Introduction
Candida albicans (C. albicans) is considered
the most common opportunistic fungal pathogen in humans due to its
widespread distribution in the environment. It has a high
propensity to cause invasive infections on mucosal surfaces and is
particularly life-threatening in individuals with immunodeficiency
and dysbacteriosis, as systemic candidiasis has a high mortality
rate in such patients (1,2). Fungi continue to vary and mutate in
response to environmental and body stimuli and the antifungal drugs
available are simply insufficient for the clinical/pharmacological
treatment of patients. In addition, the incidence of C.
albicans resistance increased rapidly due to the widespread and
long-term usage of limited antifungal drugs, causing additional
difficulty in treatment and ultimately leading to refractory fungal
infections (3-5).
One of the morphological characteristics that
distinguish C. albicans from other organisms is polymorphism
and pleomorphism. C. albicans may have three distinct
phenotypic characteristics: A yeast-like form at 25˚C, and
additional pseudohyphae and hyphae forms at 37˚C, which are
commonly referred to as filamentous morphologies (6,7). All
three forms may coexist and transform with the change of
environmental stimuli. C. albicans biofilms, as an important
barrier, are considered significant for pathogenicity and
virulence, as well as adhesion and infectivity, which are
contributing factors in the process of infecting the host and are
able to withstand the stimulation of the external environment and
drugs. Therefore, numerous clinical antifungal drugs are mainly
resistant to biofilms. The major conventional antifungal drugs
currently in clinical use are fluconazole and amphotericin B.
Studies have demonstrated that the frequent and excessive use of
antifungal drugs led to the notorious resistance of C.
albicans to a wide variety of clinical antifungal drugs,
including the aforementioned two (8,9).
The contradiction between effective antifungal
therapy and finite available clinical antifungal drugs is
increasingly prominent. Therefore, novel antifungal agents are
urgently required in order overcome the resistance of C.
albicans biofilms. Recent scientific studies have re-evaluated
tetracycline (TE) derivatives, such as minocycline (MH), TE,
doxycycline (DO), tigecycline (TGC), oxytetracycline and
demethylchlortetracycline, for their potential antifungal activity
as well as old traditional antibiotics combined with antifungal
drugs to achieve an outstanding effect against fungal infections
(10-13).
MH, a TE derivative, is a traditional antimicrobial
agent that inhibits bacterial protein synthesis, which is
extensively used in the clinic due to its large spectrum of
antibiotic activity. MH has a strong effect on gram-positive
bacteria, including Staphylococcus aureus, with the effect
on gram-negative bacilli being slightly weaker. When MH is combined
with other antifungal agents, such as fluconazole, the antifungal
effect of combination treatment is significantly stronger than that
of MH used alone (10,11,14,15).
Previous studies have reported that MH is able to mediate several
biological phenomena and is associated with the capacity of
morphological transformation from yeast to hypha in C.
albicans (14).
The RAS/cyclic (c)AMP/protein kinase A (PKA)
signaling pathway has an impact on cell growth, differentiation,
protein secretion and transport. Hogan and Sundstrom (16) and Phillips and Crowe (17) determined separately that in C.
albicans, the Ras signaling pathway mediates the morphological
transition and biofilm formation, and contains certain vital
hypha-specific genes, including ras family GTPase (RAS1),
thiamin pyrophosphokinase 1 (TPK1), enhanced filamentous growth
protein 1 (EFG1), TEC1, CAP1 (adenylyl
cyclase-associated protein 1), phosphodiesterase 2 (PDE2)
and adenylate cyclase (CDC35), as well as certain
adhesion-specific genes, including agglutinin-like protein 3
(ALS3) and hyphal wall protein 1 (HWP1). Based on the
results of these studies, MH may be a promising and effective
antifungal agent. It is able to alleviate the urgent problem of
drug resistance and short supply of antifungal drugs, which may
provide a major breakthrough in the clinical treatment of fungal
infections. However, in order to understand the antifungal activity
of MH thorough and systematic evaluation is required and the effect
of MH on C. albicans biofilms and the mechanisms underlying
this require further investigation. In the present study, it was
hypothesized that the antifungal effect of MH on C. albicans
is mainly associated with biofilm formation and its regulation is
closely associated with the RAS/cAMP/PKA pathway. In the present
study, the in vitro antifungal activity of the widely used
antimicrobial MH in C. albicans was assessed, and whether
its potential antifungal mechanism is associated with the Ras/cAMP
signaling pathway was systematically explored to verify the
validity of this hypothesis.
Materials and methods
Strains, antibacterial agents and
growth conditions
A total of four TE derivatives were examined in the
present study: MH, TE, DO and TGC. The TH drug susceptibility test
(TE, 30 µg; DO, 30 µg; TGC, 15 µg) was obtained from BioMérieux and
MH drug papers (30 µg) from Merck KGaA. MH was dissolved in DMSO. A
high concentration of the original solution was prepared, stored at
-20˚C away from light and then diluted to the desired final
concentration prior to use. The final content of DMSO was <2%
and had no effect on cells in the present study.
The reference strain C. albicans ATCC 90028
was a gift obtained from Renmin Hospital of Wuhan University. All
clinically isolated yeast strains used in the present study were
isolated at the People's Hospital of China Three Gorges University
(Yichang, China) and stored in the central laboratory. All strains
used in the present study were identified according to the standard
morphological criteria (18).
Strains were routinely cultured on Sabouraud dextrose agar (SDA)
plates at 28˚C for 24-48 h, then streaked out onto Yeast
Extract-Peptone-Dextrose (YPD) liquid medium and grown at 30˚C in a
shaking incubator for 24-48 h until they reached the logarithmic
growth phase and the absorbance was measured at 600 nm. Hyphal
inductions were performed at 37˚C in liquid medium, and Spider and
RPMI-1640 medium were prepared as previously described (19).
Effect of TE derivatives on the growth
of yeast strains (Kirby-Bauer disc diffusion method)
All strains were adjusted to 0.5 McFarland bacterial
suspensions and then evenly spread to a Mueller-Hinton agar plate
(Thermo Fisher Scientific, Inc.). Once the surface of the plate was
dry, sensitivity test papers containing one of the 4 TE antibiotics
were applied. All plates were incubated for 24 h at 28˚C and the
diameter of the inhibition ring was measured.
Effect of MH on planktonic cells in C.
albicans
Antifungal susceptibility testing was performed in
96-well tissue culture plates (Corning, Inc.) according to the
protocol of the Clinical and Laboratory Standards Institute
(20). In brief, C. albicans
ATCC 90028 was prepared with initial cells at 1x105
CFU/ml and cultured in YPD liquid medium for 48 h at 37˚C. To
evaluate the effect of MH on the cell growth in C. albicans
ATCC 90028, an inoculum concentration of yeast cells at
~1x106 cells/ml was added into liquid RPMI 1640 medium
containing MH at concentrations of 0, 125, 250, 500 and 1,000
µg/ml, incubated in 96-well tissue culture plates at 37˚C and
shaken at 200 rpm.(2.2 x g). The cells were monitored every 4 h and
counted at the predetermined time-points (0, 4, 8, 12, 16, 20, 24,
28 and 32 h) using a Multiskan GO Microplate reader (Thermo Fisher
Scientific, Inc.) to measure the absorption/optical density (OD) at
600 nm, and the background optical densities were already
subtracted. A total of three independent experiments were performed
(21).
Hyphal formation assessment
To further estimate whether the MH affects
bud-to-hypha transition, a fixed amount of C. albicans cells
(1x106 cells/ml) was cultured in several nutritious
hypha transition media. The spider and RPMI 1640 media (Sangon
Biotech Co., Ltd.) were supplemented with 20% fetal bovine serum
(FBS; Cytiva) in 24-well tissue culture plates containing various
concentrations of MH and incubated at 37˚C for 3 h, and hyphal
formation was evaluated microscopically to count 200 cells and
calculate the percentage of hyphal formation, as previously
described (14).
Antifungal activity against mature
biofilms
The C. albicans cells in the logarithmic
growth phase were collected and centrifuged, RPMI 1640 medium was
added to adjust the concentration to 1x106 CFU/ml and
100 µl bacterial liquid was added to the 96-well culture plate for
90 min of adhesion at 37˚C. The medium was gently sucked off with a
sample gun, non-adherent planktonic cells were removed, 100 µl
fresh RPMI 1640 medium was added and the plate was incubated at
37˚C for 24 h until the mature biofilm was formed (20). The biofilm supernatant was then
aspirated, RPMI 1640 with MH was added to for a 24-h incubation,
the growth of the biofilm was observed under a microscope and an
XTT kit (Sangon Biotech Co., Ltd.) was used to evaluate the formed
mature biofilm. Similarly, in order to detect the effect of MH on
the formation of biofilms, MH was added to fresh RPMI 1640 after 90
min of adhesion; subsequent methods were described above until
biofilm formation to observe the anti-biofilm effect of MH
(20,22).
Gene expression analysis of C.
albicans-specific genes
Following MH treatment overnight, C. albicans
cells were washed and collected, and a Fungal RNA out kit (Sangon
Biotech Co, Ltd.) was used to extract and isolate the total RNA of
C. albicans according to the manufacturer's protocol and the
standard method. The OD was measured at 260-280 nm using a Nanodrop
2000 Ultraviolet spectrophotometer (Thermo Fisher Scientific, Inc.)
to test the nucleic acid concentration and purity of the RNA. The
integrity of the RNA was assessed using denatured electrophoresis
through a 1% agarose-3-(N-morpholino) propanesulfonic acid gel
(Sangon Biotech Co., Ltd.). A reverse transcription kit (Takara
Bio, Inc.) was used to reverse-transcribe single-stranded RNA (1
µg) into cDNA. Next, reverse transcription-quantitative PCR
(RT-qPCR) was performed using a 7500 real-time PCR system (Thermo
Fisher Scientific, Inc.); hyphal-specific and biofilm-associated
genes of C. albicans, as well as their transcriptional
regulators, were analyzed in the present study, and the primers are
listed in Table I. The associated
genes were as follows: RAS1, a major transcription factor involved
in the MAPK pathway; EFG1, transcription factor in the cAMP/PKA
pathway; GAP1, which encodes a general amino acid permease; ALS3
and HWP1, which encode agglutinin-like protein 3 and hyphal wall
protein 1, respectively, and are activated by EFG1; CDC35 (coding
for adenylyl cyclase); TPK1 (PKA catalytic subunits); PDE2
(high-affinity phosphodiesterase gene); TEC1, a transcription
factor; ACT1, a housekeeping gene that has the ability to express
stably and consistently in organisms, and was chosen as an
endogenous control gene for correcting the loading inaccuracies
(23). The final values were
calculated by the 2-ΔΔCq method (24).
 | Table IPCR primers used for detection of
gene expression in Candida albicans. |
Table I
PCR primers used for detection of
gene expression in Candida albicans.
| Primer | Sequence
(5'-3') |
|---|
| CAP1-F |
GAACCACCATCAACATCA |
| CAP1-R |
CAACCCATCAATATCAAGT |
| TPK1-F |
AGAAGTTCAAGATGTGACTTAT |
| TPK1-R |
CATCATCAGAACCACCTTGT |
| CDC35-F |
TCCATGTCAAATATGCCAACG |
| CDC35-R |
CTTCACATCCCAACTTTCAGG |
| EFG1-F |
TCCATGTCAAATATGCCAACG |
| EFG1-R |
TGGATTCATACCGTATTGGTCATTA |
| TEC1-F |
TGGATTCATACCGTATTGGTCATTA |
| TECI-R |
TCGGGCAATCCTTTGAATAAA |
| RAS1-F |
GTGGTGGTGTTGGTAAATC |
| RAS1-R |
TTCTTGTCCAGCAGTATCT |
| PDE2-F |
TGCTGTGGGACATTGGAG |
| PDE2-R |
GGCGGAAATTATGGAACG |
| ALS3-F |
GTGATGCTGGATCTAACGGTATTG |
| ALS3-R |
GTCTTAGTTTTGTCGCGGTTAGG |
| HWP1-F |
CGGAATCTAGTGCTGTCGTCTCT |
| HWP1-R |
CGACACTTGAGTAATTGGCAGATG |
| ACT 1-F |
TTGATTTGGCTGGTAGAG |
| ACT 1-R |
ATGGCAGAAGATTGAGA |
The PCR conditions were programmed and divided into
three steps according to the experimental conditions and purposes.
The first step was initial denaturation at 95˚C for 5 min, where
double-stranded DNA templates break hydrogen bonds under thermal
action to form single-stranded DNA. The second step was annealing
at 55-65˚C, with the continuous gradient temperature set at a
heating rate of 0.1˚C/sec to match the appropriate suitable
solution curve temperature, the system temperature was lowered and
the primer bound to the DNA template to form a local double strand.
This step eventually went through 40 cycles of amplification and
quantification at 95˚C for 10 sec, 55˚C for 30 sec and 72˚C for 15
sec with a continuous single fluorescence measurement and end. The
last step was a final extension maintained at 72˚C for 5-10 min.
dNTP was used as raw material and primers started from the 3'end,
extending from the 5' to 3' end, to synthesize DNA strands
complementary to the template, followed by a cooling step at
42˚C.
Statistical analysis
Values are expressed as the mean fold change ±
standard deviation of three independent experiments. Statistical
analyses were performed using GraphPad Prism 5.0 (GraphPad
Software, Inc.). One-way analysis of variance with Tukey's post hoc
test was used to evaluate the statistical significance of
differences among the treatment groups. P<0.05 were considered
to indicate a statistically significant difference.
Results
Inhibitory effects of TE derivatives
on yeasts
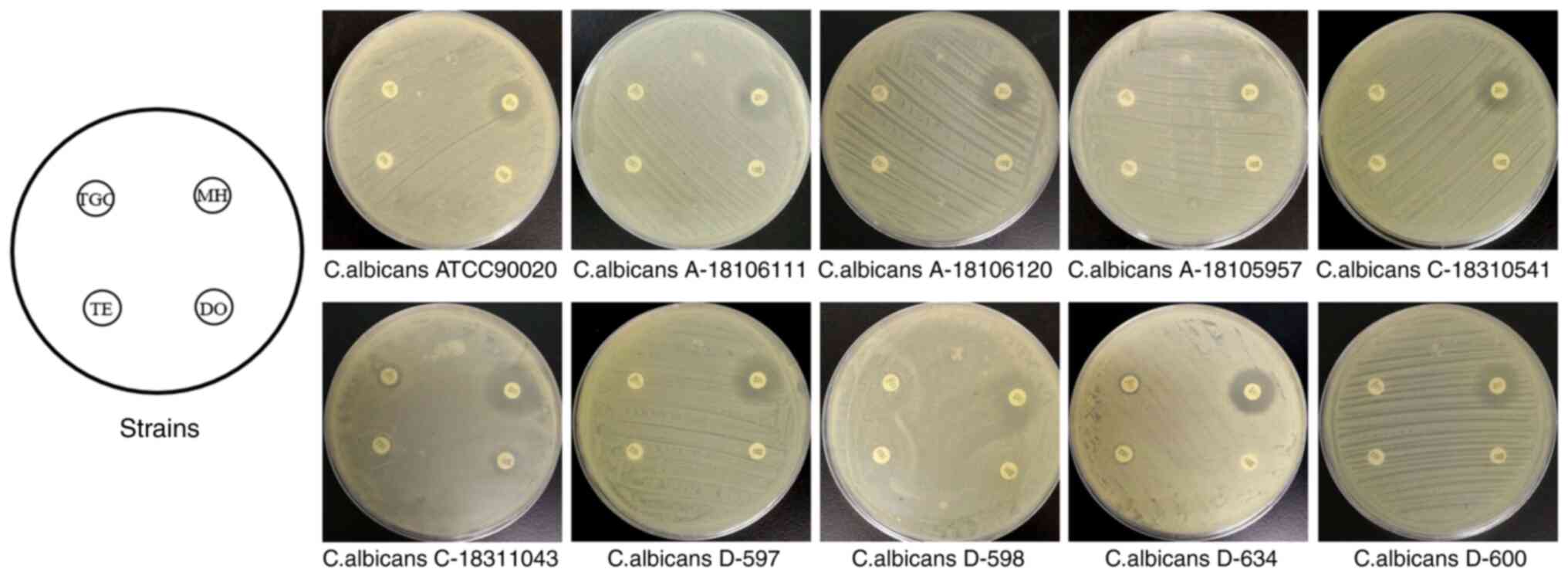
The results of the drug sensitivity test are
presented in Fig. 1. MH had the
most obvious inhibition circle among the four TE derivatives (ME,
TE, DO and TGC). By measurement and quantification, the ratio of
the inhibition zone diameter ≥14 mm for MH on C. albicans
was determined to be 80%, that on C. tropicalis was 96% and
that on C. Portuguese was 87%. TE, DO and TGC exhibited no
sufficient antifungal activity against C. albicans with the
inhibition zone diameter being 6-10 mm, <14 mm in all
strains.
Effects of MH on C. albicans
growth
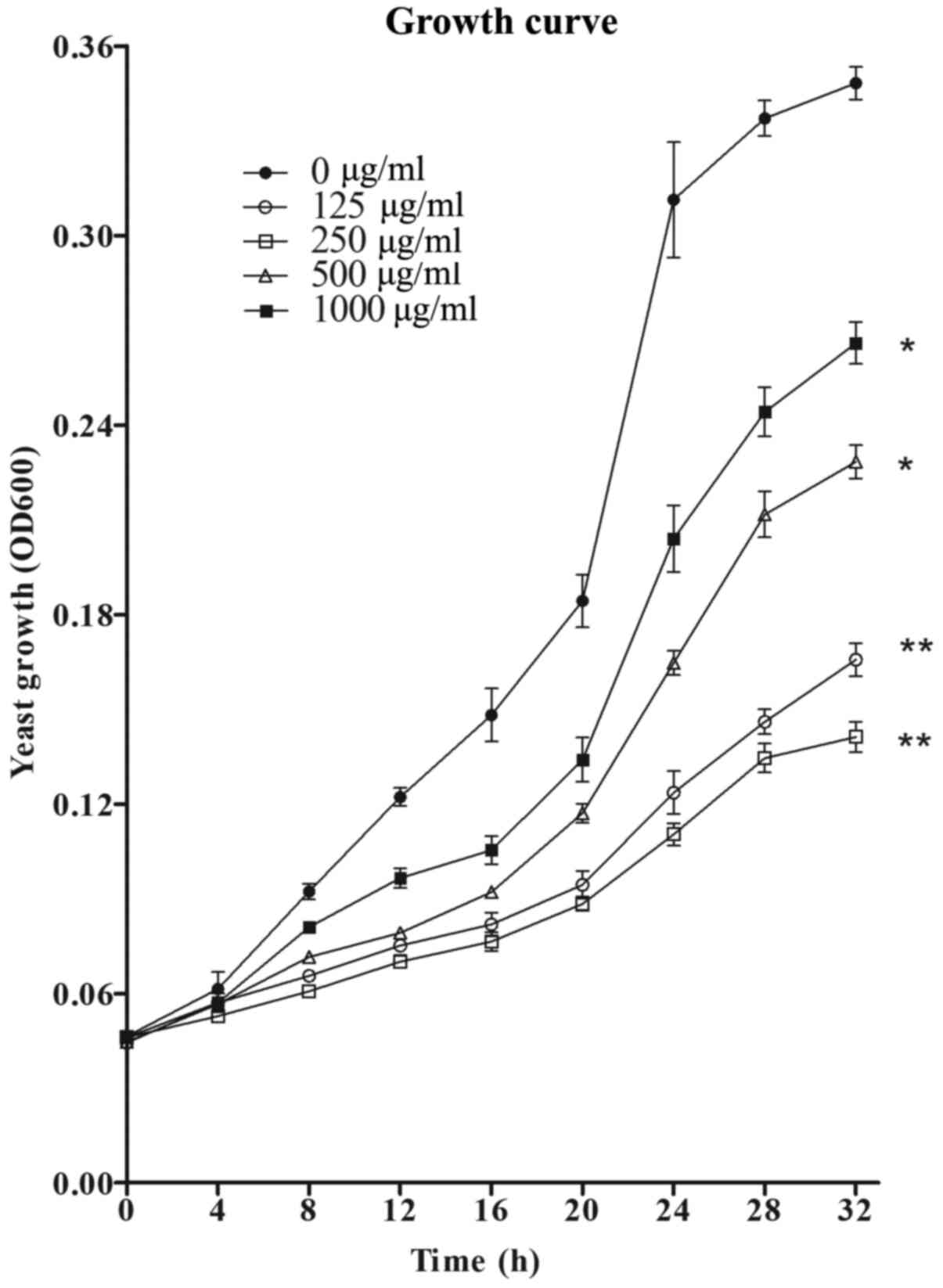
In the presence of MH at the concentrations of 125,
250, 500 and 1,000 µg/ml, a significant growth inhibition of C.
albicans was observed in the treatment group as compared to the
control group without MH treatment (Fig. 2). At 250 µg/ml, MH exhibited the
most potent inhibitory activity on C. albicans growth among
all concentrations used. After 8 h of incubation in liquid culture,
the growth curves of C. albicans began to diverge and
exhibit visible differences. After 32 h, the growth was
significantly restricted in RPMI-1640 medium with MH at 125 µg/ml,
leading to a 59.9% reduction (P<0.01), and in the presence 250
µg/ml MH, the maximum inhibition was 68.0% (P<0.01) as compared
to the control group. However, with further increases in the
concentration of MH, the growth inhibition of planktonic C.
albicans decreased.
MH regulates the hyphal formation
process of C. albicans
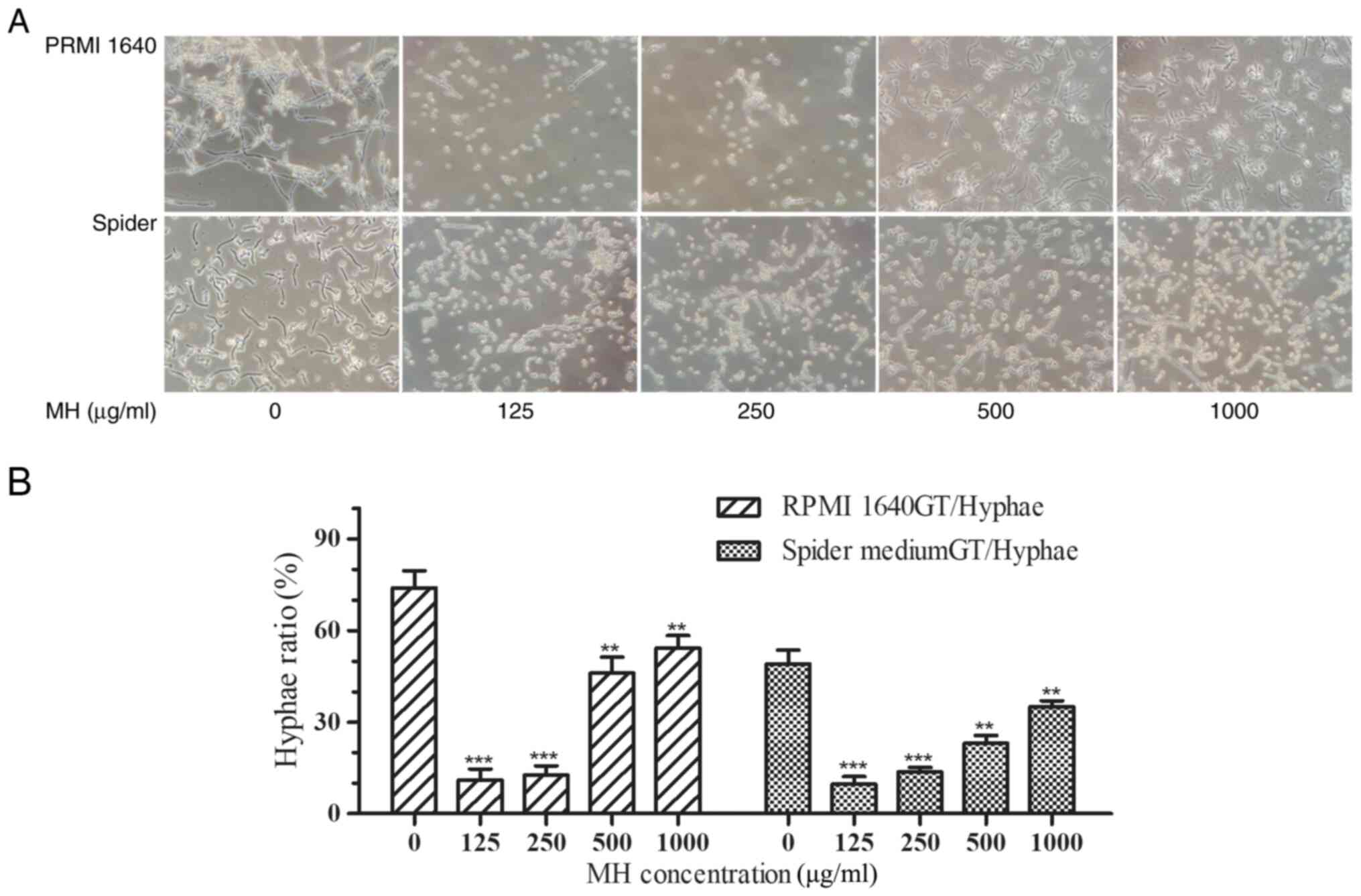
Morphological changes were observed under light
microscopy following incubation for 4 h and the quantitative
results suggested that MH inhibited hyphal growth induction at all
tested concentrations. More specifically, the maximum hypha
formation inhibition occurred at the lowest concentration of MH
(125 µg/ml) in both RPMI-1640 and Spider media and the degree of
hypha inhibition slightly decreased with the increase of the MH
concentration (Fig. 3A). The hypha
rate of ~200 cells for each sample was determined using light
microscopy (Fig. 3B). In drug-free
medium, >70% of C. albicans cells converted to the hyphal
morphology after 4 h of incubation, with only a small percentage
retaining their yeast form. However, following incubation with 125
and 250 µg/ml MH, the hypha formation rate was only 11.75±2.86%
with a significant inhibition of hyphae in all media (P<0.001).
Following incubation with 500 µg/ml MH, the hypha rates were
46.00±5.29% in RPMI-1640 medium and 23.00±2.64% in Spider medium,
both were significantly different from the control group
(P<0.001). Following incubation with 1,000 µg/ml MH, the hypha
rates were 54.33±4.04% in RPMI-1640 medium and 35.33±2.51% in
Spider medium, both were significantly different from the control
group (P<0.01).
MH inhibits C. albicans biofilms
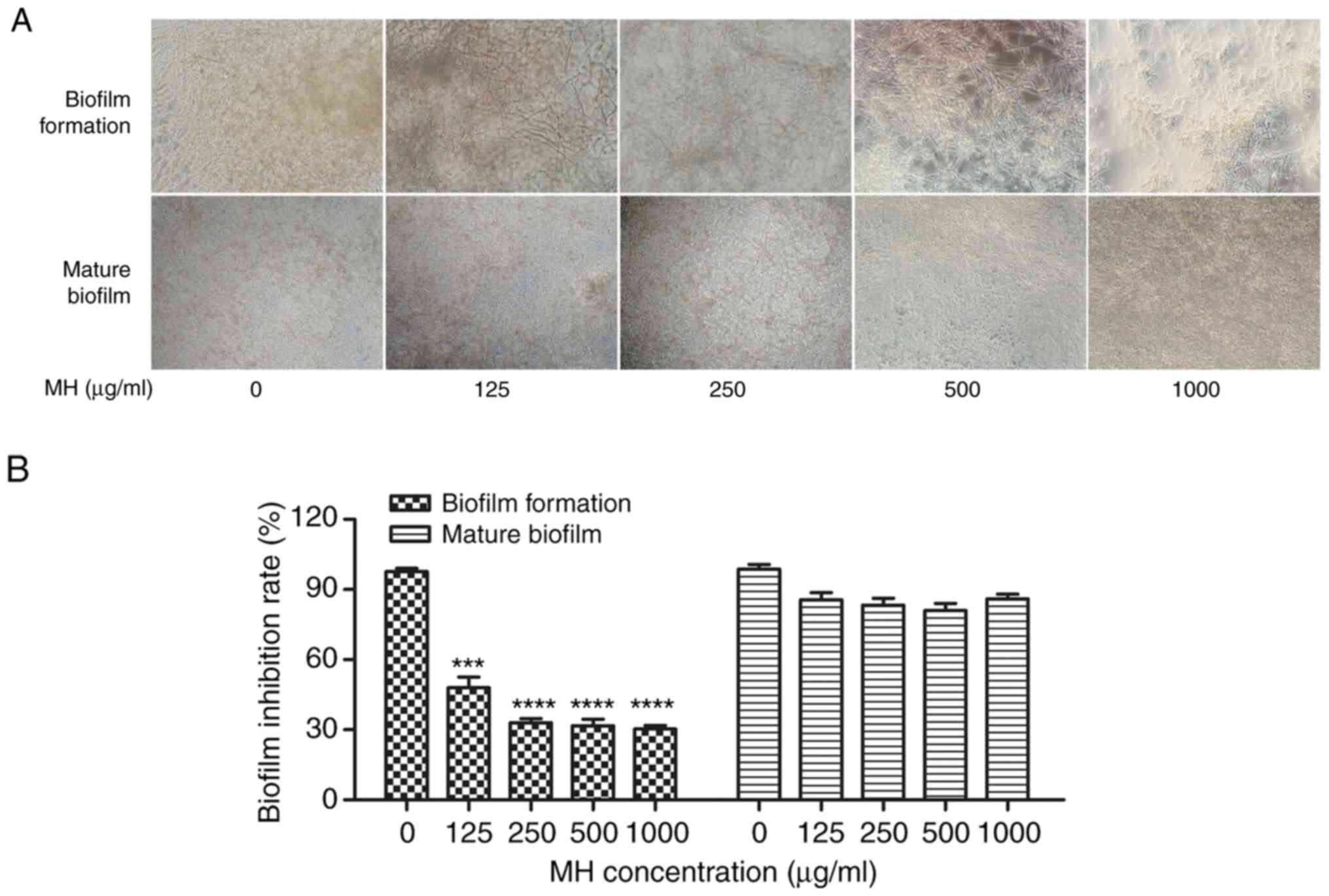
An XTT reduction assay was performed to examine the
potential of MH to inhibit biofilm formation in vitro. As
presented in Fig. 4, the biofilms
formed by C. albicans decreased in an MH dose-dependent
manner in terms of their density and thickness. More specifically,
MH inhibited biofilm formation by 52% at 125 µg/ml (P<0.001 vs.
control), and with the increase of the MH concentration, the
inhibitory effect became more pronounced. MH inhibited biofilm
formation by 67% at 250 µg/ml (P<0.001 vs. control). However, MH
had no marked activity against the mature biofilms, which exhibited
no obvious/significant changes following treatment with any of the
MH concentrations. In conclusion, the tendency of MH to inhibit
biofilm formation was confirmed.
MH regulates the expression of cAMP
pathway genes
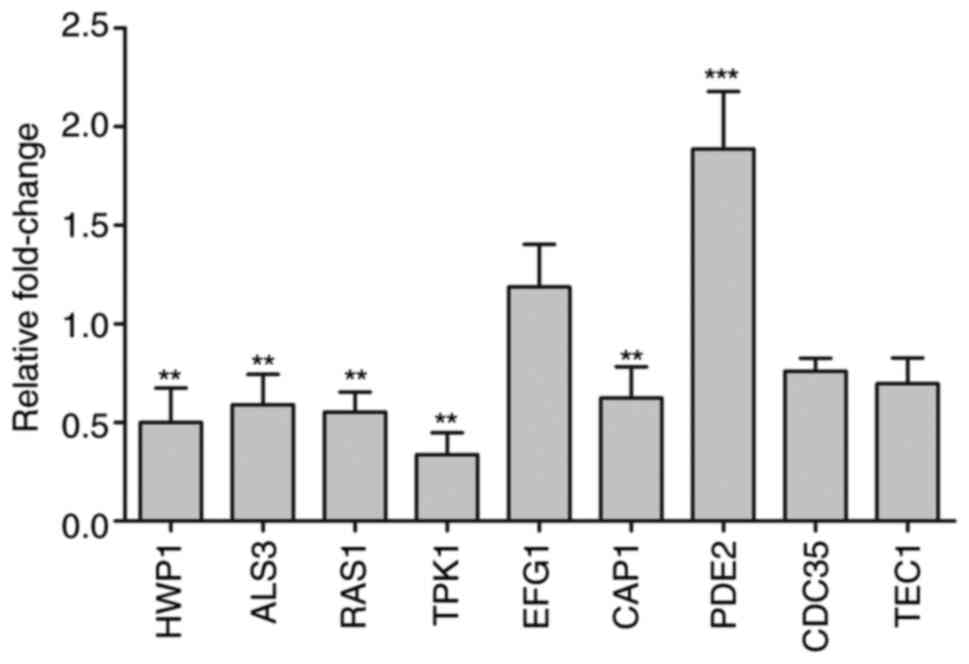
The expression levels of 9 genes were monitored by
RT-qPCR. Hypha-specific genes from the adhesion process,
HWP1 and ALS3, which are downstream components of the
cAMP/PKA pathway, are positively regulated by EFG1(25), and they were downregulated by 0.48-
and 0.59-fold, respectively, following 250 µg/ml MH treatment.
Certain hyphal transcriptional regulation genes from the cAMP/PKA
pathway, including RAS1, TPK1, EFG1,
CAP1, PDE2 and CDC35, were also examined
(Fig. 5). The upregulated genes
included EFG1 and PDE2. The expression levels of EFG1
were slightly, though not significantly upregulated by MH treatment
in the present study, while the upstream components RAS1,
TPK1 and CAP1 were downregulated by 0.55-, 0.33- and
0.62-fold, respectively. CDC35 and TEC1 were
downregulated by 0.76- and 0.69-fold, respectively. Taken together,
the results suggested that genes downregulated following MH
treatment, including HWP1, ALS3, RAS1 and
TPK1, were all associated with the AMP pathway and regulated
by the Ras/cAMP pathway (16).
Discussion
The results of the present study demonstrated that
the antifungal activity of MH was mediated by regulatory factors in
the Ras/cAMP pathway, which was conducive to the reduction of
biofilm formation and limiting of hyphae formation and long-term
maintenance of C. albicans.
It was indicated that MH had different inhibitory
effects on different strains of fungus. It had a strong antifungal
activity on yeasts such as C. albicans, C. tropicalis
and C. lusitaniae, with the inhibition zones being >14 mm
and demonstrating a significant activity against C.
albicans. MH had no inhibitory activity against C.
pseudosmooth, C. glabrata and C. parapsilosis,
while TE, DO and TGC had no significant antifungal properties. The
antifungal effect of MH was marked as compared to other TE
derivative agents. This may be linked to the morphological
transformation and virulence in fungi (26); further research is required in the
future to confirm this.
In addition, several studies have verified that the
length and thickness during the growth of filamentous fungi
directly affects the pathogenicity of hyphae, with the two being
positively correlated (27,28). The yeast form of C. albicans
is less pathogenic and has a better adhesion capacity than the
hyphae form due to its small and single form, at least in part
because the yeast form is more easily eliminated by the defense
system of the host than the hyphae form (29). In Spider and RPMI-1640 media,
nutrient-rich conditions were added with 20% FBS in order for C.
albicans to switch to the hyphae form more rapidly. However, in
the present study, MH was able to maintain the yeast form for
longer and inhibited the formation of hyphae, while MH clearly
affected the hypha growth of C. albicans at the
concentrations of 125 and 250 µg/ml. In RPMI-1640 medium and Spider
medium, the hypha rate (%) of 125 MH treatment was the most
significant, 11.00±3.61 and 9.67±2.51, respectively. These results
are consistent with those by Shi et al (10), who reported that when MH was
combined with fluconazole, the minimum inhibitory concentration on
C. albicans was decreased as compared with that of
fluconazole used alone. Although there have been numerous reports
on the phenomenon of antifungal agents used alone or combined with
antimicrobial agents through C. albicans biofilms (13-15),
few have investigated the particular antifungal characteristics of
antimicrobial agents in C. albicans. To explore the
potential mechanisms, the interference ability of MH to penetrate
biofilms was first assessed.
C. albicans biofilms have a complex and
intricate three-dimensional network architecture, with the barrier
characteristics of C. albicans biofilms mainly reflected in
an increased tolerance to traditional antifungal therapy. An XTT
reduction assay was performed to demonstrate the suppressive
function of MH on biofilms, and it was indicated that MH had a
distinct inhibitory effect against C. albicans biofilms. The
results suggested that 250 µg/ml MH significantly depressed biofilm
formation by 67% (P<0.001). With the increase in concentration,
the inhibitory activity of MH tended to be stable and changes
showed no significant difference, indicating that the optimal
inhibitory effect had been achieved or approached at 250 µg/ml. Of
note, it exhibited no activity to impair the maintenance of mature
biofilms. In conclusion, MH had a strong antibiofilm effect against
C. albicans in vitro, which appears to be attributable to
its anti-morphological transition activities.
According to the inhibition of C. albicans
filamentous growth and biofilm formation, the transcription levels
of genes associated with mycelium growth, adhesion and biofilm
formation in C. albicans were determined to further clarify
the molecular mechanism of the anti-biofilm effects of MH. The
analysis revealed that several important genes were differentially
expressed following MH treatment. CAP1, TPK1, CDC35, TECI, RAS1,
ALS3 and HWP1 were downregulated, and EFG1 and
PDE2 were upregulated following treatment with 250 µg/ml MH.
In this assay, MH was applied at 250 µg/ml, since at this
concentration had a significant antibiofilm activity. It should be
noted that these downregulated genes are all important components
of the RAS1/cAMP/PKA signaling pathway; the present hypothesis that
a direct interaction between the pathways occurs at the
transcriptional level was confirmed. Previous studies have reported
that several signaling pathways are linked to morphogenesis and
biofilms in C. albicans, with the MAPK and cAMP/PKA
signaling pathways being the most prominent (16,30,31).
The cAMP/PKA pathway has attracted extensive attention in fungal
pathogens such as C. albicans, Trichophyton,
Cryptococcus neoformans and Aspergillus fumigatus
(32,33), due to its vital function in the
specific morphology development of fungal biofilms. The Ras/AMP/PKA
pathway is involved in the regulation of various traits of C.
albicans and responds to specific environmental stimuli and
cell subtype combinations (30).
In the present study, the most significantly
downregulated genes were RAS1 and TPK1 (0.55- and 0.33-fold,
respectively). RAS1 has a vital role in the morphological switch
and biofilm formation of yeast and hypha (34,35).
It is a soluble small GTPase that transmits signals to stimulate
filamentous growth in the Ras1/cAMP/PKA signaling pathway as a
transcription factor (36-38).
Despite RAS1 gene mutation and deletion of alleles, which resulted
in serious hyphal growth defects, such as deformation and
irregularity in response to serum, and inhibition of pseudohyphal
development (34), studies have
indicated that the addition of sufficient exogenous cAMP or
overexpressed components of MAP kinase cascade in the growth media
is able to correct the above morphological defects and make
mycelium return to normal (35,39).
Evidence suggested that the RAS1 gene is an intermediate factor in
the anti-fungal regulation of MH, which is involved in signaling
pathway regulation through the RAS1 gene to interfere with
biofilms (14).
Recent research has demonstrated that the deletion
of TPK1 results in hyphal formation defects on solid media in C.
tropicalis (40). In
GlcNAc-inducing liquid medium, TPK1 was associated with the
appearance of germ-tube, with the mRNA levels of TPK1
increasing gradually with cell growth and peaking at the onset of
germ-tube emergence (41). Lin and
Chen (40) reported that PDE2
tightly regulated the intracellular cAMP level. Deficiency of PDE2
led to the activation of cAMP, which decreased the thickness of the
cell wall and cell membrane of the fungus, therefore reducing the
virulence of the fungus and making cells more sensitive to
antifungal agents. High expression of PDE2 inhibited cAMP synthesis
and limited hypha production (42),
while all the defects observed in hyphal morphogenesis in
vitro were reversible and rectifiable. The present results
suggested that the increase in PDE2 mRNA following MH intervention
was most significant, which was consistent with the results of
previous study (43), which further
explained the inhibition of PDE2 on hypha and biofilm growth in
C. albicans.
EFG1 is a well-characterized transcription factor
and important component of the Ras/cAMP pathway, which may both
bidirectionally regulate genes that participate in the essential
processes and stages of cellular growth, such as white-opaque
transition in morphogenesis, and glycolysis and respiration in cell
metabolism (44,45). Furthermore, EFG1 acts upstream of
TEC1 and is located in the middle stream of the RAS1/cAMP-PKA
signaling pathway, which regulates and controls the upstream and
downstream components, including HWP1, TCE1 and ALS3(42). Mutational analysis suggested that
deficiency of the EFG1 gene disrupted normal morphological
transformation, impeded the growth of mycelium, induced hyphae in a
pseudorevertant strain and was dependent on the EFG1 transcription
factor that has a regulatory role in the downstream of protein
kinase A (46). TEC1, located
downstream of EFG1, also regulates the growth of hyphae-specific
genes, with the EFG1/efg1 mutants resulting in the overexpression
of TEC1(47). Deletion of TEC1
resulted in thinner biofilms and the observed disordered growth of
short spores on their surface. In the present study, a slight
elevation of EFG1 was observed and the modulating effect of MH on
EFG1 was not significant. In the present results, most aberrant
genes were downregulated genes, which were all important components
of the RAS1/cAMP/PKA signaling pathway, while fewer genes were
upregulated. These results suggested that the signaling pathways of
MH may be dominated by negative regulation and that the
RAS1/cAMP/PKA signaling pathway may have a crucial role in the
regulation of hypha and biofilm by MH.
However, the present study had certain limitations.
For instance, due to the limited resources and funding available,
no resistant strain analysis was performed and the effects of MH on
drug-resistant strains of C. albicans were not analyzed. In
addition, animal studies are required to verify the drug toxicity
and feasible doses of MH. Further experiments, such as western blot
analysis to detect changes in the protein levels, may be performed,
or changes in electrolytes in the biofilm, such as ionic calcium,
may be assessed in order to investigate the mechanism of action of
MH. Validation using clinical samples is also required. However,
despite these limitations, the results of the present study may
provide novel insight that may lead to the development of clinical
medication and treatment.
In conclusion, the present study demonstrated that
MH exhibits a strong anti-biofilm effect against C. albicans
selectively, and the predominant mechanisms are associated with the
Ras/cAMP pathway. Therefore, the hypothesis of the present study,
that the antifungal effect of MH on C. albicans is mainly
associated with biofilm formation and its regulation is closely
associated with the RAS/cAMP/PKA pathway was validated. These
results indicated that MH may be an effective antifungal agent for
C. albicans infections. However, clinical application is not
yet possible, as more prospective studies using strains and animal
models are required, and treatment strategies for drug-resistant
C. albicans infection require to be developed to further
verify the clinical applicability of MH. This is the direction of
our future research.
Acknowledgements
Not applicable.
Funding
This work was supported by grants from the Health Committee
Union Foundation of Hubei Province (grant no. WJ2019H512) and the
Youth Science Foundation of Three Gorges University (grant no.
KJ2018A007).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
LZ and QD designed the experiments and drafted the
manuscript. ZM performed the experiments and analyzed the data. TG
and BZ performed the RT-qPCR experiments. QD reviewed the
manuscript. All authors read and approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Gow NA, van de Veerdonk FL, Brown AJ and
Netea MG: Candida albicans morphogenesis and host defence:
Discriminating invasion from colonization. Nat Rev Microbiol.
10:112–122. 2011.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Ferreira AV, Prado CG, Carvalho RR, Dias
KS and Dias AL: Candida albicans and non-C. albicans
Candida species: Comparison of biofilm production and metabolic
activity in biofilms, and putative virulence properties ofisolates
from hospital environments and infections. Mycopathologia.
175:265–272. 2013.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Chandra J, Mukherjee PK, Leidich SD,
Faddoul FF, Hoyer LL, Douglas LJ and Ghannoum MA: Antifungal
resistance of candidal biofilms formed on denture acrylic in vitro.
J Dent Res. 80:903–908. 2001.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Karagoz E, Ugan RA, Duzgun E, Cadirci E,
Keles S, Uyanik MH, Yavan I and Turhan V: Comparative study of the
effects of intravitreal anidulafungin, voriconazole, and
amphotericin B in an experimental Candida endophthalmitis
model. Curr Eye Res. 42:225–232. 2017.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Pfaller MA, Rhomberg PR, Messer SA, Jones
RN and Castanheira M: Isavuconazole, micafungin, and 8 comparator
antifungal agents' susceptibility profiles for common and uncommon
opportunistic fungi collected in 2013: Temporal analysis of
antifungal drug resistance using CLSI species-specific clinical
breakpoints and proposed epidemiological cutoff values. Diagn
Microbiol Infect Dis. 82:303–313. 2015.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Noble SM, Gianetti BA and Witchley JN:
Candida albicans cell-type switching and functional
plasticity in the mammalian host. Nat Rev Microbiol. 15:96–108.
2017.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Sudbery PE: Growth of Candida
hyphae. Nat Rev Microbiol. 9:737–748. 2011.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Li DD, Xu Y, Zhang DZ, Quan H, Mylonakis
E, Hu DD, Li MB, Zhao LX, Zhu LH, Wang Y and Jiang YY: Fluconazole
assists berberine to kill fluconazole-resistant Candida
albicans. Antimicrob Agents Chemother. 57:6016–6027.
2013.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Nett JE, Sanchez H, Cain MT, Ross KM and
Andes DR: Interface of Candida albicans biofilm
matrix-associated drug resistance and cell wall integrity
regulation. Eukaryot Cell. 10:1660–1669. 2011.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Shi W, Chen ZZ, Chen X, Cao L, Liu P and
Sun S: The combination of minocycline and fluconazole causes
synergistic growth inhibition against Candida albicans: An
in vitro interaction of antifungal and antibacterial agents. FEMS
Yeast Res. 10:885–893. 2010.PubMed/NCBI View Article : Google Scholar
|
|
11
|
de Oliveira LF, Jorge AC and Santos SF: In
vitro minocycline activity on superinfecting microorganisms
isolated from chronic periodontitis patients. Braz Oral Res.
20:202–206. 2006.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Raad I, Darouiche R, Hachem R, Sacilowski
M and Bodey GP: Antibiotics and prevention ofmicrobial colonization
of catheters. Antimicrob Agents Chemother. 39:2397–2400.
1995.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Gao Y, Li H, Liu S, Zhang X and Sun S:
Synergistic effect of fluconazole and doxycycline against
Candida albicans biofilms resulting from calcium fluctuation
and downregulation of fluconazole-inducible efflux pump gene
overexpression. J Med Microbiol. 63:956–961. 2014.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Kurakado S, Takatori K and Sugita T:
Minocycline inhibits Candida albicans budded-to-hyphal-form
transition and biofilm formation. Jpn J Infect Dis. 70:490–494.
2017.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Li H, Liu P, Liu W, Gao Y and Sun S: In
vitro interactions between fluconazole and minocycline against
mixed cultures of Candida albicans and Staphylococcus
aureus. J Microbiol Immunol Infect. 48:655–661. 2015.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Hogan DA and Sundstrom P: The Ras/cAMP/PKA
signaling pathway and virulence in Candida albicans. Future
Microbiol. 4:1263–1270. 2009.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Phillips AJ, Crowe JD and Ramsdale M: Ras
pathway signaling accelerates programmed cell death in the
pathogenic fungus Candida albicans. Proc Natl Acad Sci USA.
103:726–731. 2006.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Li DD, Zhao LX, Mylonakis E, Hu GH, Zou Y,
Huang TK, Yan L, Wang Y and Jiang YY: In vitro and in vivo
activities of pterostilbene against Candida albicans
biofilms. Antimicrob Agents Chemother. 58:2344–2355.
2014.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Gimeno CJ, Ljungdahl PO, Styles CA and
Fink GR: Unipolar cell divisions in the yeast S. cerevisiae lead to
filamentous growth: Regulation by starvation and RAS. Cell.
68:1077–1090. 1992.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Zhong H, Hu DD, Hu GH, Su J, Bi S, Zhang
ZE, Wang Z, Zhang RL, Xu Z, Jiang YY and Wang Y: Activity of
sanguinarine against Candida albicans biofilms. Antimicrob
Agents Chemother. 61:02259–16. 2017.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Ma C, Du F, Yan L, He G, He J, Wang C, Rao
G, Jiang Y and Xu G: Potent activities of roemerine against
Candida albicans and the underlying mechanisms. Molecules.
20:17913–17928. 2015.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Ku TS, Palanisamy S K and Lee SA:
Susceptibility of Candida albicans biofilms to azithromycin,
tigecycline and vancomycin and the interaction between tigecycline
and antifungals. Int J Antimicrob Agents. 36:441–446.
2010.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Cao S, Zhang X, Ye N, Fan X, Mou S, Xu D,
Liang C, Wang Y and Wang W: Evaluation of putative internal
reference genes for gene expression normalization in
Nannochloropsis sp. by quantitative real-time RT-PCR. Biochem
Biophys Res Commun. 424:118–123. 2012.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Langford ML, Hargarten JC, Patefield KD,
Marta E, Blankenship JR, Fanning S, Nickerson KW and Atkina AL:
Candida albicans Czf1 and Efg1 coordinate the response to
farnesol during quorum sensing, white-opaque thermal dimorphism,
and cell death. Eukaryot Cell. 12:1281–1292. 2013.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Ramírez-Zavala B, Weyler M, Gildor T,
Schmauch C, Kornitzer D, Arkowitz R and Morschhäuser J: Activation
of the Cph1-dependent MAP kinase signaling pathway induces
white-opaque switching in Candida albicans. PLoS Pathog.
9(e1003696)2013.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Berman J and Sudbery PE: Candida
albicans: A molecular revolution built on lessons from budding
yeast. Nat Rev Genet. 3:918–930. 2002.PubMed/NCBI View
Article : Google Scholar
|
|
28
|
Biswas S, Dijck PV and Datta A:
Environmental sensing and signal transduction pathways regulating
morphopathogenic determinants of Candida albicans. Microbiol
Mol Biol Rev. 71:348–376. 2007.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Huh WK and Kang SO: Molecular cloning and
functional expression of alternative oxidase from Candida
albicans. J Bacteriol. 181:4098–4102. 1999.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Huang G, Huang Q, Wei Y, Wang Y and Du H:
Multiple roles and diverse regulation of the Ras/cAMP/protein
kinase A pathway in Candida albicans. Mol Microbiol.
111:6–16. 2019.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Stoldt VR, Sonneborn A, Leuker CE and
Ernst JF: Efg1p, an essential regulator of morphogenesis of the
human pathogen Candida albicans, is a member of a conserved
class of bHLH proteins regulating morphogenetic processes in fungi.
EMBO J. 16:1982–1991. 1997.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Fuller KK and Rhodes JC: Protein kinase A
and fungal virulence: A sinister side to a conserved nutrient
sensing pathway. Virulence. 3:109–121. 2012.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Cloutier M, Castilla R, Bolduc N, Zelada
A, Martineau P, Bouillon M, Magee BB, Passeron S, Giasson L and
Cantoreb ML: The two isoforms of the cAMP-dependent protein kinase
catalytic subunit are involved in the control of dimorphism in the
human fungal pathogen Candida albicans. Fungal Genet Biol.
38:133–141. 2003.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Feng Q, Summers E, Guo B and Fink G: Ras
signaling is required for serum-induced hyphal differentiation in
Candida albicans. J Bacteriol. 181:6339–6346.
1999.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Leberer E, Harcus D, Dignard D, Johnson L,
Ushinsky S, Thomas DY and Schröppel K: Ras links cellular
morphogenesis to virulence by regulation of the MAP kinase and cAMP
signalling pathways in the pathogenic fungus Candida
albicans. Mol microbiol. 42:673–687. 2001.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Cullen PJ and Sprague GJ Jr: The
regulation of filamentous growth in yeast. Genetics. 190:23–49.
2012.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Lu Y, Su C and Liu H: Candida
albicans hyphal initiation and elongation. Trends Microbiol.
22:707–714. 2014.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Davis-Hanna A, Piispanen AE, Stateva LI
and Hogan DA: Farnesol and dodecanol effects on the Candida
albicans Ras1-cAMP signalling pathway and the regulation of
morphogenesis. Mol Microbiol. 67:47–62. 2008.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Zhu Y, Fang HM, Wang YM, Zeng GS, Zheng XD
and Wang Y: Ras1 and Ras2 play antagonistic roles in regulating
cellular cAMP level, stationary-phase entry and stress response in
Candida albicans. Mol Microbiol. 74:862–875. 2009.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Lin CJ, Wu CY, Yu SJ and Chen YL: Protein
kinase A governs growth and virulence in Candida tropicalis.
Virulence. 9:331–347. 2018.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Souto G, Giacometti R, Silberstein S,
Giasson L, Cantore ML and Passeron S: Expression of TPK1 and TPK2
genes encoding PKA catalytic subunits during growth and
morphogenesis in Candida albicans. Yeast. 23:591–603.
2006.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Lin CJ and Chen YL: Conserved and
divergent functions of the cAMP/PKA signaling pathway in Candida
albicans and Candida tropicalis. J Fungi (Basel).
4(68)2018.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Bahn YS, Staab J and Sundstrom P:
Increased high-affinity phosphodiesterase PDE2 gene expression in
germ tubes counteracts CAP1-dependent synthesis of cyclic AMP,
limits hypha production and promotes virulence of Candida
albicans. Mol Microbiol. 50:391–409. 2003.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Nobile CJ, Fox EP, Nett JE, Sorrells TR,
Mitrovich QM, Hernday AD, Tuch BB, Andes DR and Johnson AD: A
recently evolved transcriptional network controls biofilm
development in Candida albicans. Cell. 148:126–138.
2012.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Tebarth B, Doedt T, Krishnamurthy S, Weide
M, Monterola F, Dominguez A and Ernst JF: Adaptation of the Efg1p
morphogenetic pathway in Candida albicans by negative
autoregulation and PKA-dependent repression of the EFG1 gene. J Mol
Biol. 329:949–962. 2003.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Parrino SM, Si H, Naseem S, Groudan K,
Gardin J and Konopka JB: cAMP-independent signal pathways stimulate
hyphal morphogenesis in Candida albicans. Mol Microbiol.
103:764–779. 2017.PubMed/NCBI View Article : Google Scholar
|
|
47
|
Lopes da Rosa J and Kaufman PD:
Chromatin-mediated Candida albicans virulence. Biochim
Biophys Acta. 1819:349–355. 2012.PubMed/NCBI
|