurrence,
development and outcome of a variety of cancer types (14,16,20,21).
However, the role of COL4A1 in the diagnosis and prognosis of
glioma has remained to be determined.
In the present study, thousands of glioma tissue
samples were collected through a variety of different detection
techniques [such as sequencing data, chip data, reverse
transcription-quantitative (RT-q)PCR, in situ hybridization
and immunology] to reveal that COL4A1 expression was not only
increased at the mRNA level, but also markedly higher at the
protein level in comparison with normal brain tissue. It was
attempted to determine the relationship between COL4A1 expression
levels and the clinical characteristics of glioma, and also the
possible role of COL4A1 in the pathological process of gliomas. The
present study was the first to report that high expression of
COL4A1, as a novel oncogene, is significantly associated with the
prognosis of gliomas. Furthermore, by performing gene-set
enrichment analysis (GSEA) analysis, the possible carcinogenic
pathways of COL4A1 were identified. Therefore, it may be indicated
that COL4A1 is a valuable potential biomarker for the diagnosis and
treatment of gliomas.
Materials and methods
Data collection
The glioma RNA sequencing (RNA-seq) dataset and
matched clinical information were downloaded from The Cancer Genome
Atlas (TCGA; https://portal.gdc.cancer.gov/) and the Chinese Glioma
Genome Atlas (CGGA; http://www.cgga.org.cn/) databases. In the CGGA, the
information of 1,018 glioma patients was collected. Clinical
information included gender, age, grade, Primary, Recurrent,
Secondary (PRS) type, radiotherapy status, chemotherapy status and
histological grade of glioma, while gene expression information
included 1p19 co-deletion status and isocitrate dehydrogenase (IDH)
mutation status. Samples with incomplete clinical information were
excluded. The sequencing data of 749 glioma samples from CGGA were
further analyzed. Furthermore, sequencing data of 655 glioma
samples from TCGA and microarray data of 268 glioma samples from
CGGA, which were separate from the 749 samples aforementioned, were
used for verification of the association between the expression of
COL4A1 and the prognosis of glioma.
In addition, the datasets of the four glioma gene
chips were obtained from Gene Expression Omnibus database (GEO;
https://www.ncbi.nlm.nih.gov/geo/),
namely GSE2223, GSE4290, GSE50161 and GSE116520. The GSE2223
dataset included 50 glioma tissue samples and 4 normal brain tissue
samples; the GSE4290 dataset comprised 77 glioma tissue samples and
23 normal brain tissue samples; the GSE50161 dataset included 34
glioma tissue samples and 13 normal brain tissue samples; and the
GSE116520 dataset comprised 34 glioma tissue samples and 8 normal
brain tissue samples. The four glioma datasets included a total of
195 glioma tissue samples and 48 normal brain tissue samples. These
samples were used to detect changes in the expression levels of
COL4A1 in tumor tissues compared with normal samples.
Certain online data analysis platforms were also
used in the present study. Gene expression profiling interactive
analysis (GEPIA; http://gepia.cancer-pku.cn/) is an online public data
analysis platform that was used to detect changes in the expression
levels of COL4A1 in various human malignant tumor tissues. The
Human Protein Atlas (https://www.proteinatlas.org/) is a proteomics
database that was used to detect changes in protein expression
levels of COL4A1 in gliomas vs. normal brain tissues. The database
was used to evaluate immunohistochemical results for the protein
expression of COL4A1 in 2 normal brain tissues and 11 glioma
samples. Images of immunohistochemical sections of 2 normal brain
tissues and 2 glioma tissues were obtained for display. The Ivy
Glioblastoma Atlas (http://glioblastoma.alleninstitute.org/) is a database
focused on glioma research, which contains multiple data types such
as in situ hybridization (ISH) and H&E staining. The Ivy
Glioblastoma Atlas database was used by the present study to detect
the expression levels of COL4A1 in gliomas. From this database, 8
glioma samples with ISH staining for COL4A1 containing tumor
boundaries were obtained. H&E staining and annotated images of
anatomical boundaries were also obtained for the corresponding
samples.
GSEA
Gene set enrichment analysis (GSEA) is a
bioinformatics analysis tool that is widely used to annotate and
predict gene functions. To reveal the potential impact of COL4A1 on
the prognosis of gliomas, an enrichment analysis was performed
using GSEA version 3.0 software (https://www.gsea-msigdb.org/gsea/index.jsp). The
important pathways that may be involved in the pathological
mechanisms of glioma in the group with high and low COL4A1
expression were elucidated. In this analysis, 1,000 was set as the
number of gene set permutations. The gene sets with a normal
P<0.05 and false discovery rate (FDR) <0.25 were considered
to be significantly enriched. FDR indicates false positive
discovery rate that may be included and the nominal P-value
describes the statistical significance of the enrichment score
obtained for a subset of functional genes (22).
RT-qPCR analysis
To confirm COL4A1 expression in glioma patients and
healthy controls, RT-qPCR analysis was performed. Tissues of glioma
patients who underwent surgical resection in Henan Provincial
People's Hospital (Zhengzhou, China) from March 2017 to June 2020
were collected. A total of 30 glioma tissues and 15 normal brain
tissues were collected. Normal brain tissue was obtained from
patients who underwent surgical resection for primary epilepsy. All
patients signed an informed consent to use their organization for
relevant research.
Three glioma cell lines (U87, U251 and LN229) and
human-derived astrocytes (HA) were used to detect changes in the
expression levels of COL4A1 by RT-qPCR. Total RNA was isolated from
the tissues using Tri®Reagent (Sigma-Aldrich; Merck
KGaA). RNA quality and quantity were determined using the NanoDrop
One spectrophotometer (Thermo Fisher Scientific, Inc.).
Complementary DNA (cDNA) was prepared using the Transcriptor First
Stand cDNA Synthesis Kit (Roche Diagnostics) under the protocol of
55˚C for 30 min and 85˚C for 5 min. FastStart™ SYBR®
Green (Roche Diagnostics) was used for qPCR experiment according to
the manufacturer's protocols in the QuantStudio™ 3 Real-Time PCR
System (Applied Biosystems; Thermo Fisher Scientific, Inc.). The
thermocycling conditions were as follows: Initial denaturation at
95˚C for 10 min, followed by 40 cycles of denaturation at 95˚C for
10 sec, annealing and extension at 60˚C for 30 sec. GAPDH was used
as the internal reference and the primer sequence was
5'-CAAGGTCATCCATGACAACTTTG-3' (forward) and
5'-GTCCACCACCCTGTTGCTGTAG-3' (reverse). The primer sequence for
COL4A1 was 5'-CTGCCTGGAGGAGTTTAGAAG-3' (forward) and
5'-GAACATCTCGCTCCTCTCTATG-3' (reverse). The relative COL4A1
expression levels were determined using the 2-∆ΔCq
method (23). An unpaired t-test
was performed for analyzing the data of the two groups and a
P-value <0.05 was considered to indicate statistical
significance.
Cell culture and transfection
The human glioma cell line lines (U87-MG, U251 and
LN229) and human-derived astrocytes (HA, cat. no.
3111C0001CCC000525; http://www.cellresource.cn/fdetail.aspx?id=2446) were
purchased from the Chinese Academy of Sciences. Of these, U87-MG is
a glioblastoma cell line of unknown origin (cat. no. TCHu138). All
cells were authenticated by STR analysis and were cultured in DMEM
plus 10% FBS (Gibco; Thermo Fisher Scientific, Inc.). All cells
were cultured at 37˚C in a humidified atmosphere with 5% carbon
dioxide in an incubator. Subsequent cell passages were performed
every 2-3 days. The short interfering (si)RNA to COL4A1 was
purchased from Shanghai GenePharma Co., Ltd. The siRNA sequence was
as follows: sense, 5'-CCCACCUGGAAUUGUUAUATT-3' and antisense,
5'-UAUAACAAUUCCAGGUGGGTT-3'. The negative control (NC) siRNA
sequence was sense, 5'-UUCUCCGAACGUGUCACGUTT-3' and antisense,
5'-ACGUGACACGUUCGGAGAATT-3'. Cells were transfected using
Lipofectamine 2000 reagent (Invitrogen; Thermo Fisher Scientific,
Inc.) following the manufacturer's protocol. After 24 h of
transfection, the knockdown efficiency was detected by RT-qPCR
technology.
Cell viability assay
Cell proliferation was quantified by an MTT assay.
The transfected cells were plated onto 96-well plates (2,000
cells/well). Cell proliferation assays were performed at 12, 24, 48
and 72 h according to the manufacturer's protocol. First, 20 µl of
MTT (5 mg/ml) was added to each well and after 4 h of incubation,
the cell supernatant was discarded. Subsequently, 150 µl of DMSO
was added to each well and the plate was agitated evenly for 15 min
at room temperature. Finally, the absorbance of the solution in
each well was measured at 490 nm on a spectrophotometer.
Cell immunofluorescence staining
The treated cells were fixed with 4%
paraformaldehyde for 30 min, and subsequently, 0.1% Triton X-100
was used for cell permeation for 10 min at room temperature. After
blocking with 5% BSA (Beijing Solarbio Science & Technology
Co., Ltd.) for 30 min, cells were incubated with a primary antibody
against Ki-67 (cat. no. ab15580; 1:1,000 dilution; Abcam) at 37˚C
for 2 h. The cells were washed three times with PBS for 5 min each
time and then incubated with DyLight 594-conjugated AffiniPure Goat
Anti-rabbit IgG (cat. no. BA1142; 1:500 dilution; Boster Biological
Technology) in a humidified atmosphere at room temperature in the
dark for 1 h. Subsequently, DAPI was used to stain the nuclei.
Finally, images were acquired under a fluorescence microscope.
Wound-healing assay
The LN229 cells were seeded in a six-well plate at
5x105 cells/well. Wounding was achieved by performing a
linear scratch using a 200-µl sterile pipette tip. Cells that did
not reattached were thoroughly washed away with PBS. After adding 2
ml of serum-free medium, the cells were incubated in a humid
environment at 37˚C and 5% CO2 to allow the cells to
migrate into the scratched area for 48 h. The periphery of the
wound was observed under a general light microscope and images are
acquired at 0, 24 and 48 h. The wound areas were analyzed using
Image J software (v.1.52r; National Institutes of Health).
Calculation formula of healing rate: Healed area/total wound
area.
Statistical analysis
Data were analyzed by using SPSS version 22.0 (IBM
Corp.) and R software (v.3.6.1). Student's t-test was used to
compare the two groups. A Mann-Whitney U-test was used to determine
the COL4A1 expression levels in the glioma and healthy brain tissue
samples. Uni- and multivariate logistic Cox regression and the
Kaplan-Meier method were used to determine the influence on COL4A1
expression levels on the overall survival rate. ROC method was used
by R software (v.3.6.1) to detect whether COL4A1 may be used as an
independent prognostic factor for glioma. Mann-Whitney U-test or
Kruskal-Wallis test was used to examine the association between
COL4A1 expression levels and the clinical data of glioma patients
obtained from the databases. P<0.05 was considered to indicate a
statistically significant difference.
Results
COL4A1 is abnormally highly expressed
in gliomas
In order to detect changes in the expression levels
of COL4A1 in tumor tissues, data on COL4A1 expression were first
retrieved from the GEPIA database. The database contained 163
glioblastoma multiforme (GBM) tissue samples, 518 low-grade glioma
(LGG) tissue samples and 207 normal brain tissues. It was revealed
that the expression levels of COL4A1 were abnormally increased in
various human tumor tissue types, including GBM and LGG (Fig. 1A).
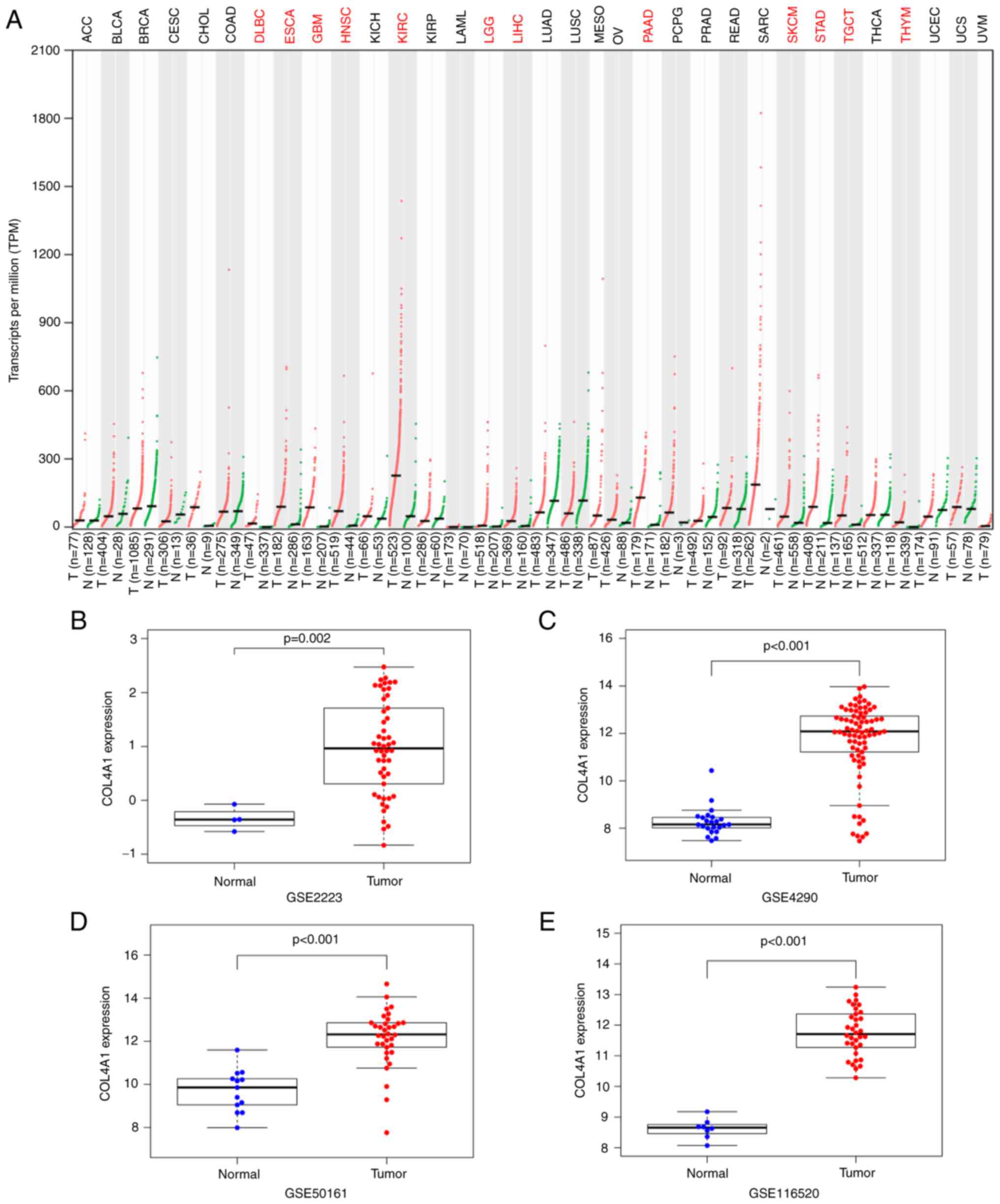 | Figure 1Expression levels of COL4A1 in glioma
compared to normal brain tissue. (A) COL4A1 expression in various
tumor types based on the GEPIA database. Red indicates that the
expression level of COL4A1 in tumor tissues is significantly higher
than that in normal tissues. Black indicates no difference in
expression between tumor and normal tissues. Expression of COL4A1
in the Gene Expression Omnibus datasets (B) GSE2223, (C) GSE4290,
(D) GSE50161, (E) GSE116520. COL4A1, collagen α-1 (IV) chain; T,
tumor samples; N, normal samples; DLBC, lymphoid neoplasm diffuse
large b-cell lymphoma; ESCA, esophageal carcinoma; GBM,
glioblastoma multiforme; HNSC, head and neck squamous cell
carcinoma; KIRC, kidney renal clear cell carcinoma; LGG, brain
lower grade glioma; LIHC, liver hepatocellular carcinoma; PAAD,
pancreatic adenocarcinoma; SKCM, skin cutaneous melanoma; STAD,
stomach adenocarcinoma; TGCT, testicular germ cell tumors; THYM,
thymoma. |
In order to improve the reliability of the results,
four glioma gene chip datasets (GSE2223, GSE4290, GSE50161 and
GSE116520) were further obtained from the GEO database to detect
changes in the expression levels of COL4A1. The results indicated
that the expression levels of COL4A1 were indeed abnormally
increased in gliomas, as presented in Fig. 1B-E. The expression levels of COL4A1
were also detected in the glioma cell lines LN229, U87 and U251 and
compared with those in HA cells by using RT-qPCR, and it was
revealed that the expression levels of COL4A1 in the three glioma
cell lines was also significantly increased compared with that in
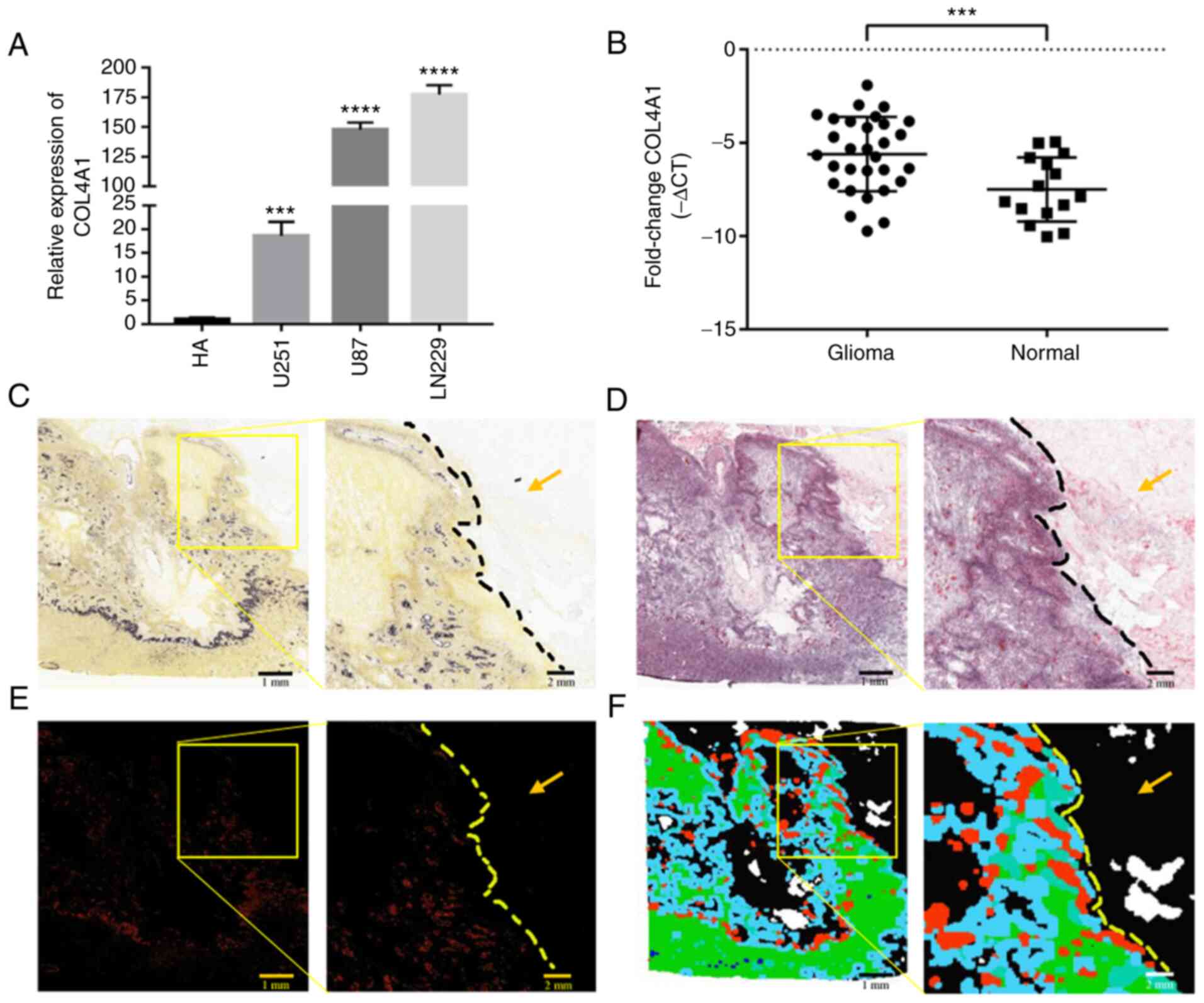
HA cells (Fig. 2A). Furthermore,
the expression levels of COL4A1 were detected in 30 glioma samples
and 15 healthy brain samples by RT-qPCR and it was indicated that
COL4A1 was significantly increased in the glioma samples (Fig. 2B). The glioma group consisted of 16
males and 14 females, aged 23-72 years, with an average age of
48.51±13.77 years. Normal brain tissues from epileptic patients
were obtained from 8 males and 7 females, aged 7-66 years, with an
average age of 32.2±17.37 years.
Data aforementioned confirmed that the expression
levels of COL4A1 were indeed increased in glioma. In order to make
the data of this study more objective and comprehensive, microscopy
images on the changes in the expression levels of COL4A1 in GBM
were further obtained from the Ivy Glioblastoma Atlas. As presented
in Fig. 2C-F, Fig. 2C and E shows the in situ hybridization
experiment and related features. The expression of COL4A1 in the
tumor area is higher than that in the non-tumor infiltrated area.
Fig. 2D and F shows the image and anatomical tumor
characteristics of H&E-stained tissue sections. In Fig. 2D, the tissue structure in the
non-tumor infiltrated area is clear without obvious abnormality,
whilst in the tumor area, the cell density is high and the
morphology is fusiform. According to the anatomical region
annotations in Fig. 2F, it can be
determined that the upper right corner of the sample is the region
of non-tumor cell infiltration and the left side is the tumor
region. As shown in the figure, the expression level of COL4A1 is
higher in tumor areas than in non-tumor cell infiltration
areas.
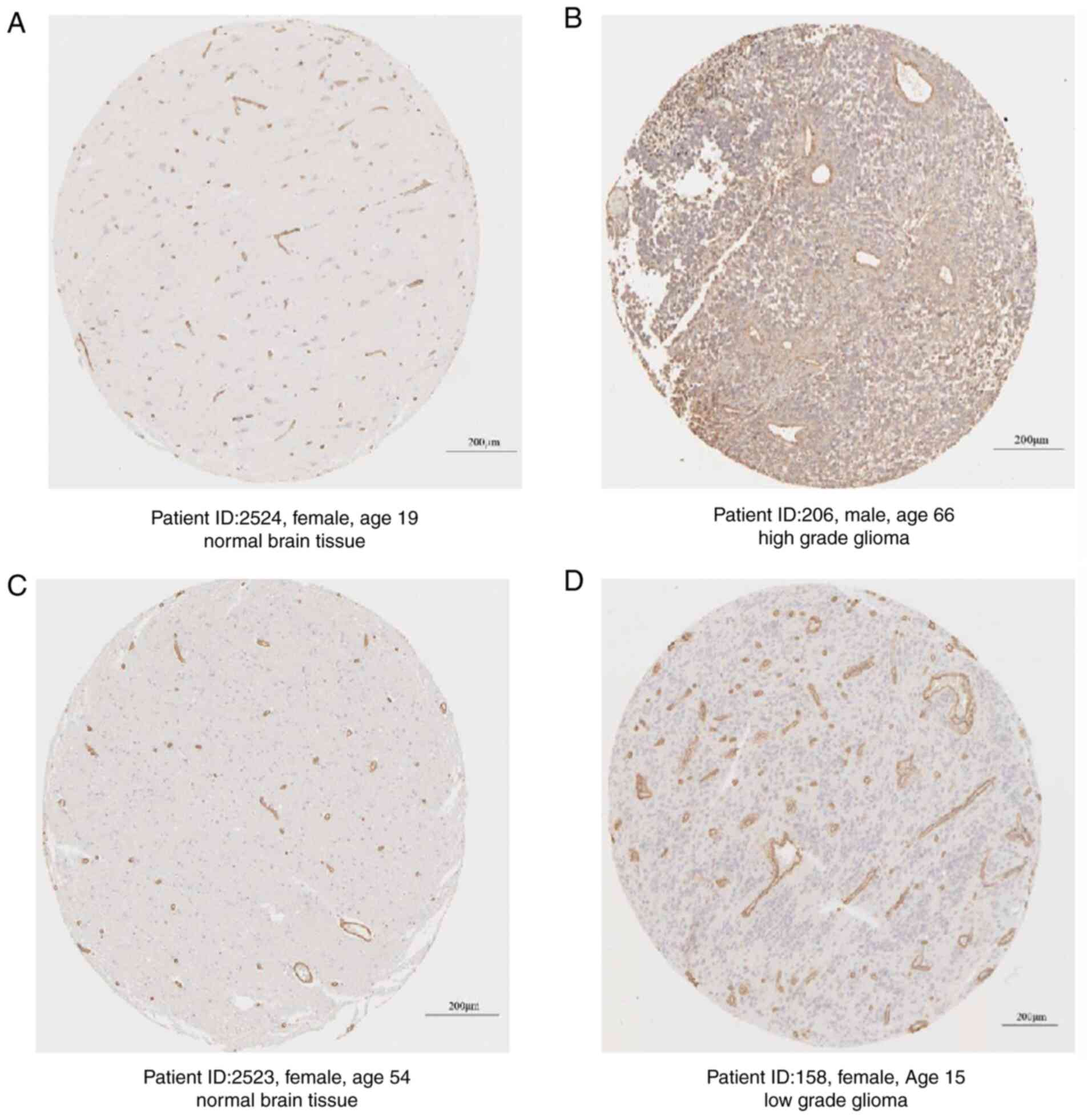
Furthermore, the protein expression levels of COL4A1
in glioma tissues and normal brain tissues were obtained from The
Human Protein Atlas and the results also suggested that the
expression of COL4A1 at the protein level was markedly higher than
that in normal brain tissues from non-glioma patients (Fig. 3).
Association between COL4A1 expression
and clinicopathological variables
As presented in Fig.
4A-G, COL4A1 expression was significantly correlated with the
clinical data obtained from the CGGA database. Increased expression
levels of COL4A1 were positively associated with the histological
type (P<0.001) and histological grade (P<0.001) of the tumor,
patient age (P<0.001), PRS type (P<0.001) and chemotherapy
status (P<0.001), while it was negatively associated with IDH
mutation (P<0.001) and 1p19q co-deletion (P<0.001). The
specific clinical information of the patients is presented in
Table SI.
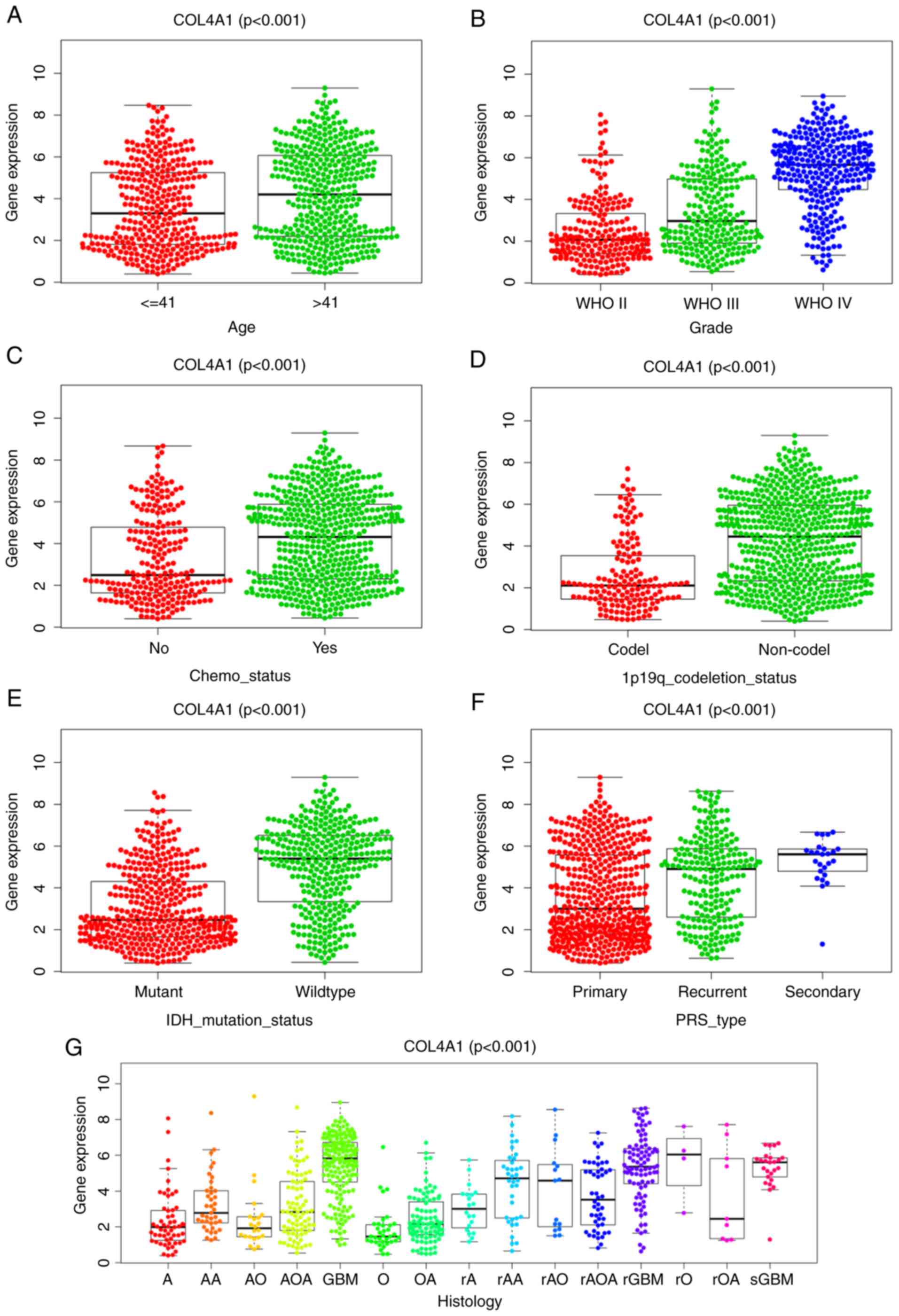 | Figure 4In the CGGA database, COL4A1
expression in glioma according to different clinicopathological
characteristics. Association of COL4A1 expression with (A) patient
age (years), (B) tumor grade, (C) chemotherapy status, (D) 1p19q
codel status, (E) IDH mutation status, (F) PRS type and (G)
Histology. COL4A1, collagen α-1 (IV) chain; IDH, isocitrate
dehydrogenase; codel, codeletion; WHO, World Health Organization;
PRS, Primary, Recurrent, Secondary; A, strocytoma; AA, anaplastic
astrocytoma; AO, anaplastic oligodendroglioma; AOA, anaplastic
oligoastrocytoma; GBM, glioblastoma multiforme; O,
oligodendroglioma; OA, oligoastrocytoma; rA, recurrence of
strocytoma; rAA, recurrence of anaplastic astrocytoma; rAO,
recurrence of anaplastic oligodendroglioma; rAOA, recurrence of
anaplastic oligoastrocytoma; rGBM, recurrence of glioblastoma
multiforme; rO, recurrence of oligodendroglioma; rOA, recurrence of
oligoastrocytoma; sGBM, secondary glioblastoma. |
Survival analysis and diagnostic
value
Although it has been clarified that the expression
of COL4A1 was significantly increased in gliomas and has a close
association with various clinical features of patients, the
prognostic impact of COL4A1 on glioma patients remained elusive.
Therefore, in order to investigate the effect of COL4A1 on the
overall survival of glioma patients, three sets of data were
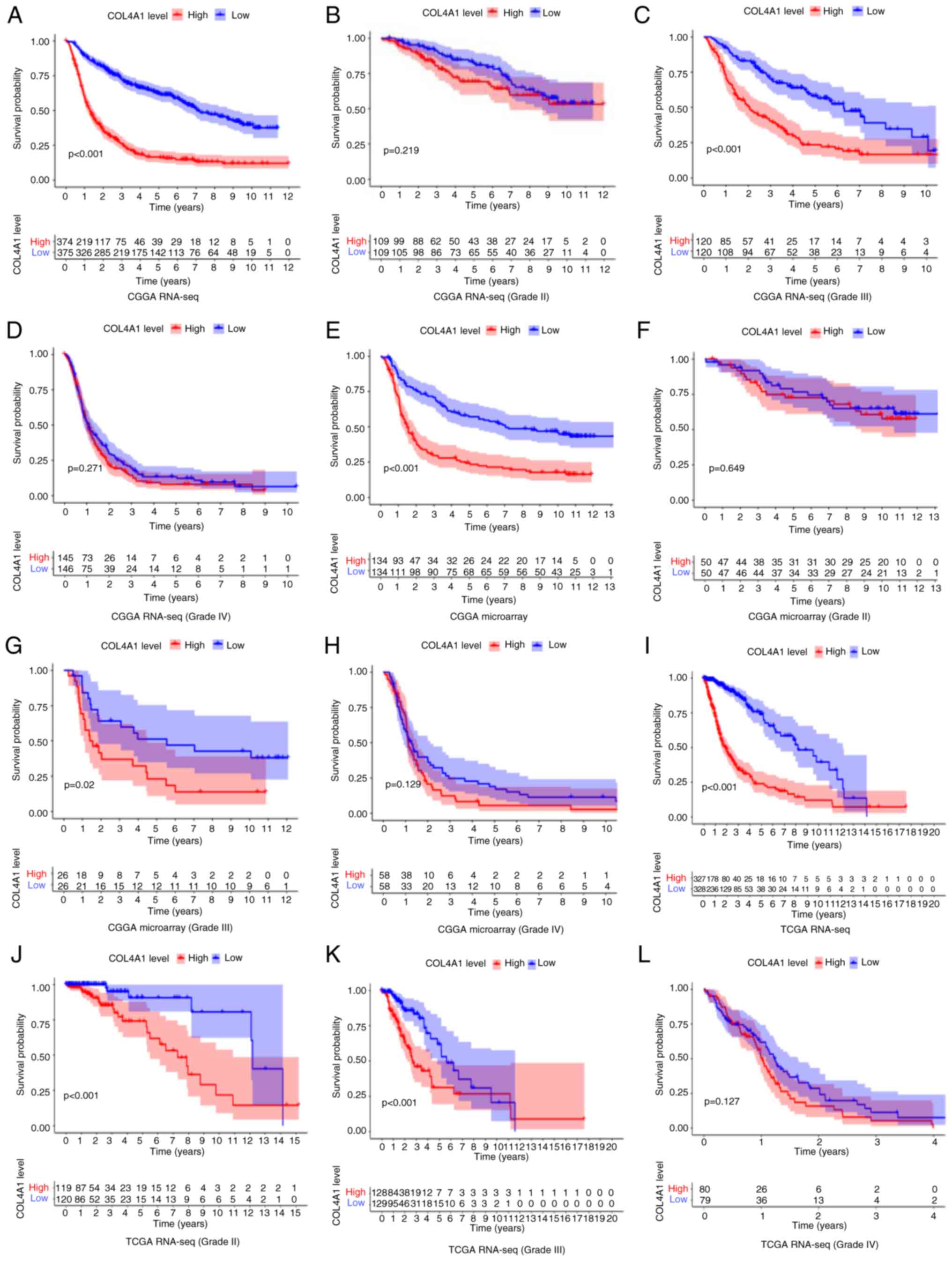
analyzed to determine this (Fig.
5). The overall survival analysis of glioma patients of all
grades in the CGGA RNA-seq, CGGA microarray and TCGA RNA-seq
datasets is provided in Fig. 5A,
E and I, respectively. The results obtained with
the three datasets all suggested that the survival time of patients
was significantly shortened in the COL4A1 high expression group
(P<0.001). In the World Health Organization (WHO) grade III
category, the three sets of data had consistent results in terms of
high expression of COL4A1 being associated with a reduction in
patient survival (Fig. 5C, G and K).
Among the remaining data, only the TCGA RNA-seq (Grade II) results
indicated that high expression of COL4A1 is associated with poor
prognosis of patients (Fig. 5J),
while the results obtained with the other datasets for Grade II and
all datasets for Grade IV were not statistically significant
(Fig. 5B, D, F,
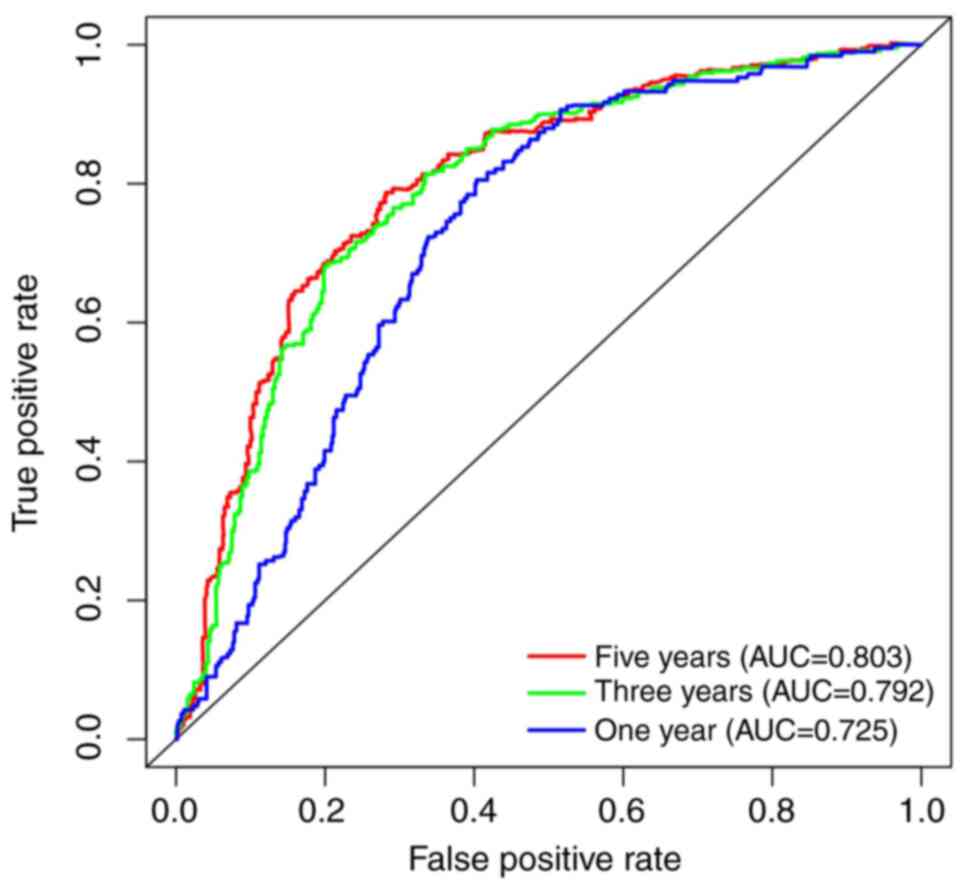
H and L). To further evaluate the diagnostic
value of COL4A1 for patients with glioma, receiver operating
characteristic curves were plotted based on the CGGA RNA-seq data
(Fig. 6), using survival time and
survival status as cutoff levels. The area under the curve values
for COL4A1 to predict one-, three- and five-year survival were
0.725, 0.792 and 0.803, respectively, further reiterating the
observation that the prognosis of patients with glioma and high
COL4A1 expression levels was poor.
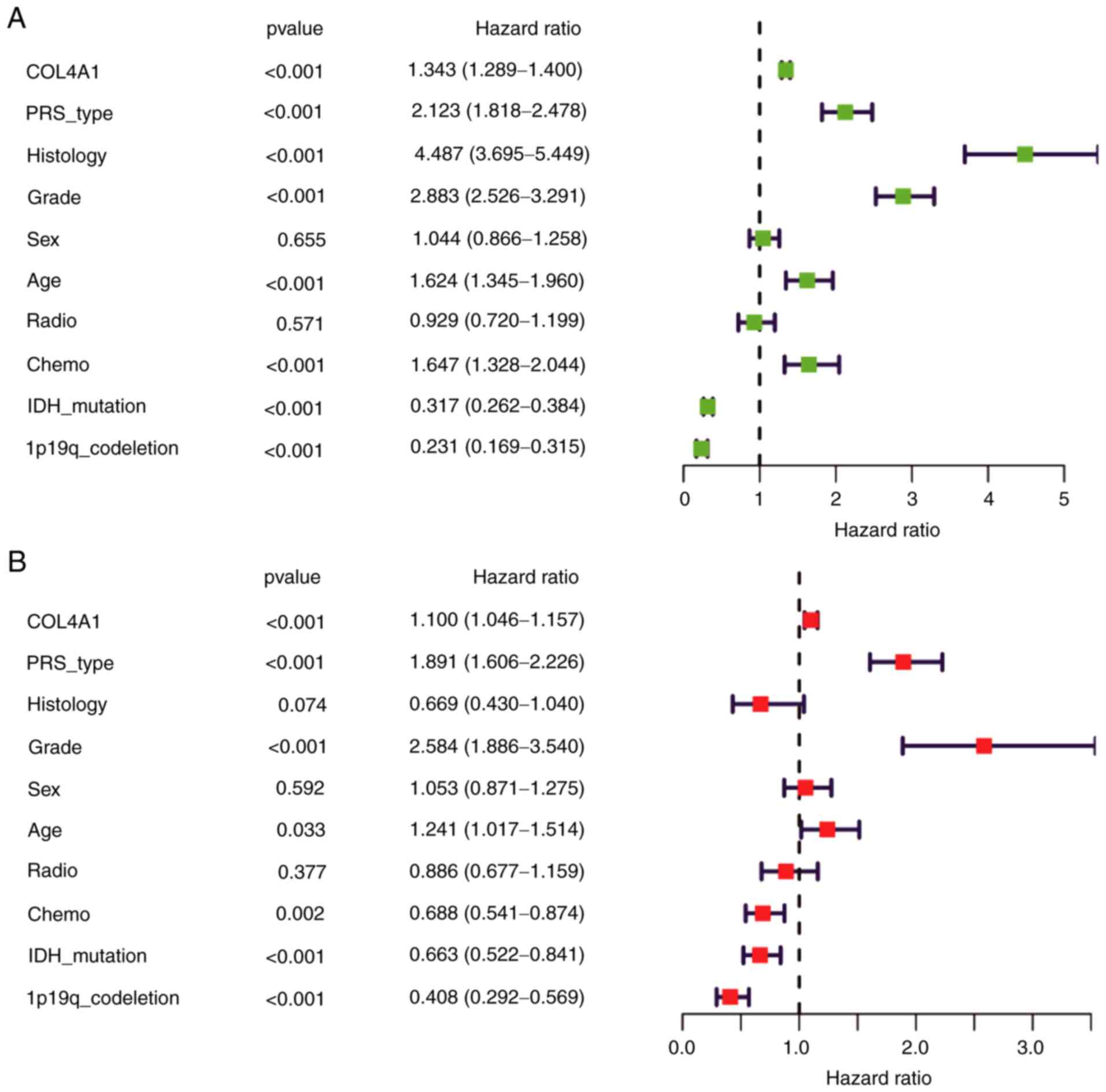
Univariate and multivariate
analyses
As presented in Fig.
7A, univariate regression analysis confirmed the result that
prognosis was poor in patients with high COL4A1 expression levels
in their glioma tissues [hazard ratio (HR): 1.343, 95% CI:
1.289-1.400, P<0.001]. Furthermore, PRS-type (HR: 2.123, 95% CI:
1.818-2.478, P<0.001), Histology (HR: 4.487, 95% CI:
3.695-5.449, P<0.001), high grade (HR: 2.883, 95% CI:
2.526-3.291, P<0.001), age (HR=1.624, 95% CI: 1.345-1.960,
P<0.001), chemotherapy (HR: 1.647, 95% CI: 1.328-2.044,
P<0.005), IDH mutation status (HR: 0.317, 95% CI: 0.262-0.384,
P<0.001) and patients with 1p19q codeletion status (HR: 0.231,
95% CI: 0.169-0.315, P<0.001) significantly influenced the
survival prognosis. Multivariate regression analysis was also
performed (Fig. 7B), and the
results indicated that COL4A1 expression levels (HR: 1.100, 95% CI:
1.046-1.157, P<0.001), PRS-type (HR: 1.891, 95% CI: 1.606-2.226,
P<0.001), high grade (HR: 2.584, 95% CI: 1.886-3.540,
P<0.001), age (HR: 1.241, 95% CI: 1.017-1.514, P=0.033),
chemotherapy (HR: 0.688, 95% CI: 0.541-0.874, P=0.002), IDH
mutation status (HR: 0.663, 95% CI: 0.522-0.841, P<0.001) and
1p19q codeletion status (HR: 0.408, 95% CI: 0.292-0.569,
P<0.001) were independently associated with survival. These
results indicated that COL4A1 is a potential prognostic factor for
glioma and increased expression levels of COL4A1 may lead to poor
prognosis.
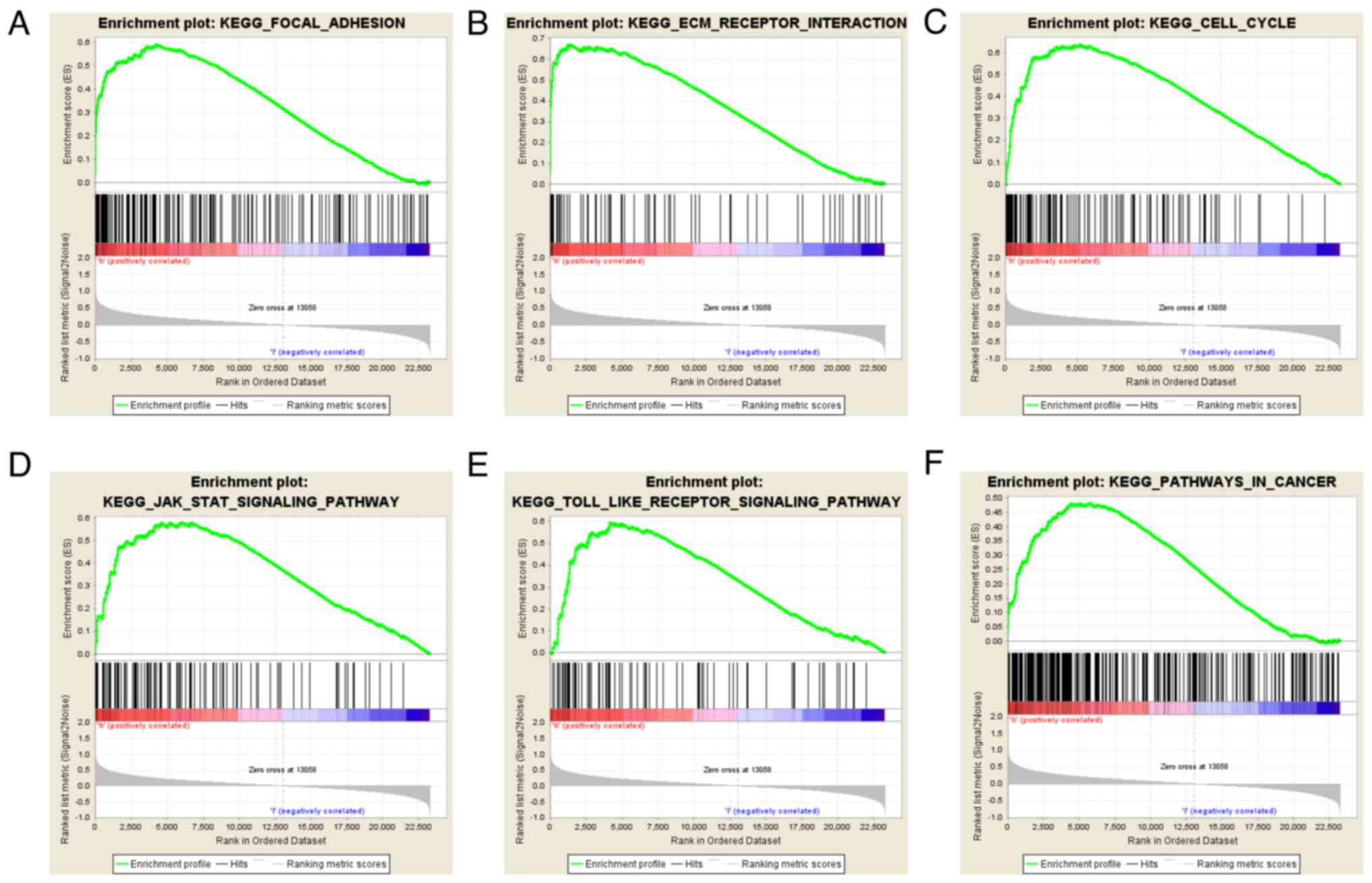
GSEA identifies a COL4A1-associated
signaling pathway
GSEA is a commonly used gene enrichment analysis
tool, and in the present study, GSEA was performed to identify
signaling pathways associated with glioma development and compare
datasets with low and high COL4A1 expression. An FDR value of
<0.05 and a NOM P-value of <0.05 were considered to indicate
statistical significance. Among the factors analyzed, extracellular
matrix (ECM) receptor interaction, JAK/STAT signaling pathway, cell
cycle, focal adhesion and Toll-like receptor signaling pathway
exhibited significant enrichment in the COL4A1 high expression
group (Fig. 8; Table I). These results suggested that
COL4A1 has an important role in the pathogenesis of glioma through
these biological pathways.
 | Table IGene set enrichment in the high
collagen α-1 (IV) chain expression phenotype. |
Table I
Gene set enrichment in the high
collagen α-1 (IV) chain expression phenotype.
| Gene set name | NES | NOM Q-value | FDR Q-value |
|---|
|
KEGG_FOCAL_ADHESION | 2.09 | <0.01 | <0.01 |
|
KEGG_ECM_RECEPTOR_INTERACTION | 2.02 | <0.01 | <0.01 |
|
KEGG_CELL_CYCLE | 1.93 | <0.01 | 0.02 |
|
KEGG_JAK_STAT_SIGNALING_PATHWAY | 1.83 | <0.01 | 0.05 |
|
KEGG_TOLL_LIKE_RECEPTOR_SIGNALING_PATHWAY | 1.79 | <0.01 | 0.05 |
|
KEGG_PATHWAYS_IN_CANCER | 1.81 | <0.01 | 0.06 |
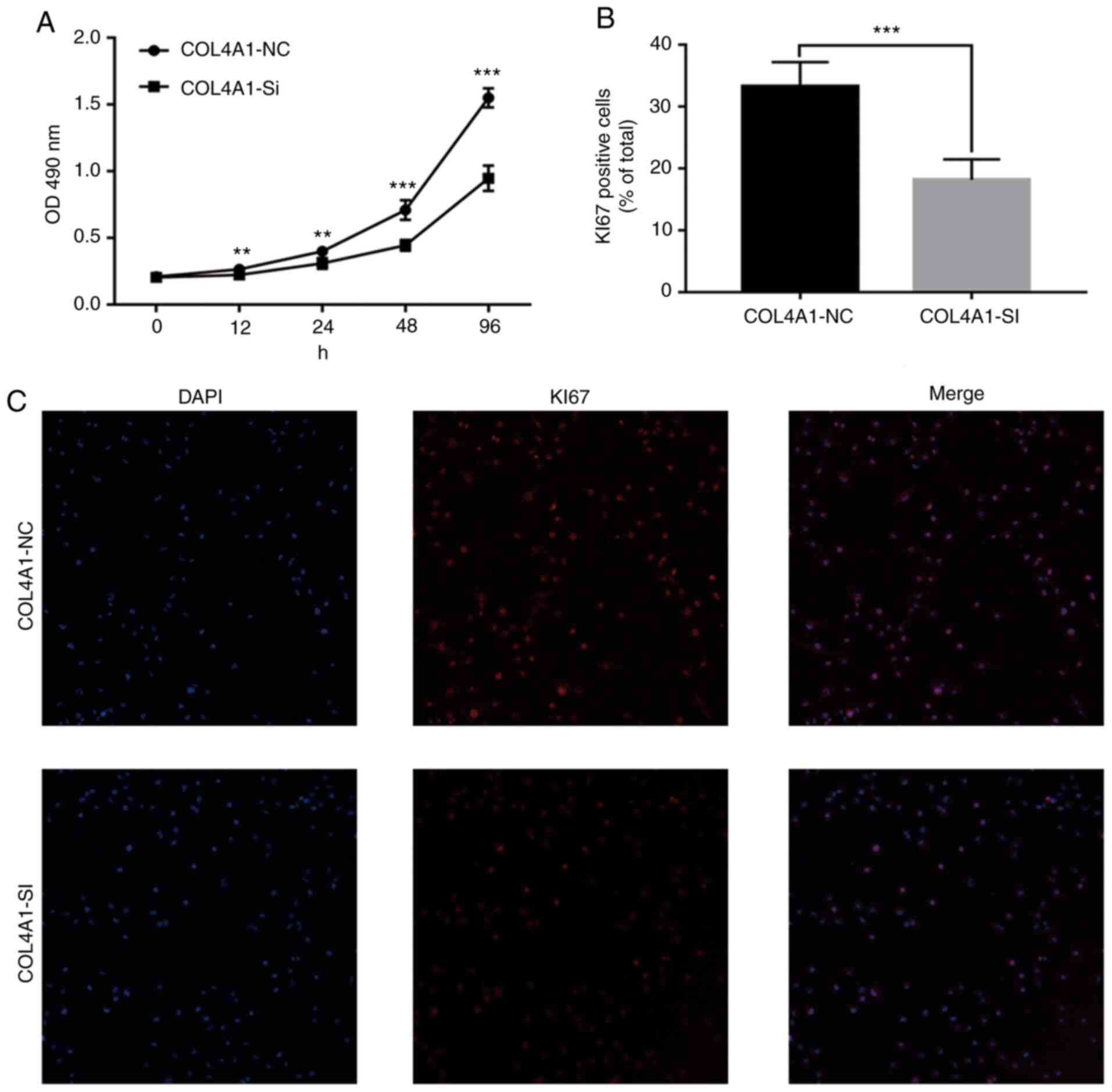
Knockdown of COL4A1 inhibits the
malignant behavior of LN229 cells
To assess the effect of COL4A1 on the malignant
behavior of glioma cell lines, siRNA was used to interfere with the
expression of COL4A1. The knockdown efficacy is provided in
Fig. S1. An MTT assay then
indicated that the proliferation ability of cells in the
experimental group was significantly reduced (Fig. 9A). Furthermore, immunofluorescence
staining suggested that the expression levels of the cell
proliferation marker KI67 were significantly lower than those in
the control group (Fig. 9B and
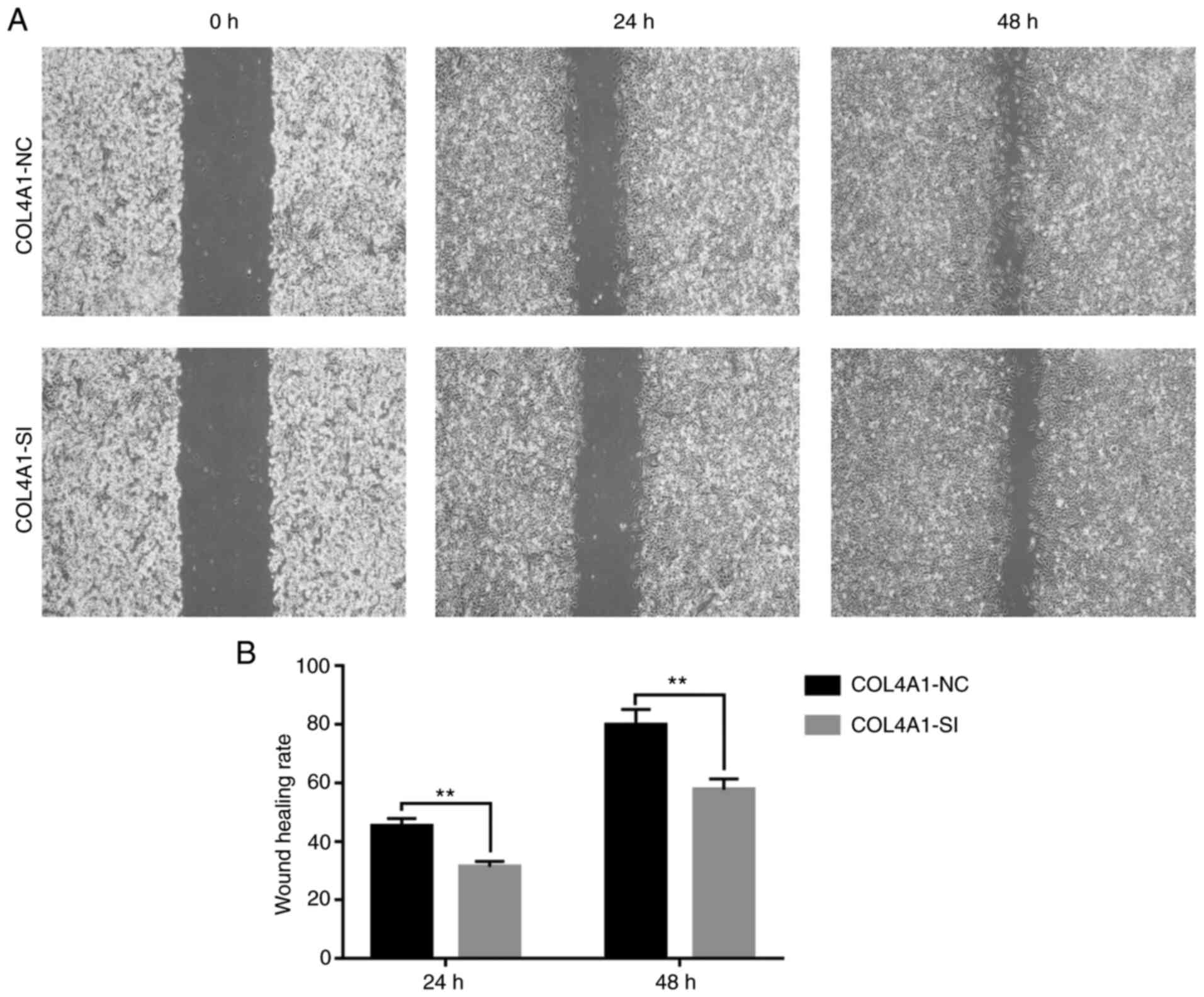
C). Finally, a wound-healing assay
indicated that compared with the control group, the scar healing
ability in the experimental group was significantly reduced
(Fig. 10).
Discussion
Various studies have indicated that COL4A1 is
involved in the pathological process of a variety of cancer types
and has a key role in their occurrence, development and metastasis
(24-26).
In addition, certain studies suggested that COL4A1 is a novel
oncogenic transcription regulator that is closely associated with
the prognosis of glioma (27). The
present study aimed to determine the role of COL4A1 in glioma using
a variety of different data types and experimental techniques,
including sequencing data, chip data, RT-PCR, in situ
hybridization and immunohistochemistry. The results indicated that
the expression levels of COL4A1 in glioma tissues were
significantly higher than those in normal brain tissues. This is
supported by previous reports that COL4A1 expression is increased
in malignant tumors (16,24,25,28).
For instance, COL4A1 expression is also increased in numerous types
of malignant tumor tissue (such as esophageal squamous cell
carcinoma, renal cell carcinoma, urothelial carcinoma of the
bladder and breast cancer) and has an important impact on the
occurrence and development of cancer (16,20,24,28).
Of note, COL4A1 has also been reported to be
increased in LGG. Jiang et al (27) indicated that COL4A1 has synergistic
effects with COL1A2, COL3A1, COL1A1, COL4A2 and COL5A2 and is
associated with poor prognosis of patients. However, that study
only relied on a single data source and only revealed the
expression changes of COL4A1 in LGG and the prognosis of patients,
but did not fully explain the role of COL4A1 in different grades of
gliomas. In addition, since Jiang et al (27) did not explore the clinical
diagnostic value of COL4A1, whether COL4A1 was associated with
clinical characteristics of glioma patients remains unknown.
Finally, the six genes reported by the authors were suggested to
have regulatory effects in LGG, but in the mechanistic analysis,
certain results indicated no association with the occurrence of
cancer, and further investigation is required to empirically
determine their effect. Although the present study and that
performed by Jiang et al (27) are based on the analysis of public
data, the present study is the first to explore the role of COL4A1
in glioma. In addition, the present study also performed
experiments using glioma cells and clinical samples.
The present study objectively confirmed the
increased expression level of COL4A1 in glioma tissues through
public database data from GEPIA, GEO, Ivy and HPA in addition to
tissue and cell lines. The relationship between COL4A1 and the
prognostic diagnostic value and clinical characteristics of glioma
patients also warrants further exploration. Therefore, the CGGA
database was used to obtain clinically relevant traits of glioma
patients and the association of the expression of COL4A1 with
clinicopathological features was assessed through a Mann-Whitney
U-test or Kruskal-Wallis test. Of note, the expression level of
COL4A1 increased with the WHO grade and was significantly increased
in recurrent and secondary glioma. Previous studies have also
indicated that abnormally high expression of COL4A1 in gastric
carcinoma may lead to decreased overall survival of patients and
has a close relationship with tumor recurrence (16).
Furthermore, the present study revealed for the
first time that COL4A1 expression, particularly in WHO grade III
tumors, is negatively associated with patient prognosis, which
confirms that COL4A1 is associated with poor prognosis in glioma
patients. In addition, COL4A1 was revealed to be an independent
risk factor leading to poor prognosis in the multivariate analysis.
The above results suggest that COL4A1 may be involved in the
malignant progression of glioma. Therefore, an MTT assay, KI67
staining and wound-healing assay further confirmed that knockdown
of COL4A1 inhibited the proliferation and migration ability of
glioma cells. Taken together, the present results suggest that
COL4A1 is indeed an oncogene in gliomas, and a similar role has
been reported in breast cancer (14).
To further understand the role of COL4A1 in the
development and progression of gliomas and the cellular pathways
involved, a functional annotation by GSEA was performed. The
results suggested that in the high-expression COL4A1 group, focal
adhesion, ECM receptor interaction, JAK/STAT signaling pathway,
Toll-like receptor signaling pathway and cell cycle were
significantly enriched. Focal adhesion is overexpressed in numerous
tumor types and has an important role in cell cycle regulation,
adhesion, migration and formation through various signaling
pathways (29-31).
Compared with normal brain tissue, FAK expression was higher in
glioma cells, and it was negatively correlated with survival time
(32). Some studies have shown that
focal adhesion proteins and proline-containing proteins were linked
together by a tyrosine phosphatase to form a complex, which in turn
controls the invasion of glioblastoma cells (33). Another study has also indicated that
focal adhesion kinase promotes the formation of small blood vessels
in high-grade malignant gliomas (34). The occurrence, development, invasion
and metastasis of malignant tumors are frequently accompanied by
changes in the ECM and its cell surface receptor expression.
Previous studies have indicated that ECM components are connected
to glioma cells through hyaluronic acid receptor molecules, and
their interaction is mainly controlled by glycosidase and protease.
These enzymes regulate the adhesion of the ECM to the intracellular
matrix and also activate growth molecules and chemokines in glioma
cells (35). The JAK/STAT signaling
pathway is a common signal transduction pathway for numerous
cytokines. It is widely involved in cell proliferation,
differentiation, apoptosis and inflammation, and promotes the
occurrence and development of various diseases, including
inflammatory diseases, lymphoma, leukemia and formation of solid
tumors. Henrik Heiland et al (36) indicated that JAK/STAT pathway
activation was significantly increased in glioma samples compared
to healthy tissue samples, which is consistent with the present
results. Furthermore, several studies suggested that inhibition of
the JAK/STAT signaling pathway may inhibit the proliferation,
migration and invasion of glioma cells (37-39).
Toll-like receptor signaling is a well-known cancer pathway.
Numerous human tumors and tumor cell lines express Toll-like
receptors, which have an important role in the progression from
precancerous lesions to tumors. Hu et al (40) indicated that versican released by
glioma cells promotes tumor proliferation through Toll-like
receptor 2 signal transduction and expression of membrane type-1
matrix metalloproteinase. It has been suggested that deregulation
of miRNAs in glioma cells may promote tumor proliferation by
directly acting on key cell-cycle regulators. For instance, lncRNA
breast cancer anti-estrogen resistance 4, the earliest
anti-estrogen resistance found in breast cancer, was reported to
accelerate the progression of glioma cells by affecting the cell
cycle and inhibiting apoptosis (41). Therefore, COL4A1 promoting the
activation of oncogenic pathways may be a major cause of poor
prognosis in glioma patients.
Although a large amount of data was analyzed in the
present study to demonstrate that COL4A1, as a novel oncogene, is
involved in the pathological processes of glioma, there are various
limitations. First, there are certain shortcomings in the public
database platform, including the lack of clinical characteristics
for various patients, and it was not possible to include the
details of clinical treatment for all patients. However, the
database has unique advantages, such as multi-center cohorts, a
large sample size and ethnic diversity, which may provide
information that may improve the prediction, diagnosis, treatment
and monitoring of tumors. Furthermore, COL4A1 as a novel oncogene
in glioma may be involved in its molecular mechanisms. By using
GSEA, it was indirectly revealed that multiple signaling pathways
of COL4A1 are involved in the pathological processes of glioma.
However, it is well known that a gene may have a variety of roles
in organisms. The present study provides an index for further
research to study the role of COL4A1 as an oncogene in the
pathological processes of glioma.
In conclusion, through a series of analytical
methods, the present study confirmed that abnormally high
expression of COL4A1 is an independent predictor of poor prognosis
in glioma patients, and possible molecular pathways underlying its
oncogenic effect were elucidated. The present study revealed part
of the complex pathological process of glioma from the perspective
of molecular biology and provided COL4A1 as an effective potential
diagnostic and therapeutic target.
Supplementary Material
Confirmation of knockdown of COL4A1
expression by reverse transcription-quantitative PCR.
***P<0.001 vs. COL4A1-NC. COL4A1, collagen α-1 (IV)
chain; MOCK, mock transfection; NC, negative control; SI, short
interfering RNA.
Characteristics of patients with
glioma based on the Chinese Glioma Genome Atlas.
Acknowledgements
Not applicable.
Funding
Funding: This study was supported by the 2018 Henan Provincial
Medical Science and Technology Tackling Program
Provincial-ministerial Co-construction Project (grant no.
SBGJ2018076) and the Central Plains Thousand Talents Plan of Henan
Province (grant no. ZYQR201912122).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
HBW, YZG and ZDL conceived and designed the study.
AL is responsible for collecting clinical samples and collating raw
data from public databases. JLW, JTL, BFL and XYL performed the
experiments. BZ, BP and LYL analyzed the data and prepared figures
and/or tables. YZG contributed reagents/materials/analysis tools,
wrote and reviewed drafts of the manuscript and approved the final
manuscript. HBW and ZDL can authenticate the raw data in the study.
All authors read and approved the final manuscript.
Ethics approval and consent to
participate
All procedures performed in the study were in
accordance with the ethical standards of the Ethical Committee of
the Henan Provincial People's Hospital (Zhengzhou, China). The
experimental scheme was also approved by the Ethics Committee of
Henan Provincial People's Hospital (Zhengzhou, China). All patients
signed an informed consent and agreed to use their glioma tissues
or normal brain tissues after surgical resection for relevant
studies.
Patient consent for publication
Not applicable.
Competing interests
The authors declare they have no competing
interests.
References
|
1
|
Ostrom QT, Gittleman H, Farah P, Ondracek
A, Chen Y, Wolinsky Y, Stroup NE, Kruchko C and Barnholtz-Sloan JS:
CBTRUS statistical report: Primary brain and central nervous system
tumors diagnosed in the United States in 2006-2010. NeuroOncol. 15
(Suppl 2):ii1–ii56. 2013.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Ohgaki H and Kleihues P: Epidemiology and
etiology of gliomas. Acta Neuropathol. 109:93–108. 2005.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Ostrom QT, Bauchet L, Davis FG, Deltour I,
Fisher JL, Langer CE, Pekmezci M, Schwartzbaum JA, Turner MC, Walsh
KM, et al: The epidemiology of glioma in adults: A ‘state of the
science’ review. Neuro Oncol. 16:896–913. 2014.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Claus EB and Black PM: Survival rates and
patterns of care for patients diagnosed with supratentorial
low-grade gliomas: Data from the SEER program, 1973-2001. Cancer.
106:1358–1363. 2006.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Yang P, Wang Y, Peng X, You G, Zhang W,
Yan W, Bao Z, Wang Y, Qiu X and Jiang T: Management and survival
rates in patients with glioma in China (2004-2010): A retrospective
study from a single-institution. J Neurooncol. 113:259–266.
2013.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Delgado-López PD and Corrales-García EM:
Survival in glioblastoma: A review on the impact of treatment
modalities. Clin Transl Oncol. 18:1062–1071. 2016.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Omuro A and DeAngelis LM: Glioblastoma and
other malignant gliomas: A clinical review. JAMA. 310:1842–1850.
2013.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Lohr J, Ratliff T, Huppertz A, Ge Y,
Dictus C, Ahmadi R, Grau S, Hiraoka N, Eckstein V, Ecker RC, et al:
Effector T-cell infiltration positively impacts survival of
glioblastoma patients and is impaired by tumor-derived TGF-β. Clin
Cancer Res. 17:4296–4308. 2011.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Murat A, Migliavacca E, Gorlia T, Lambiv
WL, Shay T, Hamou MF, de Tribolet N, Regli L, Wick W, Kouwenhoven
MC, et al: Stem cell-related ‘self-renewal’ signature and high
epidermal growth factor receptor expression associated with
resistance to concomitant chemoradiotherapy in glioblastoma. J Clin
Oncol. 26:3015–3024. 2008.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Hale JS, Otvos B, Sinyuk M, Alvarado AG,
Hitomi M, Stoltz K, Wu Q, Flavahan W, Levison B, Johansen ML, et
al: Cancer stem cell-specific scavenger receptor CD36 drives
glioblastoma progression. Stem Cells. 32:1746–1758. 2014.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Pollner R, Schmidt C, Fischer G, Kühn K
and Pöschl E: Cooperative and competitive interactions of
regulatory elements are involved in the control of divergent
transcription of human Col4A1 and Col4A2 genes. FEBS Lett.
405:31–36. 1997.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Yurchenco PD: Basement membranes: Cell
scaffoldings and signaling platforms. Cold Spring Harb Perspect
Biol. 3(a004911)2011.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Labelle-Dumais C, Schuitema V, Hayashi G,
Hoff K, Gong W, Dao DQ, Ullian EM, Oishi P, Margeta M and Gould DB:
COL4A1 mutations cause neuromuscular disease with tissue-specific
mechanistic heterogeneity. Am J Hum Genet. 104:847–860.
2019.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Jin R, Shen J, Zhang T, Liu Q, Liao C, Ma
H, Li S and Yu Z: The highly expressed COL4A1 genes contributes to
the proliferation and migration of the invasive ductal carcinomas.
Oncotarget. 8:58172–58183. 2017.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Kitzler TM, Schneider R, Kohl S,
Kolvenbach CM, Connaughton DM, Dai R, Mann N, Nakayama M, Majmundar
AJ, Wu CW, et al: COL4A1 mutations as a potential novel cause of
autosomal dominant CAKUT in humans. Hum Genet. 138:1105–1115.
2019.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Li F, Wang NN, Chang X, Wang SL, Wang LS,
Yao J, Li ZS and Bai Y: Bioinformatics analysis suggests that
COL4A1 may play an important role in gastric carcinoma recurrence.
J Dig Dis. 20:391–400. 2019.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Lin S, Xia C, He S, Yang J, Li H, Zheng J,
Liu M and You C: Genetic variations of the COL4A1 gene and
intracerebral hemorrhage risk: A case-control study in a Chinese
han population. World Neurosurg. 112:e527–e533. 2018.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Raza ST, Abbas S, Eba A, Karim F, Wani IA,
Rizvi S, Zaidi A and Mahdi F: Association of COL4A1 (rs605143,
rs565470) and CD14 (rs2569190) genes polymorphism with coronary
artery disease. Mol Cell Biochem. 445:117–122. 2018.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Saskin A, Sillon G, Palfreeman N and Buhas
D: COL4A1/2 CNVs and cerebral small vessel disease: Narrowing in on
the critical chromosomal region. Neurology. 90:1026–1028.
2018.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Chen FF, Zhang SR, Peng H, Chen YZ and Cui
XB: Integrative genomics analysis of hub genes and their
relationship with prognosis and signaling pathways in esophageal
squamous cell carcinoma. Mol Med Rep. 20:3649–3660. 2019.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Pan Z, Li L, Fang Q, Zhang Y, Hu X, Qian Y
and Huang P: Analysis of dynamic molecular networks for pancreatic
ductal adenocarcinoma progression. Cancer Cell Int.
18(214)2018.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Subramanian A, Kuehn H, Gould J, Tamayo P
and Mesirov JP: GSEA-P: A desktop application for gene set
enrichment analysis. Bioinformatics. 23:3251–3253. 2007.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) methods. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Wang SM, Chen PM, Sung YW, Huang WC, Huang
HS and Chu PY: Effect of COL4A1 expression on the survival of
neoadjuvant chemotherapy breast cancer patients. J Oncol.
2020(5209695)2020.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Wang T, Jin H, Hu J, Li X, Ruan H, Xu H,
Wei L, Dong W, Teng F, Gu J, et al: COL4A1 promotes the growth and
metastasis of hepatocellular carcinoma cells by activating FAK-Src
signaling. J Exp Clin Cancer Res. 39(148)2020.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Wu F, Li F, Lin X, Xu F, Cui RR, Zhong JY,
Zhu T, Shan SK, Liao XB, Yuan LQ and Mo ZH: Exosomes increased
angiogenesis in papillary thyroid cancer microenvironment. Endocr
Relat Cancer. 26:525–538. 2019.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Jiang Y, He J, Guo Y, Tao H, Pu F and Li
Y: Identification of genes related to low-grade glioma progression
and prognosis based on integrated transcriptome analysis. J Cell
Biochem. 121:3099–3111. 2020.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Miyake M, Morizawa Y, Hori S, Tatsumi Y,
Onishi S, Owari T, Iida K, Onishi K, Gotoh D, Nakai Y, et al:
Diagnostic and prognostic role of urinary collagens in primary
human bladder cancer. Cancer Sci. 108:2221–2228. 2017.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Aboubakar Nana F, Vanderputten M and Ocak
S: Role of focal adhesion kinase in small-cell lung cancer and its
potential as a therapeutic target. Cancers (Basel).
11(1683)2019.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Giaginis CT, Vgenopoulou S, Tsourouflis
GS, Politi EN, Kouraklis GP and Theocharis SE: Expression and
clinical significance of focal adhesion kinase in the two distinct
histological types, intestinal and diffuse, of human gastric
adenocarcinoma. Pathol Oncol Res. 15:173–181. 2009.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Ozkal S, Paterson JC, Tedoldi S, Hansmann
ML, Kargi A, Manek S, Mason DY and Marafioti T: Focal adhesion
kinase (FAK) expression in normal and neoplastic lymphoid tissues.
Pathol Res Pract. 205:781–788. 2009.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Ding L, Sun X, You Y, Liu N and Fu Z:
Expression of focal adhesion kinase and phosphorylated focal
adhesion kinase in human gliomas is associated with unfavorable
overall survival. Transl Res. 156:45–52. 2010.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Chen Z, Morales JE, Guerrero PA, Sun H and
McCarty JH: PTPN12/PTP-PEST regulates phosphorylation-dependent
ubiquitination and stability of focal adhesion substrates in
invasive glioblastoma cells. Cancer Res. 78:3809–3822.
2018.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Haskell H, Natarajan M, Hecker TP, Ding Q,
Stewart J Jr, Grammer JR and Gladson CL: Focal adhesion kinase is
expressed in the angiogenic blood vessels of malignant astrocytic
tumors in vivo and promotes capillary tube formation of brain
microvascular endothelial cells. Clin Cancer Res. 9:2157–2165.
2003.PubMed/NCBI
|
|
35
|
Ferrer VP, Moura Neto V and Mentlein R:
Glioma infiltration and extracellular matrix: Key players and
modulators. Glia. 66:1542–1565. 2018.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Henrik Heiland D, Ravi VM, Behringer SP,
Frenking JH, Wurm J, Joseph K, Garrelfs NWC, Strähle J, Heynckes S,
Grauvogel J, et al: Tumor-associated reactive astrocytes aid the
evolution of immunosuppressive environment in glioblastoma. Nat
Commun. 10(2541)2019.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Wang P, Peng X, Zhang J, Wang Z, Meng J,
Cen B, Ji A and He S: LncRNA-135528 inhibits tumor progression by
up-regulating CXCL10 through the JAK/STAT pathway. Apoptosis.
23:651–666. 2018.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Xu CH, Liu Y, Xiao LM, Chen LK, Zheng SY,
Zeng EM, Li DH and Li YP: Silencing microRNA-221/222 cluster
suppresses glioblastoma angiogenesis by suppressor of cytokine
signaling-3-dependent JAK/STAT pathway. J Cell Physiol.
234:22272–22284. 2019.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Zhang P, Chen FZ, Jia QB and Hu DF:
Upregulation of microRNA-133a and downregulation of connective
tissue growth factor suppress cell proliferation, migration, and
invasion in human glioma through the JAK/STAT signaling pathway.
IUBMB Life. 71:1857–1875. 2019.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Hu F, Dzaye O, Hahn A, Yu Y, Scavetta RJ,
Dittmar G, Kaczmarek AK, Dunning KR, Ricciardelli C, Rinnenthal JL,
et al: Glioma-derived versican promotes tumor expansion via
glioma-associated microglial/macrophages Toll-like receptor 2
signaling. Neuro Oncol. 17:200–210. 2015.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Wei L, Yi Z, Guo K and Long X: Long
noncoding RNA BCAR4 promotes glioma cell proliferation via
EGFR/PI3K/AKT signaling pathway. J Cell Physiol. 234:23608–23617.
2019.PubMed/NCBI View Article : Google Scholar
|