Introduction
Intervertebral disc degeneration (IVDD) is one of
the most common musculoskeletal disorders observed in clinical
practice. Substantial evidence has demonstrated that lower back
pain and spinal cord compression nerve pain are prevalent clinical
and public health concerns caused by IVDD (1,2). The
etiology of IVDD is multifactorial, including aging, genetic
predisposition, high mechanical stress, metabolic disorders,
insufficient nutrition and neurogenic inflammation (3,4). At
present, the therapeutic strategies mainly focus on pain relief
through injections, physical therapy and activity modification or
surgical interventions, such as disc decompression, spinal fusion
and IVD replacement (5,6). However, to date, there is no
effective method available to repair IVDD (7). Consequently, it is necessary to
explore the pivotal pathogenesis of IVDD and identify available
treatment methods.
Although various factors are relevant to the
occurrence of IVDD, nutritional dysregulation is regarded to be the
ultimate common pathway of IVDD, since the IVD is the largest
non-vascular organ in the human body (8,9). The
material exchange between the IVD and the vertebral body mainly
relies on the cranial and caudal endplates (EPs) (10,11),
which contain bone marrow contact channels (12). Studies have demonstrated that the
nutrition acquisition of IVD is dependent on EP permeability and
that there is a significant association between the permeability of
the EP and the porosity of the bone marrow contact channel
(13,14). The calcification of the
cartilaginous EP leads to the obstruction of the bone marrow
contact channel, which may induce and accelerate the occurrence of
IVDD; thus, the alteration of EPs is a critical feature of the
physiopathology of IVDD (15).
Type IX collagen (Col9) is a heterogeneous collagen
composed of three different α chains (α1, α2 and α3), which is
predominantly expressed in cartilage (16,17).
It is also a major component of IVDs, vertebral EPs and developing
vertebral bodies (18,19). As a fiber-associated collagen with
interrupted triple helices, Col9 is assembled with type II collagen
(Col2) to form heterogeneous fibers. It is crosslinked with Col2
located on the surface of fibrils and its non-collagenous NC4
domain projecting out (20).
Numerous interactions with other cartilage matrix proteins have
been allocated to this domain, indicating that Col9 functions as an
adaptor between the collagenous network and other extracellular
matrix (ECM) superstructures (21). The Col9 polymorphism has been
reported to be a risk factor for the occurrence of IVDD,
particularly with advancing age. As one of the branches of Col9,
previous studies have suggested that the α2 chain of Col9 (Col9a2)
is closely associated with degenerative lumbar spinal stenosis and
spondylolisthesis (22,23).
However, it remains to be determined whether the
deletion of Col9α2 causes IVDD primarily by affecting EPs. Based on
these observations, it was hypothesized that the absence of Col9α2
may lead to IVDD through the modic change in EPs and other tissues
in IVDs. Therefore, the present study examined the effects of
Col9a2 knockout on EP structure, ECM deposition, matrix degradation
and chondrocyte hypertrophy-related gene and protein expression
alterations in IVD tissues.
Materials and methods
Animals
Mice with a deficiency in the Col9a2 gene
(Col9a2-/-) were provided by the Nanjing Biomedical
Research Institute of Nanjing University. The DNA from tail
biopsies of mice was genotyped using PCR. All mice were housed at a
constant room temperature of 20±2˚C and humidity of 50-60% with a
12-h light/dark cycle and free access to water and standard food. A
total of 36 male mice at 4 (12-15 g body weight), 8 (20-23 g body
weight) and 12 (26-29 g body weight) weeks of age were used for
further analysis. At each time-point, the Col9a2-/-
group was compared with the wild-type (WT) control group (n=6 per
group). All mice were humanely euthanized via intraperitoneal
injection of pentobarbital sodium (150 mg/kg). Animal death was
confirmed by observation of cessation of heartbeat and respiration.
The present study was approved by the Ethics Committee of Zhejiang
Chinese Medical University (Hangzhou, China; approval no.
20190401-10) according to the National Institutes of Health Guide
for the Care and Use of Laboratory Animals.
Micro-CT (µCT)
The lower thoracic and whole lumbar vertebrae of
mice were fixed in 4% neutral formaldehyde solution at 4˚C for 48
h, transferred to 70% anhydrous ethanol and then scanned and
examined using a high-resolution µCT scanner (Skyscan 1176; Bruker)
with the following settings: Isotropic voxel size resolution, 9 µm;
energy, 45 kV; current, 500 µA; and integration time, 780 msec. The
lower thoracic ribs were included for the identification of L4-L5
IVD localization. Image reconstruction and analysis was performed
using NRecon v1.6 (Bruker) and CTAn v1.9 (Bruker), respectively,
followed by analysis of the L4-L5 IVD parameters using
three-dimensional model visualization software (CTVolx, v3.0;
Bruker). The IVD volume was defined by the region of interest (ROI)
to cover the entire invisible space between the L4-L5 vertebrae and
the cartilage EP volume was defined as covering the visible bone
plate close to the vertebrae. A total of 50 consecutive ROI images
were used to display the three-dimensional reconstruction of the EP
and IVD space. The average of the volume and the height of the IVD
were calculated.
Histochemistry and
histomorphometry
The specimens were fixed in neutral formaldehyde
solution at a volume fraction of 4% (0.1 mol/l, pH 7.4) for 3 days
and then decalcified in 10% EDTA (buffered with pure water, pH 7.4)
for 2 weeks. The samples were embedded in paraffin, cut into 3
µm-thick sections and stained with Alcian blue hematoxylin/orange G
(ABH/OG) and safranin O and fast green. A total of six different
mice from each group at each time-point were selected for the
assessment of EP, annulus fibrosus (AF) and nucleus pulposus (NP)
using a light microscope (Carl Zeiss AG). The EP score was obtained
as previously described and used to evaluate the degeneration of
the EP, including structural disorganization, clefts and bony
sclerosis (24,25).
Immunohistochemistry (IHC) and
immunofluorescence (IF)
IHC was performed using a standard protocol. In
brief, 3-µm-thick paraffin-embedded sections were incubated at 60˚C
for 4 h prior to deparaffinizing and dehydrating. The sections were
then incubated in citrate buffer (0.01 M, pH 6.0; Beijing Solarbio
Science & Technology Co., Ltd.) at 60˚C for 4 h, or in pepsinum
(OriGene Technologies, Inc.) at 37˚C for 30 min for antigen
retrieval. The samples were then treated with endogenous peroxidase
blocker (cat. no. PV-6001; OriGene Technologies, Inc.) for 10 min
and incubated with 0.3% Triton X-100 for 15 min at room
temperature. The sections were then incubated with primary
antibodies against Col9a2 (diluted 1:100; cat. no. sc398130; Santa
Cruz Biotechnology, Inc.), Col2a1 (diluted 1:1,000 for IHC and 1:50
for IF; cat. no. ab34712; Abcam), Aggrecan (diluted 1:200 for IHC
and 1:50 for IF; cat. no. NB100-74350; Novus Biologicals, LLC),
Mmp13 (diluted 1:200; cat. no. ab39012; Abcam), ADAM
metallopeptidase with thrombospondin type 1 motif 5 (Adamts5;
diluted 1:200; cat. no. ab182795; Abcam), Col10a1 (diluted 1:200;
cat. no. ab58632; Abcam) and Runx family transcription factor 2
(Runx2; diluted 1:300; cat. no. ab76956; Abcam) overnight at 4˚C.
For IHC staining, the sections were incubated with a secondary goat
anti-mouse/rabbit antibody (diluted 1:1,000; cat. no. 31234;
Invitrogen; Thermo Fisher Scientific, Inc.) for 20 min at room
temperature. Positive staining of sections was visualized using
diaminobenzidine solution (Invitrogen; Thermo Fisher Scientific,
Inc.), while hematoxylin was used for counterstaining. For the IF
assay, the slides were incubated with fluorophore-conjugated
secondary antibodies at room temperature for 30 min in the dark.
The sections were counterstained with DAPI and observed under a
fluorescence microscope (Carl Zeiss AG). A total of five images
from each section were analyzed using Image-Pro Plus 6.0 (Media
Cybernetics, Inc.). The area of Col2a1 or Col10a1-positive staining
was calculated, while the Aggrecan-, Mmp13-, Adamts5- or
Runx2-positive cells were obtained by counting the number of
positively stained cells.
Reverse transcription-quantitative PCR
(RT-qPCR)
The L4-L5 segments of the spinal cords were removed
from 12-week-old mice and the soft tissues were excised. Total RNA
was extracted using the Qiagen RNeasy Mini kit (Qiagen GmbH) and
then reverse transcribed into cDNA using the RevertAid First Strand
cDNA Synthesis kit (Invitrogen; Thermo Fisher Scientific, Inc.)
according to manufacturers' protocols. As per the manufacturer's
instructions, qPCR was performed for target genes using SYBR Premix
Ex Taq™ II (Takara Biotechnology Co., Ltd.) with a QuantStudio™ 7
Flex Real-Time PCR System (Thermo Fisher Scientific, Inc.). The
thermocycling conditions were: Pre-denaturation at 94˚C for 5 min;
followed by 40 cycles of denaturation at 94˚C for 30 sec, annealing
at 60˚C for 30 sec and extension at 72˚C for 30 sec. The primer
sequences are presented in Table I
and the relative mRNA expression was measured using the
2-ΔΔCq method (26).
 | Table IPrimer sequences for quantitative
PCR. |
Table I
Primer sequences for quantitative
PCR.
| Gene/primer
direction | Sequence |
|---|
| β-actin | |
|
Forward |
5'-GGAGATTACTGCCCTGGCTCCTA-3' |
|
Reverse |
5'-GACTCATCTACTCCTGCTTGCTG-3' |
| Col2a1 | |
|
Forward |
5'-TGGTCCTCTGGGCATCTCAGGC-3' |
|
Reverse |
5'-GGTGAACCTGCTGTTGCCCTCA-3' |
| Mmp13 | |
|
Forward |
5'-TTTGAGAACACGGGGAAGA-3' |
|
Reverse |
5'-ACTTTGTTGCCAATTCCAGG-3' |
| Aggrecan | |
|
Forward |
5'-CGCCACTTTCATGACCGAGA-3' |
|
Reverse |
5'-TCATTCAGACCGATCCACTGGTAG-3' |
| Adamts5 | |
|
Forward |
5'-CCAAATGCACTTCAGCCACGATCA-3' |
|
Reverse |
5'-AATGTCAAGTTGCACTGCTGGGTG-3' |
| Col10a1 | |
|
Forward |
5'-ACCCCAAGGACCTAAAGGAA-3' |
|
Reverse |
5'-CCCCAGGATACCCTGTTTTT-3' |
| Runx2 | |
|
Forward |
5'-GAGGGCACAAGTTCTATCTGGA-3' |
|
Reverse |
5'-GGTGGTCCGCGATGATCTC-3' |
Statistical analysis
Statistical analysis was performed using SPSS
software (version 25.0; IBM Corp). Values are expressed as the mean
± the standard error of the mean and analyzed using a one-way ANOVA
followed by Bonferroni's post-hoc test as appropriate. P<0.05
was considered to indicate a statistically significant
difference.
Results
Changes in EP and increases in
porosity lead to early-stage IVDD in Col9a2-/- mice
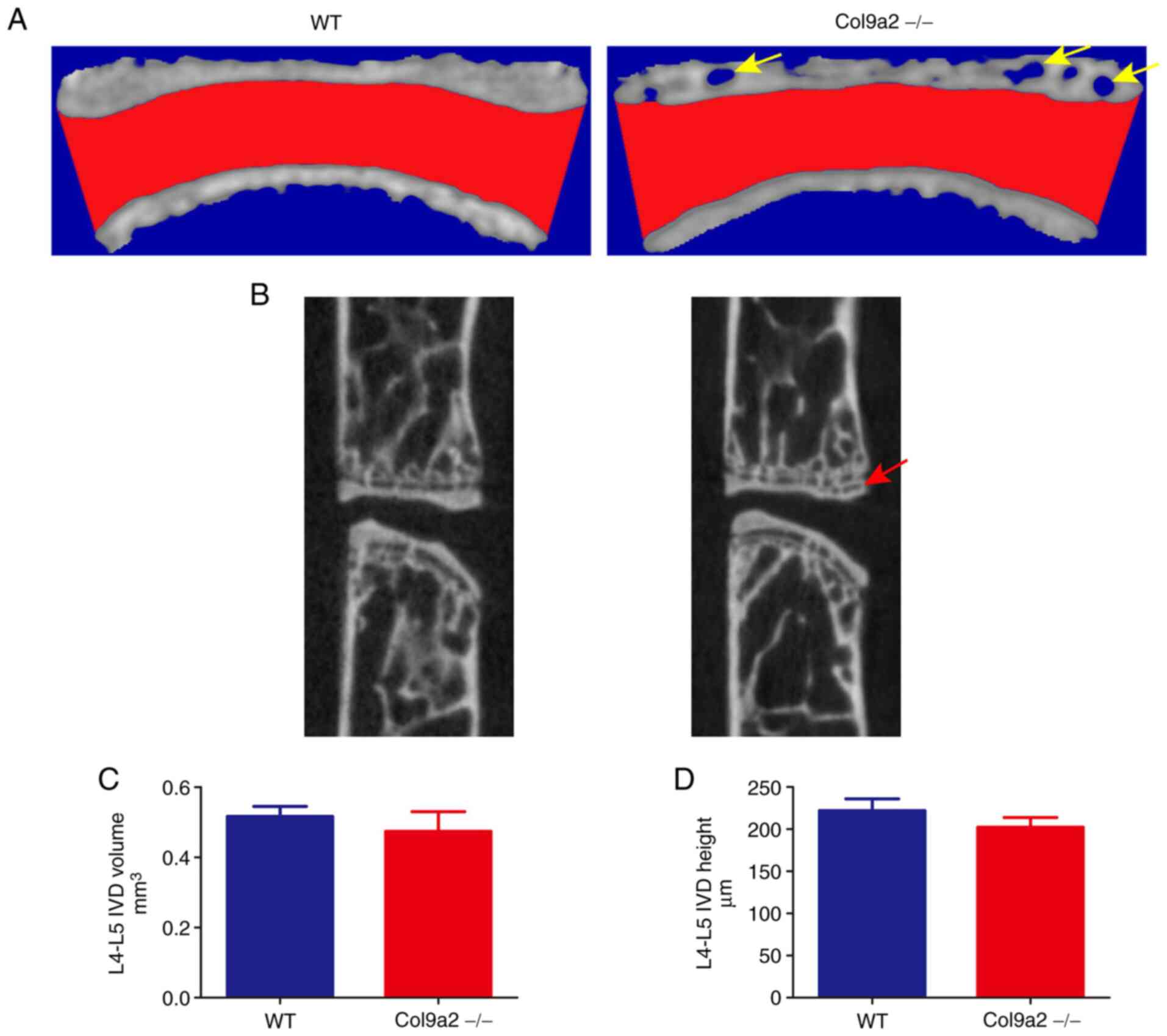
µCT scanning was implemented to qualitatively
estimate the lumbar spine of the Col9a2-/- and WT mice
at 12 weeks of age. The cavities within the EPs were significantly
increased in the CT three-dimensional images of the L4-L5 IVD
(Fig. 1A), as well as in the
median sagittal images of the CT scan (Fig. 1B). Although no statistically
significant differences were observed, the IVD volume and height of
Col9a2-/- mice were reduced compared with those of WT
mice (Fig. 1C and D), as determined through the analysis of
the three-dimensional reconstruction images of the L4-L5 IVD. Thus,
the CT data suggested that although there were no significant
differences in the L4-L5 IVD volume and height between the
Col9a2-/- and WT mice, the ossification of the EPs was
slightly increased and this is a phenotype of early-stage IVDD
(27).
Osteochondral remodeling of the EP
induces IVDD in Col9a2-/- mice
Histochemistry and histomorphometry were performed
to further observe the phenotypes of IVDD induced by Col9a2 gene
knockout. ABH/OG staining revealed that there were no obvious
histological differences between the two groups at 4 and 8 weeks,
except that the 8-week-old Col9a2-/- mice exhibited a
slight calcification of the EPs. The EPs of the
Col9a2-/- mice began to undergo significant degeneration
at 12 weeks of age compared with the WT mice, as indicated by the
apparent calcification and ossification formation of the EPs
(Fig. 2A). Furthermore, oteoglycan
loss was also present in the NP of Col9a2-/- mice, as
evidenced by less ABH/OG and safranin O and fast green staining
compared with the WT mice (Figs. 3
and 4). Furthermore, a large
number of clefts was observed in the AF of the Col9a2-/-
mice with collagen dislocation and cytopenia, and even AF rupture
(Figs. 3 and 4B). The EP score, a histological
evaluation of EP degeneration, was used to assess pathological
changes, such as the degree of osteosclerosis, structural disorders
and neovascularization. Of note, it was indicated that the EP score
was significantly increased in the Col9a2-/- mice
compared with the WT mice at 12 weeks, confirming EP degeneration
(Fig. 2B). The results of the
histochemical and histomorphometric analysis indicated that the
12-week-old Col9a2-/- mice exhibited certain other
manifestations of IVDD, such as AF rupture and proteoglycan
reduction, in addition to distinct EP degeneration, relative to WT
mice.
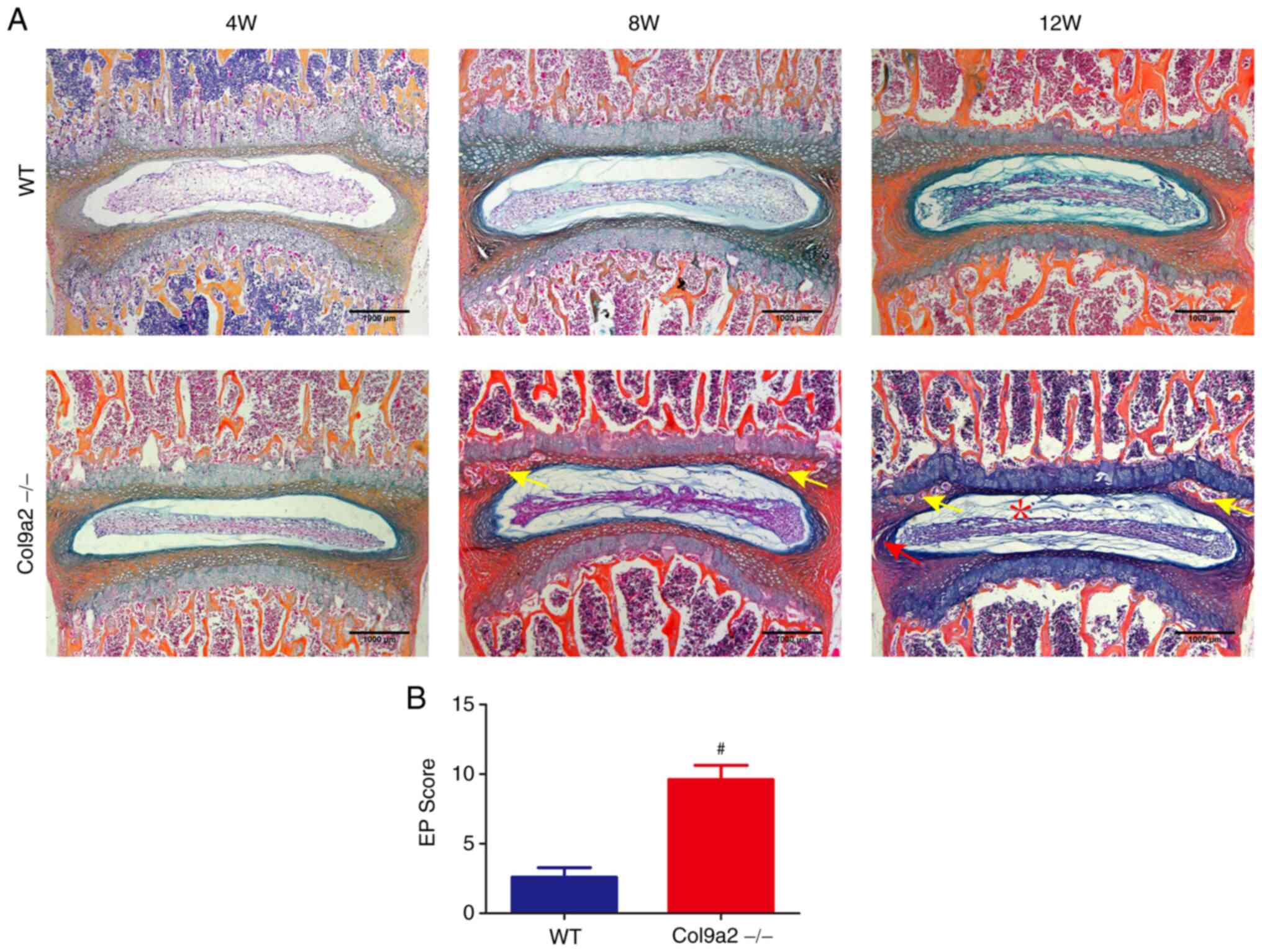 | Figure 2Accelerated osteochondral remodeling
in EPs of Col9a2-/- mice. (A) Representative ABH/OG
staining images of L4-L5 IVD in 4-, 8- and 12-week-old mice (scale
bars, 1,000 µm). (B) EP score in Col9a2-/- or WT mice as
an indication of EP degeneration. Yellow arrows indicate the
ossific endplate, the red asterisk indicates the decrease of
notochord cells and proteoglycans in the NP and the red arrow
indicates the formation of cracks in the AF. Values are expressed
as the mean ± standard deviation (n=6 per group).
#P<0.05. WT, wild-type; W, weeks; EP, endplate; NP,
nucleus pulposus; AF, annulus fibrosus; ABH/OG, Alcian
blue-hematoxylin/orange G; IVD, intervertebral disc; Col9α2, α2
chain of type IX collagen. |
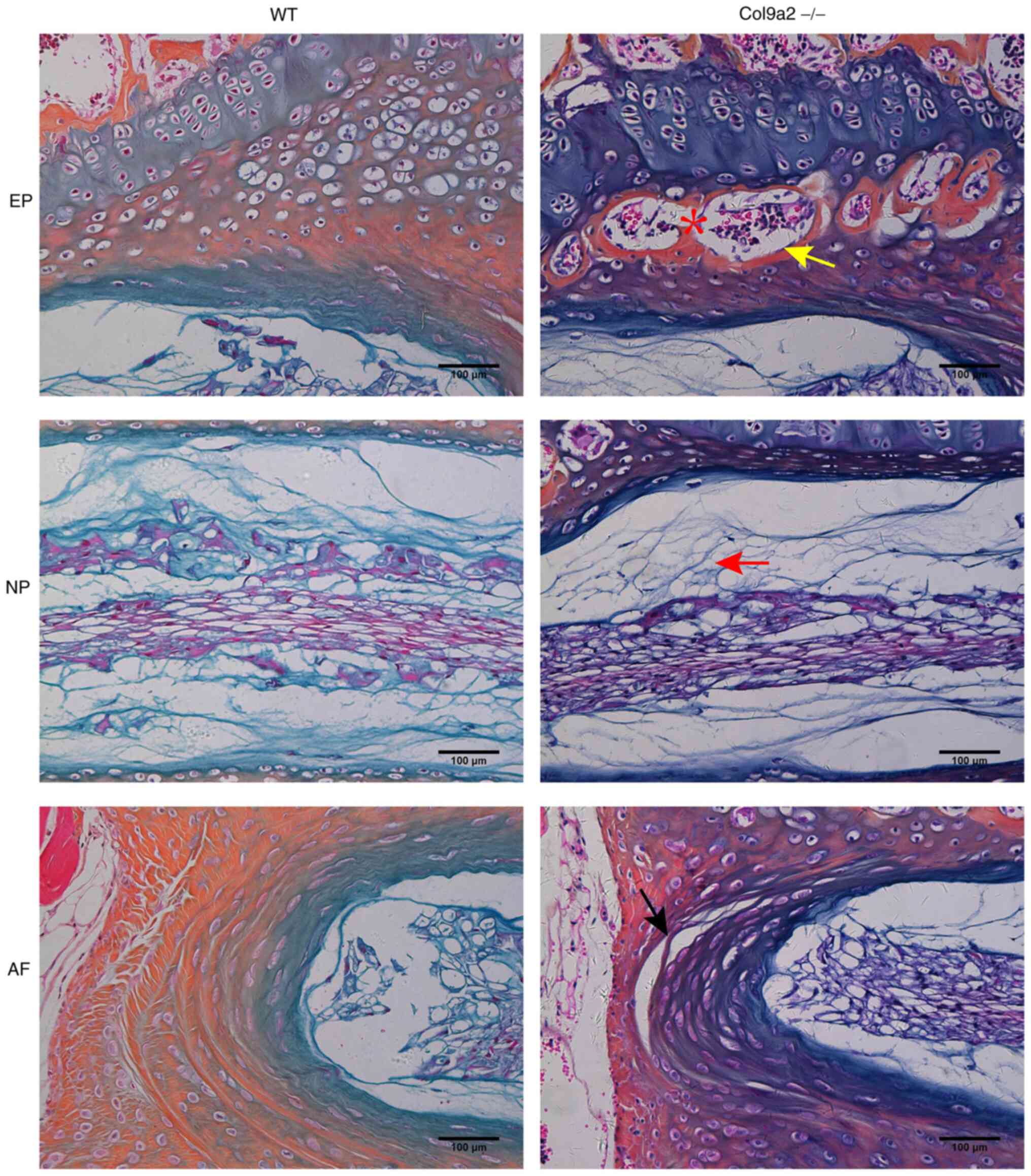 | Figure 3Representative higher-magnification
ABH/OG staining images of L4-L5 IVD of 12-week-old mice compared to
Fig. 2. Higher-magnification
images of the EP, NP and AF (scale bars, 100 µm). Ossific nodules
(yellow arrow) in the EP along with thickening of bony EP (red
asterisk) were observed in the Col9a2-/- mice.
Furthermore, a reduction of notochord cells and proteoglycan in the
NP (red arrow) as indicated by a paucity of ABH/OG staining and
crack formation within the AF (black arrow) were observed in the
Col9a2-/- mice (n=6 per group). WT, wild-type; EP,
endplate; NP, nucleus pulposus; AF, annulus fibrosus; ABH/OG,
Alcian blue-hematoxylin/orange G; IVD, intervertebral disc; Col9α2,
α2 chain of type IX collagen. |
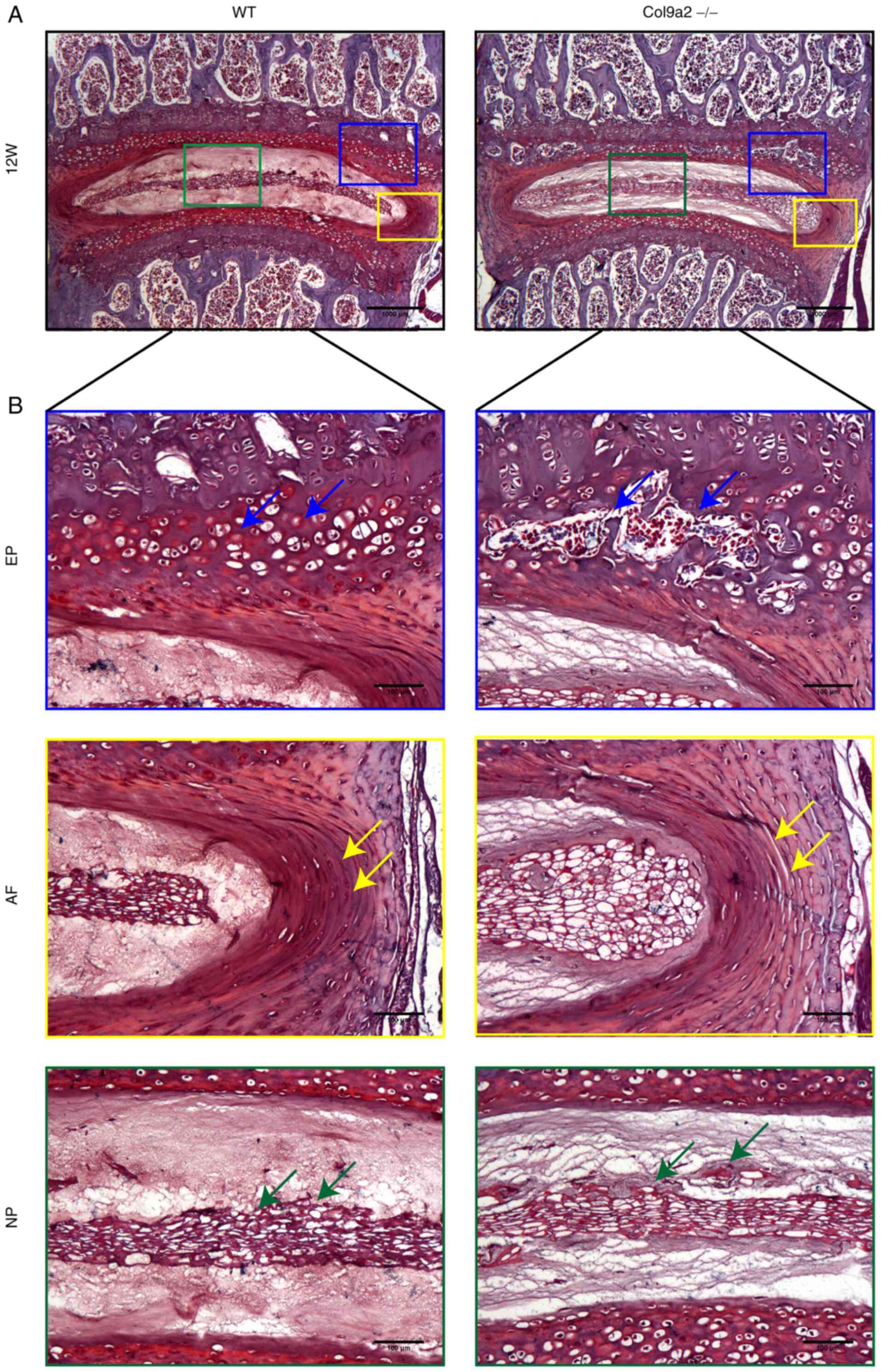 | Figure 4Safranin O and fast green staining
images of L4-L5 IVD of 12-week-old mice. (A) Representative
Safranin O and fast green staining images of L4-L5 IVD of
12-week-old WT and Col9a2-/- mice (scale bars, 1,000
µm). (B) Higher-magnification images of the EP, NP and AF (scale
bars, 100 µm). Blue arrows indicated ossific EP, yellow arrows
indicate the formation of cracks in the AF and green arrows
indicate the decrease of notochord cells and proteoglycans in the
NP (n=6 per group). WT, wild-type; EP, endplate; NP, nucleus
pulposus; AF, annulus fibrosus; IVD, intervertebral disc; Col9α2,
α2 chain of type IX collagen; W, weeks. |
Reduced expression of ECM
deposition-associated proteins and increased matrix degradation-
and chondrocyte hypertrophy-related protein levels in
Col9a2-/- mice
Further experiments were performed to determine
whether the protein levels of IVD-related proteins in
Col9a2-/- mice were also altered. IF was used to locate
the expression of Col9a2 and verify the efficiency of gene knockout
(Fig. 5A). Col9a2 was extensively
expressed in the IVDs of WT mice, while its expression was low in
the Col9a2-/- mice (Fig.
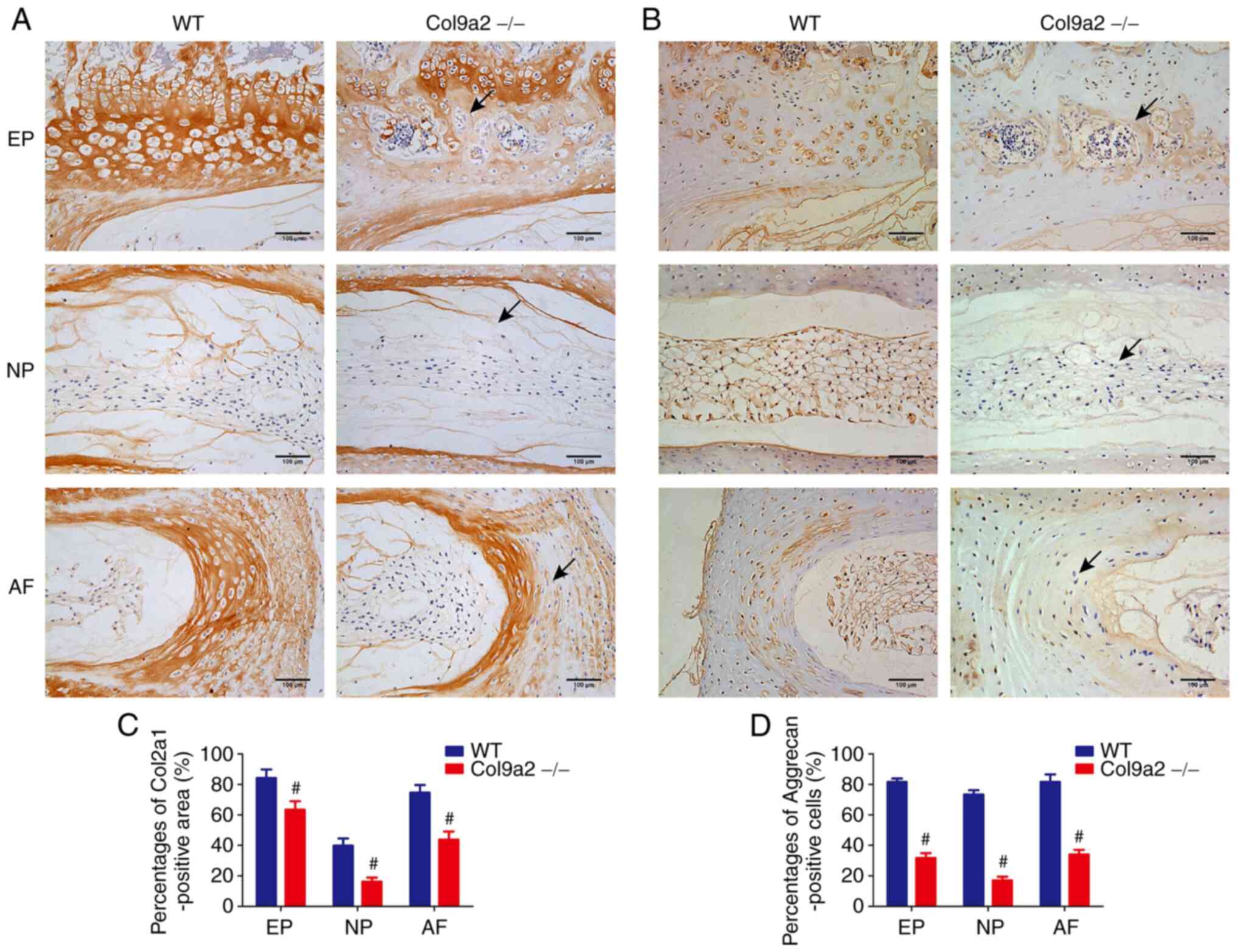
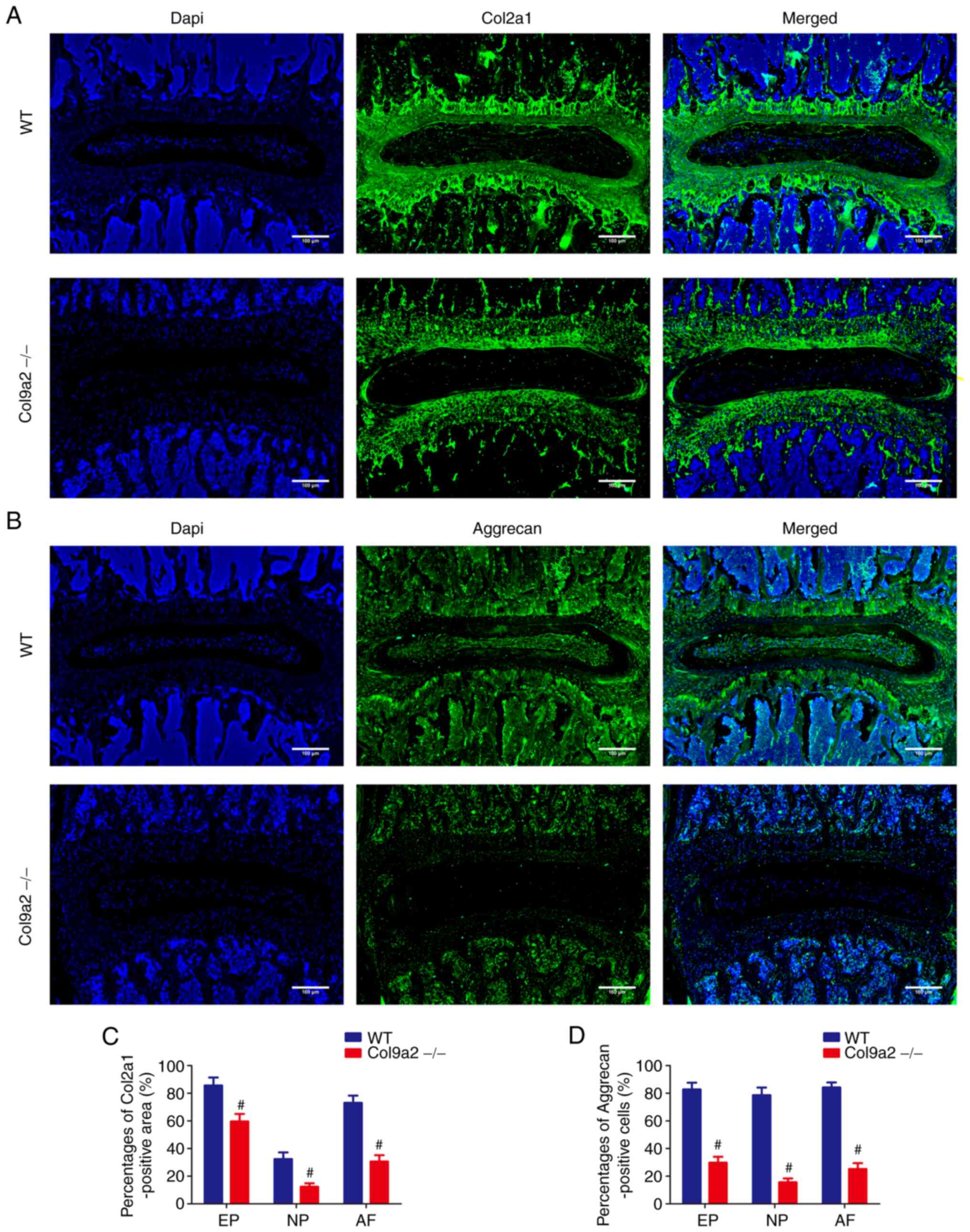
5B-D). IHC and IF analysis were then performed to assess the
expression levels of Col2a1, Aggrecan, Mmp13, Adamts5, Col10a1 and
Runx2 in the IVDs. The expression levels of Col2a1 and Aggrecan,
which indicate ECM deposition, were markedly decreased in the IVDs
of Col9a2-/- mice as compared with those in WT mice
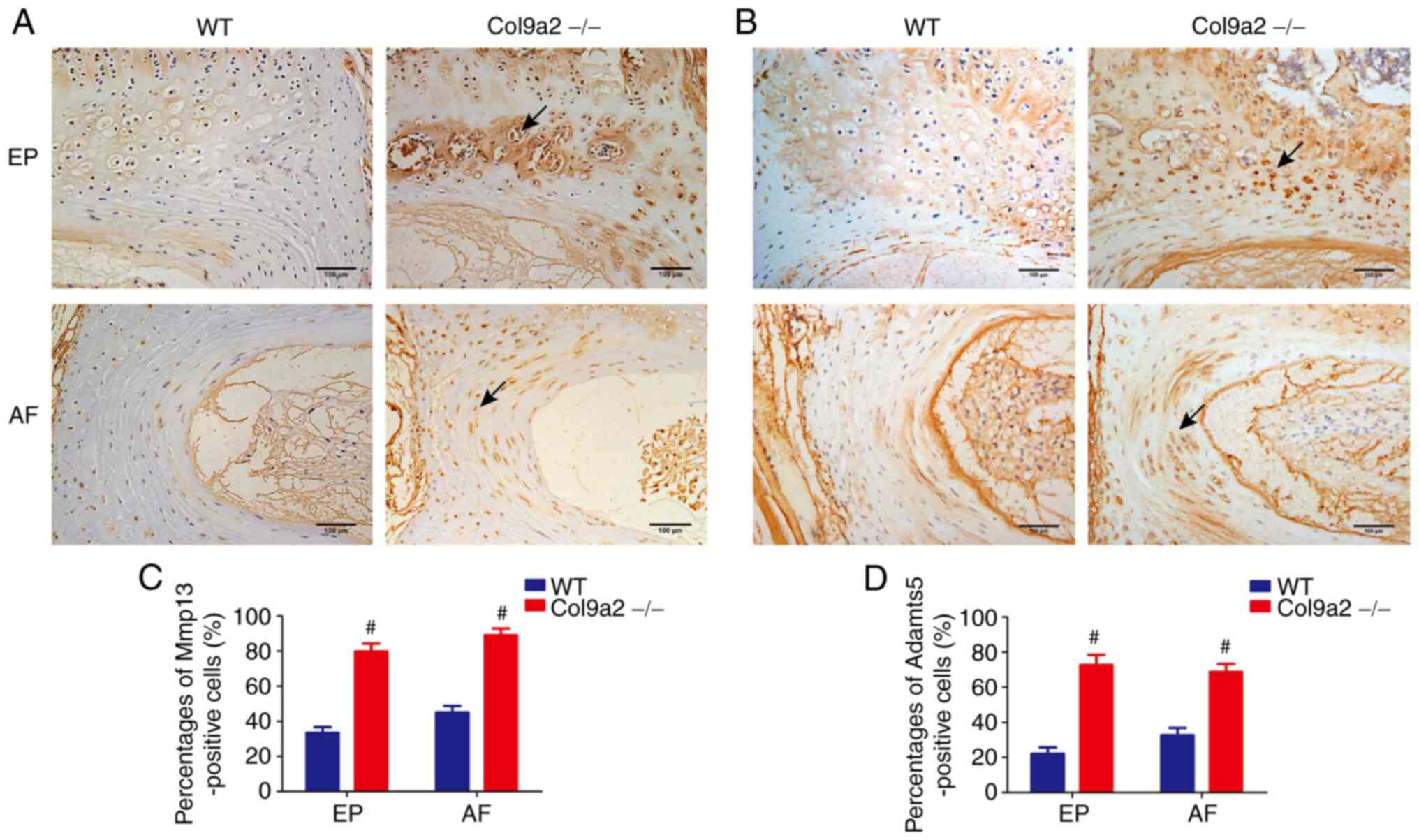
(Figs. 6 and 7). Furthermore, chondrocyte matrix
damage, as assessed by staining for Mmp13 and Adamts5 in the EPs
and AF of Col9a2-/- mice, increased significantly
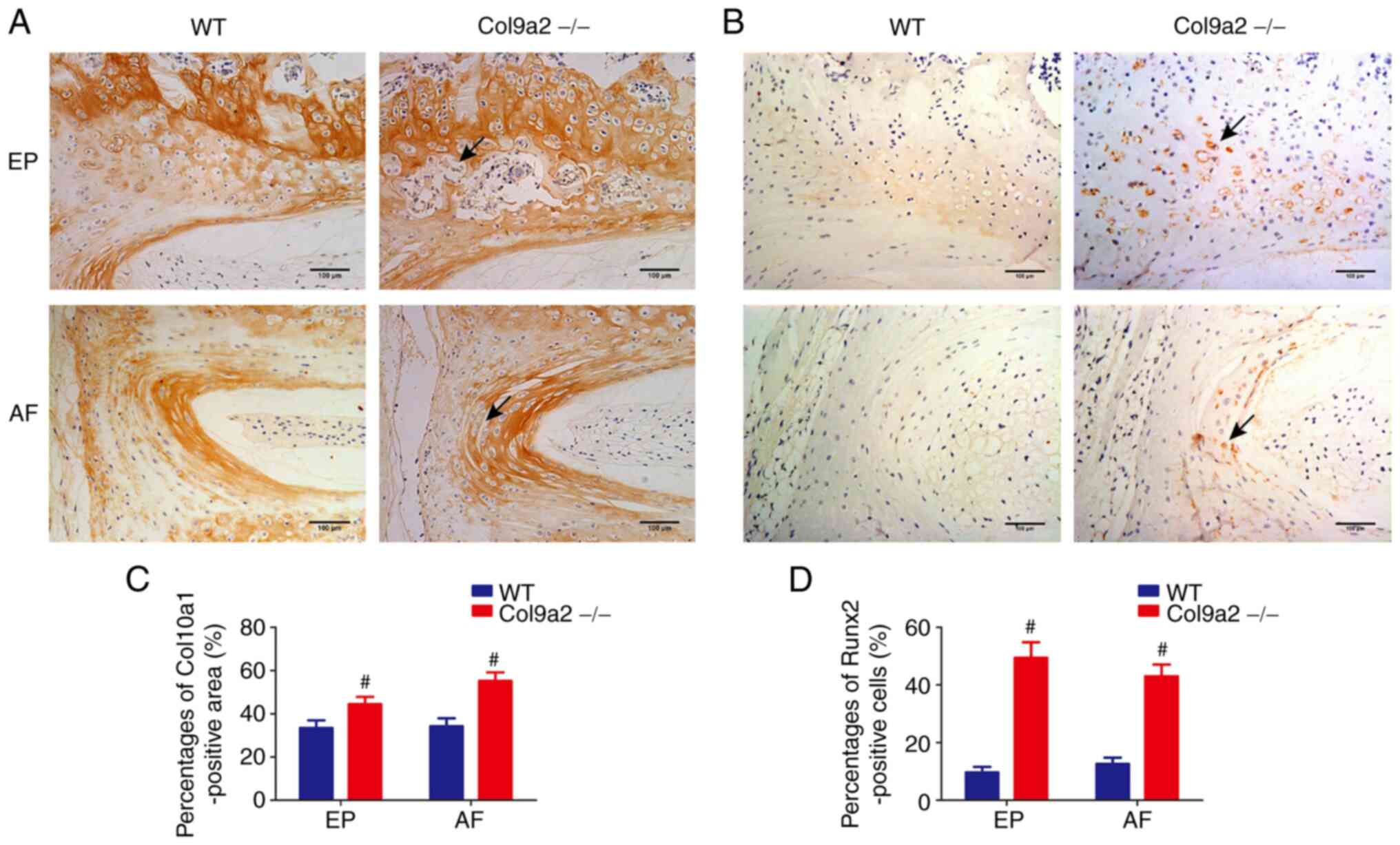
compared with that in the WT mice (Fig. 8). Furthermore, chondrocyte
hypertrophy, as evaluated by the number of Col10a1- and
Runx2-positive cells, was uniformly distributed in the EPs and AF
of WT mice, whereas in the EPs and outer layer of AF of
Col9a2-/- mice, positively stained cells were observed
at sites where the chondrocytes merged together; however, positive
cells were not present in the larger EP cavities and the expression
levels of Col10a1 and Runx2 were markedly increased in the IVDs of
Col9a2-/- mice compared with those in WT mice (Fig. 9). These results suggested that the
absence of the Col9a2 gene decreased the expression levels of ECM
deposition-associated proteins and increased that of matrix
degradation- and chondrocyte hypertrophy-related proteins in IVD
cartilage tissues, which exacerbated the occurrence and development
of IVDD.
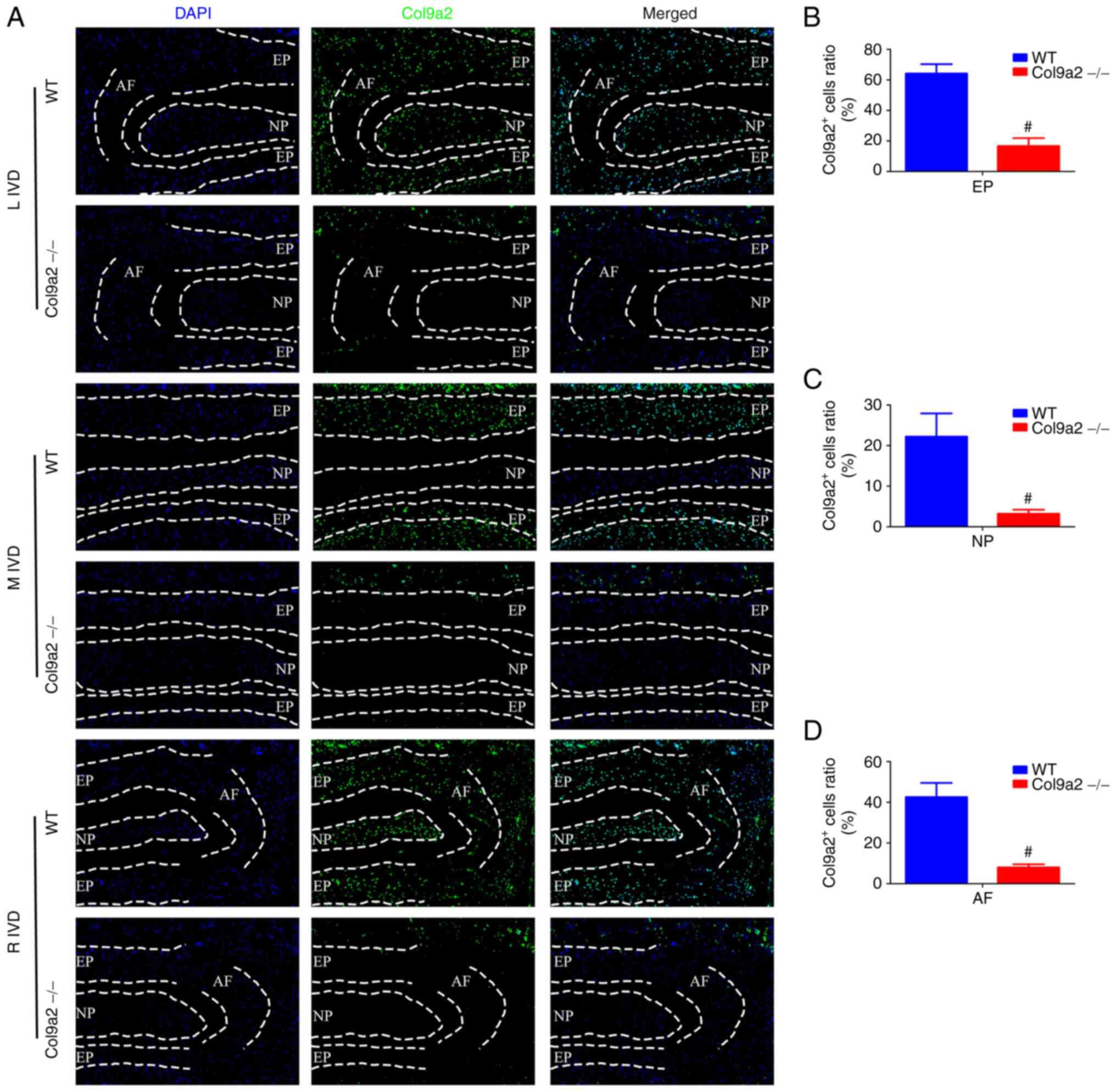 | Figure 5Col9a2 immunofluorescence staining of
IVDs of 2-week-old mice. (A) Col9a2 immunofluorescence staining of
EP, NP and AF. Nuclei were stained blue with DAPI and Col9a2
expression was detected as green (magnification, x200). (B-D)
Quantification of Co9a2 in the (B) EP, (C) NP and (D) AF, further
validating the efficiency of the gene knockout (n=6 per group).
#P<0.05 vs. WT. L, left; M, middle; R, right; WT,
wild-type; IVD, intervertebral disc; EP, endplate; NP, nucleus
pulposus; AF, annulus fibrosus; Col9α2, α2 chain of type IX
collagen. |
Diminished expression of genes
associated with ECM deposition and enhanced expression of genes
related to matrix degradation and chondrocyte hypertrophy in
Col9a2-/- mice
RT-qPCR was performed to further determine the mRNA
expression levels of ECM deposition-, matrix degradation- and
chondrocyte hypertrophy-related genes in the IVD tissue of
Col9a2-/- mice. As expected, the data demonstrated that
the mRNA expression trends of Col2a1, Aggrecan, Mmp13, Adamts5,
Col10a1 and Runx2 were similar to the results of protein expression
described above: Knockout of the Col9a2 gene diminished the mRNA
expression levels of Col2a1 and Aggrecan (Fig. 10A and B), while the mRNA expression levels of
Mmp13, Adamts5, Col10a1 and Runx2 were increased (Fig. 10C-F). These results suggested that
deletion of the Col9a2 gene suppressed ECM synthesis and
accelerated matrix degradation and chondrocyte hypertrophy in the
IVD tissue.
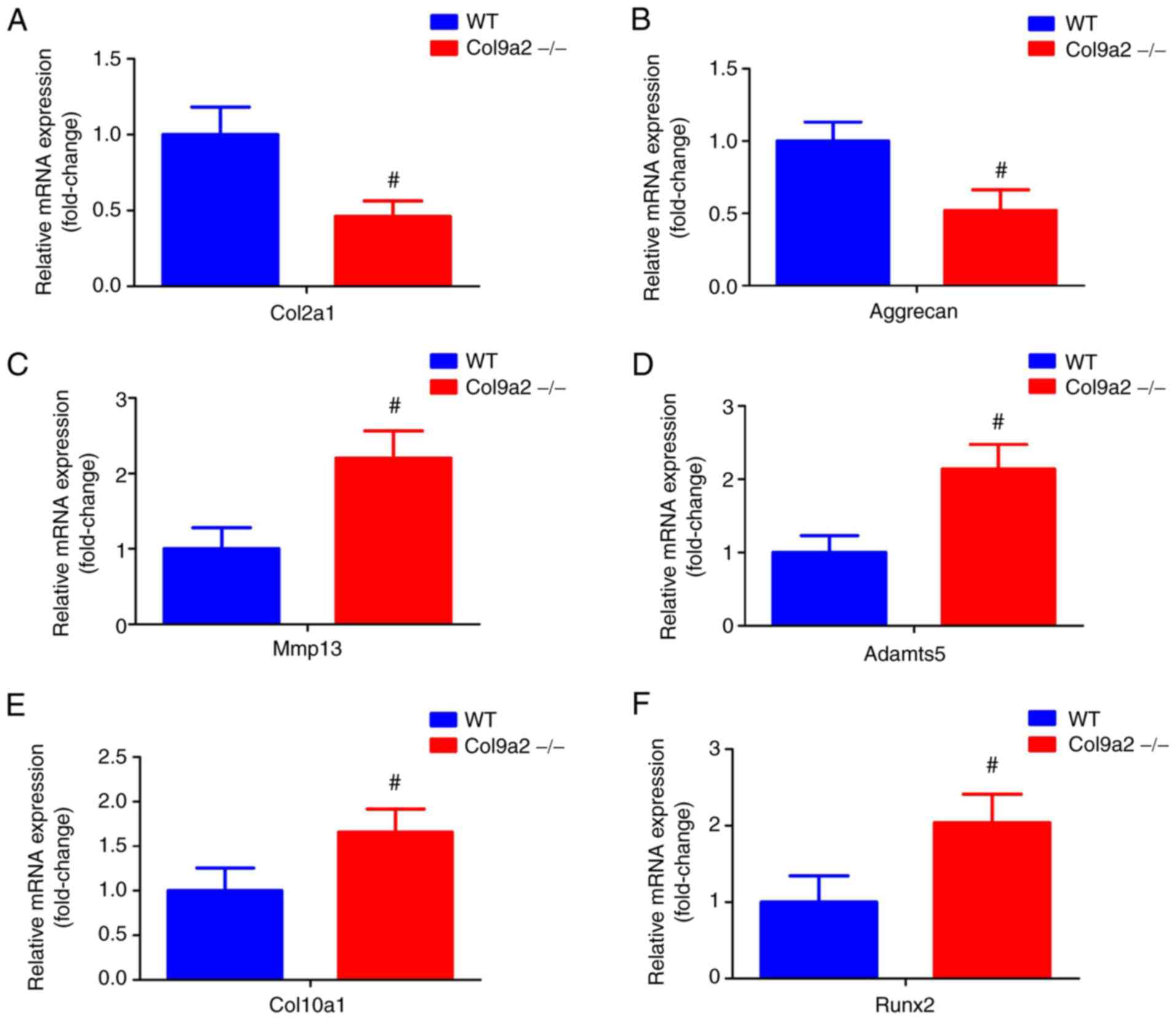 | Figure 10mRNA expression levels of Col2a1,
Aggrecan, Mmp13, Adamts5, Col10a1 and Runx2. Quantitative results
of mRNA expression analysis of (A) Col2a1, (B) Aggrecan, (C) Mmp13,
(D) Adamts5, (E) Col10a1 and (F) Runx2. Values are expressed as the
mean ± standard deviation (n=6 per group). #P<0.05
vs. WT. WT, wild-type; Col9α2, α2 chain of type IX collagen; Runx2,
Runx family transcription factor 2; Adamts5, ADAM metallopeptidase
with thrombospondin type 1 motif 5. |
Discussion
Col9 is a component of the ECM of cartilage, which
contributes to the integrity of the cartilage structure. Col9α2 is
one of the branched chains of Col9 and its deletion has been
indicated to be related to the early occurrence of IVDD. However,
its specific mechanisms of action have remained to be further
elucidated. The EP is a selective permeability barrier in IVDs and
has attracted increasing attention in recent years. Nutritional
access to the IVD is dependent on the EP. In addition, severe IVDD
is more common in patients with EP modic changes, indicating that
EP modic changes are related to the initiation and development of
IVDD (28). In the present study,
the histopathological staining results revealed that there was no
significant difference between the two groups of mice at 4 or 8
weeks; however, the Col9a2-/- mice at 12 weeks exhibited
the manifestations of early-stage IVDD, which was characterized by
EP calcification cavities, reduced proteoglycan content and AF
rupture, amongst other signs. EP degradation is considered to begin
with abnormal calcification (15).
Calcium crystalline salts are deposited into the pores of the EP,
resulting in a decrease in EP permeability. During the aging
process, the calcified EP undergoes ossification and sclerosis,
which is eventually replaced by bone (29,30).
This process is considered to reduce the transport of nutrients
from the vertebral marrow to the IVD. Thus, EP remodeling caused by
Col9a2 gene knockout may lead to increased permeability, which may
be the most important factor contributing to IVDD.
Furthermore, changes in the ECM composition of EP
chondrocytes have an important role in the occurrence and
development of IVDD. Studies have demonstrated that EPs are
critical to the nutritional supply of IVD due to the limited blood
supply (31,32). EP cartilage is a thin layer of
hyaline cartilage located between the vertebrae and IVD, mainly
composed of chondrocytes and ECM. EP chondrocytes are responsible
for the maintenance and circulation of cartilage-specific ECM
molecules. The major components of ECM within the IVDs are Col2a1
and Aggrecan, which endow EP cartilage with weight-bearing
properties and nutrient exchange abilities (33,34).
Therefore, their contents are essential for the proper function of
the IVDs, particularly in the cartilaginous EP. The excessive
destruction of ECM, and in particular, the loss of Col2a1 and
Aggrecan, may be risk factors for IVDD (35). Furthermore, Mmps and Adamts are the
main enzymes that degrade Col2a1 and Aggrecan, while Mmp13 and
Adamts5 are the hallmarks of cartilage degeneration (36). There is substantial and increasing
evidence to indicate that Mmp13 and Adamts5 expression is
upregulated in IVD tissues and cells, and are closely associated
with the process of ECM rupture and disc degeneration (37). In the present study, the results of
the IHC and RT-qPCR analysis revealed a distinct decrease in Col2a1
and Aggrecan protein and mRNA levels, while the levels of Mmp13 and
Adamts5 were markedly increased in Col9a2-/- mice
compared with WT mice. This suggests that Col9a2 gene knockout
causes excessive ECM destruction and insufficient ECM synthesis,
which further exacerbates EP calcification and ultimately
accelerates the progression of IVDD.
The PI3K/Akt signaling pathway has been reported to
be a key regulator of terminal chondrocyte differentiation in
embryonic and adult chondrogenesis (38,39),
and increased PI3K and Akt phosphorylation may enhance the levels
of Mmp13 and Adamts5, but decrease the levels of Col2a1 and
Aggrecan, ultimately exacerbating the process of IVDD (40). Furthermore, as a marker of
chondrocyte hypertrophy, the precise function of Col10a1 remains to
be determined; however, it is generally accepted that there is an
association between Col10a1 synthesis and endochondral osteogenesis
(41). Furthermore, Runx2, as a
marker of chondrocyte terminal differentiation, may alter the
phenotypic state of chondrocytes (42). In addition, it has been indicated
that the expression of Col10a1 and Runx2 in EP chondrocytes is
associated with degenerative disc disease (43,44).
It has been demonstrated that the ECM-receptor interaction
signaling pathway is dysregulated in the early stages of IVDD
(40). As components of the ECM,
Col9a2, Col2a1 and Aggrecan have important roles in the
ECM-receptor interaction signaling pathway. IVDD caused by Col9a2
gene knockout may be related to the imbalance of the ECM-receptor
interaction signaling pathway.
In conclusion, the present study provided insight
into the deteriorating effects of Col9a2 gene knockout on IVDD,
clearly demonstrating that Col9a2 gene knockout induces and
accelerates the progression of IVDD. The underlying mechanisms may
be related to the dysregulation of the PI3K/Akt and ECM-receptor
interaction signaling pathways due to the deletion of the Col9a2
gene, resulting in decreased protein and mRNA expression levels of
Col2a1 and Aggrecan, and increased expression levels of Mmp13,
Adamts5, Col10a1 and Runx2. This leads to enhanced EP
calcification, which subsequently disrupts the structural integrity
and function of the EP, and ultimately exacerbates the process of
IVDD. These changes may be a plausible explanation for IVDD
associated with the absence of the Col9a2 gene. However, the
research is still at a preliminary stage, and thus, the results
require to be further confirmed by clinical trials or other animal
experiments. The exact molecular mechanisms of IVDD and Col9a2 gene
deletion warrant further investigation.
Acknowledgements
Not applicable.
Funding
Funding: This research was funded by the Chinese National
Natural Science Foundation (grant nos. 81774346, 81873324,
81973869, 81904221 and 81904219), the Traditional Chinese Medical
Administration of Zhejiang Province (grant nos. 2021ZZ014,
2018ZA034, 2019ZQ018 and 2018ZZ011), the Health Commission of
Zhejiang Province (grant no. 2019RC225) and the Shaoxing Science
and Technology Project (grant no. 2018C30114).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
PT, HJ and CW made substantial contributions to the
conception and design of the study. HX, RD, QZ, LF and QG performed
the experiments. HX, RD, QZ, CX, PZ, SL and ZZ interpreted the data
and drafted the manuscript. PW, JL, HR and SH performed the
statistical analysis and reviewed the manuscript critically for
important intellectual content. HX, RD and QZ confirm the
authenticity of all the raw data. All authors read and approved the
final manuscript.
Ethics approval and informed consent
All animal experiments were approved by the
Institutional Ethics Committee of Zhejiang Chinese Medical
University (Hangzhou, China; no. 20190401-10).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Luoma K, Vehmas T, Kerttula L, Gronblad M
and Rinne E: Chronic low back pain in relation to modic changes,
bony endplate lesions, and disc degeneration in a prospective MRI
study. Eur Spine J. 25:2873–2881. 2016.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Izzo R, Popolizio T, D'Aprile P and Muto
M: Spinal pain. Eur J Radiol. 84:746–756. 2015.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Cazzanelli P and Wuertz-Kozak K: MicroRNAs
in intervertebral disc degeneration, apoptosis, inflammation, and
mechanobiology. Int J Mol Sci. 21(3601)2020.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Frapin L, Clouet J, Delplace V, Fusellier
M, Guicheux J and Le Visage C: Lessons learned from intervertebral
disc pathophysiology to guide rational design of sequential
delivery systems for therapeutic biological factors. Adv Drug Deliv
Rev. 149-150:49–71. 2019.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Chen BL, Guo JB, Zhang HW, Zhang YJ, Zhu
Y, Zhang J, Hu HY, Zheng YL and Wang XQ: Surgical versus
non-operative treatment for lumbar disc herniation: A systematic
review and meta-analysis. Clin Rehabil. 32:146–160. 2018.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Wu PH, Kim HS and Jang IT: Intervertebral
disc diseases PART 2: A review of the current diagnostic and
treatment strategies for intervertebral disc disease. Int J Mol
Sci. 21(2135)2020.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Makanji H, Schoenfeld AJ, Bhalla A and
Bono CM: Critical analysis of trends in lumbar fusion for
degenerative disorders revisited: Influence of technique on fusion
rate and clinical outcomes. Eur Spine J. 27:1868–1876.
2018.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Shapiro IM, Vresilovic EJ and Risbud MV:
Is the spinal motion segment a diarthrodial polyaxial joint: What a
nice nucleus like you doing in a joint like this? Bone. 50:771–776.
2012.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Chen S, Fu P, Wu H and Pei M: Meniscus,
articular cartilage and nucleus pulposus: A comparative review of
cartilage-like tissues in anatomy, development and function. Cell
Tissue Res. 370:53–70. 2017.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Kang R, Li H, Ringgaard S, Rickers K, Sun
H, Chen M, Xie L and Bünger C: Interference in the endplate
nutritional pathway causes intervertebral disc degeneration in an
immature porcine model. Int Orthop. 38:1011–1017. 2014.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Tomaszewski KA, Adamek D, Konopka T,
Tomaszewska R and Walocha JA: Endplate calcification and cervical
intervertebral disc degeneration: The role of endplate marrow
contact channel occlusion. Folia Morphol (Warsz). 74:84–92.
2015.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Mattei TA: Osteoporosis delays
intervertebral disc degeneration by increasing intradiscal
diffusive transport of nutrients through both mechanical and
vascular pathophysiological pathways. Med Hypotheses. 80:582–586.
2013.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Laffosse JM, Accadbled F, Molinier F,
Bonnevialle N, de Gauzy JS and Swider P: Correlations between
effective permeability and marrow contact channels surface of
vertebral endplates. J Orthop Res. 28:1229–1234. 2010.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Rodriguez AG, Slichter CK, Acosta FL,
Rodriguez-Soto AE, Burghardt AJ, Majumdar S and Lotz JC: Human disc
nucleus properties and vertebral endplate permeability. Spine
(Phila Pa 1976). 36:512–520. 2011.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Han Y, Li X, Yan M, Yang M, Wang S, Pan J,
Li L and Tan J: Oxidative damage induces apoptosis and promotes
calcification in disc cartilage endplate cell through
ROS/MAPK/NF-κB pathway: Implications for disc degeneration. Biochem
Biophys Res Commun. 516:1026–1032. 2019.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Paassilta P, Pihlajamaa T, Annunen S,
Brewton RG, Wood BM, Johnson CC, Liu J, Gong Y, Warman ML, Prockop
DJ, et al: Complete sequence of the 23-kilobase human COL9A3 gene.
Detection of Gly-X-Y triplet deletions that represent neutral
variants. J Biol Chem. 274:22469–22475. 1999.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Pihlajamaa T, Vuoristo MM, Annunen S,
Perala M, Prockop DJ and Ala-Kokko L: Human COL9A1 and COL9A2
genes. Two genes of 90 and 15 kb code for similar polypeptides of
the same collagen molecule. Matrix Biol. 17:237–241.
1998.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Kamper M, Paulsson M and Zaucke F: Absence
of collagen IX accelerates hypertrophic differentiation in the
embryonic mouse spine through a disturbance of the Ihh-PTHrP
feedback loop. Cell Tissue Res. 367:359–367. 2017.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Kamper M, Hamann N, Prein C,
Clausen-Schaumann H, Farkas Z, Aszodi A, Niehoff A, Paulsson M and
Zaucke F: Early changes in morphology, bone mineral density and
matrix composition of vertebrae lead to disc degeneration in aged
collagen IX -/- mice. Matrix Biol. 49:132–143. 2016.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Eyre DR, Pietka T, Weis MA and Wu JJ:
Covalent cross-linking of the NC1 domain of collagen type IX to
collagen type II in cartilage. J Biol Chem. 279:2568–2574.
2004.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Bruckner P: Suprastructures of
extracellular matrices: Paradigms of functions controlled by
aggregates rather than molecules. Cell Tissue Res. 339:7–18.
2010.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Noponen-Hietala N, Kyllonen E, Mannikko M,
Kyllönen E, Männikkö M, Ilkko E, Karppinen J, Ott J and Ala-Kokko
L: Sequence variations in the collagen IX and XI genes are
associated with degenerative lumbar spinal stenosis. Ann Rheum Dis.
62:1208–1214. 2003.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Hyun SJ, Park BG, Rhim SC, Bae CW, Lee JK,
Roh SW and Jeon SR: A haplotype at the COL9A2 gene locus
contributes to the genetic risk for lumbar spinal stenosis in the
Korean population. Spine (Phila Pa 1976). 36:1273–1278.
2011.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Boos N, Weissbach S, Rohrbach H, Weiler C,
Spratt KF and Nerlich AG: Classification of age-related changes in
lumbar intervertebral discs: 2002 volvo award in basic science.
Spine (Phila Pa 1976). 27:2631–2644. 2002.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Masuda K, Aota Y, Muehleman C, Imai Y,
Okuma M, Thonar EJ, Andersson GB and An HS: A novel rabbit model of
mild, reproducible disc degeneration by an anulus needle puncture:
Correlation between the degree of disc injury and radiological and
histological appearances of disc degeneration. Spine (Phila Pa
1976). 30:5–14. 2005.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Ruiz Wills C, Foata B, Gonzalez Ballester
MA, Karppinen J and Noailly J: Theoretical explorations generate
new hypotheses about the role of the cartilage endplate in early
intervertebral disk degeneration. Front Physiol.
9(1210)2018.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Jiang C, Guo Q, Jin Y, Xu JJ, Sun ZM, Zhu
DC, Lin JH, Tian NF, Sun LJ, Zhang XL and Wu YS: Inhibition of EZH2
ameliorates cartilage endplate degeneration and attenuates the
progression of intervertebral disc degeneration via demethylation
of Sox-9. EBioMedicine. 48:619–629. 2019.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Yuan FL, Xu RS, Ye JX, Zhao MD, Ren LJ and
Li X: Apoptotic bodies from endplate chondrocytes enhance the
oxidative stress-induced mineralization by regulating PPi
metabolism. J Cell Mol Med. 23:3665–3675. 2019.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Dudli S, Ballatori A, Bay-Jensen AC,
McCormick ZL, O'Neill CW, Demir-Deviren S, Krug R, Heggli I,
Juengel A, Karppinen J, et al: Serum biomarkers for connective
tissue and basement membrane remodeling are associated with
vertebral endplate bone marrow lesions as seen on MRI (Modic
changes). Int J Mol Sci. 21(3791)2020.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Liu Z, Easson GW, Zhao J, Makki N, Ahituv
N, Hilton MJ, Tang SY and Gray RS: Dysregulation of STAT3 signaling
is associated with endplate-oriented herniations of the
intervertebral disc in Adgrg6 mutant mice. PLoS Genet.
15(e1008096)2019.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Rade M, Määttä JH, Freidin MB, Airaksinen
O, Karppinen J and Williams FM: Vertebral endplate defect as
initiating factor in intervertebral disc degeneration: Strong
association between endplate defect and disc degeneration in the
general population. Spine (Phila Pa 1976). 43:412–419.
2018.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Herrero CF, Garcia SB, Garcia LV and
Aparecido Defino HL: Endplates changes related to age and vertebral
segment. Biomed Res Int. 2014(545017)2014.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Jackson AR, Huang CY and Gu WY: Effect of
endplate calcification and mechanical deformation on the
distribution of glucose in intervertebral disc: A 3D finite element
study. Comput Methods Biomech Biomed Engin. 14:195–204.
2011.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Li X and Yang S, Han L, Mao K and Yang S:
Ciliary IFT80 is essential for intervertebral disc development and
maintenance. FASEB J. 34:6741–6756. 2020.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Zhang T, Dai Y, Zhang L, Tian Y, Li Z and
Wang J: Effects of edible oils with different n-6/n-3 PUFA ratios
on articular cartilage degeneration via regulating the NF-κB
signaling pathway. J Agric Food Chem. 68:12641–12650.
2020.PubMed/NCBI View Article : Google Scholar
|
|
37
|
McCann MR, Veras MA, Yeung C, Lalli G,
Patel P, Leitch KM, Holdsworth DW, Dixon SJ and Séguin CA:
Whole-body vibration of mice induces progressive degeneration of
intervertebral discs associated with increased expression of Il-1β
and multiple matrix degrading enzymes. Osteoarthritis Cartilage.
25:779–789. 2017.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Kita K, Kimura T, Nakamura N, Yoshikawa H
and Nakano T: PI3K/Akt signaling as a key regulatory pathway for
chondrocyte terminal differentiation. Genes Cells. 13:839–850.
2008.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Yang S, Zhang F, Ma J and Ding W:
Intervertebral disc ageing and degeneration: The antiapoptotic
effect of oestrogen. Ageing Res Rev. 57(100978)2020.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Zhang Q, Shen Y, Zhao S, Jiang Y, Zhou D
and Zhang Y: Exosomes miR-15a promotes nucleus pulposus-mesenchymal
stem cells chondrogenic differentiation by targeting MMP-3. Cell
Signal. 86(110083)2021.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Hristova GI, Jarzem P, Ouellet JA,
Roughley PJ, Epure LM, Antoniou J and Mwale F: Calcification in
human intervertebral disc degeneration and scoliosis. J Orthop Res.
29:1888–1895. 2011.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Sun MM and Beier F: Chondrocyte
hypertrophy in skeletal development, growth, and disease. Birth
Defects Res C Embryo Today. 102:74–82. 2014.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Zhang Q, Huang M, Wang X, Xu X, Ni M and
Wang Y: Negative effects of ADAMTS-7 and ADAMTS-12 on endplate
cartilage differentiation. J Orthop Res. 30:1238–1243.
2012.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Iwata M, Aikawa T, Hakozaki T, Arai K,
Ochi H, Haro H, Tagawa M, Asou Y and Hara Y: Enhancement of Runx2
expression is potentially linked to β-catenin accumulation in
canine intervertebral disc degeneration. J Cell Physiol.
230:180–190. 2015.PubMed/NCBI View Article : Google Scholar
|