Introduction
In total, >1.8 million new colorectal cancer
cases and 881,000 deaths are estimated to have occurred in 2018,
accounting for ~1 in 10 cancer cases and deaths worldwide (1). Colorectal cancer ranks third in
incidence and second in mortality globally, and poses a serious
threat to human health, particularly in certain European countries
(e.g., Hungary, Slovenia, Slovakia, The Netherlands and Norway),
Australia/New Zealand, Northern America and Eastern Asia (including
Japan, the Republic of Korea and Singapore) (1). The currently approved therapeutic
approaches to colon adenocarcinoma include surgery, radiofrequency
ablation, chemotherapy, radiation therapy, cryosurgery and targeted
therapy (2).
Long non-coding RNAs (lncRNAs), a class of RNAs
>200 nt long and lacking protein-coding ability, have emerged to
play key roles in numerous biological processes, such as
differentiation, development, cellular address codes and
oncogenesis (3-5).
The homeobox transcript antisense intergenic RNA (HOTAIR) is a
well-known long non-coding RNA that functions as an oncogene
(6,7). Abnormal expression of HOTAIR has been
reported in multiple cancers, and may lead to abnormal cell
differentiation and proliferation, invasion and metastasis
(8-11).
Located in chromosome 12 (q13.13), homeobox C11
(HOXC11) belongs to the homeobox family of genes, which encode a
highly conserved family of transcription factors that play key
roles in embryonic implantation, evolution and morphogenesis in all
multicellular organisms (12-14).
Disorders of HOXC11 may lead to abnormal embryonal development,
endometriosis and infertility (14,15).
HOTAIR was reported to regulate the expression of homeobox genes
and is located in the antisense chain of HOXC11, where an
overlapping region between HOTAIR and HOXC11 has been identified
(16). Thus, the present study was
undertaken to explore the interaction between HOTAIR and HOXC11.
Data from The Cancer Genome Atlas (TCGA) database were analyzed,
and a co-expression trend was found between the lncRNA HOTAIR and
the transcription factor HOXC11 in several types of cancer, such as
colon adenocarcinoma, esophageal carcinoma, breast cancer and
kidney renal clear cell carcinoma, among others. Therefore, assays
with colon adenocarcinoma cells and tissues were performed to
identify the interaction between HOTAIR and HOXC11 and their
functions in colon adenocarcinoma.
Materials and methods
Cell cultures
The SW480 and HCT116 colon cancer cell lines were
purchased from the Shanghai Institute of Cell Biology (Shanghai,
China). SW480 cells were cultured in Leibovitz's L-15 Medium
(Gibco; Thermo Fisher Scientific, Inc.) supplemented with 10% FBS
(Gibco; Thermo Fisher Scientific, Inc.) at 37˚C in a humidified
atmosphere containing 5% CO2. HCT116 cells were cultured
in DMEM (Gibco; Thermo Fisher Scientific, Inc.) supplemented with
10% FBS (Gibco; Thermo Fisher Scientific, Inc.) at 37˚C in a
humidified atmosphere containing 5% CO2.
Specimens
A total of 12 paired colon carcinoma and adjacent
normal tissue samples were obtained from the Fifth Affiliated
Hospital of Sun Yat-sen University (Zhuhai, China) between 18th
February and 6th December 2018. All patients were diagnosed with
primary colon carcinoma. Ethics approval (approval no. K206-1) was
obtained from the Research Ethics Committee of the Fifth Affiliated
Hospital of Sun Yat-sen University. Informed consent for their
tissues to be used for research purposes was obtained from all
patients prior to performing this study.
Bioinformatics analysis
The gene expression data of colon carcinoma,
esophageal carcinoma, breast cancer and kidney renal clear cell
carcinoma were obtained from TCGA website (https://cancergenome.nih.gov/). Kaplan-Meier analysis
was plotted by UALCAN (17).
Overlapping region was found between HOXC11 and HOTAIR by UCSC
genome browser (assembly ID, hg38) (18).
Transfection
HOXC11 knockdown lentivirus
[pSLenti-U6-shRNA(HOXC11)-CMV-EGFP-F2A-Puro-WPRE] and negative
control (NC) lentivirus (pSLenti-U6-CMV-EGFP-F2A-Puro-WPRE) were
obtained from Shanghai OBIO Technology (Shanghai) Corp., Ltd. SW480
and HCT116 cells were transfected with lentiviruses at a
multiplicity of infection (MOI) of 10 in the presence of 4 µg/ml
puromycin. After 1 week of puromycin screening, SW480 and HCT116
cells were used for subsequent experimentation. Transfection of the
HOXC11 plasmid [pcDNA3.1(+)] was performed according to the
protocol of Lipofectamine® 3000 Reagent (Invitrogen;
Thermo Fisher Scientific, Inc.). In total, 1 µg HOXC11 or negative
control plasmid, 2 µl P3000™ Reagent, and 1.5 µl Lipofectamine™
3000 Reagent were added to 50 µl Opti-MEM medium per
1x105 cells. The medium was changed after 6 h and the
transfection effect was detected after 2 days.
RNase protection assay
The composition of the RNase digestion mixture for
RNase protection assay (RPA) was as follows: 10 mM Tris-HCl (pH
7.5), 300 mM NaCl, 5 mM EDTA (pH 7.5), and 20 µl of RNase A/T1 Mix
per 1 ml of reaction mixture. RNase A/T1 can digest single-stranded
RNAs, but not RNA duplexes. The RNA samples were incubated at 37˚C
for 60 min prior to treatment with an RNAse A+T cocktail
(Sigma-Aldrich; Merck KGaA). Subsequently, the samples were
incubated at 37˚C for 30 min after the addition of the RNAse
cocktail, and treated with proteinase K, as previously described
(19). The RNA used in the RPA was
extracted from SW480 and HCT116 cells.
RT-qPCR analysis
Total RNA from SW480 and HCT116 cells was isolated
with TRIzol® reagent (Invitrogen; Thermo Fisher
Scientific, Inc.) according to the manufacturer's protocol. Fast
All-in-One RT Kit (cat. no. ES-RT001; Shanghai Yishan
Biotechnology, Co., Ltd.; https://www.esunbio.com/search-1--/u0072/u0074/u0030/u0030/u0031.html)
was used to perform the reverse transcription using the temperature
protocol of 42˚C for 15 min. GoTaq® qPCR Master Mix
A6001 (cat. no. A6001; Promega Corporation) was used to perform
qPCR. The thermocycling conditions included an initial denaturation
step (95˚C, 2 min), followed by 40 cycles of 95˚C for 15 sec and
60˚C for 1 min. The primer sequences used were as follows: HOXC11
forward, 5'-GCTACTCCTCCTGCTATGC-3' and reverse,
5'-GACGCTGTTCTTGTTGACTG-3'; HOTAIR forward,
5'-GCCAAGCACCTCTATCTC-3' and reverse, 5'-GACACTG AACGGACTCTG-3' and
GAPDH forward, 5'-AGAAGGCT GGGGCTCATTTG-3' and reverse,
5'-AGGGGCCATCCACA GTCTTC-3' The relative expression level of mRNAs
was normalized to GAPDH and was calculated using the
2-ΔΔCq method (20).
MTT assay
The cells were inoculated into 96-well plates at a
concentration of 2,000 cells per 100 µl at 37˚C with 5%
CO2 overnight. After incubation for 4 h at 37˚C with 5%
CO2, 50 µl 1X MTT reagent (Nanjing KeyGen Biotech Co.,
Ltd.) and 150 µl DMSO were added to well. The mixture was shaken
well and the absorbance was detected at 490 nm using the
SpectraMax® 340PC38 spectrophotometer (Molecular
Devices, LLC).
Statistical analysis
Statistical analyses were performed by SPSS 20.0
(IBM Corp.). Graphs were generated by GraphPad Prism 7 (GraphPad
Software, Inc.). P<0.05 from a two-tailed test was considered to
indicate statistically significant difference. Unpaired student's
t-test was used to evaluate the differences between two groups of
independent samples. Data are presented as mean ± SD. All the
results were repeated ≥three times.
Results
HOXC11 and HOTAIR expression is
increased in multiple types of cancer
After analyzing the data of colon adenocarcinoma,
esophageal carcinoma, breast cancer and kidney renal clear cell
carcinoma, which were downloaded from TCGA, it was observed that
both HOXC11 and HOTAIR were significantly upregulated in the tumor
groups compared with the normal groups (Fig. S1). Similar expression trends were
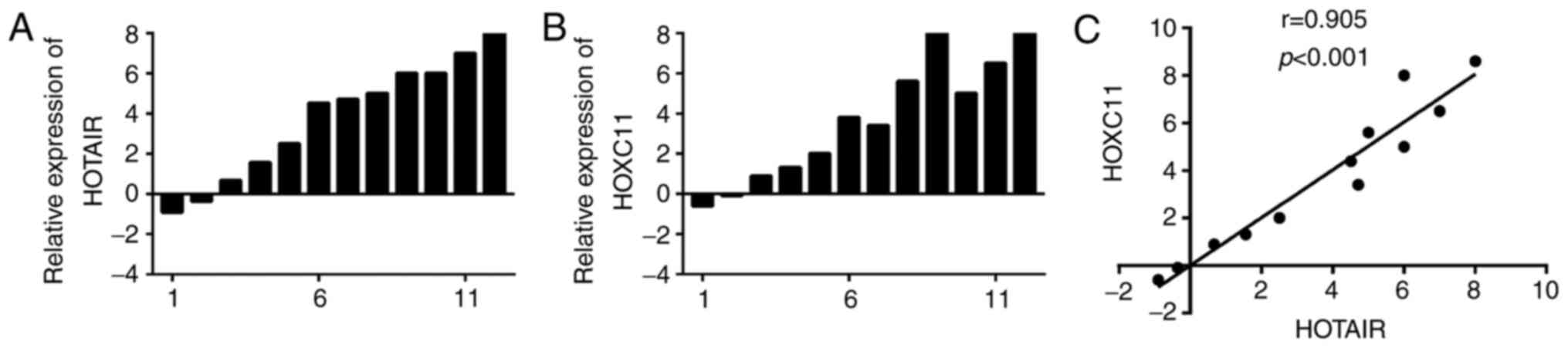
observed between HOXC11 and HOTAIR. In addition, RT-qPCR assay was
performed to detect the expression of HOXC11 and HOTAIR in 12
paired colon adenocarcinoma and adjacent normal tissue samples. It
was determined that HOTAIR and HOXC11 were upregulated in the
majority of colon adenocarcinoma tissues, which was consistent with
the results of TCGA database (Fig.
1A and B).
Co-expression of HOXC11 and
HOTAIR
A co-expression trend was observed between HOXC11
and HOTAIR in colon adenocarcinoma (r=0.797), esophageal carcinoma
(r=0.838), breast cancer (r=0.922) and kidney renal clear cell
carcinoma (r=0.719), with statistically significant differences
(supplementary Fig. S2A-D).
Similar results were obtained from colon adenocarcinoma and
adjacent normal tissues (r=0.905, P<0.001; Fig. 1C).
HOXC11 and HOTAIR expression is
associated with poor prognosis
Higher HOXC11 expression was associated with poorer
prognosis in colon adenocarcinoma (P=0.02) and kidney renal clear
cell carcinoma (P=0.006), but not in esophageal carcinoma (P=0.26)
and breast cancer (P=0.28; supplementary Fig. S3A-D). Higher expression of HOTAIR
was associated with poorer prognosis in colon adenocarcinoma
(P=0.021), breast cancer (P=0.047) and kidney renal clear cell
carcinoma (P<0.001), but there was no statistically significant
difference in the prognosis of esophageal carcinoma by HOTAIR
expression (P=0.11; supplementary Fig.
S3E-H).
Decreased expression of HOXC11
inhibits the proliferation and invasion of colon adenocarcinoma
cells
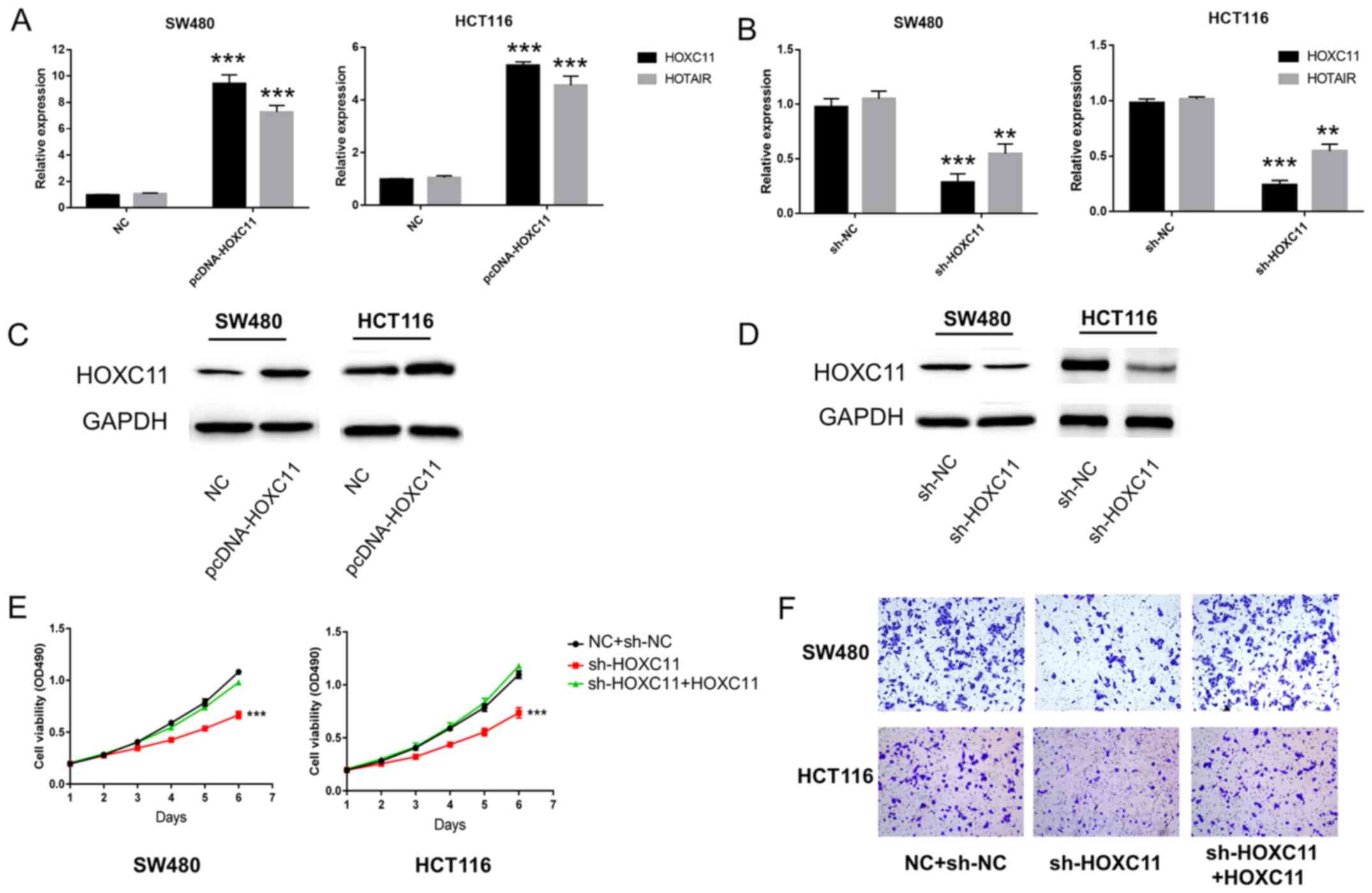
After knocking down and overexpressing HOXC11 in
SW480 cells and HCT116 cells, it was found that the expression
level of HOTAIR was modified with the changes in the expression of
HOXC11 (Fig. 2A-D). Furthermore,
decreased expression of HOXC11 inhibited the ability of
proliferation and invasion of colon adenocarcinoma cells, whereas
these effects were rescued by HOXC11 overexpression (Fig. 2E and F).
HOXC11 formed RNA duplex and increased
HOTAIR expression
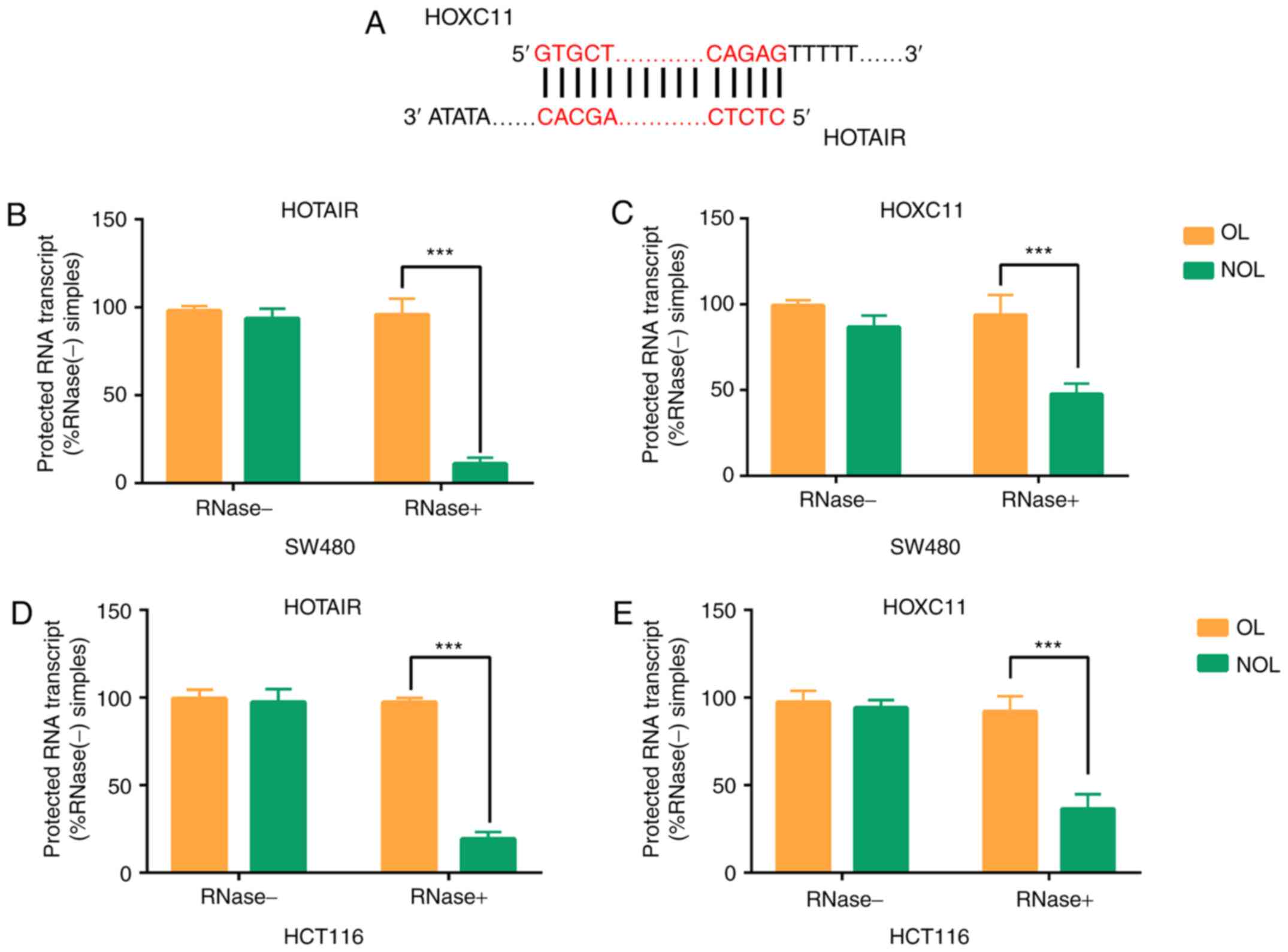
To explore the interaction between HOXC11 and
HOTAIR, HOXC11 was overexpressed and knocked down in SW480 and
HCT116 cells. It was found that HOTAIR expression changed
significantly with the changes in HOXC11 expression. According to
UCSC genome browser, an overlapping region was found between HOXC11
and HOTAIR (Fig. 3A). The results
of the RPA showed that non-overlapping regions were degraded after
adding RNAse A+T, which can digest single-stranded RNAs but not RNA
duplexes (Fig. 3B-E). Accordingly,
RNA duplex formation between HOXC11 and HOTAIR was observed.
Discussion
In the present study, it was observed that HOXC11
expression is increased in colon adenocarcinoma and may positively
regulate HOTAIR by forming an RNA duplex. The expression of HOXC11
and HOTAIR has been associated with poor prognosis in multiple
types of cancer (9,21,22).
HOXC11 belongs to the homeobox gene family, which
encodes a highly conserved family of transcription factors that
play key roles in embryo implantation, evolution and morphogenesis
in all multicellular organisms (23-26).
In recent years, there has been an increasing number of studies
indicating that HOX family genes are involved in the occurrence and
progression of several malignancies, such as gastric cancer, Wilms'
tumor and acute myeloid leukemia (27,28).
HOXC11 is reported to be involved in gene fusion, which could
promote abnormal gene expression and drive leukemogenesis (29,30).
HOXC11 was considered to play an important role in early intestinal
development during embryonic growth (31). Consistent with the present study, it
was found that the expression of HOXC11 was significantly increased
in multiple cancers, such as colon adenocarcinoma, esophageal
carcinoma, breast cancer and kidney renal clear cell carcinoma.
Furthermore, increased HOXC11 was associated with poor outcome in
colon adenocarcinoma and kidney renal clear cell carcinoma,
indicating that HOXC11 may participate in the development of these
cancers.
In recent years, lncRNAs have been reported to play
important roles in numerous biological processes, such as
development, differentiation, immunity, Alzheimer's disease,
cardiovascular disease and tumors (32,33).
HOTAIR has been extensively investigated and was reported to
participate in multiple tumors. Serum HOTAIR was considered to be a
diagnostic biomarker for esophageal squamous cell carcinoma
(34). HOTAIR may also promote the
occurrence and development of renal cell carcinoma (7). Özeş et al (35) observed that HOTAIR was implicated in
the DNA damage response, cellular senescence and chemoresistance.
Xue et al (36) reported
that HOTAIR enhances estrogen receptor signaling and confers
tamoxifen resistance in breast cancer. After analyzing the data of
multiple cancers from TCGA, the authors of the present study found
that HOTAIR was upregulated in colon adenocarcinoma, esophageal
carcinoma, breast cancer and kidney renal clear cell carcinoma, and
was associated with poor prognostic in most types of cancer.
Although HOTAIR was reported to regulate HOX gene expression, there
has been no reported research, to the authors' knowledge, on HOX
genes regulating HOTAIR expression (37). The present study was the first to
identify that HOXC11 could positively regulate HOTAIR by forming an
RNA duplex within the overlapping region.
In summary, it was herein found and validated that
the transcription factor HOXC11 and the lncRNA HOTAIR were
upregulated in colon carcinoma. In addition, the decreased
expression of HOXC11 may inhibit the proliferation and invasion
ability of colon adenocarcinoma cells. Furthermore, HOXC11 may
positively regulate HOTAIR by forming the RNA duplex. The findings
of the present study may expand our current understanding of the
functions of HOX genes and HOTAIR. Moreover, the expression of
HOXC11 and HOTAIR may be considered as a potential biomarker and
therapeutic target against colon carcinoma. However, as the main
aim of the present study was to study the association between
HOXC11 and HOTAIR, experiments in vivo were not conducted.
Therefore, the effects of HOXC11 on survival should be validated in
in vivo experiments in the future.
Supplementary Material
Increased expression of HOXC11 and
HOTAIR in multiple types of cancer. Increased expression of HOTAIR
in (A) COAD, (B) ESCA, (C) BRCA and (D) KIRC. Increased expression
of HOXC11 in (E) COAD, (F) ESCA, (G) BRCA and (H) KIRC.
***P<0.001. HOTAIR, homeobox transcript antisense
intergenic RNA; HOXC11, homeobox C11; COAD, colon adenocarcinoma;
ESCA, esophageal carcinoma; BRCA, invasive breast carcinoma; KIRC,
kidney renal clear cell carcinoma.
Co-expression of HOTAIR and HOXC11 in
(A) colon adenocarcinoma, (B) esophageal carcinoma, (C) breast
invasive carcinoma and (D) kidney renal clear cell carcinoma.
HOTAIR, homeobox transcript antisense intergenic RNA; HOXC11,
homeobox C11; N, non-tumor; T, tumor.
Association between the expression of
HOXC11 and HOTAIR and prognosis. (A-D) Induced expression of HOXC11
was related to poor prognosis in COAD and KIRC. (E-H) Induced
expression of HOTAIR was related to poor prognosis in COAD and
KIRC. HOTAIR, homeobox transcript antisense intergenic RNA; HOXC11,
homeobox C11; COAD, colon adenocarcinoma; ESCA, esophageal
carcinoma; BRCA, invasive breast carcinoma; KIRC,kidney renal clear
cell carcinoma.
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of materials and data
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
NG participated in manuscript preparation and
performed the experiments. FL and YNW performed literature search
and the collection of gene expression and survival data from TCGA
and UALCAN. WQW, ZZL and TTG performed data analysis. HTC and XJM
participated in research design and performed manuscript review. NG
and HTC confirm the authenticity of all the raw data. All authors
read and approved the final manuscript.
Ethics approval and consent to
participate
Ethics approval (approval no. K206-1) was obtained
from the Research Ethics Committee of the Fifth Affiliated Hospital
of Sun Yat-sen University. Informed consent was obtained from all
patients.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Williams C, DiLeo A, Niv Y and Gustafsson
JA: Estrogen receptor beta as target for colorectal cancer
prevention. Cancer Lett. 372:48–56. 2016.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Batista PJ and Chang HY: Long noncoding
RNAs: Cellular address codes in development and disease. Cell.
152:1298–1307. 2013.PubMed/NCBI View Article : Google Scholar
|
|
4
|
He Y, Meng XM, Huang C, Wu BM, Zhang L, Lv
XW and Li J: Long noncoding RNAs: Novel insights into hepatocelluar
carcinoma. Cancer Lett. 344:20–27. 2014.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Fatica A and Bozzoni I: Long non-coding
RNAs: New players in cell differentiation and development. Nat Rev
Genet. 15:7–21. 2014.PubMed/NCBI View
Article : Google Scholar
|
|
6
|
Kalwa M, Hänzelmann S, Otto S, Kuo CC,
Franzen J, Joussen S, Fernandez-Rebollo E, Rath B, Koch C, Hofmann
A, et al: The lncRNA HOTAIR impacts on mesenchymal stem cells via
triple helix formation. Nucleic Acids Res. 44:10631–10643.
2016.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Hong Q, Li O, Zheng W, Xiao WZ, Zhang L,
Wu D, Cai GY, He JC and Chen XM: LncRNA HOTAIR regulates HIF-1α/AXL
signaling through inhibition of miR-217 in renal cell carcinoma.
Cell Death Dis. 8(e2772)2017.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Fu WM, Zhu X, Wang WM, Lu YF, Hu BG, Wang
H, Liang WC, Wang SS, Ko CH, Waye MM, et al: Hotair mediates
hepatocarcinogenesis through suppressing miRNA-218 expression and
activating P14 and P16 signaling. J Hepatol. 63:886–895.
2015.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Gupta RA, Shah N, Wang KC, Kim J, Horlings
HM, Wong DJ, Tsai MC, Hung T, Argani P, Rinn JL, et al: Long
non-coding RNA HOTAIR reprograms chromatin state to promote cancer
metastasis. Nature. 464:1071–1076. 2010.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Zhang A, Zhao JC, Kim J, Fong KW, Yang YA,
Chakravarti D, Mo YY and Yu J: LncRNA HOTAIR enhances the
androgen-receptor-mediated transcriptional program and drives
castration-resistant prostate cancer. Cell Rep. 13:209–221.
2015.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Heubach J, Monsior J, Deenen R, Niegisch
G, Szarvas T, Niedworok C, Schulz WA and Hoffmann MJ: The long
noncoding RNA HOTAIR has tissue and cell type-dependent effects on
HOX gene expression and phenotype of urothelial cancer cells. Mol
Cancer. 14(108)2015.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Nnamani MC, Ganguly S, Erkenbrack EM,
Lynch VJ, Mizoue LS, Tong Y, Darling HL, Fuxreiter M, Meiler J and
Wagner GP: A derived allosteric switch underlies the evolution of
conditional cooperativity between HOXA11 and FOXO1. Cell Rep.
15:2097–2108. 2016.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Roumengous S, Rousset R and Noselli S:
Polycomb and Hox genes control JNK-induced remodeling of the
segment boundary during Drosophila morphogenesis. Cell Rep.
19:60–71. 2017.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Kherdjemil Y, Lalonde RL, Sheth R,
Dumouchel A, de Martino G, Pineault KM, Wellik DM, Stadler HS,
Akimenko MA and Kmita M: Evolution of Hoxa11 regulation in
vertebrates is linked to the pentadactyl state. Nature. 539:89–92.
2016.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Zanatta A, Rocha AM, Carvalho FM, Pereira
RM, Taylor HS, Motta EL, Baracat EC and Serafini PC: The role of
the Hoxa10/HOXA10 gene in the etiology of endometriosis and its
related infertility: A review. J Assist Reprod Genet. 27:701–710.
2010.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Amândio AR, Necsulea A, Joye E, Mascrez B
and Duboule D: hotair is dispensible for mouse development. PLoS
Genet. 12(e1006232)2016.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Chandrashekar DS, Bashel B, Balasubramanya
SA, Creighton CJ, Ponce-Rodriguez I, Chakravarthi BV and Varambally
S: UALCAN: A Portal for Facilitating Tumor Subgroup Gene Expression
and Survival Analyses. Neoplasia. 19:649–658. 2017.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Kent WJ, Sugnet CW, Furey TS, Roskin KM,
Pringle TH, Zahler AM and Haussler D: The human genome browser at
UCSC. Genome Res. 12:996–1006. 2002.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Faghihi MA, Modarresi F, Khalil AM, Wood
DE, Sahagan BG, Morgan TE, Finch CE, St Laurent G III, Kenny PJ and
Wahlestedt C: Expression of a noncoding RNA is elevated in
Alzheimer's disease and drives rapid feed-forward regulation of
beta-secretase. Nat Med. 14:723–730. 2008.PubMed/NCBI View
Article : Google Scholar
|
|
20
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Ali A, Creevey L, Hao Y, McCartan D,
O'Gaora P, Hill A, Young L and McIlroy M: Prosaposin activates the
androgen receptor and potentiates resistance to endocrine treatment
in breast cancer. Breast Cancer Res. 17(123)2015.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Loewen G, Jayawickramarajah J, Zhuo Y and
Shan B: Functions of lncRNA HOTAIR in lung cancer. J Hematol Oncol.
7(90)2014.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Bhatlekar S, Fields JZ and Boman BM: HOX
genes and their role in the development of human cancers. J Mol Med
(Berl). 92:811–823. 2014.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Bhatlekar S, Fields JZ and Boman BM: Role
of HOX genes in stem cell differentiation and cancer. Stem Cells
Int. 2018(3569493)2018.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Du H and Taylor HS: The role of Hox genes
in female reproductive tract development, adult function, and
fertility. Cold Spring Harb Perspect Med. 6(a023002)2015.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Luo Z, Rhie SK, Lay FD and Farnham PJ: A
Prostate cancer risk element functions as a repressive loop that
regulates HOXA13. Cell Rep. 21:1411–1417. 2017.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Zhu H, Dai W, Li J, Xiang L, Wu X, Tang W,
Chen Y, Yang Q, Liu M, Xiao Y, et al: HOXD9 promotes the growth,
invasion and metastasis of gastric cancer cells by transcriptional
activation of RUFY3. J Exp Clin Cancer Res. 38(412)2019.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Luo H, Wang F, Zha J, Li H, Yan B, Du Q,
Yang F, Sobh A, Vulpe C, Drusbosky L, et al: CTCF boundary remodels
chromatin domain and drives aberrant HOX gene transcription in
acute myeloid leukemia. Blood. 132:837–848. 2018.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Taketani T, Taki T, Shibuya N, Kikuchi A,
Hanada R and Hayashi Y: Novel NUP98-HOXC11 fusion gene resulted
from a chromosomal break within exon 1 of HOXC11 in acute myeloid
leukemia with t(11;12)(p15;q13). Cancer Res. 62:4571–4574.
2002.PubMed/NCBI
|
|
30
|
Gu BW, Wang Q, Wang JM, Xue YQ, Fang J,
Wong KF, Chen B, Shi ZZ, Shi JY, Bai XT, et al: Major form of
NUP98/HOXC11 fusion in adult AML with t(11;12)(p15;q13)
translocation exhibits aberrant trans-regulatory activity.
Leukemia. 17:1858–1864. 2003.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Mitchelmore C, Troelsen JT, Sjöström H and
Norén O: The HOXC11 homeodomain protein interacts with the
lactase-phlorizin hydrolase promoter and stimulates
HNF1alpha-dependent transcription. J Biol Chem. 273:13297–13306.
1998.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Zhang Y and Cao X: Long noncoding RNAs in
innate immunity. Cell Mol Immunol. 13:138–147. 2016.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Idda ML, Munk R, Abdelmohsen K and Gorospe
M: Noncoding RNAs in Alzheimer's disease. Wiley Interdiscip Rev
RNA. 9(9)2018.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Wang W, He X, Zheng Z, Ma X, Hu X, Wu D
and Wang M: Serum HOTAIR as a novel diagnostic biomarker for
esophageal squamous cell carcinoma. Mol Cancer.
16(75)2017.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Özeş AR, Miller DF, Özeş ON, Fang F, Liu
Y, Matei D, Huang T and Nephew KP: NF-κB-HOTAIR axis links DNA
damage response, chemoresistance and cellular senescence in ovarian
cancer. Oncogene. 35:5350–5361. 2016.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Xue X, Yang YA, Zhang A, Fong KW, Kim J,
Song B, Li S, Zhao JC and Yu J: LncRNA HOTAIR enhances ER signaling
and confers tamoxifen resistance in breast cancer. Oncogene.
35:2746–2755. 2016.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Woo CJ and Kingston RE: HOTAIR lifts
noncoding RNAs to new levels. Cell. 129:1257–1259. 2007.PubMed/NCBI View Article : Google Scholar
|