Introduction
Quercetin (Qu)
[2-(3,4-dihydroxyphenyl)-3,5,7-trihydroxy-4H-1-benzopyran-4-one]
belongs to an extensive class of polyphenolic flavonoid compounds
ubiquitous in plants and plant food sources. Qu is a beneficial
chemical compound widely dispersed in the human diet. Accordingly,
high concentrations of Qu are found in onions (391.0 mg/kg),
broccoli (74.5 mg/kg), apples (50 mg/kg) and beans (11.0 mg/kg)
(1) and intakes between 6 and 31
mg/day have been reported (2). Qu
has been used as a dietary supplement for many years, and thus, its
use does not raise concerns as regards its toxicity. Of note, when
Qu was discovered it was believed to be a mutagen, but subsequent
studies showed it to be a powerful natural anticancer agent
(3). The exact mechanism behind
the cancer preventive effect of Qu remians unknown, yet the
majority of studies have indicated that its anti-inflammatory and
antioxidant properties may be responsible for the beneficial
effects (4). Although the link
between Qu and cancer has been established in controlled scientific
settings, its efficacy in the treatment of cancer in humans is
currently unknown.
Human malignant pleural mesothelioma is a rare
tumor; however, its incidence is increasing and it has a poor
prognosis (5,6). Thus, knowledge of the mechanism of
mesothelial carcinogenesis is required to support efforts to
develop targeted treatments.
Specificity protein (Sp) is an ubiquitously
expressed transcription factor belonging to a family of 8
transcription factors (7), which
are ubiquitously expressed in a variety of mammalian cells
(8). Furthermore, Sp1 is highly
expressed in a number of cancer tissues, such as pancreatic,
thyroid, colorectal, breast, hepatocellular, prostate, gastric
cancer and lung cancer (7,9).
Sp1 was one of the first eukaryotic transactivators to be
identified (10). Sp1 proteins
have been defined as Sp/Kruppel-like transcription factors
(11), and a previous study
showed that Sp1 plays an important role in the carcinogenesis and
metastasis of human tumors by regulating growth-related signal
transductions, apoptosis, tumor suppressor genes, cell cycle
control molecules, oncogenes and angiogenesis-related factors
(12,13).
Qu is an important dietary flavonoid, found in
various vegetables, fruits, seeds, nuts, tea and red wine (14). Qu has been shown to have diverse
biological activities, including anti-inflammatory and antitumor
properties (15–17). Furthermore, Qu has been shown to
have various beneficial effects, such as antioxidant,
cardioprotective, anti-inflammatory and antitumor effects (18). However, the antitumor mechanisms
involved and the molecular targets of Qu have not been identified,
especially in human malignant pleural mesothelioma. Accordingly, in
the present study, we investigated the apoptotic effect of Qu on a
human malignant pleural mesothelioma cell line. Our results suggest
that Qu be considered as a drug or natural supplement candidate for
the prevention of malignant pleural mesothelioma.
Materials and methods
Materials
HyClone RPMI-1640 medium and fetal bovine serum
(FBS) were obtained from Thermo Scientific (Logan, UT). The
following antibodies were used: 4′-6-diamidino-2-phenylindole
(DAPI), propidium iodide (PI), anti-Sp1 (1C6), anti-poly
(ADP-ribose) polymerase (PARP) (BD Biosciences, San Diego, CA),
anti-cyclin D1 (M-20), anti-myeloid cell leukemia (Mcl)-1,
anti-survivin, anti-Bid, anti-Bax, anti-Bcl-xL (Cell Signaling
Technology, Inc., Danvers, MA), anti-caspase-3 (H-277) (Santa Cruz
Biotechnology, Inc., Santa Cruz, CA) and anti-β-actin (AC-74)
(Sigma-Aldrich, Inc. St. Louis, MO). RNase A was supplied by
Sigma-Aldrich.
Cell culture
MSTO-211H cells were obtained from the American
Tissue Culture Collection (Manassas, VA). Cells were maintained in
RPMI-1640 medium supplemented with 5% FBS and 100 U/ml each of
penicillin and streptomycin at 37°C in a 5% CO2
incubator.
MTS assay
The effects of Qu on cell viability were determined
using a
3-(4,5-dimethylthiazol-2-yl)-5-(3-carboxymethoxyphenyl)-2-(4-sulfophenyl)-2H-tetrazolium
(MTS) assay kit (Promega, Madison, WI). MSTO-211H cells were seeded
in a 96-well plate for 24 h and treated with Qu (20–80 μM)
for 24 or 48 h. MTS solution was then added for 2 h at 37°C in 5%
CO2. The absorbance at 490 nm was recorded using a
GloMax-Multi Microplate Multimode Reader (Promega).
DAPI staining
The level of nuclear condensation and fragmentation
was observed by nucleic acid staining with DAPI. MSTO-211H cells
were treated with Qu, harvested by trypsinization, and fixed in
100% methanol at room temperature for 20 min. The cells were spread
on slides, stained with DAPI solution (2 μg/ml), and
analyzed under a FluoView confocal laser microscope (Fluoview
FV10i; Olympus Corporation, Tokyo, Japan).
PI staining
Following treatment with Qu (20–80 μM) for 72
h, the detached MSTO-211H cells (floaters) were collected by
centrifugation and combined with adherent cells. The cells were
fixed with 70% ice-cold ethanol overnight at −20°C, and
subsequently treated with 150 mg/ml RNase A and 20 mg/ml PI. DNA
contents were analyzed by flow cytometry using a MACSQuant Analyzer
(Miltenyi Biotec GmbH, Bergisch Gladbach, Germany).
Western blot analysis
MSTO-211H cells treated with Qu (20–80 μM)
for 48 h were washed with phosphate-buffered saline (PBS), and were
then homogenized with PRO-PREP™ Protein Extraction Solution (Intron
Biotechnology, Korea). Extracted proteins were measured using DC
protein assay reagent (Bio-Rad Laboratories Inc., Hercules, CA).
The equal amounts of protein samples were separated by 12%
SDS-polyacrylamide gel electrophoresis and then transferred onto
membranes, which were blocked for 2 h at room temperature with 5%
non-fat dried milk in PBS containing 0.05% Tween-20, and then
incubated overnight at 4°C with specific antibodies. The protein
bands were observed after treating them with horseradish
peroxidase-conjugated secondary antibody using a Pierce ECL Western
Blotting Substrate (Thermo Scientific, Rockford, IL).
In vitro EGCG-sepharose 4B pull-down
assays
This method has been described previously (19,20). Briefly, MSTO-211H cell lysates
were reacted with sepharose 4B beads or Qu-sepharose 4B beads in
reaction buffer (50 mM Tris, pH 7.5, 5 mM EDTA, 150 mM NaCl, 1 mM
dithiothreitol, 0.01% Nonidet P-40, 2 μg/ml bovine serum
albumin, 0.02 mM phenylmethylsulfonyl fluoride and 1X proteinase
inhibitor), and washed 5 times with washing buffer (50 mM Tris, pH
7.5, 5 mM EDTA, 150 mM NaCl, 1 mM dithiothreitol, 0.01% Nonidet
P-40, 0.02 mM phenylmethylsulfonyl fluoride). Proteins bound to the
beads were analyzed by western blot analysis using anti-Sp1
antibody.
Reverse transcription-polymerase chain
reaction (RT-PCR)
Total RNA was extracted from the cells using
TRIzol® Reagent (Life Technologies, Carlsbad, CA), and 2
μg of RNA were used to synthesize cDNA using the HelixCript™
first-strand cDNA synthesis kit (NanoHelix, Korea). cDNA was
obtained by PCR amplification using β-actin-specific and
Sp1-specific primers (as described below) using the following PCR
conditions: 25 cycles of 1 min at 95°C, 1 min at 60°C and 1 min at
72°C. The β-actin primers used were: forward,
5′-GTG-GGG-CGC-CCC-AGG-CAC-CA-3′ and reverse,
5′-CTC-CTT-AAT-GTC-ACG-CAC-GAT-TTC-3′; and the Sp1 primers were:
forward, ATG CCT AAT ATT CAG TAT CAA GTA and reverse, CCC TGA GGT
GAC AGG CTG TGA. PCR products were analyzed by 2% agarose gel
electrophoresis.
Luciferase assay for cyclin D1, Mcl-1 and
survivin trans-activation
MSTO-211H cells (6x104) were added to
each well of a 24-well plate and incubated at 37°C in a humidified
atmosphere of 5% CO2 for 24 h. Transient transfection
was performed using Lipofectamine 2000 reagent (Invitrogen,
Carlsbad, CA). The survivin-269 promoter construct was kindly
provided by Dr Sung-Dae Cho (Chonbuk National University, Jeonju,
Korea). The Mcl-1 promoter was obtained from Addgene (Cambridge,
MA). Cells were transfected with 250 ng of cyclin D1 (-1745-luc),
Mcl-1 (-325-luc), or survivin (-269-luc) and 20 ng of β-gal using
Lipofectamine 2000 reagent (Invitrogen) for 24 h (21). After cafestol and kahweol
treatment for 48 h, the cells were disrupted with 100 ml of lysis
buffer [0.1 M potassium phosphate (pH 7.8), 1% Triton X-100, 1 mM
dithiothreitol (DTT), 2 mM EDTA], and firefly luciferase activity
and galactosidase were determined with a Promega luciferase assay
kit according to the manufacturer’s instructions. Relative
luciferase units were calculated by normalizing to the
galactosidase activity.
Statistical analysis
The results are presented as the means ± SD of at
least 3 independent experiments performed in triplicate. Data were
analyzed for statistical significance using one-way analysis of
variance. A P-value <0.05 was considered to indicate a
statistically significant difference.
Results
Qu reduces the viability of MSTO-211H
cells
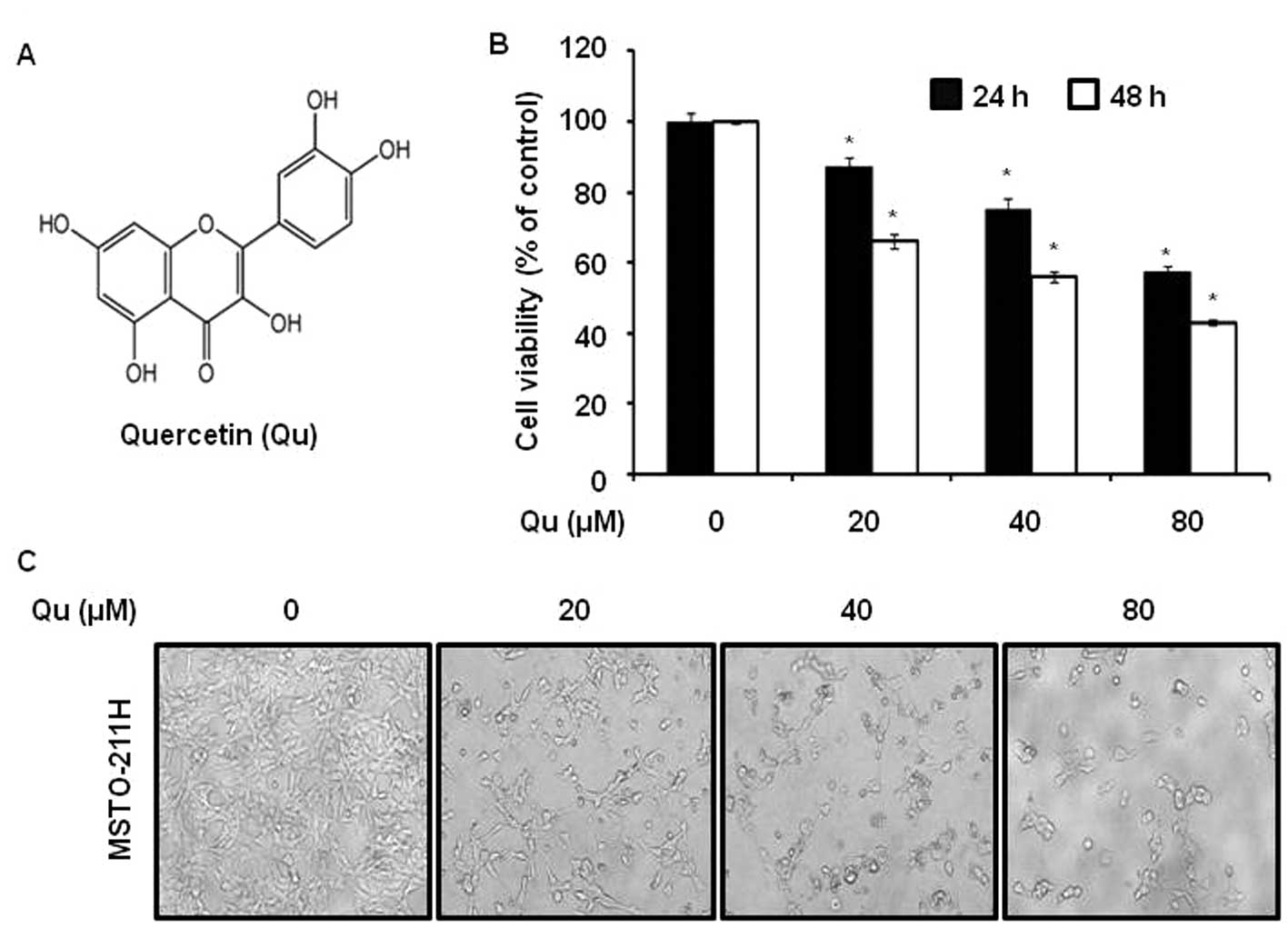
To evaluate the effects of Qu (Fig. 1A) on the viability of malignant
mesothelioma cells we used a MTS assay. It was found that Qu
suppressed cell viability with an IC50 of 58 μM
in MSTO-211H cells for 48 h (Fig.
1B). Qu treatment also resulted in a significant
concentration-dependent inhibition of cell growth with
IC50 values of approximately 73 μM in HT28 cells
(another malignant mesothelioma cell line) for 48 h (data not
shown). To investigate the morphological changes, MSTO-211H cells
were treated with various concentrations (20–80 μM) of Qu
for 48 h. The results obtained showed that the cells had decreased
in size and that they had became rounded (Fig. 1C).
Qu induces the apoptosis of MSTO-211H
cells
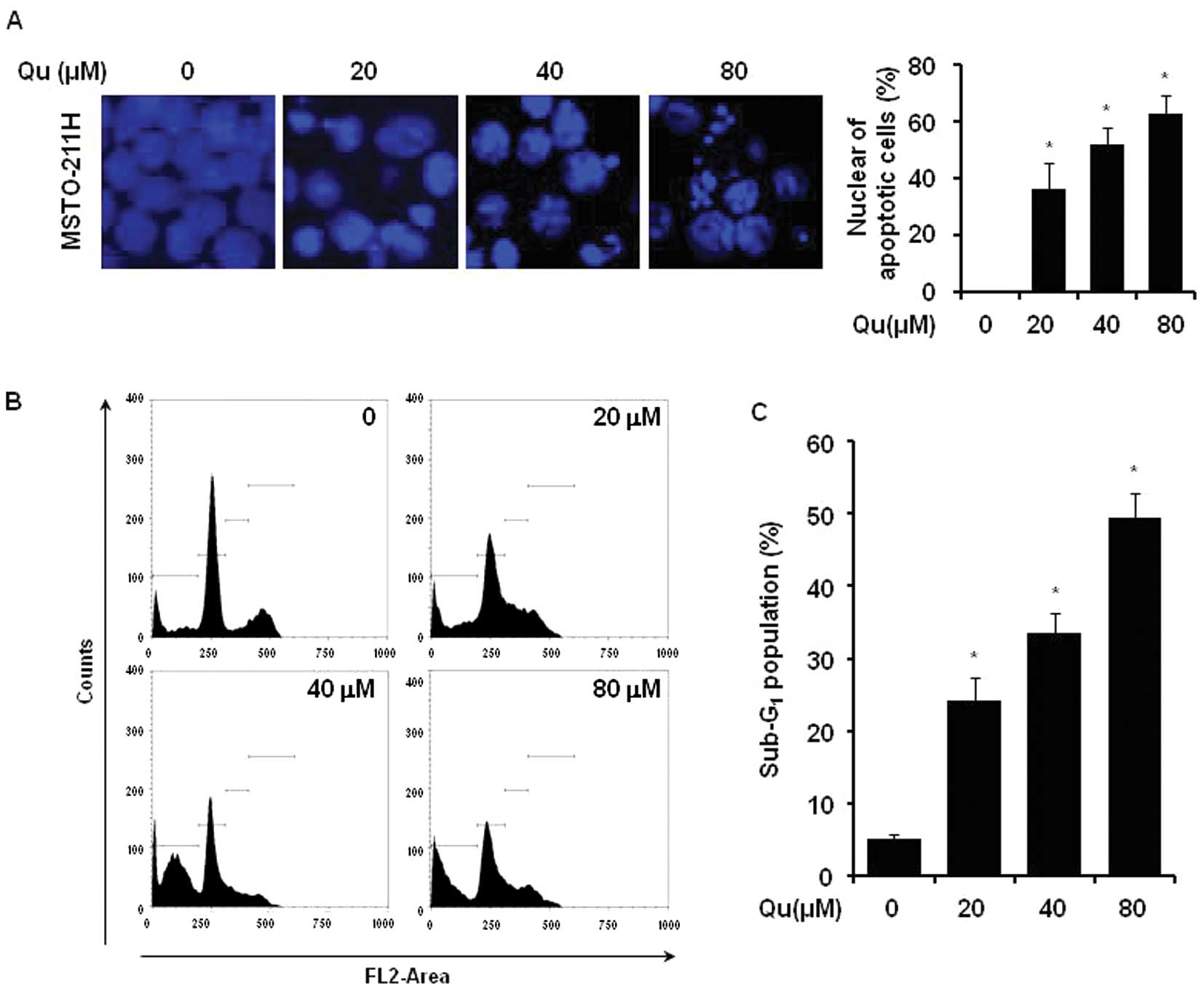
The effect of Qu treatment on the initiation of
apoptosis in MSTO-211H cells was determined by nuclear morphology
using DAPI staining. The Qu treatment of mesothelioma cells
increased nuclear condensation and fragmentation when compared to
the control group (Fig. 2A). In
order to evaluate whether the increase in the sub-G1
cell population induced by Qu was related to apoptosis, the
Qu-treated cells were used for PI staining. The number of MSTO-211H
cells in the sub-G1 phase increased from 25 to 50% in
the presence of 20–80 μM Qu (Fig. 2B and C).
Qu regulates Sp1 protein levels in
MSTO-211H cells
Sp1 contributes to cell progression and apoptotic
cell death via the regulation of the expression of a number of
genes, such as cyclin D1, Mcl-1, survivin, Bcl-xL, Bax and
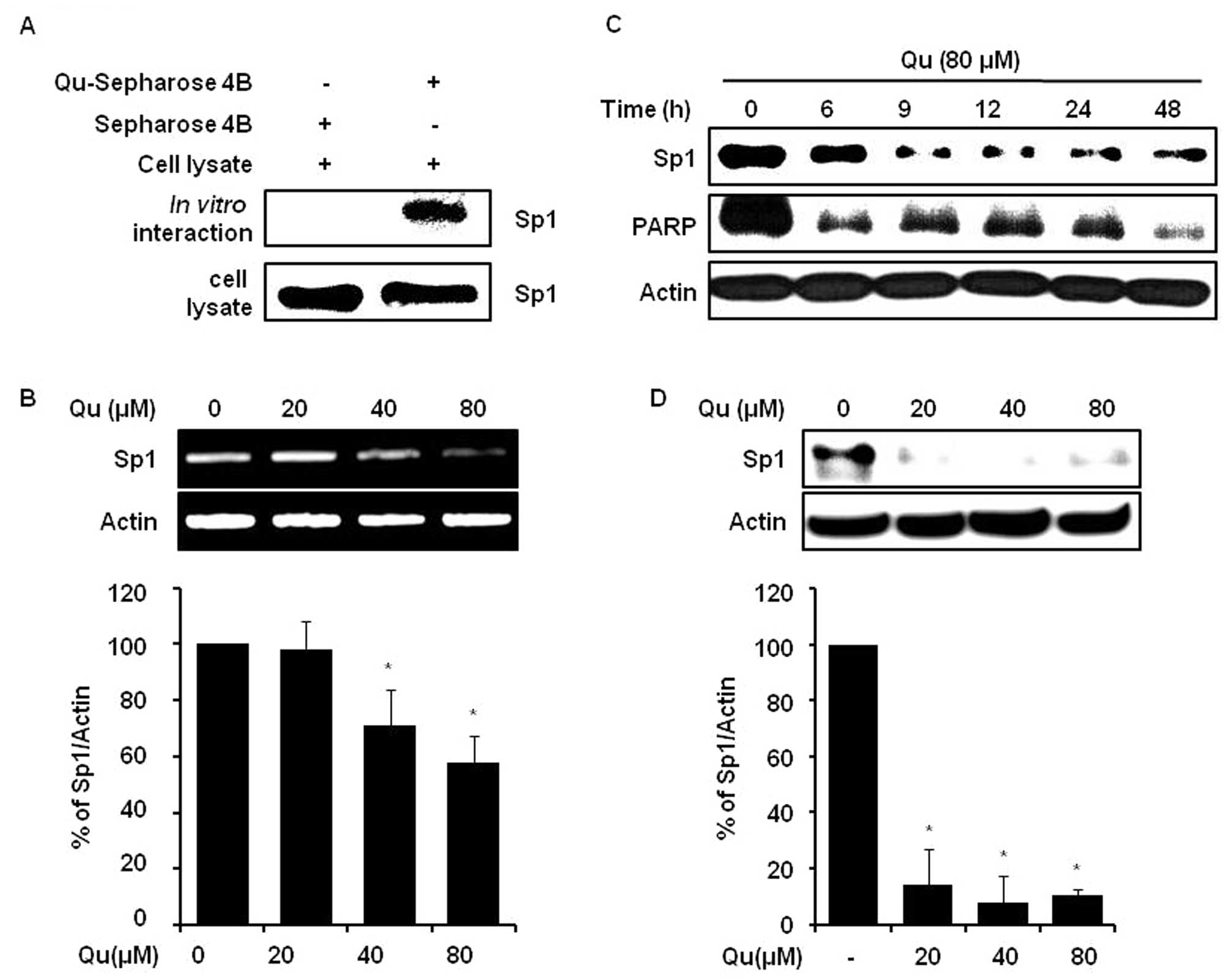
caspase-3 in various cancers (7,13,22). The interaction between Sp1 and Qu
was examined by conducting a Qu-sepharose 4B affinity experiment by
immunoblotting with anti-Sp1. The results obtained indicated that
Qu bound with Sp1 in cell lysates from human MSTO-211H cells
(Fig. 3A). Furthermore, we found
that Qu at 20, 40 and 80 μM downregulated Sp1 protein
(Fig. 3D) and mRNA levels
(Fig. 3B), and Sp1 protein
expression level monitoring showed that Qu time-dependently reduced
protein levels over 0 to 48 h (Fig.
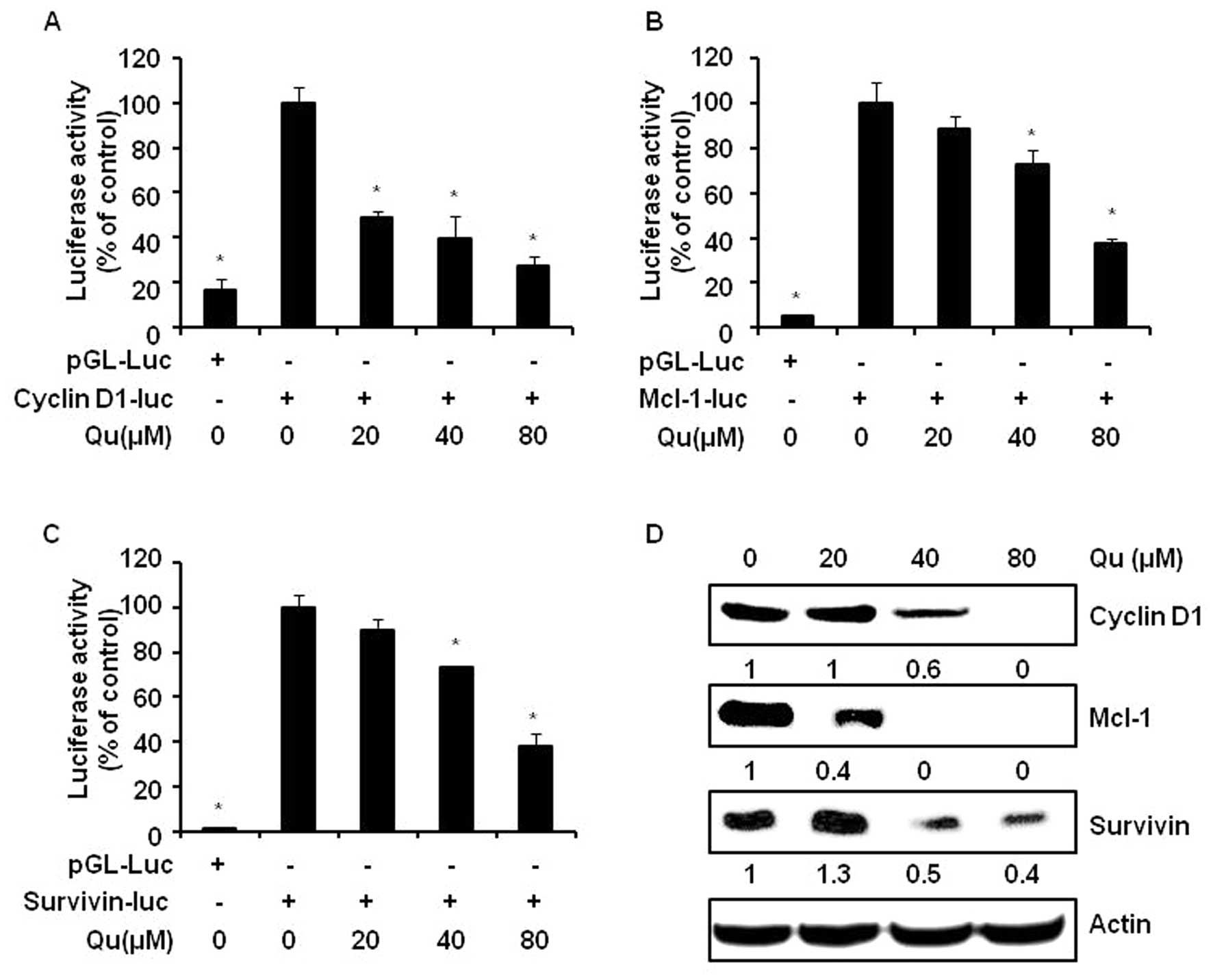
3C). Qu clearly attenuated the cyclin D1, Mcl-1 and survivin
promoter activities (Fig. 4A–C).
In addition, significant decreases in the protein levels of cyclin
D1, Mcl-1 and survivin were observed following Qu treatment
(Fig. 4D).
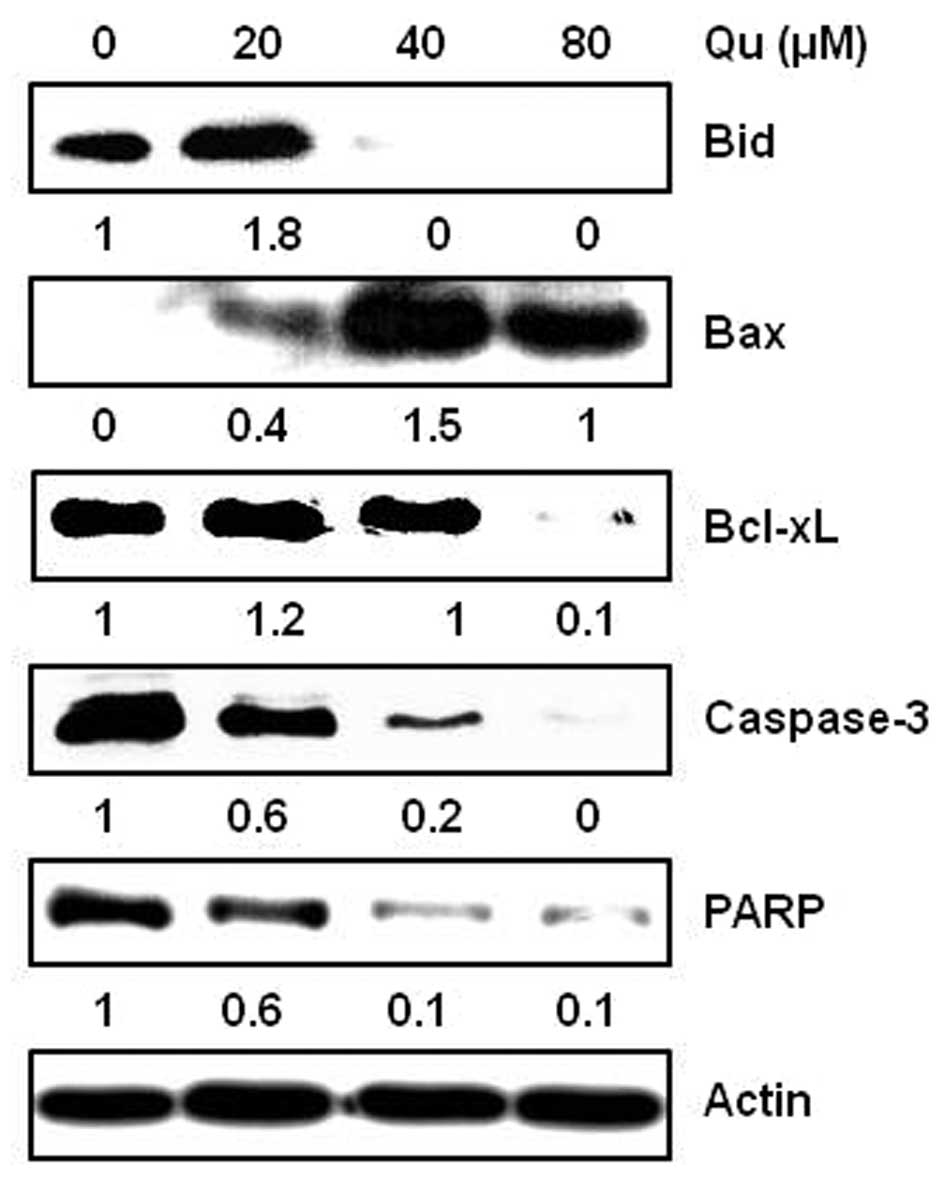
Qu regulates the expression of
anti-apoptotic and apoptotic molecules in MSTO-211H cells
The treatment of cells with Qu regulated the
expression levels of several apoptosis-related proteins. MSTO-211H
cells were treated with various concentrations (20–80 μM) of
Qu for 48 h and harvested. The protein expression levels of Bid,
Bcl-xL, caspase-3, PARP and Bax were analyzed by western blot
analysis. The results showed that Qu induced the activation of Bid,
caspase-3 and PARP, decreased Bcl-xL, and increased Bax levels in
the MSTO-211H cells (Fig. 5).
Discussion
Malignant pleural mesothelioma and the majority of
lung carcinomas are diagnosed at an advanced stage, conferring a
poor prognosis (23,24). The incidence of malignant pleural
mesothelioma is rising rapidly in many countries and it continues
to be a challenging clinical problem. Our results showed that Qu
has potential as a chemopreventive and chemotherapeutic agent for
malignant pleural mesothelioma and that Sp1 is a potential
therapeutic target of Qu.
It has previously been reported that the Sp1 protein
is overexpressed in many human tumors (25) and cancer cell lines (26–30) and a number of studies have
reported that Sp1 is highly expressed in a variety of human tumors,
such as those of the pancreas, colon, lung, breast and prostate
(9), demonstrating that the use
of natural compounds may be used to inhibit Sp1 expression in
cancer. Advances in treatment regimens have only had modest
effects, although gene therapy offers a novel therapeutic approach,
and has been evaluated in a number of clinical trials. Strategies
include the induction of apoptosis, cytokine-based therapy, suicide
gene expression, tumor suppressor gene replacement, various
vaccination approaches, and the adoptive transfer of modified
immune cells. A number of studies have considered the clinical
results, limitations and future directions of gene therapy trials
for thoracic malignancies (23,24).
Various plants and fruits have been reported to have
cancer chemopreventive properties, and thus, investigators have
sought to identify new active phytochemicals (31). Qu is an ubiquitous dietary
flavonoid that has recently been described as a potential
anticancer agent (32) due to its
ability to modulate cell proliferation, survival and
differentiation, targeting key molecules responsible for tumor cell
growth (33,34).
However, the mechanisms responsible for the
antitumor activity of Qu are not yet fully understood. The
induction of apoptosis may be one of the mechanisms as Qu has
antiproliferative effects (35,36) and can induce death via apoptosis
in leukemia (37), breast
(38), hepatoma (39), oral (40) and colon (35) cancer cells. However, no previous
study has been conducted on the effect of Qu on human mesothelioma
cells. In this study, to the best of our knowledge, we demonstrate
for the first time that Qu induces apoptotic cell death by
inhibiting Sp1 protein expression in a time- and dose-dependent
manner in MSTO-211H cells.
Previous studies have identified other flavonoids as
having similar effects as Qu. For example, mithramycin A is one of
the older chemotherapy drugs, and is known to suppress the
expression of Sp1, to inhibit Sp1 binding and to inhibit the
transcription of c-myc, p27, p21, cyclin D1, Mcl-1 and survivin
selectively (13,41–43). Remarkably, our results indicate
that Qu directly binds to Sp1, and possibly prevents the binding of
Sp1 by G-C rich promoters. Our data provide evidence that Qu
inhibits Sp1 expression at the protein and mRNA levels.
Transcriptional response targeting genes containing the Sp1 binding
site in their promoters are involved in a number of cellular
functions ranging from differentiation to cell cycle progression,
proliferation and apoptotic cell death (7). Our results showed that Qu suppressed
Sp1 downstream target genes, including cyclin D1, Mcl-1 and
survivin in MSTO-211H cells by promoter assay and western blot
analyses. The apoptotic effect of Qu on MSTO-211H cells was found
to be induced via the inhibition of Sp1 protein expression in
vitro. Our results from in vitro experiments show that
the Sp1 protein is a major factor of the antitumor effects of Qu in
mesothelioma.
In conclusion, the results from the present study
suggest that Qu has therapeutic and chemopreventative benefits and
that Sp1 be considered a therapeutic target in malignant pleural
mesothelioma and other advanced-stage cancers. Furthermore, our
data suggest that Qu be considered a drug or natural supplement
candidate for the prevention of malignant pleural mesothelioma.
Acknowledgements
This study was supported by the Basic
Science Research program through the National Research Foundation
Korea (NRF) Funded by the Ministry of Education, Science and
Technology (2012-0003226, 2010-0021532), and the Next-Generation
BioGreen 21 Program (PJ008116062011), Rural Development
Administration, Republic of Korea.
References
|
1.
|
KH MieanS MohamedFlavonoid (myricetin,
quercetin, kaempferol, luteolin, and apigenin) content of edible
tropical plantsJ Agric Food
Chem4931063112200110.1021/jf000892m11410016
|
|
2.
|
GN KimHD JangProtective mechanism of
quercetin and rutin using glutathione metabolism on HO-induced
oxidative stress in HepG2 cellsAnn NY Acad
Sci1171530537200910.1111/j.1749-6632.2009.04690.x19723100
|
|
3.
|
KV HirparaP AggarwalAJ MukherjeeN JoshiAC
BurmanQuercetin and its derivatives: synthesis, pharmacological
uses with special emphasis on anti-tumor properties and prodrug
with enhanced bio-availabilityAnticancer Agents Med
Chem9138161200910.2174/18715200978731385519199862
|
|
4.
|
L WangYC TuTW LianJT HungJH YenMJ
WuDistinctive antioxidant and antiinflammatory effects of
flavonolsJ Agric Food
Chem5497989804200610.1021/jf062071917177504
|
|
5.
|
JN JakobsenJB SorensenReview on clinical
trials of targeted treatments in malignant mesotheliomaCancer
Chemother Pharmacol68115201110.1007/s00280-011-1655-321553148
|
|
6.
|
S ToyokuniMechanisms of asbestos-induced
carcinogenesisNagoya J Med Sci711102009
|
|
7.
|
L LiJR DavieThe role of Sp1 and Sp3 in
normal and cancer cell biologyAnn
Anat192275283201010.1016/j.aanat.2010.07.01020810260
|
|
8.
|
LM KongCG LiaoF FeiX GuoJL XingZN
ChenTranscription factor Sp1 regulates expression of
cancer-associated molecule CD147 in human lung cancerCancer
Sci10114631470201010.1111/j.1349-7006.2010.01554.x20384626
|
|
9.
|
UT SankpalS GoodisonM AbdelrahimR
BashaTargeting Sp1 transcription factors in prostate cancer
therapyMed Chem7518525201110.2174/15734061179679920322022994
|
|
10.
|
MC ArcherRole of sp transcription factors
in the regulation of cancer cell metabolismGenes
Cancer2712719201110.1177/194760191142302922207896
|
|
11.
|
J WanBA CarrNS CutlerDL LanzaRN HinesGS
YostSp1 and Sp3 regulate basal transcription of the human CYP2F1
geneDrug Metab
Dispos3312441253200510.1124/dmd.105.00406915860659
|
|
12.
|
X BaiH DengResearch progress on
relationship between transcription factor Sp1 and tumorZhejiang Da
Xue Xue Bao Yi Xue Ban392152202010(In Chinese)
|
|
13.
|
C CulverA MelvinS MudieS RochaHIF-1alpha
depletion results in SP1-mediated cell cycle disruption and alters
the cellular response to chemotherapeutic drugsCell
Cycle1012491260201110.4161/cc.10.8.1532621412054
|
|
14.
|
L GibelliniM PintiM NasiQuercetin and
cancer chemopreventionEvid Based Complement Alternat
Med2011591356201121792362
|
|
15.
|
M LinsalataA OrlandoC MessaMG RefoloF
RussoQuercetin inhibits human DLD-1 colon cancer cell growth and
polyamine biosynthesisAnticancer Res3035013507201020944129
|
|
16.
|
M RussoC SpagnuoloS VolpeA MupoI TedescoGL
RussoQuercetin induced apoptosis in association with death
receptors and fludarabine in cells isolated from chronic
lymphocytic leukaemia patientsBr J
Cancer103642648201010.1038/sj.bjc.6605794
|
|
17.
|
T ThangasamyS SittadjodyGC
MitchellQuercetin abrogates chemoresistance in melanoma cells by
modulating deltaNp73BMC
Cancer10282201010.1186/1471-2407-10-28220540768
|
|
18.
|
J Gonzalez-GallegoMV Garcia-MediavillaS
Sanchez-CamposMJ TunonFruit polyphenols, immunity and
inflammationBr J Nutr104Suppl
3S15S27201010.1017/S000711451000391020955647
|
|
19.
|
JH ShimHS ChoiA
Pugliese(-)-Epigallocatechin gallate regulates CD3-mediated T cell
receptor signaling in leukemia through the inhibition of ZAP-70
kinaseJ Biol
Chem2832837028379200810.1074/jbc.M80220020018687687
|
|
20.
|
TA ZykovaF ZhuX ZhaiResveratrol directly
targets COX-2 to inhibit carcinogenesisMol
Carcinog47797805200810.1002/mc.2043718381589
|
|
21.
|
ES ChoiJH ShimJY JungApoptotic effect of
tolfenamic acid in androgen receptor-independent prostate cancer
cell and xenograft tumor through specificity protein 1Cancer
Sci102742748201110.1111/j.1349-7006.2011.01871.x21241418
|
|
22.
|
JH ShimJA ShinJY JungChemopreventive
effect of tolfenamic acid on KB human cervical cancer cells and
tumor xenograft by downregulating specificity protein 1Eur J Cancer
Prev20102111201110.1097/CEJ.0b013e328341e38f21131823
|
|
23.
|
A VachaniE MoonE WakeamSM AlbeldaGene
therapy for mesothelioma and lung cancerAm J Respir Cell Mol
Biol42385393201010.1165/rcmb.2010-0026RT20160042
|
|
24.
|
A VachaniE MoonE WakeamAR HaasDH StermanSM
AlbeldaGene therapy for lung neoplasmsClin Chest
Med32865885201110.1016/j.ccm.2011.08.006
|
|
25.
|
ES ChoiSD ChoJG JeonNP ChoThe apoptotic
effect of the hexane extract of Rheum undulatum L. in oral
cancer cells through the down-regulation of specificity protein 1
and survivinLab Anim Res271924201121826155
|
|
26.
|
E ChiefariA BrunettiF ArturiIncreased
expression of AP2 and Sp1 transcription factors in human thyroid
tumors: a role in NIS expression regulation?BMC
Cancer235200210.1186/1471-2407-2-3512475396
|
|
27.
|
Y HosoiT WatanabeK NakagawaUp-regulation
of DNA-dependent protein kinase activity and Sp1 in colorectal
cancerInt J Oncol25461468200415254745
|
|
28.
|
L WangD WeiS HuangTranscription factor Sp1
expression is a significant predictor of survival in human gastric
cancerClin Cancer Res963716380200314695137
|
|
29.
|
JC YaoL WangD WeiAssociation between
expression of transcription factor Sp1 and increased vascular
endothelial growth factor expression, advanced stage, and poor
survival in patients with resected gastric cancerClin Cancer
Res1041094117200410.1158/1078-0432.CCR-03-0628
|
|
30.
|
A ZannettiS Del VecchioMV
CarrieroCoordinate up-regulation of Sp1 DNA-binding activity and
urokinase receptor expression in breast carcinomaCancer
Res6015461551200010749121
|
|
31.
|
Y ShuklaJ GeorgeCombinatorial strategies
employing nutraceuticals for cancer developmentAnn NY Acad
Sci1229162175201110.1111/j.1749-6632.2011.06104.x21793852
|
|
32.
|
LL ZaminEC Filippi-ChielaP
Dillenburg-PillaF HornC SalbegoG LenzResveratrol and quercetin
cooperate to induce senescence-like growth arrest in C6 rat glioma
cellsCancer
Sci10016551662200910.1111/j.1349-7006.2009.01215.x19496785
|
|
33.
|
A MurakamiH AshidaJ TeraoMultitargeted
cancer prevention by quercetinCancer
Lett269315325200810.1016/j.canlet.2008.03.046
|
|
34.
|
FO RanellettiN MaggianoFG SerraQuercetin
inhibits p21-RAS expression in human colon cancer cell lines and in
primary colorectal tumorsInt J
Cancer85438445200010.1002/(SICI)1097-0215(20000201)85:3%3C438::AID-IJC22%3E3.0.CO;2-F10652438
|
|
35.
|
MJ van ErkP RoepmanTR van der
LendeIntegrated assessment by multiple gene expression analysis of
quercetin bioactivity on anticancer-related mechanisms in colon
cancer cells in vitroEur J Nutr44143156200515309432
|
|
36.
|
WH WatsonJ CaiDP JonesDiet and
apoptosisAnnu Rev Nutr20485505200010.1146/annurev.nutr.20.1.485
|
|
37.
|
SC ShenYC ChenFL HsuWR LeeDifferential
apoptosis-inducing effect of quercetin and its glycosides in human
promyeloleukemic HL-60 cells by alternative activation of the
caspase 3 cascadeJ Cell
Biochem8910441055200310.1002/jcb.1055912874837
|
|
38.
|
RL SinghalYA YehN PrajaE OlahGW Sledge JrG
WeberQuercetin down-regulates signal transduction in human breast
carcinoma cellsBiochem Biophys Res
Commun208425431199510.1006/bbrc.1995.13557887960
|
|
39.
|
YS ChiHG JongKH SonHW ChangSS KangHP
KimEffects of naturally occurring prenylated flavonoids on enzymes
metabolizing arachidonic acid: cyclooxygenases and
lipoxygenasesBiochem
Pharmacol6211851191200110.1016/S0006-2952(01)00773-011705451
|
|
40.
|
CS OngE TranTT NguyenQuercetin-induced
growth inhibition and cell death in nasopharyngeal carcinoma cells
are associated with increase in Bad and hypophosphorylated
retinoblastoma expressionsOncol Rep117277332004
|
|
41.
|
SW BlumeRC SnyderR RayS ThomasCA KollerDM
MillerMithramycin inhibits SP1 binding and selectively inhibits
transcriptional activity of the dihydrofolate reductase gene in
vitro and in vivoJ Clin
Invest8816131621199110.1172/JCI1154741834700
|
|
42.
|
S ChintharlapalliS PapineniP LeiS PathiS
SafeBetulinic acid inhibits colon cancer cell and tumor growth and
induces proteasome-dependent and -independent downregulation of
specificity proteins (Sp) transcription factorsBMC
Cancer11371201110.1186/1471-2407-11-371
|
|
43.
|
M PietrzakM
Puzianowska-Kuznickap53-dependent repression of the human MCL-1
gene encoding an anti-apoptotic member of the BCL-2 family: the
role of Sp1 and of basic transcription factor binding sites in the
MCL-1 promoterBiol Chem389383393200810.1515/BC.2008.03918208354
|