Introduction
Lung cancer is the most lethal malignancy in the
world and causes approximately 132 thousand deaths each year. A
total of 235,700 patients are diagnosed each year according to the
newest cancer statistical report by the American Cancer Society
(ACS) (1). In China, the data were
astonishing due to the large population. The average incidence was
733,300 and the number of deaths was 610,200(2). At present, the main strategy for lung
cancer is still surgical resection followed by drug chemotherapy
and/or radiation therapy (3).
However, the majority of patients do not benefit from traditional
therapy. Approximately 57% of the patients diagnosed with lung
cancer have distant metastasis while 17% have a localized
distribution (1). The patient's
survival is dependent on their cancer stage. The five-year survival
rate of patients with distant metastasis is only 6% (1). There is little time left for us to
identify more potent weapons to fight lung cancer.
Lung cancer tends to metastasize due to its high
genetic heterogeneity (4). From
the emergence of second-generation sequencing technology, a list of
genetic factors including mRNAs, long non-coding (lnc)RNAs, and
microRNAs (miRNAs) have been detected that take part in the
progression or metastasis of lung cancer. For example, miR-196b was
reported to inhibit cell metastasis of lung cancer (5). Loss of E-cadherin was found to
promote cell metastasis in lung cancer (6). However, in a cohort of lung cancer
patients, the dominant genetic factors are always different, which
suggests that it is a long journey to determine the deep underlying
mechanisms of lung cancer.
Lactamase β (LACTB) is a gene mapped to
chromosome 15q22.2 that encodes a mitochondrially localized
β-lactamase. It is reported to play suppressor roles in the
metastasis and progression of various type of cancer. For example,
LACTB was found to suppress the migration and invasion of melanoma
and to block the lung metastasis of melanoma cells (7). In glioma, LACTB inhibited the
proliferation, invasion and angiogenesis of tumor cells (8). Tumor angiogenesis is a well-known
pathway that mediates the distant metastasis of cancer cells
(9). Zeng et al
demonstrated that LACTB attenuated p53 ubiquitination and inhibited
the progression of colorectal cancer (10). However, in contrast, LACTB was
found to contribute to cancer metastasis and to correlate with the
poor prognosis of nasopharyngeal and pancreatic cancer (11,12).
These data suggest that the role of LACTB differs in a particular
cancer. Chemotherapy greatly improved the prognosis of cancer
patients in the clinic. However, drug resistance is now a great
challenge in the clinic. Drug resistance often reduces the efficacy
of chemotherapeutic drugs and results in short survival and low
quality of life of patients (13).
LACTB was shown to induce apoptosis of oxaliplatin-resistant cells
and improve the efficacy of oxaliplatin in gastric cancer (14). What is the role of LACTB in lung
cancer therapy and its underlying mechanism?
Epithelial-mesenchymal transition (EMT) is an
important cellular program and is responsible for the aggressive
behavior in cancer (15).
Activation of the EMT process confers increased metastatic
potential and drug resistance to cancer cells. In lung cancer,
activation of the EMT signaling pathway greatly promotes tumor
progression (16). At the
molecular level, LACTB was reported to inhibit EMT and suppress
tumor progression in colorectal cancer (17). Based on the information, it is
thought that LACTB might regulate progression of lung cancer
through EMT signaling.
In the present study, to explore the role of
LACTB in lung cancer, the expression of LACTB was
overexpressed/knocked down and lung cancer cells were subsequently
treated with docetaxel. Then, cell invasive ability, apoptosis and
the mechanism of action were determined.
Materials and methods
Bioinformatics analysis
The expression and methylation levels of LACTB in
lung cancer were obtained by analyzing data from The Cancer Genome
Atlas (TCGA) database (http://ualcan.path.uab.edu/analysis.html). The
association of LACTB expression with the survival of lung cancer
patients was obtained from the public database Kaplan-Meier Plotter
(http://kmplot.com/analysis/index.php?p=service&cancer=lung).
Cell culture
H1299 (non-small cell lung cancer) and H1975 (lung
adenocarcinoma) cell lines were purchased from the Chinese Academy
of Sciences (Shanghai, China) and cultured in RPMI-1640 medium
(Gibco; Thermo Fisher Scientific, Inc.) containing 10% fetal bovine
serum (FBS; Gibco; Thermo Fisher Scientific, Inc.) at 37˚C. The
culture medium was refreshed every day and the cells were passaged
at 1:3 after arriving at 100% confluence.
Reverse-transcription-quantitative
polymerase chain reaction (qPCR)
Total RNA was extracted from the treated cell lines
with RNeasy kit (Beyotime Institute of Biotechnology) according to
the manufacturer's instructions. One microgram of total RNA was
used as template for the synthesis of the first strand of cDNA with
a cDNA synthesize kit (Beyotime Institute of Biotechnology). Then 1
µl of cDNA was used as a template to quantify the expression of
target genes with SYBRGreen Mix (Beyotime Institute of
Biotechnology). The protocol was as follows: 95˚C for 5 min; (95˚C
for 15 sec; 60˚C for 15 sec) for 40 cycles. Then the
2-∆∆Cq method was used to analyze the relative
expression of target genes (18).
GAPDH was used as the internal control. The primers for qPCR
are shown in Table I.
 | Table IPrimers for quantitative polymerase
chain reaction assay. |
Table I
Primers for quantitative polymerase
chain reaction assay.
| Gene | Primers
(5'-3') |
|---|
| LACTB | |
|
Forward |
GTGGTTGGAGTTTCTGTAGATGGAA |
|
Reverse |
AGTAATCTTGTTGTGACAGAAACCT |
| GAPDH | |
|
Forward |
CGTATTGGGCGCCTGGTCAC |
|
Reverse |
ATGATGACCCTTTTGGCTCC |
Western blot analysis
Total protein was extracted from the treated cell
lines with Membrane and Cytosol Protein Extraction Kit (Beyotime
Institute of Biotechnology) according to the manufacturer's
instructions. Total protein (10 µg) was separated by SDS-PAGE
method and transferred onto PVDF membranes (Beyotime Institute of
Biotechnology). Then the PVDF membranes were blocked with 5%
non-fat milk (Beyotime Institute of Biotechnology) for 1 h at room
temperature followed by incubation with primary antibodies against
the target protein at 4˚C overnight. After washing with TBST
(Beyotime Institute of Biotechnology) for three times, the PVDF
membranes were incubated with secondary antibodies for 1 h at room
temperature. After washing with TBST for three times, the ECL kit
(Beyotime Institute of Biotechnology) was used to detect the
protein bands on the PVDF membranes. β-actin was used as the
internal control. The primary and secondary antibodies were as
below: MMP9 (cat. no. 10375-2-AP; dilution, 1:500), MMP2 (cat. no.
10373-2-AP; dilution, 1:500), N-cadherin (cat. no. 22018-1-AP;
dilution, 1:5,000), E-cadherin (cat. no. 20874-1-AP; dilution,
1:5,000), SNAIL (cat. no. 13099-1-AP; dilution: 1:500), LACTB (cat.
no. 18195-1-AP; dilution, 1:2,000), Vimentin (cat. no. 10366-1-AP;
dilution, 1:5,000), GAPDH (cat. no. 10494-1-AP; dilution, 1:10,000)
and Rabbit IgG (cat. no. SA00001-2; dilution, 1:5,000)(all
ProteinTech Group, Inc.). Adobe Photoshop CS6 (Adobe Systems, Inc.)
was used for quantification of the blots.
Synthesis of shRNA targeting LACTB,
construction of the plasmid carrying the LACTB gene and drug
administration
shRNA targeting the human LACTB gene
(shLACTB) was designed and synthesized, and the negative control
was shNC. The target sequences are listed in Table II. Then 40-80 pmol of shLACTB or
shNC was used to transfect them into the tumor cells with Lipo2000
agent (Invitrogen; Thermo Fisher Scientific, Inc.) for 6 h. Then
fresh medium was added into each well and cultured for another 48 h
followed by subsequent experiments.
 | Table IITarget sequence for LACTB
gene. |
Table II
Target sequence for LACTB
gene.
| Gene | Targets
(5'-3') |
|---|
| LACTB | |
|
shRNA-#1 |
CCTTACGTGGATAACTCCTAT |
|
shRNA-#2 |
CCTTAACACCATAGGTGCAAA |
|
shNC |
CCTTCGCCTAAGCATTAGGCAT |
A lentiviral expression plasmid carrying the coding
sequence of the human LACTB gene (pLACTB) was constructed,
confirmed by sequencing and the lentivirus-expressing LACTB was
prepared by Shanghai GeneChem. Then, 2 µl of the plasmid was added
into the cancer cells. After culture for 72 h, subsequent
experiments were carried out.
Cells were treated with the designed dosage of
docetaxel (10, 50,100 and 200 nmol/l) (QILU Pharmaceutical
Co.,Ltd.) in combination with shNC, shLACTB or empty vector pCDH
(Ctrl) or pLACTB for 48 h followed by subsequent experiments.
Cell Counting Kit 8 (CCK-8)
A total of 3x103 cells/well were seeded
into 96-well plates and treated as described above. After culture
for 72 h, 10% of CCK-8 agent (Yeasen) was added into each well and
cultured for another 1 h. Then, the absorbance at 450 nm was
detected on a microplate reader (Tecan-SPARK).
Wound-healing assay
A total of 2x105 cells/well were seeded
into 24-well plates and treated as described above. After culture
for 24 h, the confluence of 90% was achieved, and a scratch was
produced using a 10-µl tip followed by washing with PBS (Gibco;
Thermo Fisher Scientific, Inc.) two times. The width of the scratch
was determined and recorded as S0 h. After culture for
another 24 h, the width of each scratch was determined and recorded
as S48 h. The relative migration rate was calculated as
follows: Migration rate=(S0 h-S24 h)/S0
h.
Transwell assay
The insert chamber with 8-µm pores (Corning, Inc.)
was pretreated with/without 200 µl of cold Matrigel (BD,
Bioscience) at room temperature for 1 h followed by washing with
culture medium once. Then a total of 2x104 cells/well
were added to each room with 300 µl of culture medium containing no
serum and the room was inserted into 24-well plates with 300 µl of
culture medium containing 10%FBS. After culture for 24 h, cells on
the upper surface of the insert room were scraped and cells on
lower surface was fixed in 4% paraformaldehyde (Beyotime Institute
of Biotechnology) for 10 min at room temperature. After washing
with PBS, the insert chamber was stained with 0.5% crystal violet
(Beyotime Institute of Biotechnology) for 10 min at room
temperature. Then the positive cells were counted under a light
microscope (Olympus) at x100 magnification after washing with
PBS.
Flow cytometric analysis
A total of 2x105 cells/well were seeded
into 6-well plates and treated as the above. After culture for 48
h, cells were collected and washed with cold PBS once. Then cells
were suspended in staining buffer and stained with Annexin
V-PE/7-AAD reagent (Beyotime Institute of Biotechnology) at room
temperature for 15 min in the dark. The positively marked cells
were detected with BD FACSCelesta (BD Biosciences).
Statistical analysis
The data in this study were analyzed using SPSS
software (SPSS, Inc.; Version 16.0) and was expressed as the mean ±
SD. The difference between two groups was analyzed by unpaired
Student's t-test, while one-way ANOVA followed by Tukey's test was
used for more than three groups. P<0.05 indicates a significant
difference.
Results
LACTB is correlated with favorable
prognosis in lung cancer
To study the role of the LACTB gene in lung
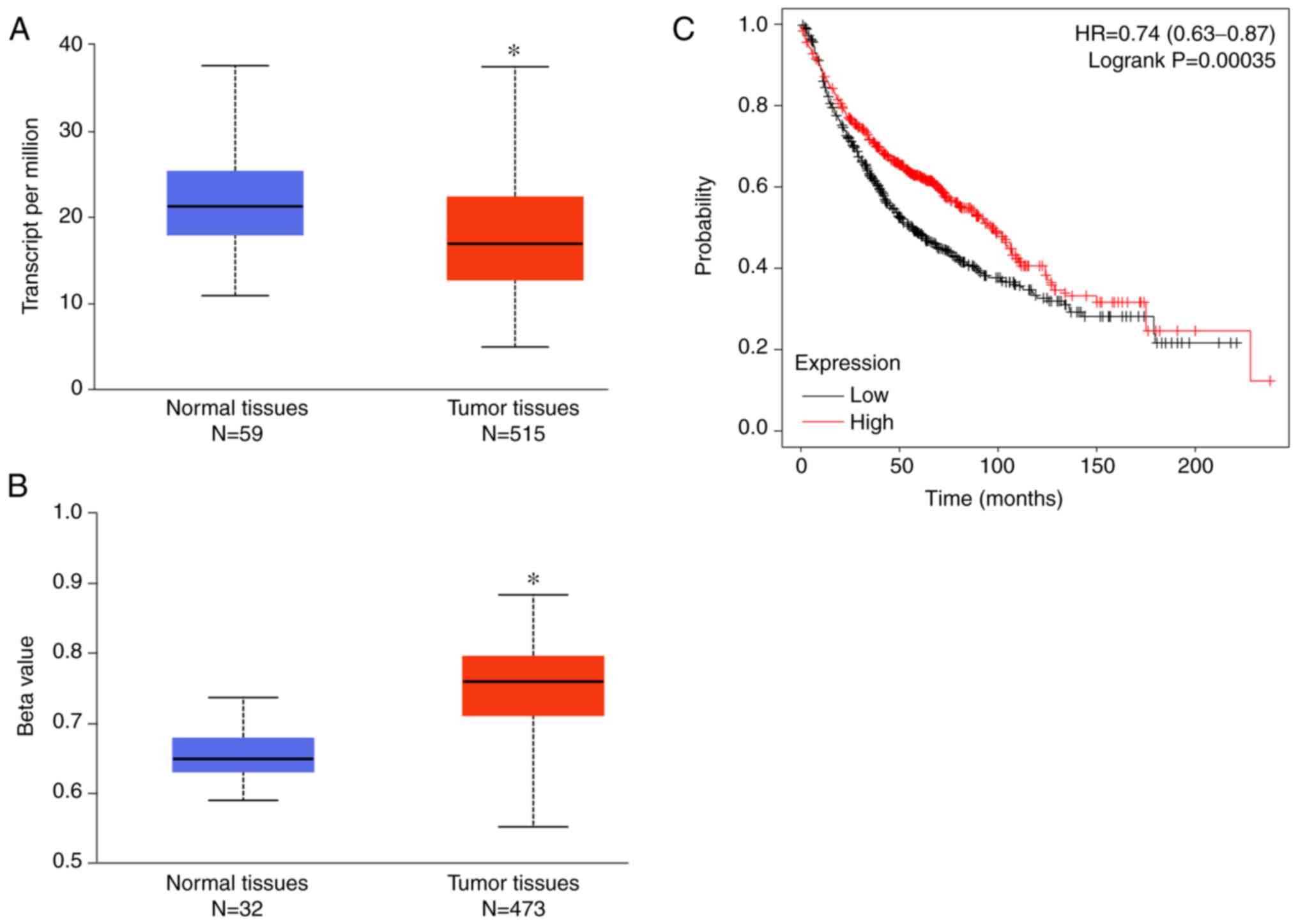
cancer, we extracted information from public databases. Based on
the TCGA database, the expression level of LACTB in 515 tissues
with lung cancer was significantly downregulated compared to that
in the normal tissues (Fig. 1A).
The methylation level of LACTB in tumor tissues was significantly
higher than in normal control tissues (Fig. 1B). Moreover, survival analysis
demonstrated that the survival probability of patients with high
LACTB expression was significantly better than that of patients
with low LACTB expression (Fig.
1C). These data suggest that LACTB might play an important role
in the progression of lung cancer.
LACTB does not affect cell survival in
lung cancer
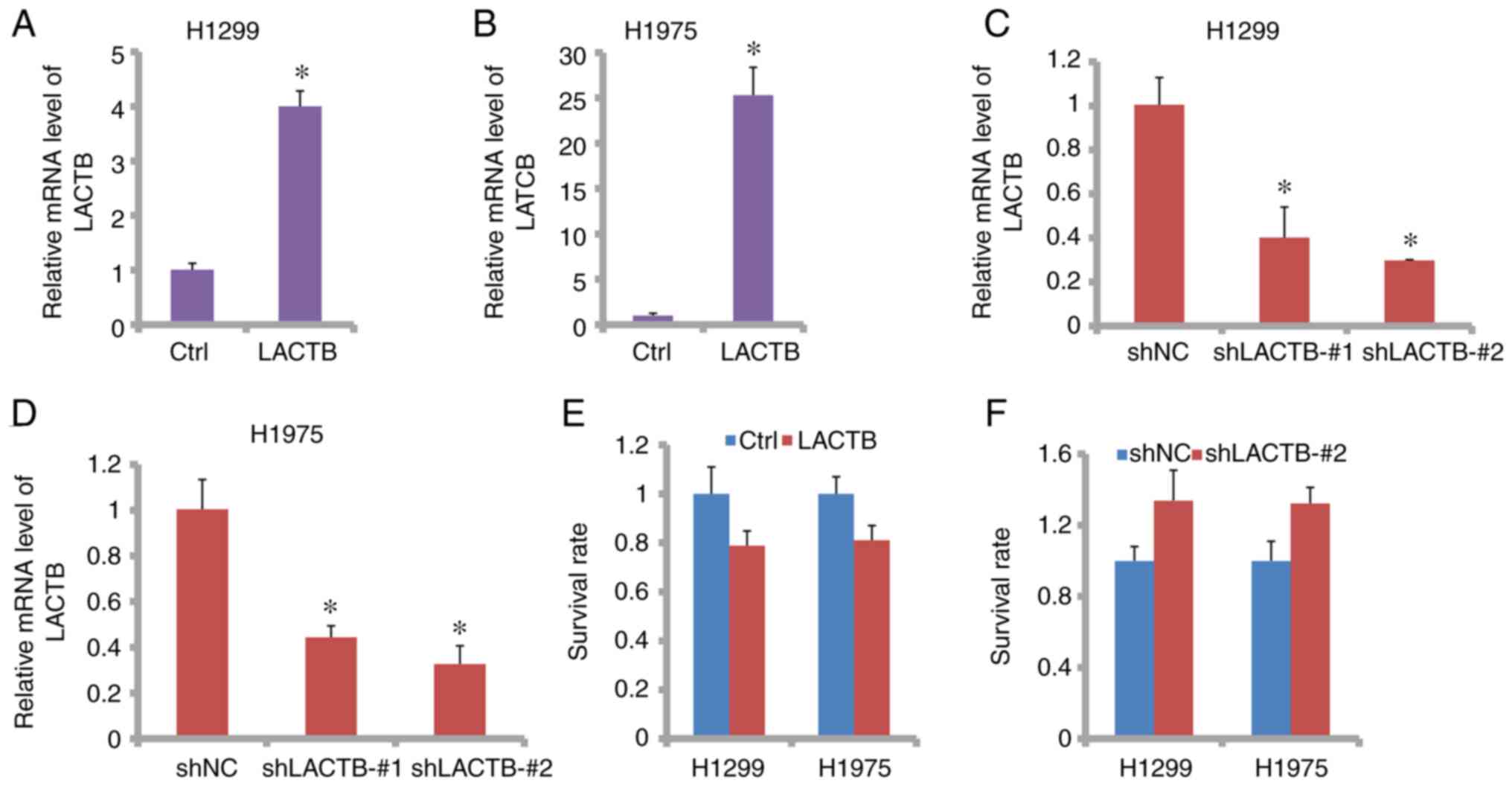
To investigate the role of LACTB in lung cancer, the
expression of LACTB was increased or decreased in the H1299 and
H1975 cell lines. As shown in Fig.
2A and B, qPCR analysis showed
that LACTB was successfully increased in the H1299 and H1975 cells.
Additionally, shRNA technology ensured that LACTB was significantly
decreased (Fig. 2C and D) in the H1299 and H1975 cells. Then by
CCK-8 assay, cell survival was shown to be minimally affected by
LACTB (Fig. 2E and F). Therefore, LACTB does not affect cell
survival in lung cancer.
LACTB negatively regulates migration
and invasion in lung cancer
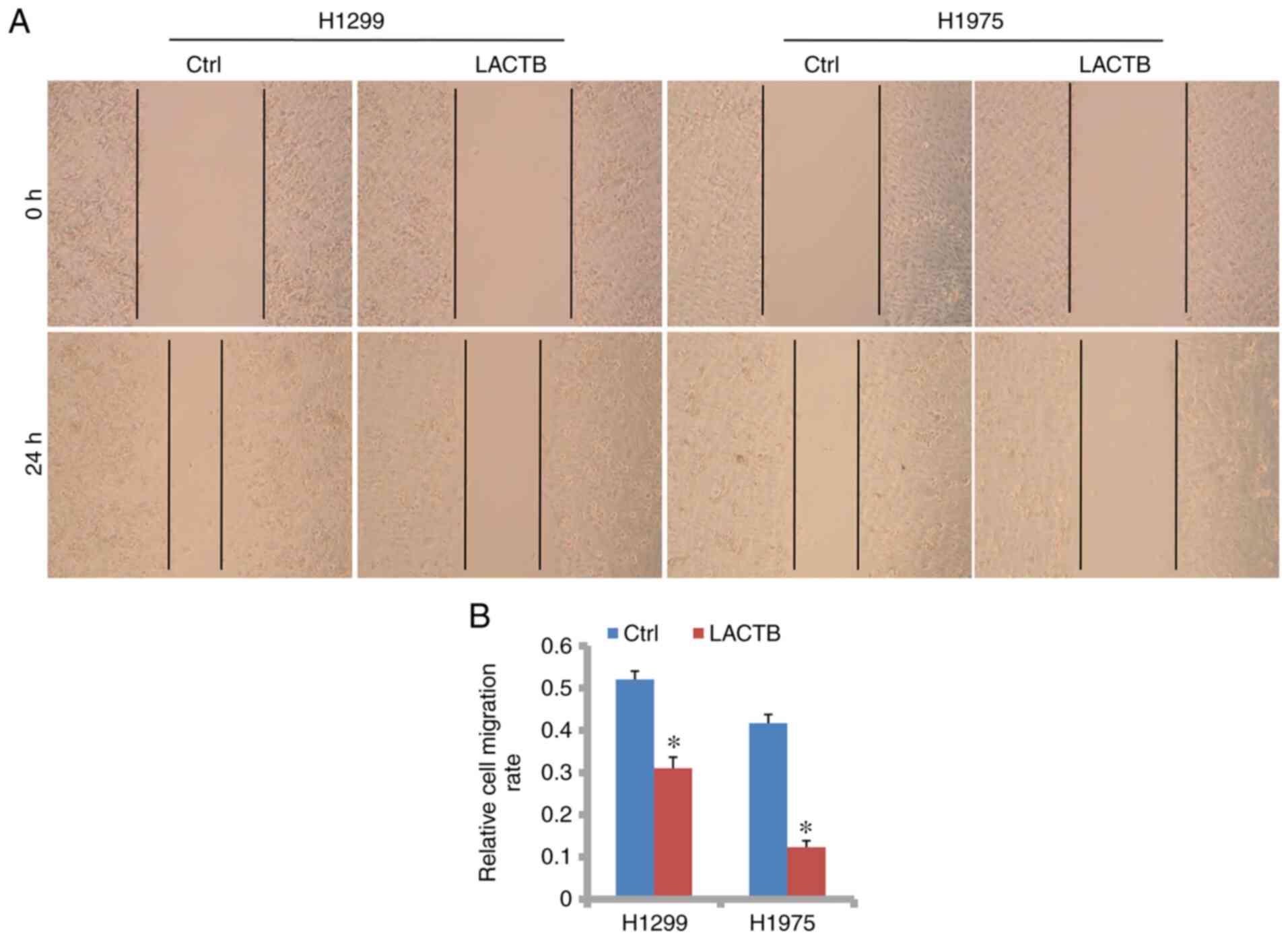
By wound-healing assay, we found that the cell
migration rate was greatly reduced by LACTB overexpression in the
H1299 and H1975 cells. As shown in Fig. 3A and B, the relative cell migration rates were
0.31 and 0.12 after LACTB overexpression in H1299 and H1975 cells,
respectively, which were significantly lower than those of the
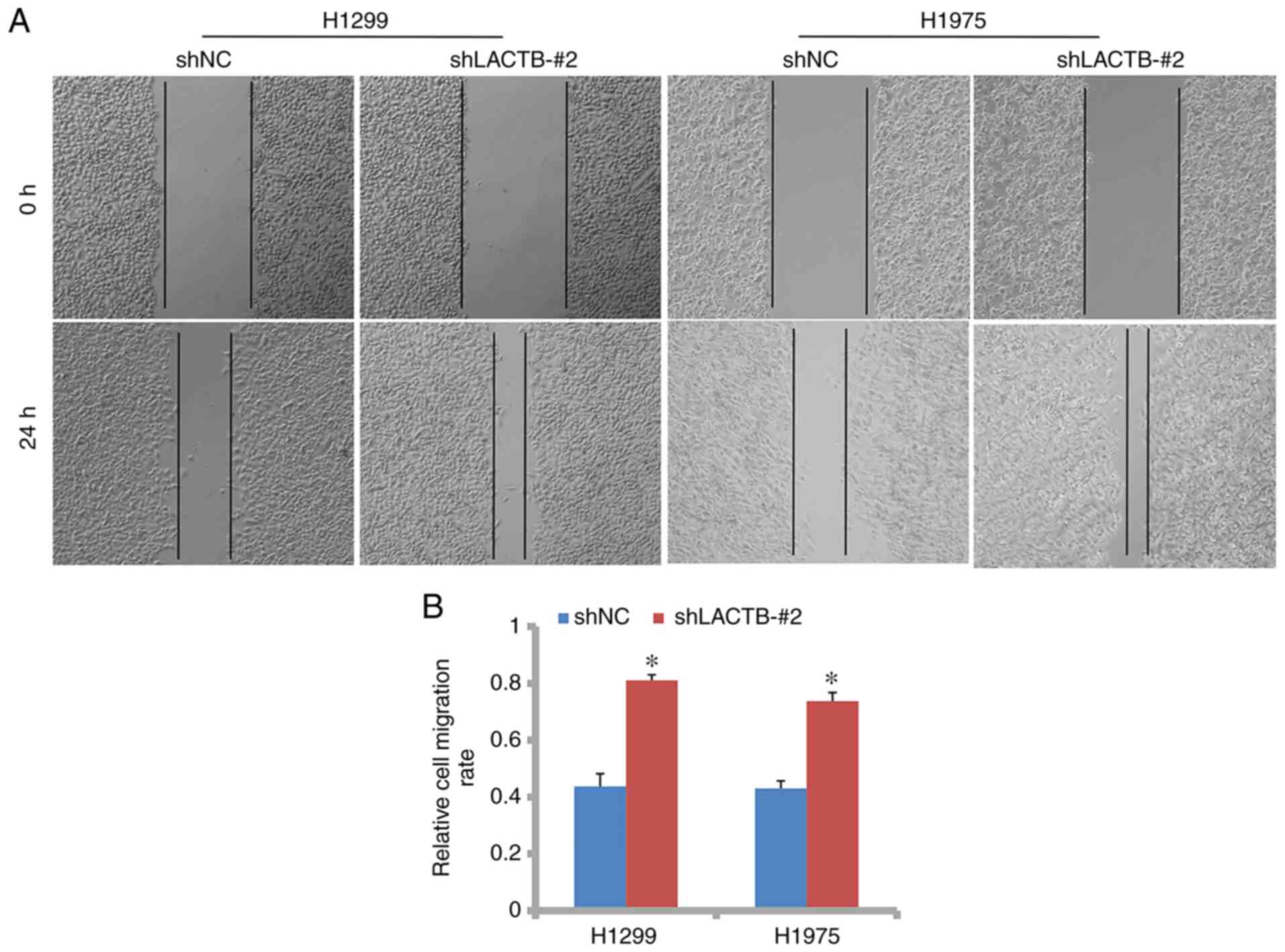
control group. However, in H1299 cells with LACTB knockdown, the
migration rate was 0.81 while it was 0.44 in the NC group (Fig. 4A and B). The same trend was observed in the
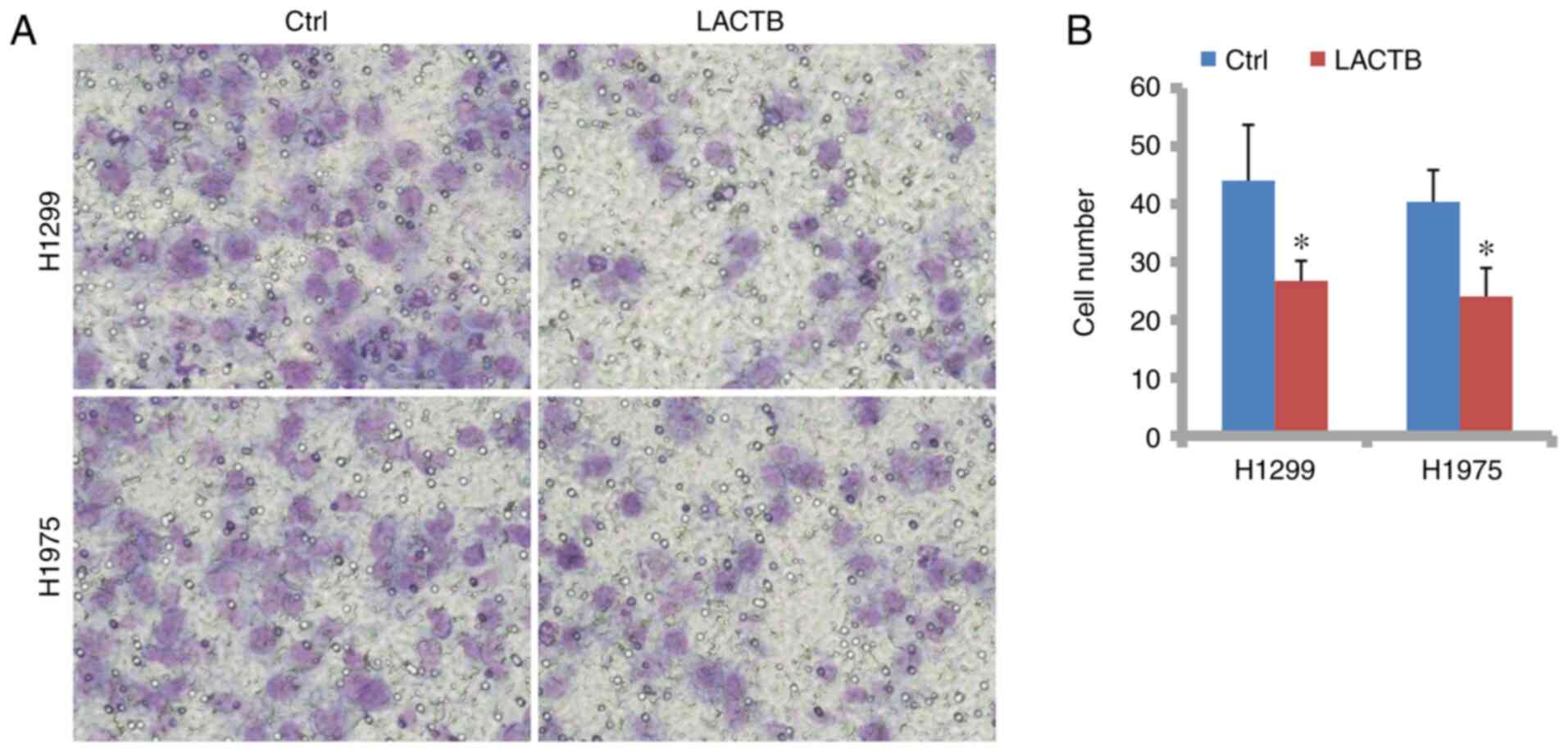
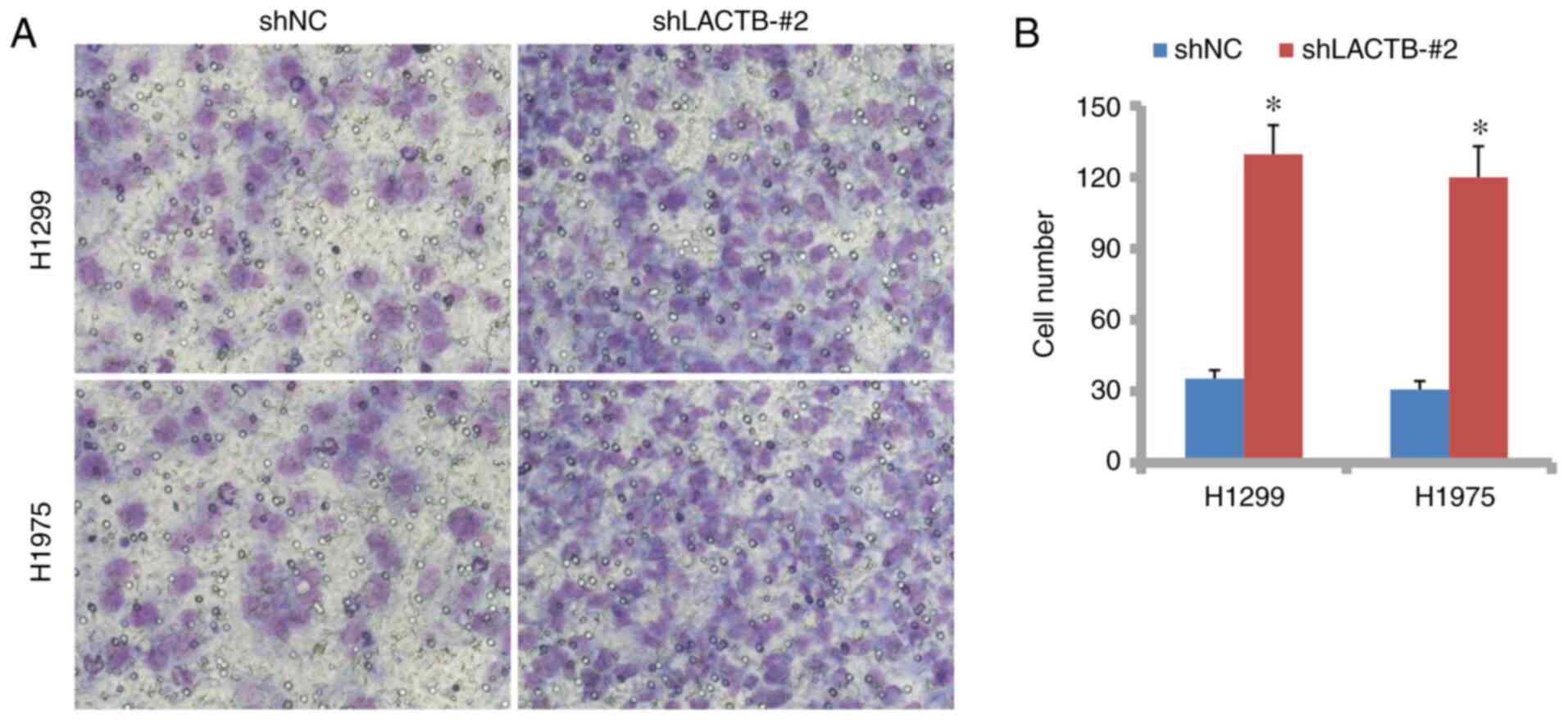
H1975 cells. In Figs. 5A and
B, and 6A and B,
Transwell assays without Matrigel demonstrated that the cell
migration was significantly suppressed by LACTB overexpression but
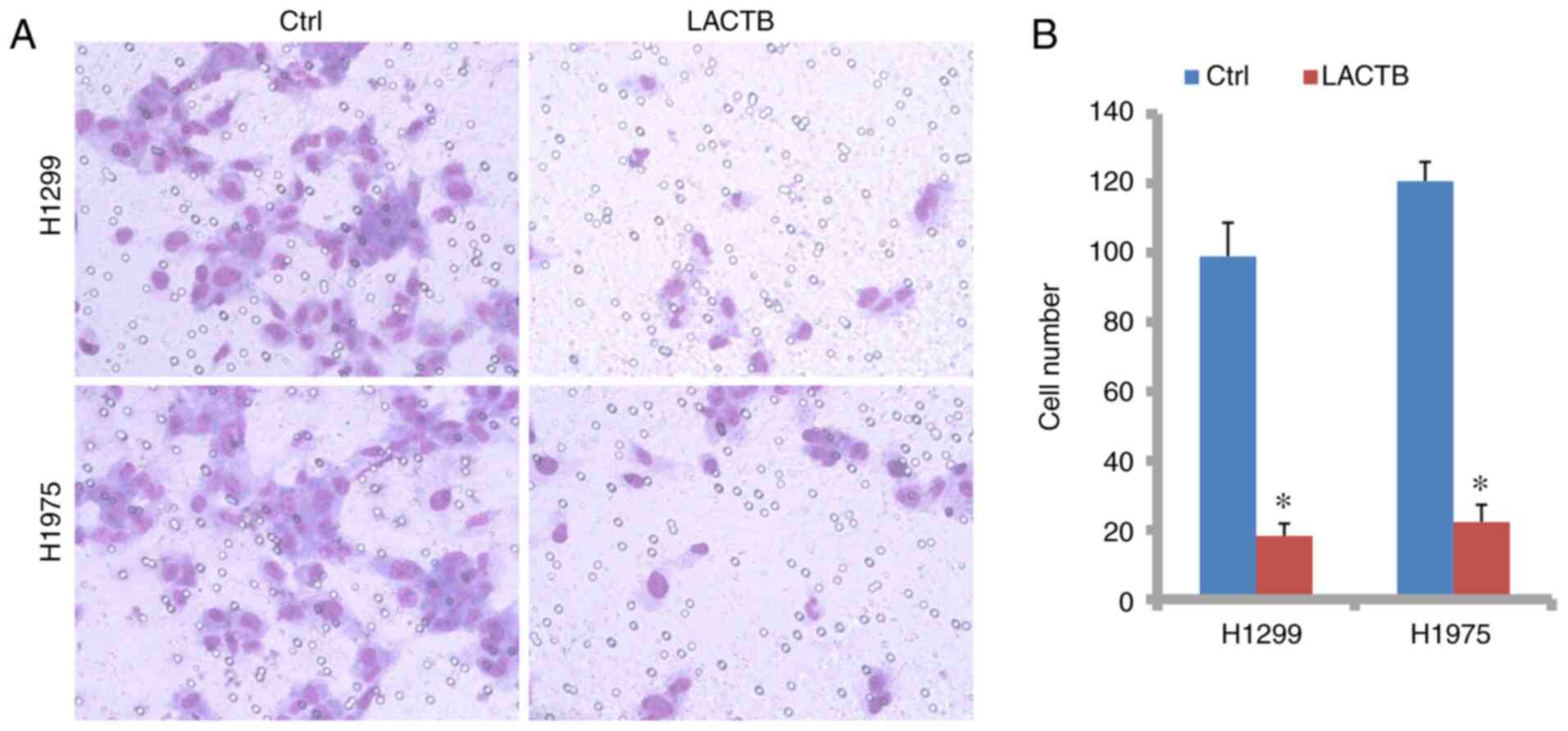
promoted by LACTB knockdown. Then, in the Transwell assay with
Matrigel, there were fewer cells penetrating through the monolayer
membrane in the LACTB overexpression group than in the control
group in H1299 cells (18 vs. 99; Fig.
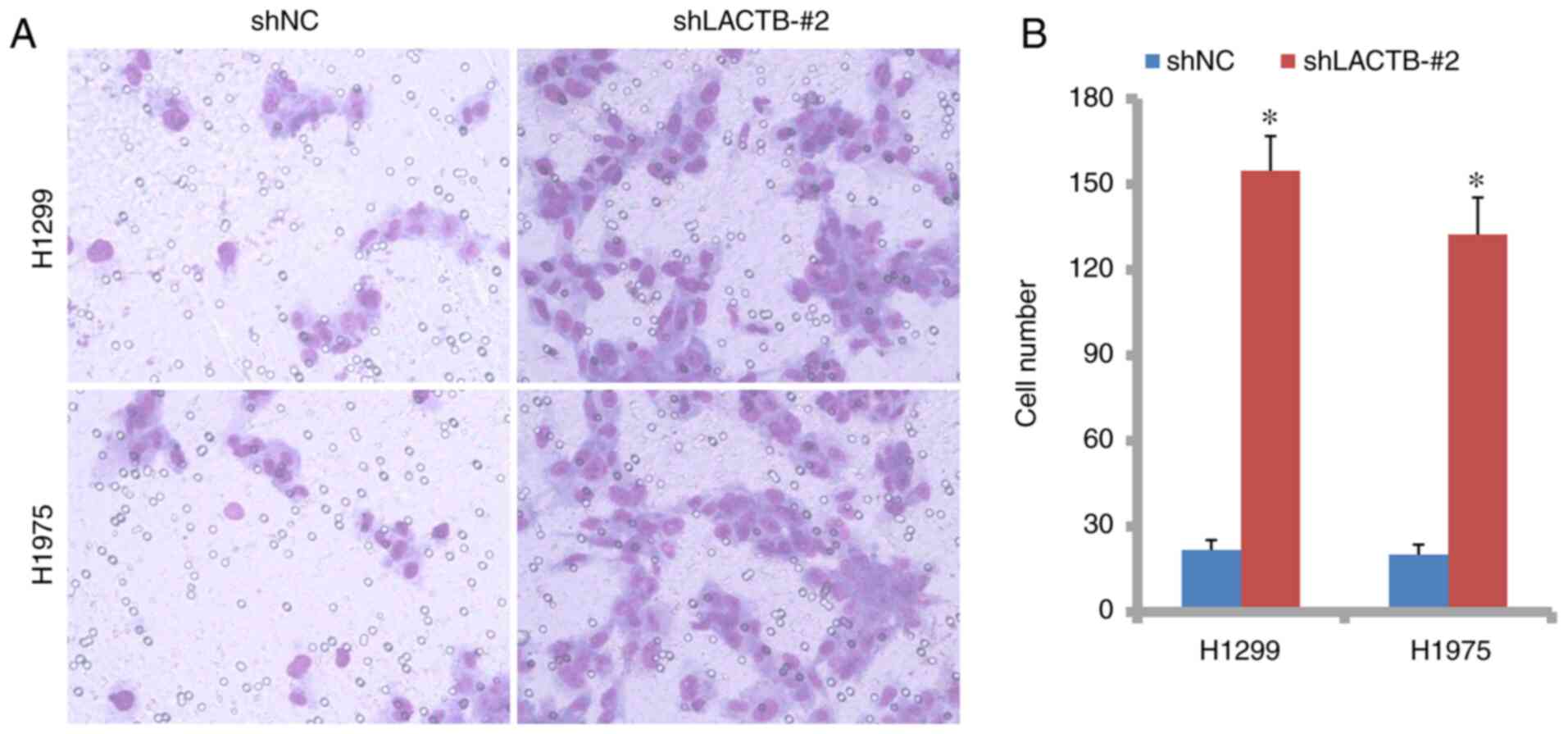
7A and B). However, in
contrast, LACTB knockdown significantly increased the cell number
(Fig. 8A and B). Therefore, LACTB negatively regulates
cell migration and invasion abilities in lung cancer cells.
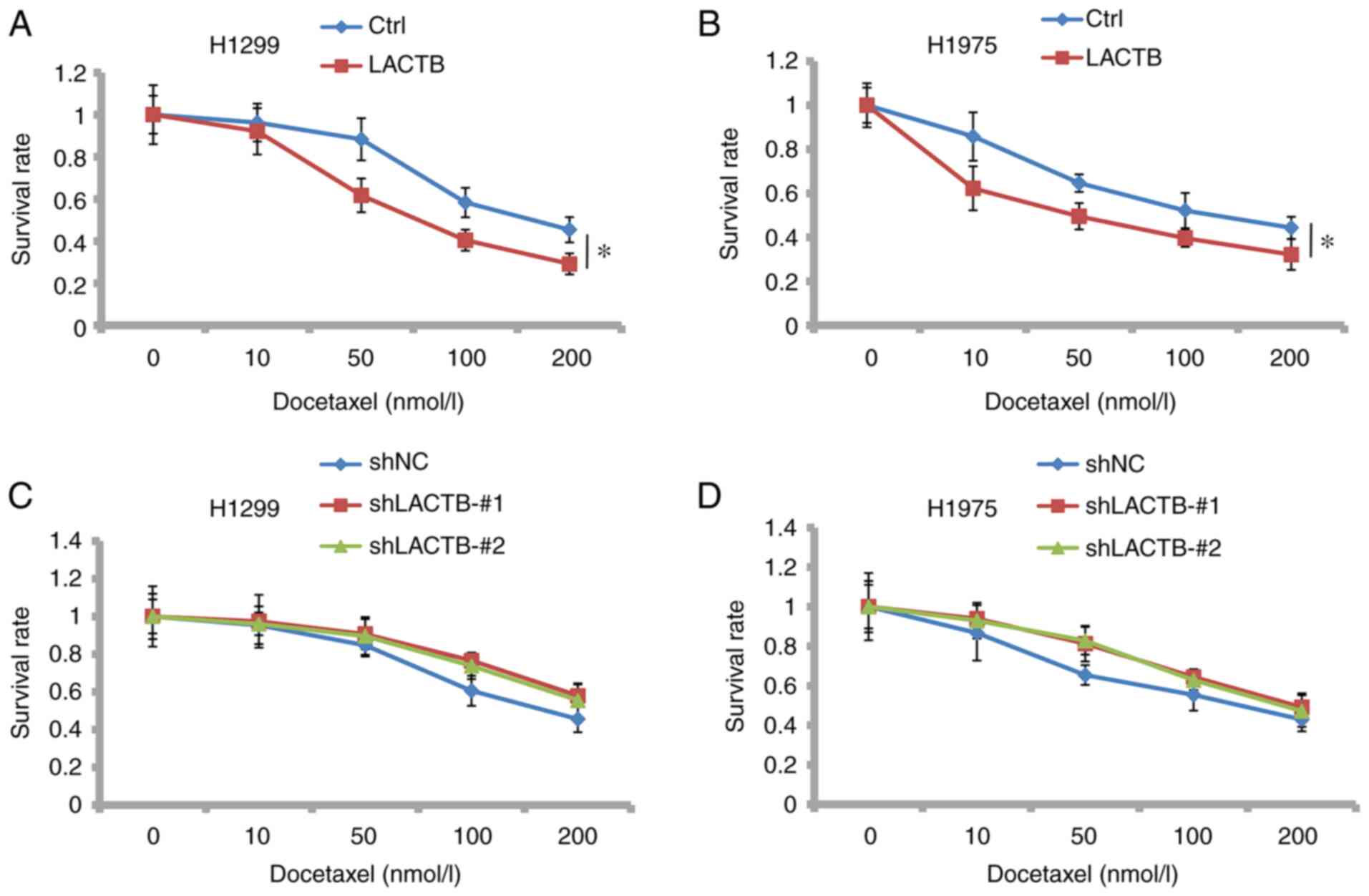
LACTB increases the inhibitory effect
of docetaxel on cell proliferation
The combination of drugs in cancer is often
surprising. Here, we found that LACTB overexpression significantly
inhibited the cell proliferation in H1299 and H1975 cells treated
with docetaxel, which indicates that LACTB has additive effects
with docetaxel (Fig. 9A and
B). In contrast, knockdown of
LACTB greatly reduced the inhibitory ability of docetaxel on lung
cancer cells (Fig. 9C and D). These data suggest that LACTB might be
applied to increase the efficacy of docetaxel in lung cancer
therapy.
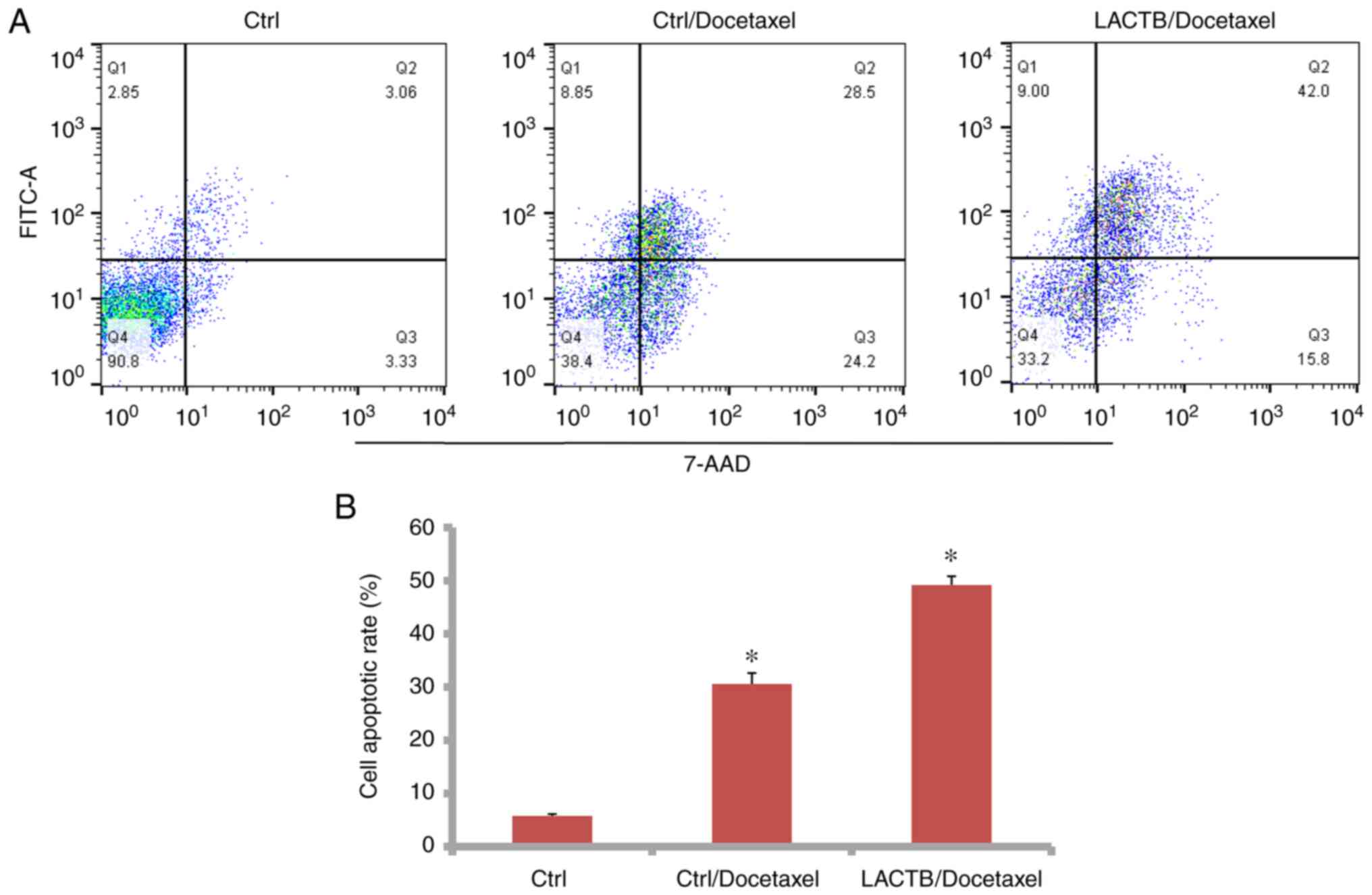
LACTB induces apoptosis of lung cancer
cells induced by docetaxel
Docetaxel induces cell apoptosis in cancer. As shown
in Fig. 10A and B, the cell apoptotic rate was 48.96% in
the H1975 cells treated with the combination of LACTB and docetaxel
while the rate was 33% in the Ctrl/docetaxel group, and the
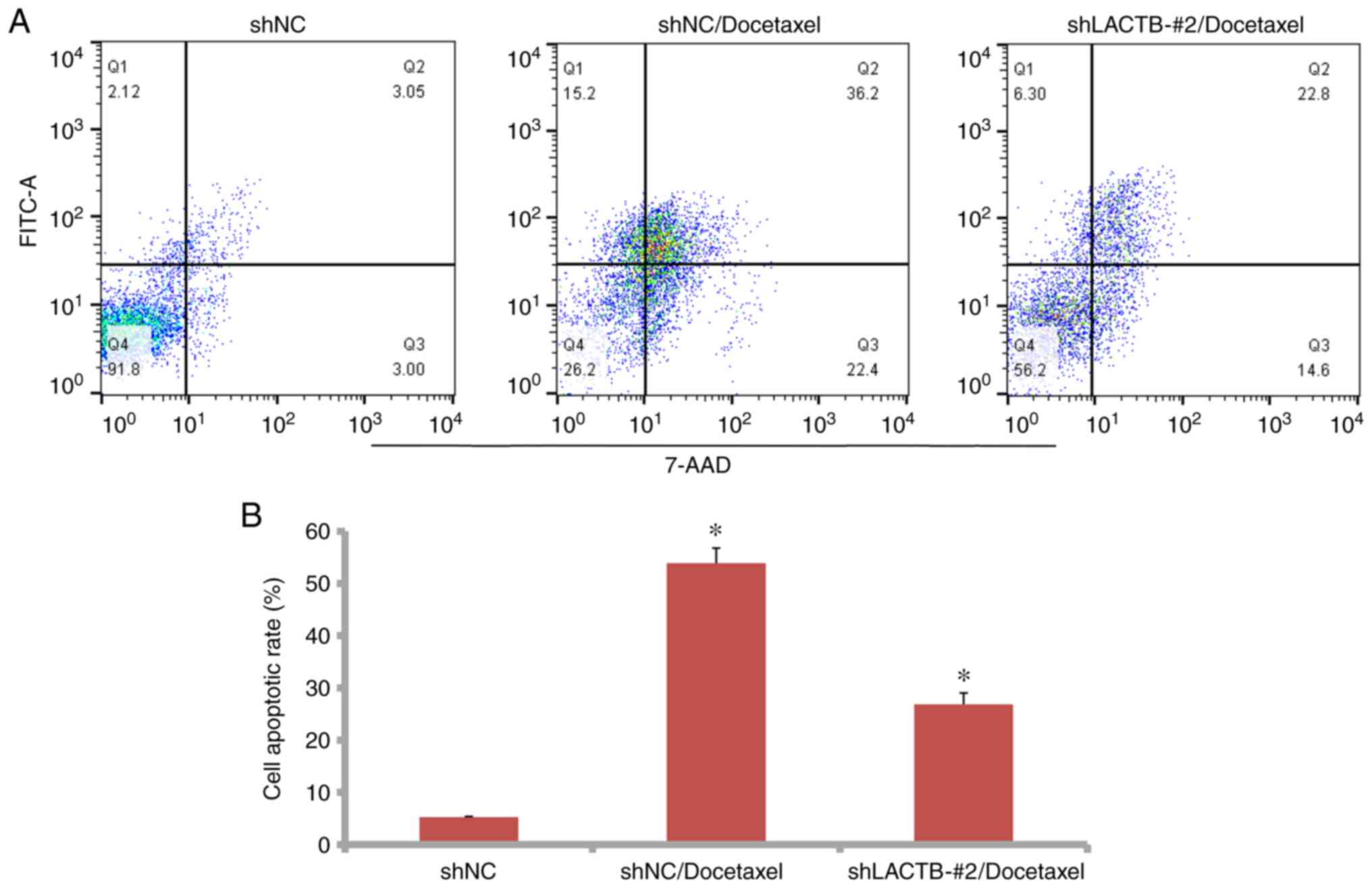
difference was significant. However, LACTB knockdown reduced the
apoptotic rate of docetaxel in the H1975 cells (Fig. 11A and B). The apoptotic rate in the
shNC/docetaxel group was 53.86% while it was 26.84% in the
shLACTB/docetaxel group. In conclusion, LACTB plays an additive
role with docetaxel in the apoptosis of lung cancer cells.
LACTB regulates the expression of
members of the EMT signaling pathway
By western blot analysis, we detected obvious
changes in typical molecules in the EMT process. As shown in
Fig. 12A and B, the expression levels of Snail, matrix
metalloproteinase (MMP2), MMP9, N-cadherin and vimentin were
significantly downregulated while E-cadherin was significantly
increased after LACTB overexpression in H1975 cells. Conversely,
knockdown of LACTB reduced E-cadherin levels while increasing the
levels of Snail, MMP2, MMP9, N-cadherin and vimentin (Fig. 12A and C). This indicates that LACTB negatively
regulates the EMT process in lung cancer.
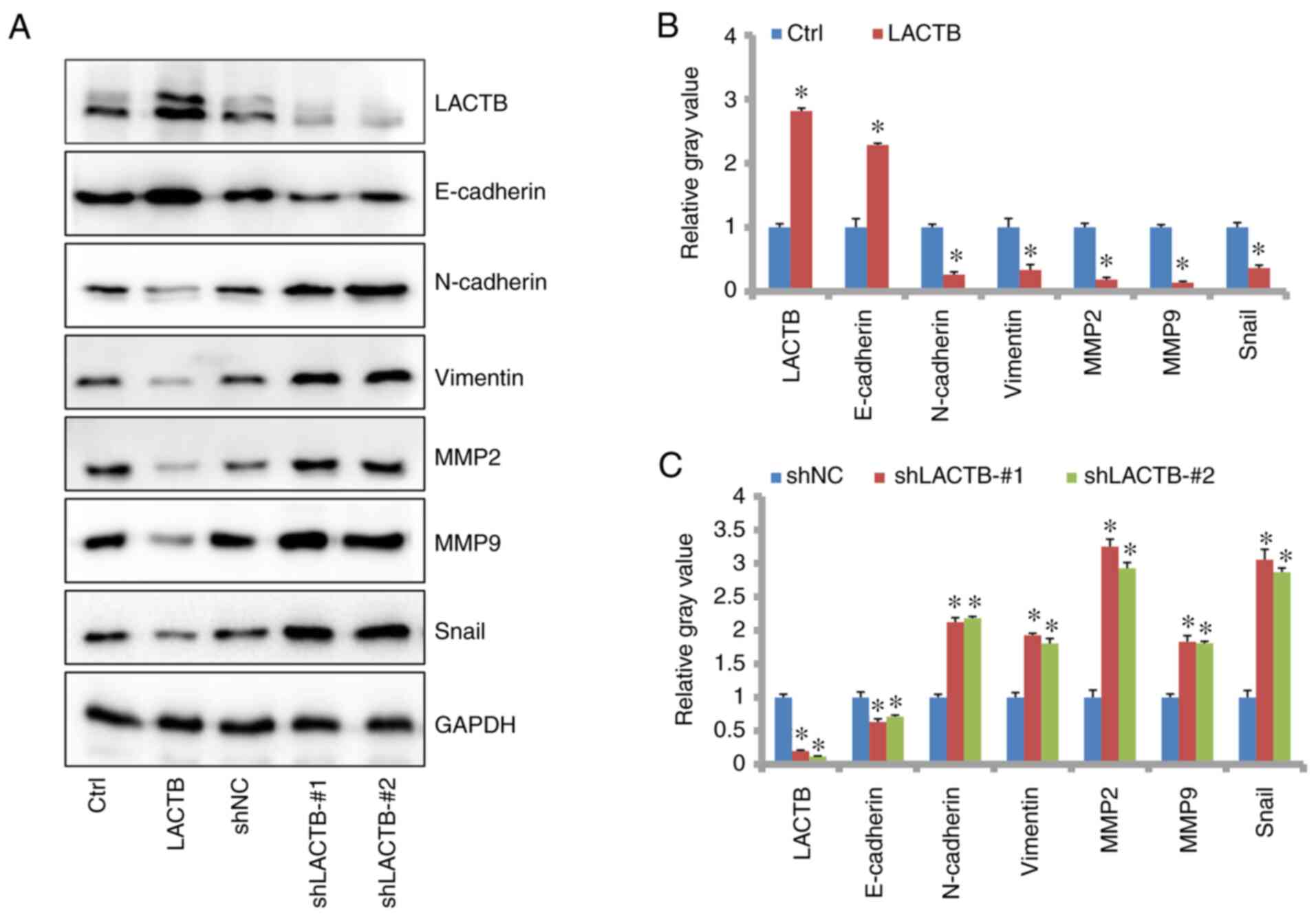 | Figure 12LACTB regulates the EMT signaling
pathway. (A) Overexpression of LACTB promoted E-cadherin expression
and reduced the level of Snail, N-cadherin, vimentin, MMP2 and MMP9
in H1975 cells. By contrast, knockdown of LACTB increased the level
of Snail, N-cadherin, vimentin, MMP2 and MMP9 while reducing
E-cadherin in H1975 cells. (B) Gray value analysis of the protein
bands (on the left) shown in in A. (C) Gray value analysis of the
protein bands (on the right) in (A). *P<0.05,
statistical difference compared to the Ctrl or shNC group. LACTB,
lactamase β; EMT, epithelial-mesenchymal transition; MMP, matrix
metalloproteinase. |
Discussion
Thousands of humans succumb to lung cancer each
year, and overall survival is a major challenge in the clinic
(1,2). This is due to the lack of knowledge
concerning lung cancer including how and when lung cancer begins or
progresses.
Based on TCGA database (19), lactamase β (LACTB) was shown
to be downregulated in lung cancer while the methylation level of
LACTB was enhanced. Methylation is an important mechanism
regulating the switch-on/off of target genes (20). Reduced expression of LACTB in lung
cancer tissues might be due to increased methylation levels of
LACTB. In colorectal cancer, LACTB was also found to be silenced by
elevated methylation of the promoter domain, which promoted
metastasis and advanced clinical stages (10). In the Kaplan-Meier Plotter
database, high LACTB expression was associated with better survival
of lung cancer patients. This is consistent with a study in
hepatocellular cancer that found that low LACTB expression was
associated with poor prognosis (21). These data suggest that LACTB might
also be a suppressor in lung cancer and a potential risk factor
when evaluating the prognosis of lung cancer patients. In the
present study, overexpression of LACTB was found to suppressed cell
proliferation, migration and invasion in lung cancer cells.
However, knockdown of LACTB reversed the inhibitory effect in lung
cancer. Generally, expansion of tumors and distant metastasis are
two typical events observed in cancer patients (22). This study confirmed the suppressor
role of LACTB in lung cancer.
Docetaxel is a specific inhibitor of the M-phase of
the cell cycle and has been successfully applied in the clinic to
treat patients with breast cancer or non-small cell lung cancer.
Docetaxel was demonstrated to increase the ratio of Bax to Bcl-2
followed by the apoptosis response when combined with Stattic in
prostate cancer (23). In another
study, docetaxel enhanced lysosomal function followed by activated
autophagy (24). In the present
study, docetaxel induced apoptosis in lung cancer and this effect
was reinforced by LACTB overexpression. This suggests that synergic
or additive effects between docetaxel and LACTB exist. LACTB might
increase the efficacy of docetaxel or may allow a decrease in the
dosage of docetaxel, which would relieve the adverse effects of
docetaxel. Thus, combination therapy could achieve much better
outcomes and fewer adverse effects. A potential combination therapy
could be the combination of immunotherapy with radiation therapy or
targeted therapy (25,26). Therefore, LACTB has the clinical
potential to increase the efficacy of docetaxel.
EMT is critical in cancer transformation and confers
aggressive ability to cancer cells (27). In lung cancer, activation of EMT
was reported to cause a series of events favorable for cancer
progression. For example, the efficacy of EGFR-targeted therapy in
lung cancer was greatly reduced through the EMT mechanism (28). T-cell infiltration in lung cancer
was negatively regulated by EMT (29). In the present study, LACTB
exhibited potent inhibition of cell migration and invasion in lung
cancer. At the molecular level, LACTB was shown to regulate the
expression levels of E-cadherin, N-cadherin, vimentin and Snail.
E-cadherin is a typical marker of epithelial cells while N-cadherin
and vimentin are typical markers of mesenchymal cells (30). Snail was reported to bind to the
E-box element in the promoter of the E-cadherin gene and suppress
the transcription of E-cadherin (31). Therefore, LACTB suppressed EMT in
lung cancer. The inhibitory role of LACTB on cell migration or
invasion might be partially due to the deficiency of EMT in lung
cancer. EMT was also shown to cause drug resistance to docetaxel in
lung cancer therapy (32). In this
study, LACTB suppressed the EMT process and induced cell apoptosis
in lung cancer. Therefore, it is possible that LACTB may suppress
EMT by inhibiting Snail and increase the sensitivity of lung cancer
cells to docetaxel. The above data suggest that LACTB is a good
candidate for lung cancer therapy. However, further experiments are
necessary before the clinical application of LACTB is realized.
In conclusion, LACTB showed a clinical correlation
with the suppressed progression in lung cancer by negatively
regulating cell migration, invasion and EMT while increasing
apoptosis. LACTB exerts additive effects with docetaxel in the
treatment of lung cancer cells. This study suggests that LACTB is a
new suppressor for lung cancer.
Acknowledgements
Not applicable.
Funding
Funding: This research was supported by grants from the Natural
Science Foundation of Shandong Province (ZR2019PC056), the
Technology Development Program of Jinan City (201907117), and the
Shandong Medical and Health Science and Technology Development Fund
(202102080591).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
HL and YW designed the study and reviewed the
manuscript. YX acquired the majority of the data and drafted the
manuscript. HS, MW and PH analyzed the data and revised the
manuscript. MX and SH confirmed the authenticity of all raw data,
interpreted the data, provided literature support and revised the
manuscript. All authors read and approved the final manuscript for
publication.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Siegel RL, Miller KD, Fuchs HE and Jemal
A: Cancer statistics, 2021. CA Cancer J Clin. 71:7–33.
2021.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Wu F, Wang L and Zhou C: Lung cancer in
China: Current and prospect. Curr Opin Oncol. 33:40–46.
2021.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Hirsch FR, Scagliotti GV, Mulshine JL,
Kwon R, Curran WJ, Yi-Long W and Paz-Ares L: Lung cancer: current
therapies and new targeted treatments. Lancet. 389:299–311.
2017.PubMed/NCBI View Article : Google Scholar
|
|
4
|
de Sousa VM and Carvalho L: Heterogeneity
in lung cancer. Pathobiology. 85:96–107. 2018.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Bai X, Meng L, Sun H, Li Z, Zhang X and
Hua S: MicroRNA-196b inhibits cell growth and metastasis of lung
cancer cells by targeting Runx2. Cell Physiol Biochem. 43:757–767.
2017.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Devlin JR and Verschuren EW: More than a
tumor suppressor: E-Cadherin loss drives lung cancer metastasis. Am
J Respir Cell Mol Biol. 59:141–142. 2018.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Ma Y, Wang L, He F, Yang J, Ding Y, Ge S,
Fan X, Zhou Y, Xu X and Jia R: LACTB suppresses melanoma
progression by attenuating PP1A and YAP interaction. Cancer Lett.
506:67–82. 2021.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Li HT, Dong DY, Liu Q, Xu YQ and Chen L:
Overexpression of LACTB, a mitochondrial protein that inhibits
proliferation and invasion in glioma cells. Oncol Res. 27:423–429.
2019.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Al-Ostoot FH, Salah S, Khamees HA and
Khanum SA: Tumor angiogenesis: Current challenges and therapeutic
opportunities. Cancer Treat Res Commun. 28(100422)2021.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Zeng K, Chen X, Hu X, Liu X, Xu T, Sun H,
Pan Y, He B and Wang S: LACTB, a novel epigenetic silenced tumor
suppressor, inhibits colorectal cancer progression by attenuating
MDM2-mediated p53 ubiquitination and degradation. Oncogene.
37:5534–5551. 2018.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Peng LX, Wang MD, Xie P, Yang JP, Sun R,
Zheng LS, Mei Y, Meng DF, Peng XS, Lang YH, et al: LACTB promotes
metastasis of nasopharyngeal carcinoma via activation of
ERBB3/EGFR-ERK signaling resulting in unfavorable patient survival.
Cancer Lett. 498:165–177. 2021.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Xie J, Peng Y, Chen X, Li Q, Jian B, Wen Z
and Liu S: LACTB mRNA expression is increased in pancreatic
adenocarcinoma and high expression indicates a poor prognosis. PLoS
One. 16(e0245908)2021.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Vasan N, Baselga J and Hyman DM: A view on
drug resistance in cancer. Nature. 575:299–309. 2019.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Yang F, Yan Z, Nie W, Cheng X, Liu Z, Wang
W, Shao C, Fu G and Yu Y: LACTB induced apoptosis of
oxaliplatin-resistant gastric cancer through regulating
autophagy-mediated mitochondrial apoptosis pathway. Am J Transl
Res. 13:601–616. 2021.PubMed/NCBI
|
|
15
|
Dibgre A and Weinberg RA: New insights
into the mechanisms of epithelial-mesenchymal transition and
implications for cancer. Nat Rev Mol Cell Biol. 20:69–84.
2019.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Jiang L, Huang J, Hu Y, Lu P, Luo Q and
Wang L: Gli promotes tumor progression through regulating
epithelial-mesenchymal transition in non-small-cell lung cancer. J
Cardiothorac Surg. 15(18)2020.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Xu W, Yu M, Qin J, Luo Y and Zhong M:
LACTB regulates PIK3R3 to promote autophagy and inhibit EMT and
proliferation through the PI3K/AKT/mTOR signaling pathway in
colorectal cancer. Cancer Manag Res. 12:5181–5200. 2020.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Livak KJ and Schmitten TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Tomczak K, Czerwińska P and Wiznerowicz M:
The cancer genome atlas (TCGA): An immeasurable source of
knowledge. Contemp Oncol (Pozn). 19:A68–A77. 2015.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Heberle E and Bardet AF: Sensitivity of
transcription factors to DNA methylation. Essays Biochem.
63:727–741. 2019.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Xue C, He Y, Zhu W, Chen X, Yu Y, Hu Q,
Chen J, Liu L, Ren F, Ren Z, et al: Low expression of LACTB
promotes tumor progression and predicts poor prognosis in
hepatocellular carcinoma. Am J Transl Res. 10:4152–4162.
2018.PubMed/NCBI
|
|
22
|
Senga SS and Grose RP: Hallmarks of
cancer-the new testament. Open Biol. 11(200358)2021.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Mohammadian J, Sabzichi M, Molavi O,
Shanehbandi D and Samadi N: Combined treatment with stattic and
docetaxel alters the Bax/Bcl-2 gene expression ratio in human
prostate cancer cells. Asian Pac J Cancer Prev. 17:5031–5035.
2016.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Zhang J, Wang J, Wong YK, Sun X, Chen Y,
Wang L, Yang L, Lu L, Shen HM and Huang D: Docetaxel enhances
lysosomal function through TFEB activation. Cell Death Dis.
9(614)2018.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Colli LM, Machiela MJ, Zhang H, Myers TA,
Jessop L, Delattre O, Yu K and Chanock SJ: Landscape of combination
immunotherapy and targeted therapy to improve cancer management.
Cancer Res. 77:3666–3671. 2017.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Asna N, Livoff A, Batash R, Debbi R,
Schaffer P, Rivkind T and Schaffer M: Radiation therapy and
immunotherapy-a potential combination in cancer treatment. Curr
Oncol. 25:e454–e460. 2018.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Pastushenko I and Blanpain C: EMT
transition states during tumor progression and metastasis. Trends
Cell Biol. 29:212–226. 2019.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Tulchinsky E, Demidov O, Kriajeska M,
Barlev NA and Imyanitov E: EMT: A mechanism for escape from
EGFR-targeted therapy in lung cancer. Biochim Biophys Acta Rev
Cancer. 1871:29–39. 2019.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Chae YK, Chang S, Ko T, Anker J, Agte S,
Iams W, Choi WM, Lee K and Cruz M: Epithelial-mesenchymal
transition (EMT) signature is inversely associated with T-cell
infiltration in non-small cell lung cancer (NSCLC). Sci Rep.
8(2918)2018.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Ribatti D, Tamma R and Annese T:
Epithelial-mesenchymal transition in cancer: A historical overview.
Transl Oncol. 13(100773)2020.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Villarejo A, Cortés-Cabrera A,
Molina-Ortiz P, Portillo F and Cano A: Differential role of Snail1
and Snail2 zinc fingers in E-cadherin repression and epithelial to
mesenchymal transition. J Biol Chem. 289:930–941. 2014.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Shen W, Pang H, Liu J, Zhou J, Zhang F and
Liu L, Ma N, Zhang N, Zhang H and Liu L: Epithelial-mesenchymal
transition contributes to docetaxel resistance in human non-small
cell lung cancer. Oncol Res. 22:47–55. 2014.PubMed/NCBI View Article : Google Scholar
|