Introduction
Postmenopausal osteoporosis (PMO) is an
aging-associated disease that manifests as degradation of bone
tissue microstructure leading to decreased bone mass and increased
bone fragility (1). Decreased
estrogen levels are the primary factor for onset of PMO (2). According to a previous study,
approximately one-third of women aged 60-70 years have from
osteoporosis worldwide and nearly one-third of women >50 years
of age develop osteoporotic fractures (3). Therefore, understanding the
pathogenesis of PMO is key for its prevention and treatment.
The primary pathogenic feature of PMO is imbalance
between bone resorption and formation, wherein the rate of bone
resorption by osteoclasts exceeds the rate of bone formation by
osteoblasts (4). Peripheral blood
mononuclear cells (PBMCs) directly participate in osteoclast
formation. PBMCs are precursors of osteoclasts (5) and secrete osteoclast-inducing factors
such as interleukin-1 (IL-1), IL-6 and tumor necrosis factor-α
(TNF-α) (6). Human PBMCs express
genes associated with osteoporosis, including annexin A1, S100
calcium-binding protein A4 and transmembrane protein 64(7). The discovery of such genes provides
novel clues to understand the pathogenesis of PMO.
To identify novel and hub genes of PMO, the present
study performed joint bioinformatics analysis including screening
of differentially expressed genes (DEGs) from Gene Expression
Omnibus (GEO) dataset and identification of targets of
bisphosphonates (a bone resorption inhibitor widely used in the
clinic) (8) from the STITCH
database. In vitro experiments were performed to verify the
role of hub genes in differentiation of mononuclear macrophage into
osteoclasts.
Materials and methods
Cell culture and induced
differentiation of osteoclasts
The human monocyte cell line THP-1 was purchased
from Santa Cruz Biotechnology, Inc. (cat. no. sc-7274, USA) and
maintained in RPMI-1640 medium (HyClone; Cytiva) supplemented with
10% FBS (Gibco; Thermo Fisher Scientific, Inc.). Cells were
cultured in an incubator under 5% CO2 at 37˚C. As
described previously (9), THP-1
cells were seeded in a 24-well plate (1x106 cells/well).
The next day, cells were treated with 1,000 units of
macrophage-colony-stimulating factor (M-CSF; R&D Systems,
Inc.), 5 ng/ml phorbol 12-myristate 13-acetate (LGC Standards Ltd.)
and 50 ng/ml soluble receptor activator of NF-κB ligand (sRANKL;
MedChemExpress). At 3 and 5 days, the formation of osteoclasts was
confirmed by morphological determination of coenocytes and
tartrate-resistant acid phosphatase staining (TRAP staining). TRAP
staining was performed according to the manufacturer's protocol
(Wuhan Servicebio Technology Co., Ltd.). Briefly, after induced
differentiation into osteoclast, the THP-1 cells were fixed using
4% paraformaldehyde for 20 min at room temperature. After washing,
THP-1 cells were treated with TRAP dyeing liquor (Wuhan Servicebio
Technology Co., Ltd.) for 2 h at room temperature. Next, the TRAP
dyeing liquor was removed and THP-1 cells were washed, then
hematoxylin dye was used for nuclear staining for 15 sec at room
temperature. Finally, the nuclear fusions were observed under a
light microscope (Nikon Eclipse E100; Nikon Corporation) and the
fusion rate of multinuclear cells that indicates the formation of
osteoclast was quantified by ImageJ software (version 1.8.0;
National Institutes of Health).
Small interfering (si)RNA synthesis
and transfection
siRNAs targeting AKT3 and RAC1 were
designed and synthesized by Jiman Biotechnology (Shanghai) Co.,
Ltd. with the following target sequences: siRNA-AKT3,
5'-CAGCAGGCACGUUAACUCGAA-3' and siRNA-RAC1,
5'-AACCUUUGUACGCUUUGCUCA-3'. All sequences of siRNAs are presented
in Table SI. Negative control
(NC) siRNAs with no homology to siRNA-AKT3 and siRNA-RAC1 were
designed and synthesized by Jiman Biotechnology (Shanghai) Co.,
Ltd. as follows: siRNA-NC, 5'-UUCUCCGAACGUGUCACGU-3'. siRNA
transfection was performed using Lipofectamine® 2000
(Invitrogen: Thermo Fisher Scientific, Inc.) according to the
manufacturer's instructions. Briefly, THP-1 cells in the
logarithmic growth phase were seeded at 5x104 cells/well
in a 24-well plate and cultured at 37˚C for 24 h. The
RPMI-1640 medium (HyClone; Cytiva) was replaced with serum-free
Opti-MEM (Thermo Fisher Scientific, Inc.) and cells were
transfected at 37˚C with lipofectamine 2000 (1:100) and 100 nmol/l
siRNA for 20 min for fluorescence-siRNA-transfection reagent
mixture formation. Subsequently, serum-free Opti-MEM was added to a
total volume of 500 µl, and THP-1 cells were cultured in an
incubator for 4-6 h. Finally, the transfection efficiency
was assessed by detecting expression of objective proteins by
western blotting. Cells treated with liposomes (cell:liposome,
1:100) and siRNA-NC at 37˚C were used as NC. At 6 h
post-transfection, the medium was replaced with RPMI-1640 (HyClone;
Cytiva) with 10% FBS (Gibco; Thermo Fisher Scientific, Inc.) and
cells were cultured at 37˚C for 48 h before being harvested for
western blotting.
Apoptosis analysis
Annexin V-FITC/PI Cell Apoptosis Detection kit
(TransGen Biotech Co., Ltd.) was used to detect apoptotic cells.
The apoptosis rate was calculated as the early apoptosis rate
(lower right quadrant) plus the late apoptosis rate (upper right
quadrant). Following bisphosphonate (10 µM) treatments for 24 h at
room temperature, THP-1 cells were detached by EDTA-free trypsin
digestion and collected by centrifugation at 4˚C at 201 x g for 5
min. The cells were re-suspended in PBS and collected by
centrifugation at 4˚C at 201 x g for 5 min. The cells were
re-suspended in 100 µl binding buffer, mixed with 5 µl Annexin
V-FITC reagent, and placed in the dark at room temperature for 15
min. Cell suspension was mixed with 5 µl PI, incubated at room
temperature for 5 min and subjected to flow cytometry analysis. The
supplier of flow cytometer (CytoFLEX S) was Beckman Coulter, Inc.,
and the analysis software was CytExpert (version 2.0; Beckman
Coulter, Inc.).
Cell viability assay
THP-1 cells were plated in a 96-well plate (8,000
cells/well) and treated with pamidronate at various concentrations
(1, 2.5, 5, 10, 20, 50 and 100 µM) at 37˚C for 24 h. The cells were
washed with PBS and incubated with 100 µl RPMI-1640 medium
(HyClone; Cytiva) containing CCK-8 reagent (Dongren Chemical
Technology) for 1-4 h. Cell viability was measured by absorbance at
450 nm using a microplate reader.
Reverse transcription-quantitative
(RT-q)PCR
Total RNA of cells was isolated using a Total RNA
Extraction kit (Beijing Solarbio Science & Technology Co.,
Ltd.) and 100 ng RNA was used for RT-qPCR assay using a Two-Step
qPCR SuperMix (Beijing Transgen Biotech Co., Ltd.). The procedure
of reverse transcription was as follows: A temperature program of
5-min priming at 25˚C followed by reverse transcription at 42˚C for
30 min and reverse transcription inactivation at 85˚C for 5 min was
run. After a final cool-down to 4˚C, the cDNA was stored at -80˚C
for subsequent use. The amplification conditions were as follows:
Pre-denaturation at 95˚C for 10 min, followed by 40 cycles of
denaturation at 95˚C for 30 sec, annealing at 56˚C for 30 sec and
extension at 72˚C for 45 sec. The qPCR data were analyzed by the
ΔΔCq method (10). GAPDH was used as an
internal reference gene as previously described (11). The
sequence information of the primers is provided in Table SII.
Retrieval of PMO gene expression
profiles from GEO database
The raw data of PMO gene expression profiles were
downloaded from National Center for Biotechnology Information-GEO
database (ncbi.nlm.nih.gov). The selection
criteria were as follows: Samples were from peripheral blood and
the object of study was PBMCs. According to these criteria, one
dataset (GEO accession no. GSE56815; ncbi.nlm.nih.gov/gds/?term=GSE56815) containing gene
expression profiles of PBMCs from a cohort of 80 females was
selected. The cohort included 40 cases each of pre- and
post-menopausal patients, with each group containing 20 cases each
of low and high bone mineral density (BMD). The gene expression
profiling of this dataset was determined using the Affymetrix
HG-133A array platform (Affymetrix; Thermo Fisher Scientific,
Inc.). A heatmap was generated to indicate hierarchical clustering
with the R language ‘pheatmap’ package (R version 4.1.2).
Prediction of bisphosphonates
targets
Using the STITCH database (stitch.embl.de/cgi), predicted bisphosphonates targets
were retrieved and a protein-protein interaction (PPI) network was
constructed. Functional enrichment, including biological processes
and KEGG pathways, of the network were analyzed using the STITCH
database. The results were plotted using the ggplot2 2.2.1 package
in RStudio (version 4.1.2) (12).
Identification of shared KEGG pathways
between bisphosphonates targets and DEGs in PMO
The enriched KEGG pathways of the
bisphosphonates-protein network were produced using STITCH
database. The enrichment analysis of KEGG pathways of DEGs in the
PMO dataset was conducted using Database for Annotation,
Visualization and Integrated Discovery Bioinformatics Resources 6.8
(13). The shared KEGG pathways between the two analyses were
obtained and displayed using Venn Diagram (version 2.1; ehbio.com/ImageGP/index.php/Home/Index/index.html).
Identification of hub genes
Cytoscape (version 3.6.0; https://cytoscape.org/) is a powerful visualization
software, including add-on modules for protein interaction visual
analysis, calculation and analysis of interaction network. The
MCODE module in Cytoscape was used to weigh the connection between
nodes. For each node in the interaction network of selected genes,
two indices were selected to calculate topological features.
‘Degree’ was defined as the number of edges to node i; ‘closeness’
was the inverse of the sum of the distance from node i to other
nodes. When applying the degree and closeness algorithm, proteins
with degree >20 and closeness >48.8 were considered to be
major hubs. Both the hub gene and network were retained,
calculation data were downloaded, and the above indicators (degree
and closeness algorithm) were sorted as previously described
(14). Finally, the genes involved
in the four common KEGG pathways, including ‘pathways in cancer’,
‘HIF-1 signaling pathway’, ‘human T-cell leukemia virus 1
infection’ and ‘viral carcinogenesis’, were filtered out and 10
genes that met the requirements (degree >20 and closeness
>48.8) were selected.
Western blotting
The THP-1 cells were collected on ice and lysed with
RIPA buffer (Roche Diagnostics) supplemented with protease
inhibitor (Complete Mini Tablets; Roche Diagnostics) for 30 min at
4˚C with shaking. The cell lysate was centrifuged at 4˚C at 16,770
x g for 15 min and supernatant was collected. Protein was
quantified using the BCA Protein Concentration Determination Kit
(cat. no. P0012S; Beyotime Institute of Biotechnology). Equal
amounts of protein (5 µg/lane) samples were mixed with
loading buffer, denatured at 100˚C for 10 min and separated by
SDS-PAGE (10%). The proteins were transferred onto PVDF membranes
(Invitrogen; Thermo Fisher Scientific, Inc.) and blocked at room
temperature with 5% non-fat dry milk for 1 h. The membranes were
incubated overnight at 4˚C in solution containing primary
antibodies (all Abcam) as follows: Anti-Akt3 (1:1,000; cat. no.
ab152157), anti-Rac1 (1:1,000; cat. no. ab155938) and anti-GAPDH
(1:2,000; cat. no. ab128915). Subsequently, blots were incubated
with horseradish peroxidase-conjugated secondary antibody (1:3,000;
cat. no. ab288151; Abcam) for 1 h at room temperature. Protein
expression was detected with SuperSignal West Pico (Thermo Fisher
Scientific, Inc.) using the ChemiDoc MP imaging system (Bio-Rad
Laboratories, Inc.). Protein densitometry was quantified using
Image Lab software (version 5.2.1; Bio-Rad Laboratories, Inc.).
Statistical analysis
GraphPad Prism 8.0 software (GraphPad Software,
Inc.) was used for all statistical analysis. Normality test was
performed before sample comparison (Shapiro-Wilk or D'Agostino and
Pearson normality test). One-way analysis of variance followed by
Tukey's post hoc test was used to compare ≥3 groups. Data are
presented as the mean ± standard deviation. Every experiment was
repeated three times. P<0.05 was considered to indicate a
statistically significant difference (15).
Results
A total of 290 DEGs were screened from
the GEO dataset in PMO
Based on the aforementioned selection criteria, the
GEO dataset GSE56815 was selected and PBMC expression profiles from
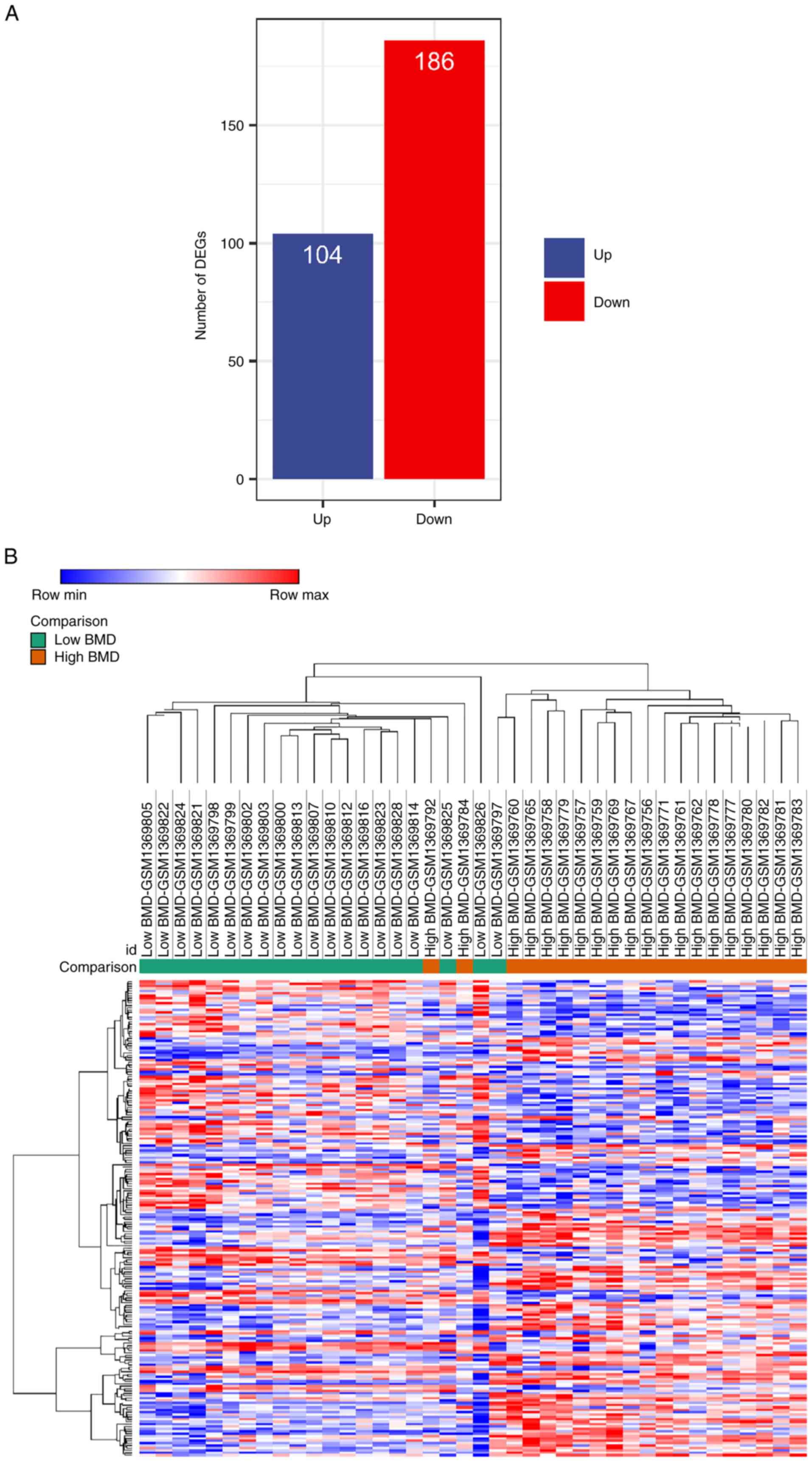
postmenopausal patients were extracted for DEG analysis. Between
patients with high and low BMD, there were 290 DEGs, of which 104
genes were up- and 186 were downregulated in the low BMD group
(Fig. 1A). The heatmap combined
with the hierarchical clustering of DEGs was shown in Fig. 1B. The results demonstrated that
most samples were discriminatively clustered as low BMD/high BMD.
Overall, the gene profile of the low BMD group was markedly
different from that in the high BMD group.
Functional annotation of DEGs in
PMO
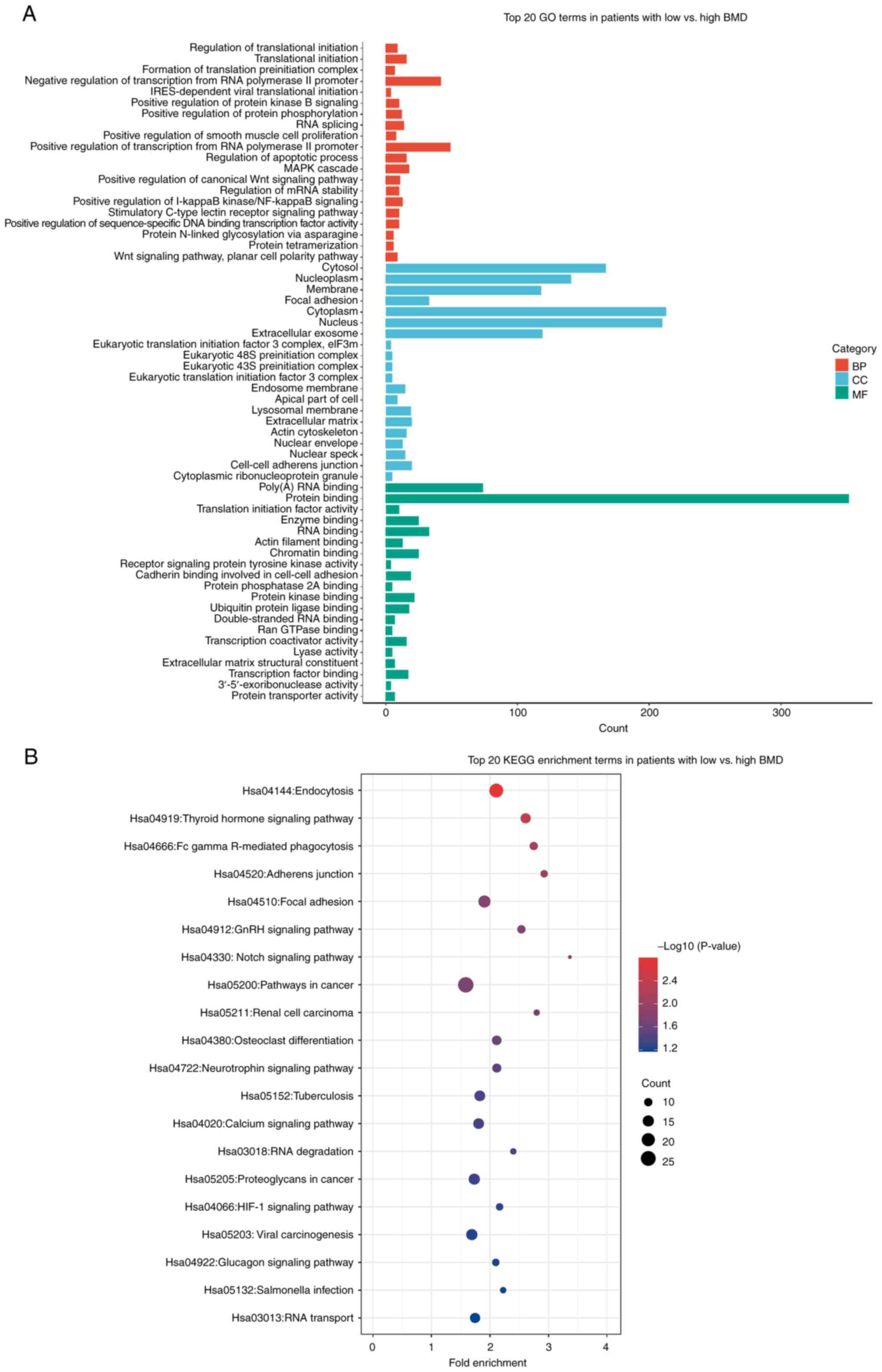
The selected 290 DEGs were subjected to functional
annotation. GO enrichment analysis of the biological process showed
that DEGs were primarily enriched for ‘positive/negative regulation
of transcription from RNA polymerase II’, ‘translational
initiation’ and ‘MAPK cascade’. The cellular components were
associated with ‘cytoplasm’ and molecular functions were associated
with ‘protein binding’ and ‘Poly(A) RNA binding’. These data
demonstrated that PMO was associated with genes involved in RNA
synthesis and regulation (Fig.
2A). In KEGG pathway analysis, the enriched pathways included
‘endocytosis’ and ‘thyroid hormone signaling’, indicating a
connection between hormonal regulation and PMO (Fig. 2B). Overall, the functional
annotation of DEGs in PMO indicated that most DEGs were closely
associated with the process of DNA reproduction.
Bisphosphonates target analysis using
the STITCH database and the interaction network
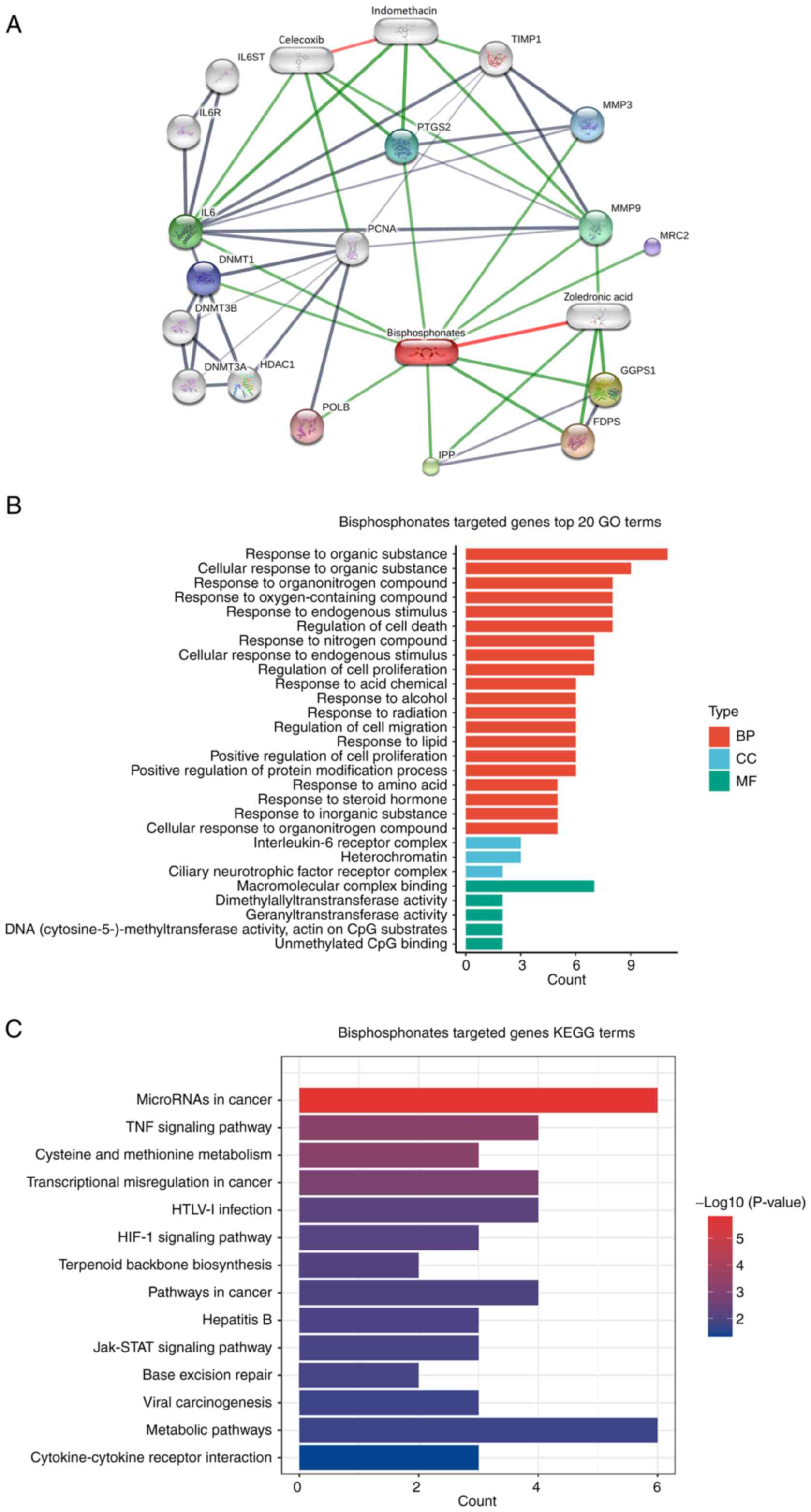
Using the STITCH database, functional partners of
bisphosphonate were analyzed, which resulted in 16 candidates. The
first cluster included 10 proteins, namely farnesyl diphosphate
synthase, geranylgeranyl diphosphate synthase 1, intracisternal A
particle-promoted polypeptide, IL6, MMP9,
prostaglandin-endoperoxide synthase 2, MMP3, DNA methyltransferase
1, mannose receptor C type 2 and DNA polymerase β. The second
cluster included IL-6ST, IL-6R, DNA methyltransferase 3α, DNA
methyltransferase 3β, histone deacetylase 1 and proliferating cell
nuclear antigen (Fig. 3A). The
enrichment analysis showed that these molecules were associated
with biological processes of ‘response to organic substance’ and
‘cellular response to organic substance’, cellular components were
associated with ‘IL-6 receptor complex’ and ‘heterochromatin’ and
molecular functions were associated with ‘macromolecular complex
binding’ (Fig. 3B). In addition,
these proteins were involved in KEGG pathways of ‘TNF-α signaling’,
‘cysteine metabolism’ and ‘methionine metabolism’ (Fig. 3C). Overall, these results indicated
a relationship between bisphosphonates and immunoregulation.
Identification of shared KEGG pathways
between bisphosphonate targets and DEGs in PMO
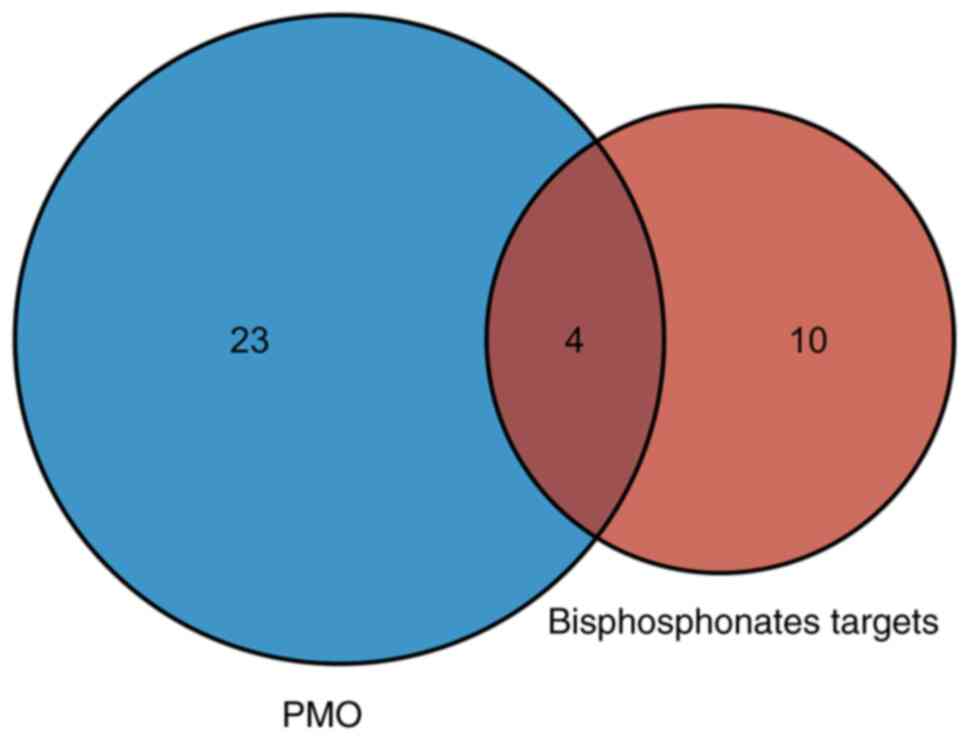
Next, common KEGG pathways between DEGs in PMO and
bisphosphonate partners were intersected, which yielded four
pathways (Fig. 4A), including
‘pathways in cancer’, ‘HIF-1 signaling pathway’, ‘human T-cell
leukemia virus 1 infection’ and ‘viral carcinogenesis’ (Fig. S1). A total of 42 DEGs in PMO were
involved in these common KEGG pathways (Table I). The shared pathways mostly
focused on cancer pathways, which suggested that PMO was related to
proliferation-related signaling pathways.
 | Table IShared Kyoto Encyclopedia of Genes
and Genomes pathway and involved genes. |
Table I
Shared Kyoto Encyclopedia of Genes
and Genomes pathway and involved genes.
| Term | P-value | Gene |
|---|
| Pathways in
cancer | 0.0189 | FH, RALB, FLT3,
LAMA4, FOXO1, EGFR, CDC42, CCND1, AKT3, DVL1, TCEB2, EP300, VHL,
RAC1, WNT1, PRKACA, APPL1, JUN, DAPK2, FLT3LG, MITF, NFKB2, BMP4,
NFKBIA, PLCB3, RARA, CTNNB1 |
| HIF-1 signaling
pathway | 0.0548 | CAMK2B, ANGPT1,
INSR, AKT3, EP300, HMOX1, TCEB2, VHL, EGFR, CAMK2B, ANGPT1, INSR,
AKT3, EP300, HMOX1, TCEB2, VHL, EGFR, |
| Viral
carcinogenesis | 0.0594 | JUN, HIST1H2BO,
SYK, ACTN2, ACTN1, RBPJ, NFKB2, CDC42, POLB, NFKBIA, CCND1,
HIST1H4G, EP300, RAC1, PRKACA |
| HTLV-I
infection | 0.0812 | MAP3K3, JUN, IL1R2,
NFKB2, POLB, NFKBIA, HLA-DMB, CCND1, AKT3, DVL1, EP300, CTNNB1,
TLN2, TCF3, SLC25A5, WNT1, PRKACA, MAP3K3, JUN, IL1R2, NFKB2, POLB,
NFKBIA, HLA-DMB, CCND1, AKT3, DVL1, EP300, CTNNB1, TLN2, TCF3,
SLC25A5, WNT1, PRKACA |
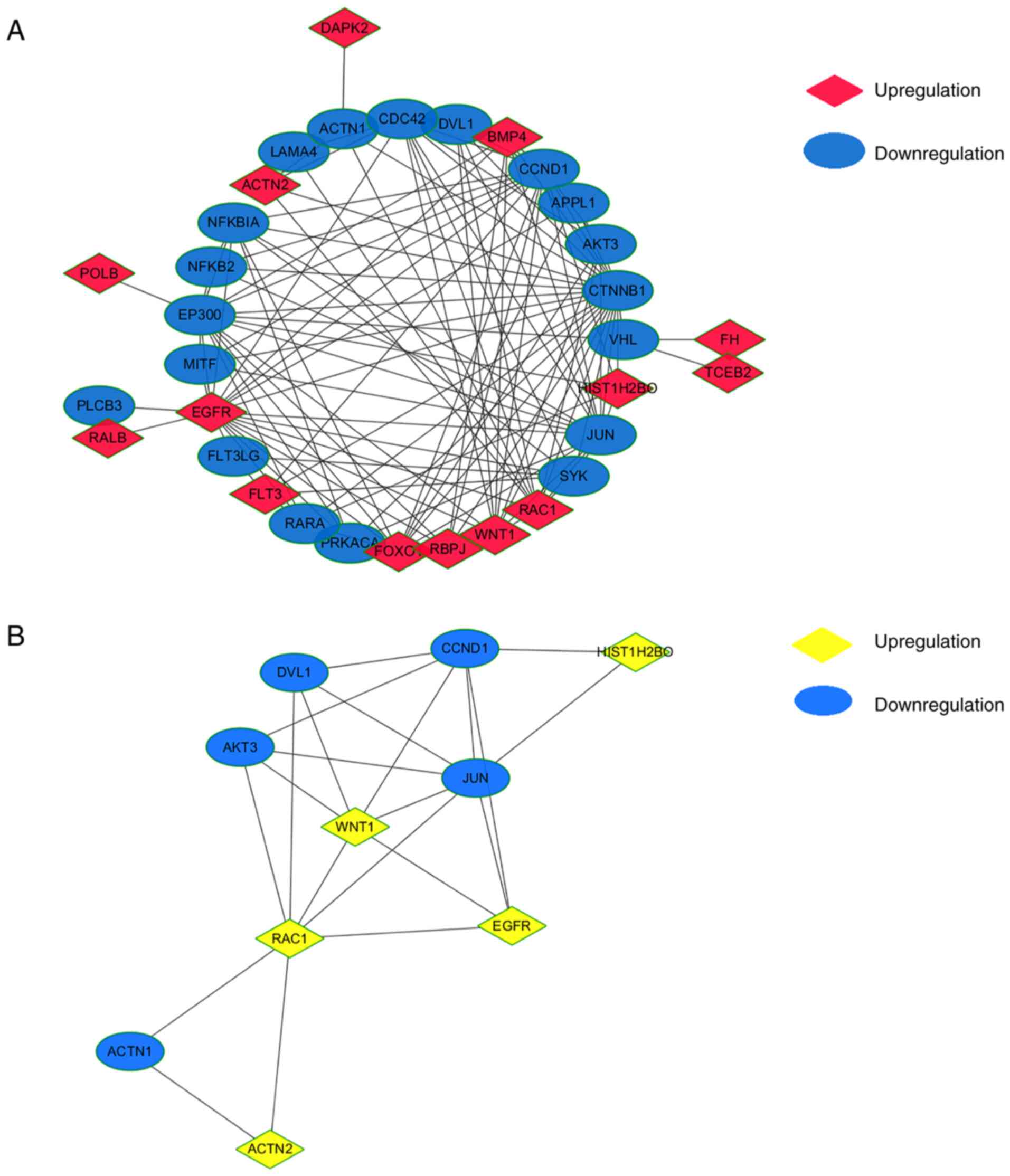
PPI network of shared KEGG pathways
and identification of hub genes
The 42 genes of the common KEGG pathways were used
to construct a PPI network using Cytoscape software (Fig. 5A). MCODE Cytoscape App was used to
integrate neighbors and density and identified 10 hub genes,
including WNT1, AKT3, disheveled segment polarity protein 1
(DVL1), cyclin D1 (CCND1), H2B clustered histone 17
(HIST1H2BO), JUN, EGFR, RAC1, actinin α1 (ACTN1) and
ACTN2 (Fig. 5B). Among
these, WNT1, RAC1, HIST1H2BO, ACTN2 and EGFR were
upregulated and AKT3, DVL1, CCND1, JUN and ACTN1 were
downregulated in PBMCs from patients with low BMD. Overall, some
hub genes were established, and further identification was
required.
AKT3 and RAC1 are critical genes in
regulation of osteoclast formation
To determine the involvement of hub genes in
osteoclast formation, THP-1 cells were treated with M-CSF and
sRANKL for 3 and 5 days to induce differentiation of osteoclasts.
When the majority of THP-1 cells formed a multi-nucleated fused
cluster, which indicated the formation of osteoclasts (Fig. 6A and B), cells were harvested for RT-qPCR
analysis. AKT3, CCND1, EGFR and RAC1 were
significantly upregulated, while WNT1, JUN and DVL1
were downregulated and ACTN1, HIST1H2BO and ACTN2
were unaffected during differentiation of osteoclasts (Fig. 6C-L). Among these, AKT3 and
RAC1 were the most differentially expressed genes and may be
involved in the regulation of osteoclast differentiation.
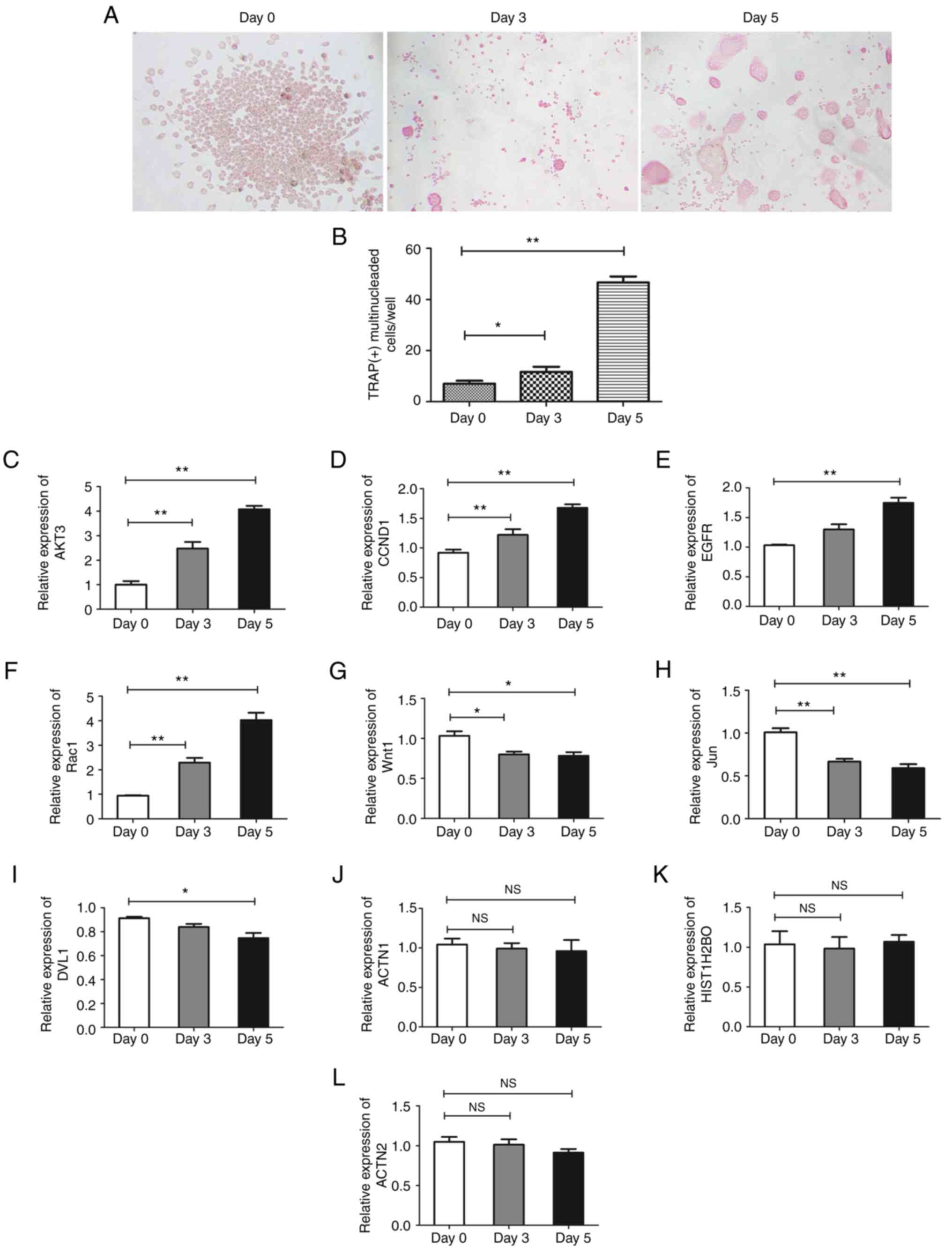 | Figure 6Expression of hub genes in
osteoclasts. THP-1 cells were induced into osteoclasts for 3 and 5
days. (A) The morphology was observed and (B) the number of nuclei
of multinuclear osteoclasts was counted to calculate the rate of
multinuclear osteoclasts.. Expression of (C) AKT3, (D)
CCND1, (E) EGFR, (F) RAC1, (G) WNT1,
(H) JUN, (I) DVL1, (J) ACTN1, (K)
HIST1H2BO and (L) ACTN2 were assessed by quantitative
PCR respectively. Magnification, x200. Data are presented as the
mean ± SD. The experiments were repeated three times.
*P<0.05, **P<0.01. CCND1, cyclin D1;
DVL1, disheveled segment polarity protein 1; ACTN, actinin α;
HIST1H2BO, H2B clustered histone 17; NS, not significant. |
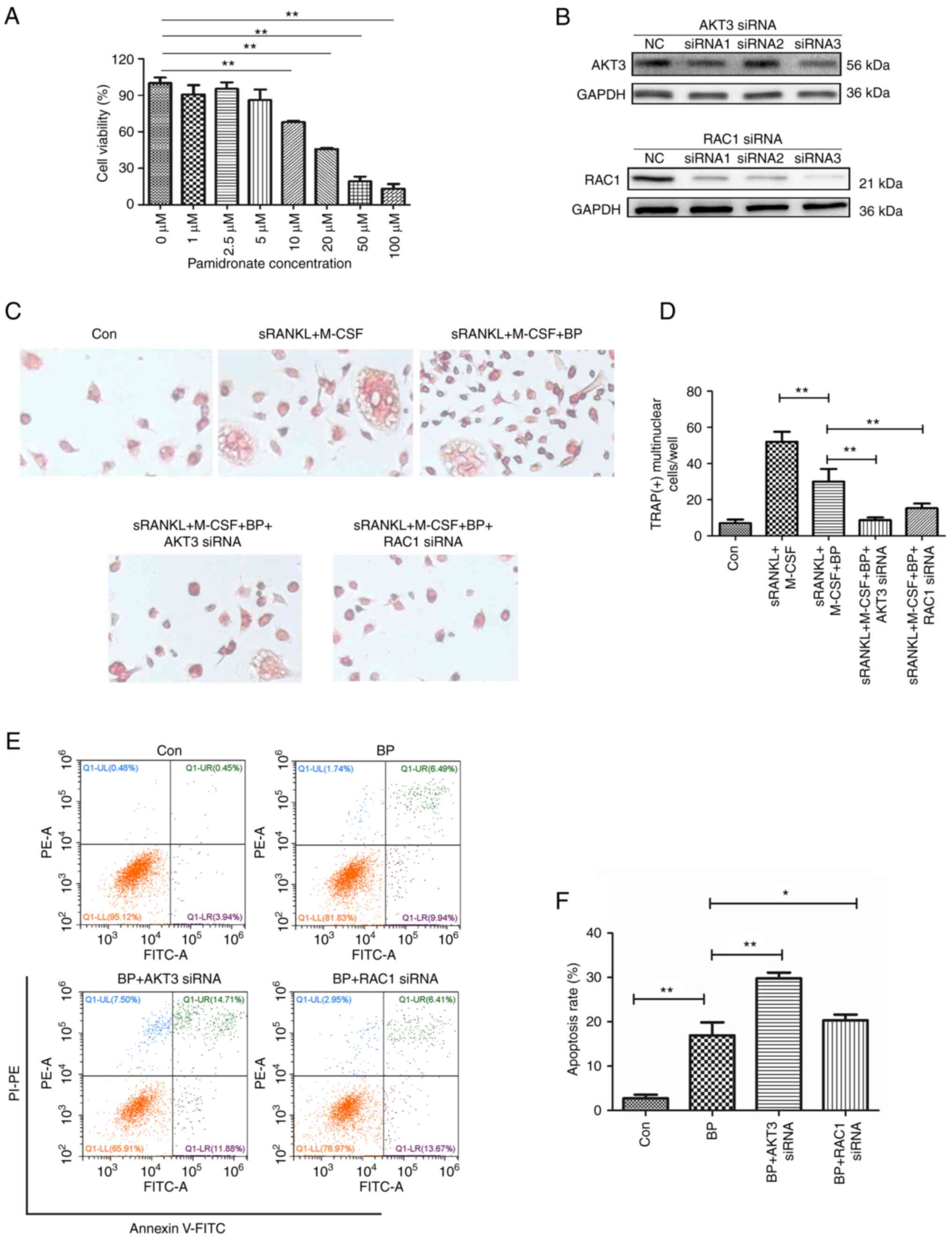
Inhibition of AKT3 and RAC1 enhances
the inhibitory effect of bisphosphonates on osteoclasts
Bisphosphonates prevent bone loss by decreasing
osteoclast activity and promoting osteoclast apoptosis (8). Cell viability assay showed that
pamidronate, a commonly used bisphosphonate, dose-dependently
inhibited viability of THP-1 cells and significant inhibition was
observed at doses ≥10 µM (Fig.
7A). Compared with the induced differentiation group (sRANKL +
M-CSF group), pamidronate at a dose of 5 µM significantly inhibited
the differentiation of THP-1 cells into osteoclasts (Fig. 7C and D). Therefore, 5 µM pamidronate was used
to inhibit osteoclast differentiation from THP-1 cells and 10 µM
was used for apoptosis analysis.
To assess whether AKT3 and RAC1 interfere with the
effect of bisphosphonates on differentiation and activity of
osteoclasts, siRNAs were used to knock down expression of AKT3 and
RAC1 in THP-1 cells prior to pamidronate treatment. The results
demonstrated that compared with NC, siRNA3 of AKT3 and siRNA1-3 of
RAC1 interfered with the expression levels of AKT3 and RAC1,
respectively. Among the three siRNAs of RAC1, siRNA3 had the most
obvious effect (Fig. 7B).
Knockdown of AKT3 and RAC1 significantly reduced the
nuclear fusion of THP-1 cells compared with pamidronate treatment
group, indicating that knockdown of AKT3 and RAC1 enhanced the
blocking effect of pamidronate on osteoclast differentiation
(Fig. 7C and D). Furthermore, siRNA of AKT3 and
RAC1 significantly enhanced pamidronate-induced apoptosis of
THP-1 cells (Fig. 7E and F). The effect of AKT3 siRNA was
notably stronger than that of RAC1 siRNA. These data
indicated that inhibition of AKT3 and RAC1 gene expression
inhibited osteoclast differentiation and promoted the
apoptosis-inducing effect of bisphosphonates.
Discussion
PMO is a common disease, and approximately one-third
of women aged 60-70 years have osteoporosis worldwide and nearly
one-third of women >50 years of age develop osteoporotic
fractures (3), but the underlying
mechanisms remain unclear. The present study performed differential
gene expression analysis using a publicly available GEO dataset and
identified DEGs between PBMCs of patients with PMO with high and
low BMD. STITCH database was used to mine proteins that interact
with bisphosphonates. Pathway enrichment data of the two analyses
was combined, common enriched signaling pathways were screened and
two key hub genes, AKT3 and RAC1, were identified.
Finally, in vitro osteoclast formation model demonstrated
that inhibiting AKT3 and RAC1 expression enhanced the
inhibitory effect of bisphosphonates on osteoclast activation and
differentiation.
Previous studies have analyzed DEGs of patients with
PMO (16,17). The present study focused on gene
expression of PBMCs in patients with PMO because osteoclasts are
differentiated from PBMCs (18).
Certain studies have shown that the RANKL signaling pathway is
highly activated in PBMCs of patients with PMO, suggesting
involvement of PBMCs in the progression of PMO (19,20).
The primary effect of bisphosphonates is to inhibit osteoclast
activation, thereby preventing bone loss (8). Therefore, it was hypothesized that
the gene expression profile of PBMCs may be used to delineate the
association between bisphosphonates and osteoporosis.
Firstly, the GEO dataset was analyzed and results
demonstrated a total of 290 DEGs between the low BMD patient and
high BMD patient. The functional annotation of DEGs indicated that
most DEGs were closely associated with hormone-related signaling
pathways, DNA replication and biosynthesis, which is in accordance
with the known pathogenesis of PMO, which involves dysregulation of
osteoclast-associated molecules and downregulation of estrogen
(21). Next, by integrating
bioinformatics data of the aforementioned databases, four common
KEGG pathways were identified; three were associated with
occurrence and development of tumors, including ‘pathways in
cancer’, ‘HIF-1 signaling pathway’ and ‘viral carcinogenesis’. This
indicated that certain activated signaling molecules involved in
occurrence and development of PMO may exhibit crosstalk with
oncogenic signaling, which has been reported in previous studies
(22,23). For example, Zhong et al
(22) showed that HIF-1 signaling
is involved in the formation of PMO and Yu et al (23) found that the tumor suppressor P53
serves a key role in the progression of osteoporosis. Here,
cancer-associated signaling pathways primarily involved biological
processes such as ‘proliferation’ and ‘differentiation’. This
suggested that the occurrence of PMO is associated with
proliferation and differentiation of osteoclasts.
Hub genes of the four common signaling pathways were
analyzed and WNT1, AKT3, DVL1, CCND1, HIST1H2BO, JUN, EGFR,
RAC1, ACTN1 and ACTN2 were screened out. These hub genes
are primarily involved in the processes of cell proliferation and
differentiation (24-31).
Some genes are also reported to be involved in the formation of
osteoclasts (32,33).
In addition to bioinformatics analysis, in
vitro experiments were performed to verify the expression of
the aforementioned hub genes during osteoclast differentiation.
AKT3, CCND1, EGFR and RAC1 were significantly
upregulated, while WNT1, JUN and DVL1 were
downregulated during differentiation of THP-1 cells into
osteoclasts. However, in the GEO dataset, expression levels of
WNT1, RAC1, HIST1H2BO, ACTN2 and EGFR were
upregulated, while those of AKT3, DVL1, CCND1, JUN and
ACTN1 were downregulated in patients with PMO with low BDM.
The present results showed that only four hub genes showed an
expression pattern consistent with that in patients with PMO,
indicating that PMO is a complex and dynamic process (34).
AKT3 and RAC1, which are upregulated
during osteoclast differentiation, were selected for functional
analysis. The inhibitory effects of bisphosphonates on osteoclasts
were significantly enhanced when AKT3 and RAC1
expression was knocked down, which not only decreased
differentiation of osteoclasts but also increased apoptosis of
monocytes. Therefore, AKT3 and RAC1 may be promising
targets for enhancing the therapeutic effect of bisphosphonates on
PMO.
The association between Rac1 and PMO has previously
been reported (35,36). Multiple studies have shown that
activation of Rac1 promotes osteoclastogenesis (37-39);
therefore, Rac1 may be an effective target for the prevention and
treatment of PMO. The present results are consistent with previous
findings (37-39),
indicating that screening hub genes by bioinformatics combined with
target prediction of drugs is feasible. To the best of our
knowledge, there are no previous reports on the association between
Akt3 and PMO or osteoclast activation. Therefore, the present study
provided novels insights into the molecular mechanism of PMO.
The present study had certain limitations. First,
only one dataset was used in the analysis, which may not reflect
the gene expression pattern of PBMCs in PMO. Second, only an in
vitro osteoclast differentiation model was used; the present
findings need to be validated in vivo.
In summary, the present study identified AKT3
and RAC1 as two novel key genes in PMO via combined analysis
of a GEO dataset of patients with PMO and the STITCH database. The
present data provided a new avenue for understanding the mechanism
of PMO and improving the therapeutic efficacy of
bisphosphonates.
Supplementary Material
Shared KEGG pathways. Shared KEGG
pathways, including (A) ‘pathways in cancer’, (B) ‘human T-cell
leukemia virus 1 infection’, (C) ‘HIF-1 signaling pathway’ and (D)
‘viral carcinogenesis’. HIF, hypoxia-inducible factor. Pentagrams
represent the genes that were involved in the DEGs, rectangles
refer to gene products, circles generally refer to compounds, +p
indicates phosphorylated, -p indicates dephosphorylation, solid
arrows represent activation, dotted arrows represent indirect
effects, dotted lines represent state changes, the straight line
represents the combination, and four rectangles represent the
complex. KEGG, Kyoto Encyclopedia of Genes and Genomes.
siRNA sequences.
Primer sequences of hub genes.
Acknowledgements
Not applicable.
Funding
Funding: This study was supported by The Tribology Science Fund
of State Key Laboratory of Tribology (project name, Preparation and
Tribological Properties of self-healing hydrogel materials for
artificial joint interface; grant no. SKLTKF21B04).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
SX and YW designed the study. LZ performed
bioinformatics analysis and wrote the manuscript. XL performed
experiments in vitro and took part in the manuscript
writing. CW, WD, JZ and LF performed experiments. QF analyzed the
experimental data. SX and YW confirm the authenticity of all the
raw data. All authors have read and approved the final
manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Arceo-Mendoza RM and Camacho PM:
Postmenopausal osteoporosis: Latest guidelines. Endocrinol Metab
Clin North Am. 50:167–178. 2021.PubMed/NCBI View Article : Google Scholar
|
|
2
|
McNamara LM: Osteocytes and estrogen
deficiency. Curr Osteoporos Rep. 19:592–603. 2021.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Baccaro LF, Conde DM, Costa-Paiva L and
Pinto-Neto AM: The epidemiology and management of postmenopausal
osteoporosis: A viewpoint from Brazil. Clin Interv Aging.
10:583–591. 2015.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Manolagas SC: Birth and death of bone
cells: Basic regulatory mechanisms and implications for the
pathogenesis and treatment of osteoporosis. Endocr Rev. 21:115–137.
2000.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Fujikawa Y, Quinn JM, Sabokbar A, McGee JO
and Athanasou NA: The human osteoclast precursor circulates in the
monocyte fraction. Endocrinology. 137:4058–4060. 1996.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Cohen-Solal ME, Graulet AM, Derme MA,
Gueris J, Baylink D and de Vernejoul MC: Peripheral monocyte
culture supematants of menopausal women can induce bone resorption:
Involvement of cytokines. J Clin Endocrinol Metab. 77:1648–1653.
1993.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Dera AA, Ranganath L, Barraclough R,
Vinjamuri S, Hamill S, Mandourah AY and Barraclough DL: Altered
levels of mRNAs for calcium-binding/associated proteins, Annexin
A1, S100A4, and TMEM64, in peripheral blood mononuclear cells are
associated with osteoporosis. Dis Markers.
2019(3189520)2019.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Endo Y, Funayama H, Yamaguchi K, Monma Y,
Yu Z, Deng X, Oizumi T, Shikama Y, Tanaka Y, Okada S, et al: Basic
studies on the mechanism, prevention, and treatment of
osteonecrosis of the jaw induced by bisphosphonates. Yakugaku
Zasshi. 140:63–79. 2020.PubMed/NCBI View Article : Google Scholar : (Article in
Japanese).
|
|
9
|
Ihn HJ, Lee D, Lee T, Kim SH, Shin HI, Bae
YC, Hong JM and Park EK: Inhibitory effects of kp-a159, a
thiazolopyridine derivative, on osteoclast differentiation,
function, and inflammatory bone loss via suppression of
rankl-induced map kinase signaling pathway. PLoS ONE.
10(e0142201)2015.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Dutta P, Zhang L, Zhang H, Peng Q,
Montgrain PR, Wang Y, Song Y, Li J and Li WX: Unphosphorylated
STAT3 in heterochromatin formation and tumor suppression in lung
cancer. BMC Cancer. 20(145)2020.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Lithgow KV, Church B, Gomez A, Tsao E,
Houston S, Swayne LA and Cameron CE: Identification of the
neuroinvasive pathogen host target, LamR, as an endothelial
receptor for the treponema pallidum adhesin Tp0751. mSphere.
5:e00195–20. 2020.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Huang da W, Sherman BT and Lempicki RA:
Systematic and integrative analysis of large gene lists using DAVID
bioinformatics resources. Nat Protoc. 4:44–57. 2009.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Wang J, Liu H, Xie G, Cai W and Xu J:
Identification of hub genes and key pathways of dietary advanced
glycation end products-induced non-alcoholic fatty liver disease by
bioinformatics analysis and animal experiments. Mol Med Rep.
21:685–694. 2020.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Guimaraes HI, Santana RH, Silveira R,
Pinto OH, Quirino BF, Barreto CC, Bustamante MM and Krüger RH:
Seasonal variations in soil microbiota profile of termite
(syntermes wheeleri) mounds in the Brazilian tropical savanna.
Microorganisms. 8(1482)2020.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Yang C, Ren J, Li B, Jin C, Ma C, Cheng C,
Sun Y and Shi X: Identification of gene biomarkersin patients with
postmenopausal osteoporosis. Mol Med Rep. 19:1065–1073.
2019.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Liu T, Huang J, Xu D and Li Y: Identifying
a possible new target for diagnosis and treatment of postmenopausal
osteoporosis through bioinformatics and clinical sample analysis.
Ann Transl Med. 9(1154)2021.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Salamanna F, Maglio M, Borsari V,
Giavaresi G, Aldini NN and Fini M: Peripheral blood mononuclear
cells spontaneous osteoclastogenesis: Mechanisms driving the
process and clinical relevance in skeletal disease. J Cell Physiol.
231:521–530. 2016.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Jin H, Yao L, Chen K, Liu Y, Wang Q, Wang
Z, Liu Q, Cao Z, Kenny J, Tickner J, et al: Evodiamine inhibits
RANKL-induced osteoclastogenesis and prevents ovariectomy-induced
bone loss in mice. J Cell Mol Med. 23:522–534. 2019.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Chen X, Li J, Ye Y, Huang J, Xie L, Chen
J, Li S, Chen S and Ge J: Association of cardiotrophin-like
cytokine factor 1 levels in peripheral blood mononuclear cells with
bone mineral density and osteoporosis in postmenopausal women. BMC
Musculoskelet Disord. 22(62)2021.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Li L and Wang Z: Ovarian aging and
osteoporosis. Adv Exp Med Biol. 1086:199–215. 2018.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Zhong H, Cao C, Yang J and Huang Q:
Research on relationship of HIF-1 signaling pathway and
postmenstrual osteoporosis. Sichuan Da Xue Xue Bao Yi Xue Ban.
48:862–868. 2017.PubMed/NCBI(In Chinese).
|
|
23
|
Yu T, You X, Zhou H, Kang A, He W, Li Z,
Li B, Xia J, Zhu H, Zhao Y, et al: p53 plays a central role in the
development of osteoporosis. Aging (Albany NY). 12:10473–10487.
2020.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Wei W, He HB, Zhang WY, Zhang HX, Bai JB,
Liu HZ, Cao JH, Chang KC, Li XY and Zhao SH: miR-29 targets Akt3 to
reduce proliferation and facilitate differentiation of myoblasts in
skeletal muscle development. Cell Death Dis. 4(e668)2013.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Castro-Piedras I, Sharma M, den Bakker M,
Molehin D, Martinez EG, Vartak D, Pruitt WM, Deitrick J, Almodovar
S and Pruitt K: DVL1 and DVL3 differentially localize to CYP19A1
promoters and regulate aromatase mRNA in breast cancer cells.
Oncotarget. 9:35639–35654. 2018.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Wang LJ and Cai HQ: Let-7b downgrades
CCND1 to repress osteogenic proliferation and differentiation of
MC3T3-E1 cells: An implication in osteoporosis. Kaohsiung J Med
Sci. 36:775–785. 2020.PubMed/NCBI View Article : Google Scholar
|
|
27
|
He Y, Cao Y, Wang X, Jisiguleng W, Tao M,
Liu J, Wang F, Chao L, Wang W, Li P, et al: Identification of hub
genes to regulate breast cancer spinal metastases by bioinformatics
analyses. Comput Math Methods Med. 2021(5548918)2021.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Koyuturk M, Ersoz M and Altiok N:
Simvastatin induces proliferation inhibition and apoptosis in C6
glioma cells via c-jun N-terminal kinase. Neurosci Lett.
370:212–217. 2004.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Abe S, Ueno M, Nishitani M, Akamatsu T,
Sato T, Shimoda M, Kanaoka H, Nii Y, Yamasaki H and Yuasa K: Citrus
sudachi peel extract suppresses cell proliferation and promotes the
differentiation of keratinocytes through inhibition of the EGFR-ERK
signaling pathway. Biomolecules. 10(1468)2020.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Gahankari A, Dong C, Bartoletti G, Galazo
M and He F: Deregulated Rac1 activity in neural crest controls cell
proliferation, migration and differentiation during midbrain
development. Front Cell Dev Biol. 9(704769)2021.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Peng W, Liu Y, Qi H and Li Q:
Alpha-actinin-4 is essential for maintaining normal trophoblast
proliferation and differentiation during early pregnancy. Reprod
Biol Endocrinol. 19(48)2021.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Song C, Guo Y, Chen F and Liu W: LncRNA
MALAT1 promotesosteogenic differentiation through the miR-217/AKT3
axis: A possible strategy to alleviate osteoporosis. J Gene Med.
24(e3409)2022.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Sharma A, McAfee J, Wang L, Cook E,
Ababneh E and Bergfeld WF: Utility of Cyclin D1 immunostaining in
cutaneous xanthogranuloma. Am J Dermatopathol. 43:e141–e145.
2021.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Korsić M: Pathophysiology of
postmenopausal osteoporosis. Reumatizam. 53:32–35. 2006.PubMed/NCBI(In Croatian).
|
|
35
|
Li J, Li X, Liu D, Hamamura K, Wan Q, Na
S, Yokota H and Zhang P: eIF2α signaling regulates autophagy of
osteoblasts and the development of osteoclasts in OVX mice. Cell
Death Dis. 10(921)2019.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Magalhaes JK, Grynpas MD, Willett TL and
Glogauer M: Deleting Rac1 improves vertebral bone quality and
resistance to fracture in a murine ovariectomy model. Osteoporos
Int. 22:1481–1492. 2011.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Gao L, Kong L and Zhao Y: The regulatory
role of Rho GTPases and their substrates in osteoclastogenesis.
Curr Drug Targets. 22:1064–1070. 2021.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Wang Y, Belsham DD and Glogauer M: Rac1
and Rac2 in osteoclastogenesis: A cell immortalization model.
Calcif Tissue Int. 85:257–266. 2009.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Wang Y, Lebowitz D, Sun C, Thang H,
Grynpas MD and Glogauer M: Identifying the relative contributions
of Rac1 and Rac2 to osteoclastogenesis. J Bone Miner Res.
23:260–270. 2008.PubMed/NCBI View Article : Google Scholar
|