Introduction
In the present study, we established an in
vitro model in which primary cultures of fetal fibroblasts from
murine origin (PFC) were subjected for 65 h to simulated
microgravity, chronic irradiation or a combination. Genome-wide
gene expression changes were thereafter assessed by microarrays.
For microgravity simulation, we used the random positioning machine
(RPM), which is one of the most widely used instruments for this
purpose and has proven valuable in many cell types (1–6).
As far as cosmic radiation is concerned, simulating the wide
variety of ions ranging from low to very high energies encountered
in space is problematic, particularly if irradiation is combined
with microgravity simulation models. At present, no facility offers
the possibility of producing chronic exposures of very high-energy
beams consisting of multiple charged particles. We therefore used a
source of californium Cf-252 for low-dose rate long-term exposure
consisting of a mixture of high-linear energy transfer (LET)
neutrons and low-LET gamma-rays (7).
The large amount of data generated with a
high-throughput technology such as microarrays constitutes a
double-edged sword: whole expression pattern may be recorded, but
extracting the relevant information becomes more challenging
(8,9). To overcome this problem, analysis
tools have been developed, such as single gene statistical analysis
methods (SGA), which are widely used to determine the
differentially expressed genes, and the gene set enrichment
analysis (GSEA), which aims to identify gene expression differences
in groups of genes, for instance in those acting synergistically in
a cell process (9,10). The two analytical methods were
used concomitantly in this study.
Materials and methods
Cell culture
All the animals were handled following the Belgian
legislation after approval by the appropriate Ethics Committees
(agreement number 08-002). BALB/cJ Rj (Janvier Laboratories,
Saint-Berthevin, France) fetuses (three males and three females)
originating from two different litters were dissected 17 days
post-conception (day 0 being the fertilization day). Their skin was
harvested and mechanically dissociated. The obtained tissue was
enzymatically digested for 1 h at 37°C in phosphate-buffered saline
(PBS; N.V. Invitrogen SA, Merelbeke, Belgium) solution containing 1
mg/ml of collagenase/dispase (Roche, Mannheim, Germany) and 5 mg/ml
of trypsin 2,000 E/g (Merck KGaA, Darmstadt, Germany). The
enzymatic reaction was subsequently stopped by adding fetal bovine
serum (FBS; N.V. Invitrogen SA). The obtained cell suspension was
subsequently centrifuged for 10 min at 350 × g and the cells were
seeded in 6-well plates in F12 medium supplemented with 20% FBS and
1% penicillin/streptomycin (both from N.V. Invitrogen SA), one
fetus skin in each well. The cells were allowed to grow for up to 3
or 4 passages at 37°C (5%, CO2) and were subsequently
frozen in FBS with 10% dimethyl sulfoxide (Sigma-Aldrich, St.
Louis, MO, USA). The primary cultures were then thawed and allowed
to grow for two weeks. The cells were seeded at a density of
×105 cells in 12.5 cm2 flasks and allowed to
adhere for 24 h prior to treatment.
Simulation of space conditions
Exposure to simulated space conditions included
microgravity simulation using the desktop RPM (Dutch Space, Leiden,
The Netherlands) and ionizing radiation (IR) (7). The exposure lasted for a period of
65 h. Four treatment conditions were used: controls (CTRL),
microgravity simulation (RPM), irradiation and a combination of the
treatment methods (RPM and IR). For microgravity simulation, the
flasks were completely filled with medium, sealed and placed on the
RPM at a rotational velocity between 55 and 65°/sec. Direction,
speed and interval were set as random. The CTRL were placed in the
same incubator under the same conditions as the treated samples.
For chronic low-dose irradiation, the cells were exposed to a
mixture of neutrons (98.2%) and gamma-rays (1.8%) directly or
indirectly originating from a Cf-252 source were placed at 4.13 m
from the incubator. The dosimetry was performed with bubble
detectors as previously described (11) for neutron irradiation and with 600
cc ionization chamber (NE) coupled with a Farmer electrometer for
gamma-rays. The total dose received was 55.94±19.70 mSv (862
μSv/h), which approximately corresponds to 35 times the dose rate
measured on the International Space Station (ISS) (12), the total dose corresponding
approximately to a stay of 100 days in the ISS.
RNA extraction
Immediately after treatment, adherent cells were
washed in PBS, lysed in 350 ml of AllPrep DNA/RNA/Protein Mini kit
lysis buffer (Qiagen, Hilden, Germany) and frozen at −80°C. RNA was
extracted using the same kit and its concentration was measured
using the Nanodrop spectrophotometer (Thermo Scientific, Waltham,
MA, USA) while its quality (RNA integrity number, RIN) was
determined with Agilent’s lab-on-chip Bioanalyzer 2100 (Agilent
Technologies, Inc., Palo Alto, CA, USA). All the RNA samples had a
RIN value of >9.0.
Affymetrix microarrays and data
analysis
The RNA was treated using the GeneChip WT cDNA
Synthesis and Amplification kit (Affymetrix, Santa Clara, CA, USA)
according to the manufacturer’s instructions. The resulting RNA was
hybridized onto Affymetrix Mouse Gene 1.0 ST arrays.
Raw data (.cel-files) were imported at exon level in
Partek Genomics Suite v6.5 (Partek Incorporated, St. Louis, MO,
USA). Briefly, robust Multi-array Average (RMA) background
correction was applied, data were normalized by quantile
normalization and probe set summarization was performed using the
median polish method. Gene summarization was performed using
One-Step Tukey’s Biweight method. These data were further analyzed
with the Partek Genomics Suite software for SGA and by the GSEA
software (v2.0, Broad Institute of Harvard and MIT, Cambridge, MA,
USA).
For the single gene method, taking into
consideration the scan date (also available for the litter), the
fetus, the gender and the treatment as factors, a four-way ANOVA
was performed to determine the genes that had a significantly
altered expression for different conditions. For the pathway
analysis, KEGG and PathArt databases were analyzed with ArrayTrack
v3.3.0 (National Center for Toxicological Research, Jefferson, AR,
USA).
For the GSEA, a selection of 144 gene sets from gene
ontology (GO) databases was based on biological relevance (Table I). Gene sets were considered to be
significantly differently regulated with a false discovery rate
(FDR) when q<0.05.
 | Table IList of the 144 gene sets selected for
GSEA. |
Table I
List of the 144 gene sets selected for
GSEA.
| Gene set
description | Gene Ontology |
|---|
| Actin binding | GO:0003779 |
| Actin
cytoskeleton | GO:0015629 |
| Activation of JNK
activity | GO:0007257 |
| Activation of MAPK
activity | GO:0000187 |
| Adherens
junction | GO:0005912 |
| Anti-apoptosis | GO:0006916 |
| Antioxidant
activity | GO:0016209 |
| Apoptosis GO | GO:0006915 |
| Base excision
repair | GO:0006284 |
| Calcium ion
binding | GO:0005509 |
| Calcium ion
transport | GO:0006816 |
| Caspase
activation | GO:0006919 |
| Cell-cell
adhesion | GO:0016337 |
| Cell-cell
signaling | GO:0007267 |
| Cell cycle
arrest | GO:0007050 |
| Cell cycle | GO:0007049 |
| Cell cycle
process | GO:0022402 |
| Cell junction | GO:0030054 |
| Cell matrix
adhesion | GO:0007160 |
| Cellular
respiration | GO:0045333 |
| Centrosome | GO:0005813 |
| Chaperone
binding | GO:0051087 |
| Chromatin | GO:0000785 |
| Chromosome | GO:0005694 |
| Collagen | GO:0005581 |
| Cortical
cytoskeleton | GO:0030863 |
| Cytokine
activity | GO:0005125 |
| Cytoskeletal
protein binding | GO:0008092 |
| Cytoskeleton | GO:0005856 |
| DNA damage
checkpoint | GO:0000077 |
| DNA integrity
checkpoint | GO:0031570 |
| DNA repair | GO:0006281 |
| Double-strand break
repair | GO:0006302 |
| Electron
transport | GO:0006118 |
| Embryonic
development | GO:0009790 |
| Endoplasmic
reticulum | GO:0005783 |
| Excretion | GO:0007588 |
| Extracellular
matrix | GO:0031012 |
| Focal adhesion | GO:0005925 |
| G-protein coupled
receptor activity | GO:0004930 |
| G-protein coupled
receptor protein signaling pathway | GO:0007186 |
| G-protein signaling
coupled to IP3 second messenger phospholipase C activating | GO:0007200 |
| G1 phase | GO:0051318 |
| G1/S transition of
mitotic cell cycle | GO:0000082 |
| G2/M transition of
mitotic cell cycle | GO:0000086 |
| Glutathione
transferase activity | GO:0004364 |
| Golgi
apparatus | GO:0005794 |
| GTPase regulator
activity | GO:0030695 |
| Histone
modification | GO:0016570 |
| Hormone
activity | GO:0005179 |
| Inositol or
phosphatidylinositol kinase activity | GO:0004428 |
| Inositol or
phosphatidylinositol phosphatase activity | GO:0004437 |
| Inositol or
phosphatidylinositol phosphodiesterase activity | GO:0004434 |
| Insulin receptor
signaling pathway | GO:0008286 |
| Integrin
binding | GO:0005178 |
| Intercellular
junction | GO:0005911 |
| Ion channel
activity | GO:0005216 |
| JAK/STAT
cascade | GO:0007259 |
| JNK cascade | GO:0007254 |
| Lamellipodium | GO:0030027 |
| Lipid binding | GO:0008289 |
| M phase | GO:0000279 |
| Magnesium ion
binding | GO:0000287 |
| MAP kinase
activity | GO:0004707 |
| MAPKKK cascade | GO:0000165 |
| Microtubule | GO:0005874 |
| Microtubule
cytoskeleton | GO:0015630 |
| Mitochondrial inner
membrane | GO:0005743 |
| Mitochondrial
respiratory chain | GO:0005746 |
| Mitochondrion | GO:0005739 |
| Motor activity | GO:0003774 |
| Negative regulation
of apoptosis | GO:0043066 |
| Negative regulation
of cell adhesion | GO:0007162 |
| Negative regulation
of cell cycle | GO:0045786 |
| Negative regulation
of cell proliferation | GO:0008285 |
| Negative regulation
of cellular metabolic process | GO:0031324 |
| Negative regulation
of signal transduction | GO:0009968 |
| Negative regulation
of transcription | GO:0016481 |
| Negative regulation
of translation | GO:0017148 |
| Nuclear pore | GO:0005643 |
| Nucleolus | GO:0005730 |
| Nucleus | GO:0005634 |
| Oligosaccharide
metabolic process | GO:0009311 |
|
Phosphoinositide-mediated signaling | GO:0048015 |
| Phospholipase
activity | GO:0004620 |
| Phospholipid
binding | GO:0005543 |
|
Phosphorylation | GO:0016310 |
| Positive regulation
of caspase activity | GO:0043280 |
| Positive regulation
of cell adhesion | GO:0045785 |
| Positive regulation
of cell cycle | GO:0045787 |
| Positive regulation
of cell proliferation | GO:0008284 |
| Positive regulation
of JNK activity | GO:0043507 |
| Positive regulation
of MAP kinase activity | GO:0043406 |
| Positive regulation
of protein metabolic process | GO:0051247 |
| Positive regulation
of signal transduction | GO:0009967 |
| Positive regulation
of transcription | GO:0045941 |
| Positive regulation
of translation | GO:0045727 |
| Post-translational
protein modification | GO:0043687 |
| Potassium ion
transport | GO:0006813 |
| Programmed cell
death | GO:0012501 |
| Protein
folding | GO:0006457 |
| Protein kinase
activity | GO:0004672 |
| Protein kinase
cascade | GO:0007243 |
| Protein metabolic
process | GO:0019538 |
| Protein
modification process | GO:0006464 |
| Protein/RNA complex
assembly | GO:0022618 |
| Protein
serine/threonine kinase activity | GO:0004674 |
| Protein
ubiquitination | GO:0016567 |
| Proteolysis | GO:0006508 |
| RAS GTPase
activator activity | GO:0005099 |
| RAS GTPase
binding | GO:0017016 |
| Receptor
binding | GO:0005102 |
| Regulation of
apoptosis | GO:0042981 |
| Replication
fork | GO:0005657 |
| Respiratory chain
complex I | GO:0045271 |
| Response to DNA
damage stimulus | GO:0006974 |
| Response to
ionizing radiation | GO:0010212 |
| Response to
radiation | GO:0009314 |
| Response to
stress | GO:0006950 |
| RHO GTPase
activator activity | GO:0005100 |
| RHO protein signal
transduction | GO:0007266 |
| Rhodopsin-like
receptor activity | GO:0001584 |
| RNA helicase
activity | GO:0003724 |
| RNA processing | GO:0006396 |
| RNA splicing | GO:0008380 |
| Ruffle | GO:0001726 |
| S phase | GO:0051320 |
| Second
messenger-mediated signaling | GO:0019932 |
| Small conjugated
protein ligase activity | GO:0019787 |
| Small
GTPase-mediated signal transduction | GO:0007264 |
| Sodium channel
activity | GO:0005272 |
| Spindle | GO:0005819 |
| Spliceosome | GO:0005681 |
| Structural
constituent of cytoskeleton | GO:0005200 |
| Structural
constituent of ribosome | GO:0003735 |
| Tight junction | GO:0005923 |
| Transcription | GO:0006350 |
| Translation | GO:0006412 |
| Transmembrane
receptor protein kinase activity | GO:0019199 |
| Transmembrane
transporter activity | GO:0022857 |
| T-RNA metabolic
process | GO:0006399 |
| Ubiquitin
cycle | GO:0006512 |
| Ubiquitin protein
ligase activity | GO:0004842 |
| Voltage-gated
channel activity | GO:0022832 |
Results
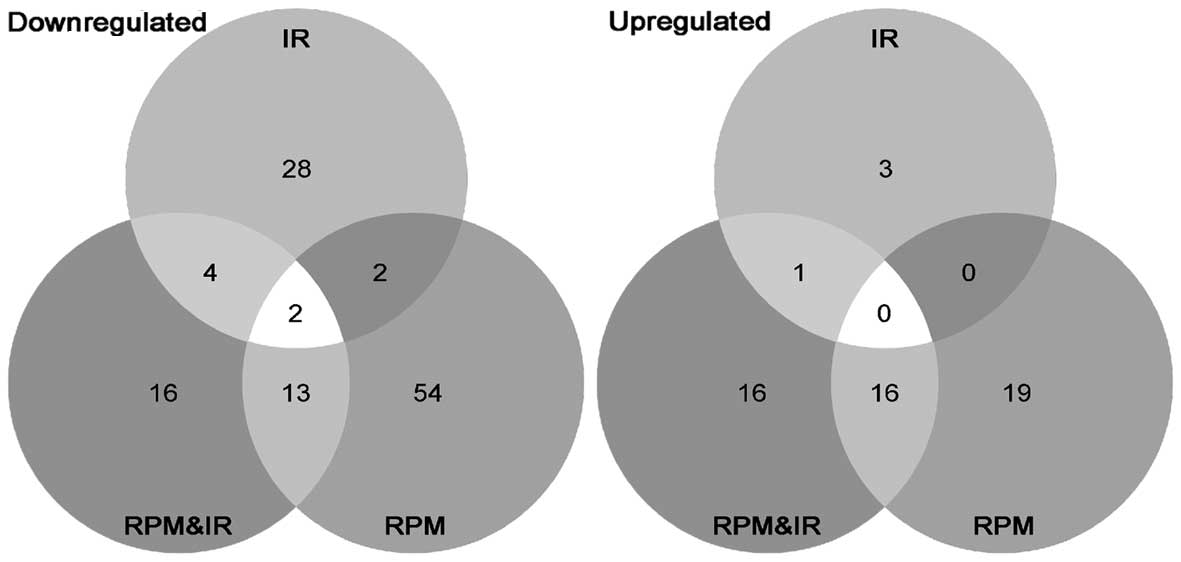
Single gene analysis revealed that 119 genes were
downregulated and 55 genes were upregulated by >1.5-fold change
(unadjusted p-value <0.01) across all the treatments (Fig. 1 and an exhaustive list of the
differentially expressed genes can be found in Table II). KEGG and PathArt databases
indicated that the 54 genes that were downregulated only by RPM
treatment were mostly involved in cell cycle regulation (p53- and
p21-mediated pathways), in cytoskeleton modeling, cell junctions
and cell signaling via integrins, IL-1, and TGF-β. Within the list
of individual genes that were downregulated after IR or RPM and IR
treatments, no clear pathway was found. On the other hand, in the
52 genes that were upregulated following RPM and RPM and IR
treatments, interleukin signaling (IL-11 and MMP) and glutathione
metabolism were the most prominent pathways affected. Some genes
were differentially expressed by RPM and RPM and IR, however, only
a few genes were common between IR and RPM and IR. Six genes were
upregulated (S1p3, Rab11b, Ptger3, Vldlr, Cnn1 and Serping1) and
only one predicted gene of unknown function was downregulated
(Gm13668) in both irradiated treatments (IR and RPM and IR). The
upregulated genes were mostly membrane proteins, G-protein coupled
(S1p3 and Ptger3) or involved in ligand endocytosis (Rab11b and
Vldlr). Cnn1 and Serping1, involved in cytoskeleton organization
and peptidase inhibition, respectively, were both upregulated in
all the treatments, including RPM.
 | Table IIDown- and upregulated genes following
IR, RPM or RPM and IR treatments (p<0.001, fold change
>1.5). |
Table II
Down- and upregulated genes following
IR, RPM or RPM and IR treatments (p<0.001, fold change
>1.5).
| A, Down- and
upregulated genes following IR |
|---|
|
|---|
| Gene symbol | GenBank | p-value | FC |
|---|
| Rab11b | NM_008997 | 5,48E-03 | −2,026 |
| Csgalnact1 | NM_172753 | 9,04E-03 | −1,989 |
| Smarca5 | NM_053124 | 4,08E-03 | −1,986 |
| Tceb3 | NM_013736 | 6,85E-03 | −1,953 |
| Serping1 | NM_009776 | 1,39E-03 | −1,948 |
| Ppp1r2 | NM_025800 | 9,20E-03 | −1,801 |
| Ptgfrn | NM_011197 | 4,38E-03 | −1,801 |
| Dnaja1 | NM_008298 | 6,13E-03 | −1,772 |
| Arhgap24 | NM_029270 | 6,13E-03 | −1,767 |
| Thra | NM_178060 | 3,67E-04 | −1,728 |
| Itga8 | NM_001001309 | 9,74E-03 | −1,718 |
| Gpr108 | NM_030084 | 2,40E-03 | −1,696 |
| Zfp346 | NM_012017 | 1,54E-04 | −1,672 |
| Rbmx | NM_011252 | 1,04E-03 | −1,655 |
| B4galt6 | NM_019737 | 9,25E-03 | −1,638 |
| BC003331 | NM_145511 | 5,04E-03 | −1,637 |
| Vldlr | NM_013703 | 3,68E-03 | −1,636 |
| Unc93b1 | NM_019449 | 5,57E-04 | −1,625 |
| Pip4k2a | NM_008845 | 3,79E-04 | −1,622 |
| Mgll | NM_001166251 | 2,52E-03 | −1,620 |
| BC005624 | NM_144885 | 2,10E-03 | −1,619 |
| S1pr3 | NM_010101 | 2,64E-03 | −1,613 |
| Prkcd | NM_011103 | 3,32E-03 | −1,583 |
| Cnn1 | NM_009922 | 4,26E-03 | −1,575 |
| P2ry2 | NM_008773 | 6,80E-03 | −1,566 |
| Saps1 | NM_172894 | 8,65E-03 | −1,566 |
| Casc4 | NM_177054 | 4,45E-03 | −1,559 |
| Opa1 | NM_133752 | 7,47E-03 | −1,552 |
| Emb | NM_010330 | 5,84E-04 | −1,551 |
| Cyb5d1 | NM_001045525 | 5,23E-03 | −1,549 |
| Ptger3 | NM_011196 | 1,41E-03 | −1,549 |
| Usp30 | NM_001033202 | 1,36E-03 | −1,543 |
| Tbc1d2b | NM_194334 | 6,00E-03 | −1,539 |
| Cyld | NM_001128169 | 2,57E-03 | −1,530 |
| Trip4 | NM_019797 | 8,21E-03 | −1,520 |
| Luzp1 | NM_024452 | 9,75E-03 | −1,502 |
| Gm13668 | XR_032757 | 6,87E-04 | 1,856 |
| Hist1h2ao | NM_001177544 | 3,69E-03 | 1,710 |
| Bmyc | NM_023326 | 3,25E-03 | 1,575 |
| 4930458L03Rik | NM_030047 | 1,32E-03 | 1,523 |
|
| B, Down- and
upregulated genes following RPM |
|
| Dmpk | NM_032418 | 2,73E-05 | −2,522 |
| Myh10 | NM_175260 | 1,51E-03 | −2,485 |
| Myh9 | NM_022410 | 2,08E-03 | −2,432 |
| Maob | NM_172778 | 1,48E-04 | −2,335 |
| Slc38a4 | NM_027052 | 9,81E-04 | −2,270 |
| Cnn1 | NM_009922 | 4,53E-05 | −2,147 |
| Adh1 | NM_007409 | 2,01E-04 | −2,126 |
| Serping1 | NM_009776 | 5,85E-04 | −2,095 |
| Actg2 | NM_009610 | 3,08E-05 | −2,089 |
| Ccnb2 | NM_007630 | 2,94E-04 | −2,071 |
| Kif20a | NM_001166406 | 1,35E-04 | −2,023 |
| Gjb2 | NM_008125 | 1,16E-04 | −2,014 |
| Anln | NM_028390 | 8,66E-04 | −2,001 |
| Nfix | NM_001081981 | 3,95E-03 | −1,964 |
| Itga8 | NM_001001309 | 2,19E-03 | −1,963 |
| Pygb | NM_153781 | 1,46E-03 | −1,913 |
| Bub1 | NM_001113179 | 3,71E-05 | −1,881 |
| Ly6c1 | NM_010741 | 9,89E-04 | −1,879 |
| ND4L |
ENSMUST00000084013 | 2,03E-05 | −1,843 |
| Myl9 | NM_172118 | 1,95E-04 | −1,830 |
| Actn4 | NM_021895 | 8,48E-03 | −1,819 |
| Itgbl1 | NM_145467 | 8,49E-03 | −1,814 |
| Efemp1 | NM_146015 | 6,17E-04 | −1,801 |
| D17H6S56E-5 | L78788 | 2,29E-07 | −1,791 |
| Plk1 | NM_011121 | 1,55E-03 | −1,774 |
| ND4L |
ENSMUST00000084013 | 3,76E-05 | −1,750 |
| Susd2 | NM_027890 | 2,62E-04 | −1,736 |
| Ly6c2 | NM_001099217 | 7,26E-04 | −1,731 |
| Ucp2 | NM_011671 | 4,37E-04 | −1,717 |
| Cenpa | NM_007681 | 3,25E-03 | −1,713 |
| Nuf2 | NM_023284 | 6,69E-04 | −1,711 |
| Rbmx | NM_011252 | 7,19E-04 | −1,692 |
| Kif2c | NM_134471 | 2,28E-03 | −1,687 |
| Rpl22l1 | NM_026517 | 9,88E-03 | −1,678 |
| Ly6a | NM_010738 | 8,47E-03 | −1,671 |
| Pkp2 | NM_026163 | 1,21E-04 | −1,667 |
| Tgfb1i1 | NM_009365 | 6,54E-03 | −1,652 |
| Acta1 | NM_009606 | 7,19E-06 | −1,644 |
| Gas2l3 | NM_001033331 | 5,26E-04 | −1,643 |
| Lrrc17 | NM_028977 | 4,37E-03 | −1,642 |
| 2810417H13Rik | NM_026515 | 3,58E-03 | −1,640 |
| Lpar4 | NM_175271 | 3,20E-03 | −1,639 |
| Dlgap5 | NM_144553 | 1,76E-03 | −1,622 |
| Hgf | NM_010427 | 1,71E-03 | −1,611 |
| Trp53inp2 | NM_178111 | 1,30E-03 | −1,605 |
| Cyb5r3 | NM_029787 | 1,06E-03 | −1,603 |
| Mfap2 | NM_008546 | 6,77E-04 | −1,600 |
| Cyp1b1 | NM_009994 | 5,70E-03 | −1,597 |
| Trpv2 | NM_011706 | 4,75E-03 | −1,596 |
| Kif23 | NM_024245 | 1,56E-03 | −1,591 |
| Sh3pxd2a | NM_008018 | 1,37E-03 | −1,566 |
| ND2 |
ENSMUST00000082396 | 1,35E-03 | −1,564 |
| Tgfb3 | NM_009368 | 1,15E-03 | −1,562 |
| Scd2 | NM_009128 | 5,24E-03 | −1,554 |
| Dner | NM_152915 | 1,80E-03 | −1,546 |
| Pdgfrl | NM_026840 | 4,88E-04 | −1,543 |
| Cenpm | NM_025639 | 6,47E-03 | −1,539 |
| Ppp1r3c | NM_016854 | 1,04E-03 | −1,536 |
| Fam114a1 | NM_026667 | 1,68E-03 | −1,533 |
| D2Ertd750e | NM_026412 | 7,48E-04 | −1,533 |
| Nkd2 | NM_028186 | 7,87E-03 | −1,531 |
| Nov | NM_010930 | 9,41E-03 | −1,529 |
| Tgm2 | NM_009373 | 2,62E-03 | −1,525 |
| Nucb2 | NM_001130479 | 7,36E-03 | −1,518 |
| 5730469M10Rik | BC056635 | 1,03E-03 | −1,516 |
| Ccna2 | NM_009828 | 8,65E-03 | −1,514 |
| Maged2 | NM_030700 | 6,56E-03 | −1,512 |
| Eif4b | NM_145625 | 7,83E-03 | −1,512 |
| Sepx1 | NM_013759 | 2,65E-04 | −1,506 |
| Shisa4 | NM_175259 | 5,60E-03 | −1,503 |
| St3gal5 | NM_011375 | 8,29E-03 | −1,502 |
| Fhl5 | NM_021318 | 2,32E-04 | −1,502 |
| Serpinb9e | NM_011456 | 2,02E-03 | 2,514 |
| Gstα1 | NM_008181 | 1,59E-05 | 2,232 |
| Taf1d | BC056964 | 1,32E-03 | 2,223 |
| Gstα1 | NM_008181 | 2,15E-05 | 2,210 |
| Prl2c3 | NM_011118 | 2,70E-05 | 2,209 |
| Snhg1 | AK051045 | 5,45E-06 | 2,192 |
| Prl2c5 | NM_181852 | 2,79E-04 | 2,179 |
| Malat1 | NR_002847 | 1,90E-04 | 2,139 |
| Il1rl1 | NM_001025602 | 2,81E-04 | 2,061 |
| Snhg1 | AK051045 | 1,78E-05 | 1,967 |
| Gm10639 | NM_001122660 | 2,00E-04 | 1,908 |
| Sema7a | NM_011352 | 5,57E-04 | 1,870 |
| Lce1h | NM_026335 | 6,44E-03 | 1,841 |
| Taf1d | BC056964 | 7,18E-04 | 1,812 |
| Crct1 | NM_028798 | 3,49E-04 | 1,802 |
| Gm8074 | XM_983501 | 3,90E-04 | 1,799 |
| Lsm1 | NM_026032 | 1,31E-03 | 1,794 |
| 2310002L13Rik |
ENSMUST00000025390 | 4,85E-04 | 1,771 |
| Sirt7 | NM_153056 | 4,15E-04 | 1,760 |
| Serpinb9b | NM_011452 | 4,07E-06 | 1,734 |
| Snord14e | NR_028275 | 7,22E-04 | 1,704 |
| Gstα2 | NM_008182 | 7,50E-07 | 1,700 |
| Ppbp | NM_023785 | 5,04E-03 | 1,691 |
| Hsd3b6 | NM_013821 | 1,21E-04 | 1,684 |
| Snord14d | NR_028274 | 7,27E-04 | 1,679 |
| Hmox1 | NM_010442 | 5,24E-06 | 1,678 |
| Clcf1 | NM_019952 | 1,97E-04 | 1,671 |
| Snord14d | NR_028274 | 7,39E-04 | 1,667 |
| Procr | NM_011171 | 2,00E-03 | 1,649 |
| Hist1h4i | NM_175656 | 4,77E-03 | 1,635 |
| Dusp4 | NM_176933 | 4,07E-03 | 1,626 |
| Mmp10 | NM_019471 | 6,51E-04 | 1,587 |
| Cops3 | NM_011991 | 1,24E-03 | 1,584 |
| Gas5 | NR_002840 | 3,87E-03 | 1,573 |
| Chrna1 | NM_007389 | 1,19E-03 | 1,565 |
| Ifrd1 | NM_013562 | 1,44E-03 | 1,556 |
| D4Wsu53e | BC043057 | 2,60E-03 | 1,515 |
| S100a7a | NM_199422 | 8,26E-03 | 1,513 |
| Scarna17 | NR_028560 | 6,25E-04 | 1,512 |
| Scarna17 | NR_028560 | 6,25E-04 | 1,512 |
|
| C, Down- and
upregulated genes following RPM and IR |
|
| Cnn1 | NM_009922 | 6,32E-06 | −2,494 |
| Serping1 | NM_009776 | 1,65E-04 | −2,337 |
| Dmpk | NM_032418 | 1,36E-04 | −2,206 |
| Actg2 | NM_009610 | 1,44E-05 | −2,203 |
| Adh1 | NM_007409 | 2,24E-04 | −2,107 |
| Rab11b | NM_008997 | 4,88E-03 | −2,051 |
| Itgbl1 | NM_145467 | 2,48E-03 | −2,041 |
| Gjb2 | NM_008125 | 1,20E-04 | −2,009 |
| Srpx | NM_016911 | 1,74E-05 | −1,882 |
| Myl9 | NM_172118 | 1,72E-04 | −1,844 |
| S1pr3 | NM_010101 | 4,04E-04 | −1,824 |
| Maob | NM_172778 | 2,99E-03 | −1,814 |
| Tmem45a | NM_019631 | 3,16E-04 | −1,793 |
| Pdgfrl | NM_026840 | 3,28E-05 | −1,7722 |
| Nov | NM_010930 | 1,36E-03 | −1,749 |
| Pigc | NM_026078 | 5,96E-03 | −1,691 |
| Il1r1 | NM_008362 | 5,64E-03 | −1,690 |
| Vldlr | NM_013703 | 2,37E-03 | −1,687 |
| Susd2 | NM_027890 | 6,17E-04 | −1,652 |
| Ptger3 | NM_011196 | 4,43E-04 | −1,651 |
| Lysmd3 | NM_030257 | 2,93E-03 | −1,650 |
| Fhl1 | NM_001077361 | 1,90E-08 | −1,629 |
| Cyp1b1 | NM_009994 | 4,48E-03 | −1,625 |
| Plk1 | NM_011121 | 6,41E-03 | −1,600 |
| St3gal5 | NM_011375 | 3,71E-03 | −1,583 |
| Rab13 | NM_026677 | 7,09E-04 | −1,581 |
| Snta1 | NM_009228 | 4,11E-05 | −1,577 |
| Aqp1 | NM_007472 | 5,14E-03 | −1,556 |
| Cpa6 | NM_177834 | 4,01E-03 | −1,554 |
| Nosip | NM_025533 | 1,35E-03 | −1,540 |
| Pla2g16 | NM_139269 | 5,13E-03 | −1,532 |
| Lmod1 | NM_053106 | 3,01E-03 | −1,523 |
| Zcchc17 | NM_153160 | 4,13E-03 | −1,519 |
| Islr | NM_012043 | 1,27E-03 | −1,509 |
| 6330406I15Rik | BC116246 | 1,22E-04 | −1,502 |
| Serpinb9e | NM_011456 | 7,91E-04 | 2,818 |
| Slc40a1 | NM_016917 | 1,83E-05 | 2,670 |
| Taf1d | BC056964 | 2,85E-04 | 2,595 |
| Snhg1 | AK051045 | 8,57E-06 | 2,126 |
| Prl2c3 | NM_011118 | 8,34E-05 | 2,037 |
| Gstα1 | NM_008181 | 1,21E-04 | 1,938 |
| Procr | NM_011171 | 1,83E-04 | 1,935 |
| Gstα1 | NM_008181 | 1,60E-04 | 1,921 |
| Mmp13 | NM_008607 | 5,88E-04 | 1,894 |
| Malat1 | NR_002847 | 1,05E-03 | 1,873 |
| Serpinb9g | NM_011455 | 5,62E-03 | 1,865 |
| Snhg1 | AK051045 | 5,03E-05 | 1,848 |
| Serpinb9g | NM_011455 | 5,24E-03 | 1,801 |
| 310002L13Rik |
ENSMUST00000025390 | 4,28E-04 | 1,785 |
| Gm8074 | XM_983501 | 6,00E-04 | 1,750 |
| Myc | NM_010849 | 1,98E-05 | 1,748 |
| Peg10 | NM_130877 | 1,83E-03 | 1,746 |
| Mamdc2 | NM_174857 | 7,44E-04 | 1,744 |
| Il11 | NM_008350 | 9,82E-03 | 1,728 |
| Mmp3 | NM_010809 | 8,35E-03 | 1,724 |
| Fabp7 | NM_021272 | 2,79E-03 | 1,693 |
| Serpinb9b | NM_011452 | 7,99E-06 | 1,681 |
| Taf1d | BC056964 | 2,40E-03 | 1,667 |
| Gmnn | AF068780 | 7,71E-03 | 1,620 |
| Gm10639 | NM_001122660 | 2,64E-03 | 1,610 |
| Dusp4 | NM_176933 | 5,31E-03 | 1,596 |
| Bcl2l11 | NM_207680 | 9,00E-03 | 1,595 |
| Ctu1 | NM_145582 | 1,01E-03 | 1,594 |
| Gstα2 | NM_008182 | 3,74E-06 | 1,591 |
| Hsd3b6 | NM_013821 | 5,25E-04 | 1,561 |
| Gm13668 | XR_032757 | 8,44E-03 | 1,553 |
| Ang2 | NM_007449 | 1,54E-03 | 1,548 |
| Scarna17 | NR_028560 | 3,97E-04 | 1,545 |
| Scarna17 | NR_028560 | 3,97E-04 | 1,545 |
| Hmox1 | NM_010442 | 3,84E-05 | 1,541 |
| Serpinb9f | NM_183197 | 9,11E-04 | 1,540 |
| Opa3 | NM_207525 | 1,23E-03 | 1,540 |
| Ormdl3 | NM_025661 | 8,94E-03 | 1,505 |
In contrast to the results obtained by SGA, GSEA
revealed a high impact of IR on coordinately differentially
expressed genes. A total of 63 gene sets were significantly
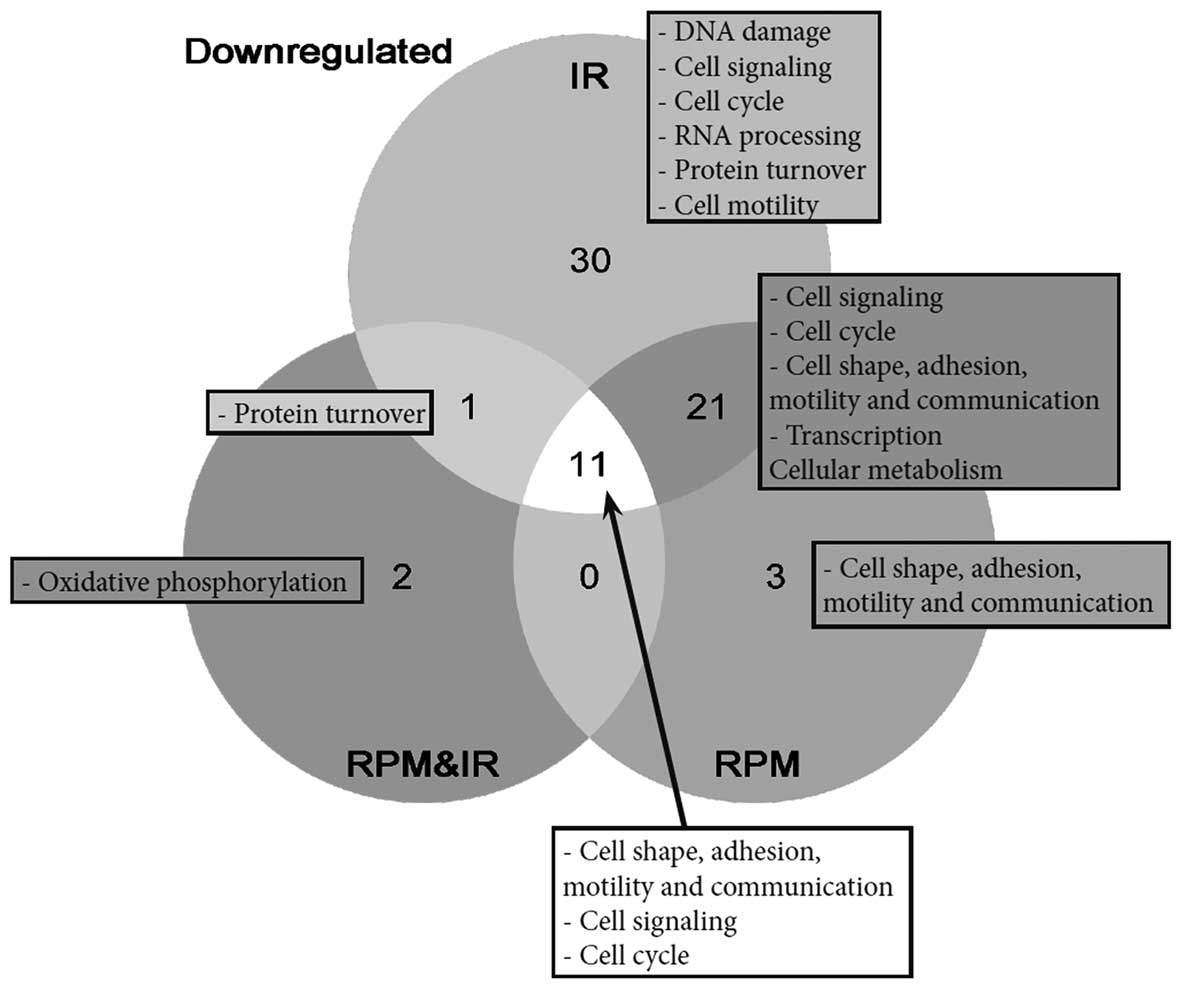
downregulated following chronic low-dose irradiation. Of the 63
genes, 30 were exclusively enriched in irradiated samples (Fig. 2), although this number may be an
overestimation due to redundancy between some of the gene sets. The
gene sets that were specifically downregulated after irradiation
conditions are mostly involved in DNA damage response, cell
signaling, cell cycle, RNA processing and protein turnover
(Table III). Moreover, we
detected significantly downregulated gene sets involved in cell
signaling, cell cycle, transcription, protein turnover, cell shape,
adhesion, motility and communication for all the treatments. Of
note, two gene sets involved in oxidative phosphorylation were
significantly downregulated solely in the RPM and IR samples. No
gene set was significantly upregulated in any of the
treatments.
 | Table IIIDownregulated gene sets revealed by
GSEA, based on the list of gene sets provided by Fig. 2. |
Table III
Downregulated gene sets revealed by
GSEA, based on the list of gene sets provided by Fig. 2.
| Treatment | Cell process | Gene set (GO) |
|---|
| IR | DNA damage | DNA damage
checkpoint |
| DNA repair |
| Histone
modification |
| Response to DNA
damage stimulus |
| Response to
radiation |
| Response to
stress |
| Cell signaling | Negative regulation
of signal transduction |
| Inositol or
phosphatidylinositol kinase activity |
| Ras GTPase
binding |
| Positive regulation
of JNK activity |
| RHO GTPase
activator activity |
| Protein kinase
cascade |
| Magnesium ion
binding |
| Protein
serine/threonine kinase activity |
|
Phosphorylation |
| Cell cycle | Cell cycle arrest
(GO 0007050) |
| Negative regulation
of cell cycle |
| RNA processing | RNA processing |
| RNA splicing |
| Spliceosome |
| Nuclear pore |
| Protein
turnover | tRNA metabolic
process |
| Post translational
protein modification |
| Endoplasmic
reticulum |
| Golgi
apparatus |
| Portein
ubiquitination |
| Ubiquitin
cycle |
| Ubiquitin protein
ligase activity |
| Small conjugating
protein ligase activity |
| Cell motility | Lamellipodium |
| IR + RPM | Cell signaling | G protein signaling
coupled to IP3 |
|
Phosphoinositide-mediated signaling |
| RAS GTPase
activator activity |
| GTPase regulator
activity |
| Small
GTPase-mediated signal transduction |
| Transmembrane
receptor |
| protein kinase
activity |
| Protein kinase
activity |
| Cell cycle | Cell cycle (GO
0007049) |
| Centrosome |
| Cell shape,
adhesion, motility and communication | Microtubule |
| Cytoskeletal
protein binding |
| Ruffle |
| Cell junction |
| Collagen |
| Extracellular
matrix |
| Transcription | Positive regulation
of transcription |
| Negative regulation
of transcription |
| RNA helicase
activity |
| Chromosome |
| Nucleolus |
| Cellular
metabolism | Negative regulation
of cellular metabolic process |
| IR + RPMIR + RPM
and IR | Protein
turnover | Protein/RNA complex
assembly |
| IR + RPM + RPM +
IR | Cell shape,
adhesion, motility and communication | Cytoskeleton |
| Actin binding |
| Actin
cytoskeleton |
| Adherens
junction |
| Cell matrix
adhesion |
| Motor activity |
| Microtubule
cytoskeleton |
| Cell signaling | Insulin receptor
signaling pathway |
| Cell cycle | Cell cycle
process |
| M phase |
| Spindle |
| RPM | Cell shape,
adhesion, motility and communication | Structural
constituent of cytoskeleton |
| Integrin
binding |
| Receptor
binding |
| RPM + IR | Oxidative
phosphorylation | Electron transport
(GO 0006118) |
| Mitochondrion |
Discussion
In this study, primary cultures of murine fetal
fibroblasts were chronically exposed (65 h) to simulated space
conditions including simulated microgravity via RPM and a low-dose
mixture of neutrons and gamma-rays (IR). The duration of the
experiment was chosen to allow cellular adaptation to the simulated
microgravity environment for instance for cytoskeleton remodeling
(13,14), in order to decrease the primary
stress response mechanisms and to better characterize the effects
of chronic exposure to these conditions. Microarrays were performed
on RNA harvested from CTRL, IR, RPM and RPM and IR conditions.
Microarrays generate a substantial amount of information on the
gene expression pattern of cells subjected to a defined treatment.
However, a <2-fold difference in the gene expression is often
not sufficient to meet the requirements for statistical
significance (8). Identification
of moderate gene expression differences in groups of genes acting
together in a cell process can nevertheless be achieved by means of
GSEA. For this reason, we analyzed our microarray output data using
the single gene analysis method as well as GSEA.
The RPM has a dominant impact on single
gene expression
The SGA method revealed a significant impact of 65 h
of simulated microgravity on gene expression in murine fetal
fibroblasts. The combination of RPM and IR triggered a differential
expression of fewer genes than RPM alone. Only a few genes had an
altered expression in IR samples, suggesting that such a low dose
of radiation exerted a moderate impact on the expression of
individual genes. It was also noted that only a few genes were
commonly differentially expressed in all irradiated treatments (IR
and RPM and IR), of which there were only six known genes, all
upregulated (S1p3, Rab11b, Ptger3, Vldlr, Cnn1 and Serping1), with
most of them being involved in cell signaling. No explanation can
be provided for the fact that few genes were commonly up- or
downregulated in the irradiated treatments (with or without RPM).
However the strong effect of RPM may have concealed a more subtle
effect of IR, making it statistically less significant.
Among the upregulated genes following RPM treatment,
glutathione-S-transferases α 1 and 2 (Gstα1 and
Gstα2) were prominent enzymes for the detoxification of
breakdown products of oxidative stress (15). However, since the Affymetrix
arrays cannot distinguish between the two isoforms due to their
very high sequence homology (97%), we cannot dismiss the
possibility that only one of the two isoforms was actually affected
by the treatment. The modifier subunit of glutathione-cysteine
ligase (Gclm) was significantly upregulated as well. The
protein encoded by this gene was shown to play an important role in
controlling the rate of glutathione synthesis in murine fetal
fibroblasts (16). We also report
upregulation of the heme oxygenase 1 (Hmox1), a
cytoprotective enzyme against oxidative stress (17). In murine fibroblasts, its
upregulation by curcumin was found to block radiation-induced
reactive oxygen species (ROS) generation (18). Notably, these three genes are
targets of the nuclear factor-erythroid 2 p45-related factor 2
(Nrf2) which induces transcription of cytoprotective genes
containing antioxidant response elements (19). The transcription factor Nrf2 may
therefore play a cytoprotective role against a possible oxidative
stress induced by the RPM, which is in line with previous
observations of increased oxidative stress in simulated
microgravity (20–22).
After RPM treatment, two members of the actin
filament family, Actg2 and Acta1 were downregulated.
These genes were described in smooth (23) or skeletal muscles (24), respectively. Calponin 1
(Cnn1), a gene coding for a protein involved in the
cytoskeleton organization (25),
and four and a half LIM domains 1 (Fhl1), which functions in
adherens junctions signaling to the cytoskeleton (26), were also downregulated. Notably,
the four genes were shown to be regulated by the serum response
factor (SRF). SRF was shown to be mediated by the Rho signaling
pathway (25–27), which may have been triggered by
the RPM. Rho signaling is believed to be an important pathway for
focal adhesion assembly and cytoskeleton remodeling in response to
cellular tension stress (28) and
has been suggested to play a role in the microgravity response
(21,29–31). Furthermore, Rho GTPase activities
were shown to be increased in dermal fibroblasts subjected to
simulated microgravity for 30 and 120 min, thereafter decreasing to
reach similar values to those of the CTRL at 48 h of treatment
(32). Our hypothesis is that a
65-h exposure to RPM induced downregulation of the Rho signaling
pathway, which decreased the activity of the transcription factor
SRF, decreasing in turn the expression of genes involved in
cytoskeleton organization (Cnn1) and adherens junctions
(Fhl1).
IR has a dominant effect on gene
sets
At the gene set level, GSEA did not detect any
upregulation, except for the structural constituents of the
ribosome in IR-treated samples. This result is noteworthy as it did
not occur with SGA. Since SGA and GSEA are purely statistical
methods, it is unlikely that this result originates from an
experimental issue, which may have affected both methods. We also
examined the gene set selection, however, a screening of all the
gene sets of GO provided the same result. Since the experimental
design involved long-term irradiation, it is possible that a
feedback loop occurred and decreased the expression pattern of the
gene sets.
We identified a significant downregulation of 63
gene sets in response to low-dose IR, although single gene analysis
did not reveal any important effects. Of the 63 gene sets, 30 were
specifically enriched in IR-treated samples (Fig. 2). These latter gene sets are
involved in DNA damage response, cell signaling, cell cycle, RNA
processing, protein turnover or cell motility. Of note, the DNA
damage response gene sets were downregulated, which may be
explained by the long duration of continuous irradiation at an
extremely slow-dose rate. It is possible that an adaptation
mechanism of the cells to irradiation triggered a feedback loop to
decrease the expression of these pathways, as was observed at the
gene level (SGA) for SRF responsive genes in response to the RPM.
Various other gene sets involved in the same cell processes were
also enriched in the RPM, and RPM and IR treatments.
Many of the downregulated gene sets are involved in
cell signaling, including Rho and Ras GTPases, inositol and
phosphatidylinositol, JNK and insulin receptor-mediated pathways.
The downregulation of these signaling pathways may lead to an
alteration of the cell cycle (33). In addition to its major role in
the cell response to radiation (34,35), the regulation of the cell cycle
has been shown to be affected by simulated microgravity (36). GSEA revealed that gene sets
involved in the positive regulation of the cell cycle were
downregulated in all treatments. However, cells that were only
irradiated exhibited a significant downregulation of gene sets
involved in cell cycle arrest, indicating no trend towards a pro-
or anti-proliferative expression profile, while both RPM and RPM
and IR showed an anti-proliferative expression profile. We suggest
that all the treatments may have induced a general stress response
that decreased the expression of cell cycle progression pathways,
while irradiation alone also reduced the expression of genes
involved in cell cycle arrest. This hypothesis is in agreement with
the decreased expression of DNA damage response pathways that we
also detected. In RPM and IR, the effect of the RPM may have
concealed the cell cycle arrest gene set downregulation.
In addition, many gene sets involved in the
composition of the cytoskeleton (actin and microtubule) and inter-
(cell junctions) and extracellular connections (extracellular
matrix) were affected by all the treatments. While it has been
shown in various cell types that cytoskeleton remodeling starts
immediately after exposure to simulated or real microgravity
(21,29–31), few studies investigated the
effects of IR on the cytoskeleton. However, therapeutic doses of
irradiation were shown to affect cell permeability of microvascular
endothelial cells through Rho-mediated cytoskeleton remodeling
(37). More recently,
Rho-mediated focal adhesion and fibronectin adhesion were shown to
be increased in endothelial cells in response to radiation
(38). As Rho GTPases intervene
in a number of additional cell pathways (e.g., cell cycle arrest,
and regulation of apoptosis) (39), Rho GTPases potentially play a
pivotal role in the cell response to simulated space conditions. In
agreement with this hypothesis, GSEA revealed that Rho GTPases
activity was downregulated in IR-treated samples. Notably, gene
sets involved in integrin and receptor binding were specifically
downregulated following treatment using the RPM. The results of
this study confirm therefore that integrins play a significant role
in the cellular response to simulated microgravity.
In conclusion, this study has shown that continuous
exposure to simulated microgravity affects fetal murine
fibroblasts, especially at the single gene level, by increasing the
expression of oxidative stress responsive genes and decreasing the
expression of genes involved in cytoskeleton remodeling. As far as
irradiation is concerned, we detected a decreased expression of
gene sets involved in cytoskeleton mechanisms, in cell signaling
and DNA damage response after a chronic low-dose rate of
irradiation, particularly at the gene set level. The results
indicate that the effects of the combination of the two treatments
did not result in a synergism between the two separate effects,
since many genes or gene sets that were altered by RPM or IR
treatment, were not changed by the combined treatment (RPM and
IR).
Acknowledgements
This study was supported by the ESA Topical Team on
‘Developmental Biology in Vertebrates’ and 4 PRODEX/ESA contracts
[C90-303, C90-380, C90-391 and 42-000-90-380].
References
|
1
|
Huijser RH: Desktop RPM: new small size
microgravity simulator for the bioscience laboratory. Fokker Space
FS-MG-R00-017. 1–5. 2000.
|
|
2
|
van Loon JJWA: Some history and use of the
random positioning machine, RPM, in gravity related research. Adv
Space Res. 39:1161–1165. 2007.
|
|
3
|
Borst A and van Loon J: Technology and
developments for the random positioning machine, RPM. Microgravity
Sci Technol. 21:287–292. 2009. View Article : Google Scholar
|
|
4
|
Kraft TF, van Loon JJ and Kiss JZ: Plastid
position in arabidopsis columella cells is similar in microgravity
and on a random-positioning machine. Planta. 211:415–422. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Villa A, Versari S, Maier JA and
Bradamante S: Cell behavior in simulated microgravity: a comparison
of results obtained with RWV and RPM. Gravit Space Biol Bull.
18:89–90. 2005.
|
|
6
|
Grimm D, Bauer J, Ulbrich C, et al:
Different responsiveness of endothelial cells to vascular
endothelial growth factor and basic fibroblast growth factor added
to culture media under gravity and simulated microgravity. Tissue
Eng Part A. 16:1559–1573. 2010. View Article : Google Scholar
|
|
7
|
Mastroleo F, Van Houdt R, Leroy B, et al:
Experimental design and environmental parameters affect
Rhodospirillum rubrum S1H response to space flight. ISME J.
3:1402–1419. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Shi J and Walker MG: Gene set enrichment
analysis (GSEA) for interpreting gene expression profiles. Curr
Bioinform. 2:133–137. 2007. View Article : Google Scholar
|
|
9
|
Subramanian A, Tamayo P, Mootha VK, et al:
Gene set enrichment analysis: A knowledge-based approach for
interpreting genome-wide expression profiles. Proc Natl Acad Sci
USA. 102:15545–15550. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
El-Saghire H, Thierens H, Monsieurs P,
Michaux A, Vandevoorde C and Baatout S: Gene set enrichment
analysis highlights different gene expression profiles in whole
blood samples X-irradiated with low and high doses. Int J Radiat
Biol. 89:628–638. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Vanhavere F, Loos M, Plompen AJM,
Wattecamps E and Thierens H: A combined use of the BD-PND and BDT
bubble detectors in neutron dosimetry. Radiat Meas. 29:573–577.
1998. View Article : Google Scholar
|
|
12
|
Cucinotta FA, Kim MH, Willingham V and
George KA: Physical and biological organ dosimetry analysis for
international space station astronauts. Radiat Res. 170:127–138.
2008. View
Article : Google Scholar : PubMed/NCBI
|
|
13
|
Crawford-Young SJ: Effects of microgravity
on cell cytoskeleton and embryogenesis. Int J Dev Biol. 50:183–191.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Meloni MA, Galleri G, Pani G, Saba A,
Pippia P and Cogoli-Greuter M: Space flight affects motility and
cytoskeletal structures in human monocyte cell line J-111.
Cytoskeleton (Hoboken). 68:125–137. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Hayes JD and McLellan LI: Glutathione and
glutathione-dependent enzymes represent a co-ordinately regulated
defence against oxidative stress. Free Radic Res. 31:273–300. 1999.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Chen Y, Johansson E, Fan Y, et al: Early
onset senescence occurs when fibroblasts lack the
glutamate-cysteine ligase modifier subunit. Free Radic Biol Med.
47:410–418. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Haines DD, Lekli I, Teissier P, Bak I and
Tosaki A: Role of haeme oxygenase-1 in resolution of oxidative
stress-related pathologies: Focus on cardiovascular, lung,
neurologic and kidney disorders. Acta Physiol (Oxf). 204:487–501.
2012. View Article : Google Scholar
|
|
18
|
Lee JC, Kinniry PA, Arguiri E, et al:
Dietary curcumin increases antioxidant defenses in lung,
ameliorates radiation-induced pulmonary fibrosis, and improves
survival in mice. Radiat Res. 173:590–601. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Hayes JD and McMahon M: NRF2 and KEAP1
mutations: permanent activation of an adaptive response in cancer.
Trends Biochem Sci. 34:176–188. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Wang J, Zhang J, Bai S, et al: Simulated
microgravity promotes cellular senescence via oxidant stress in rat
PC12 cells. Neurochem Int. 55:710–716. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Nikawa T, Ishidoh K, Hirasaka K, et al:
Skeletal muscle gene expression in space-flown rats. FASEB J.
18:522–524. 2004.PubMed/NCBI
|
|
22
|
Liu Y and Wang E: Transcriptional analysis
of normal human fibroblast responses to microgravity stress.
Genomics Proteomics Bioinformatics. 6:29–41. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Carson JA, Culberson DE, Thompson RW,
Fillmore RA and Zimmer W: Smooth muscle gamma-actin promoter
regulation by RhoA and serum response factor signaling. Biochim
Biophys Acta. 1628:133–139. 2003. View Article : Google Scholar
|
|
24
|
Philippar U, Schratt G, Dieterich C, et
al: The SRF target gene Fhl2 antagonizes RhoA/MAL-dependent
activation of SRF. Mol Cell. 16:867–880. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Beamish JA, He P, Kottke-Marchant K and
Marchant RE: Molecular regulation of contractile smooth muscle cell
phenotype: implications for vascular tissue engineering. Tissue Eng
Part B Rev. 16:467–491. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Olson EN and Nordheim A: Linking actin
dynamics and gene transcription to drive cellular motile functions.
Nat Rev Mol Cell Biol. 11:353–365. 2010. View Article : Google Scholar
|
|
27
|
Sun Q, Chen G, Streb JW, et al: Defining
the mammalian CArGome. Genome Res. 16:197–207. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Ingber DE: Tensegrity II. How structural
networks influence cellular information processing networks. J Cell
Sci. 116:1397–1408. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Meloni MA, Galleri G, Pippia P and
Cogoli-Greuter M: Cytoskeleton changes and impaired motility of
monocytes at modelled low gravity. Protoplasma. 229:243–249. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Servotte S, Zhang Z, Lambert CA, et al:
Establishment of stable human fibroblast cell lines constitutively
expressing active rho-GTPases. Protoplasma. 229:215–220. 2006.
View Article : Google Scholar
|
|
31
|
Nichols HL, Zhang N and Wen X: Proteomics
and genomics of microgravity. Physiol Genomics. 26:163–171. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Loesberg WA, Walboomers XF, van Loon JJWA
and Jansen JA: Simulated microgravity activates MAPK pathways in
fibroblasts cultured on microgrooved surface topography. Cell Motil
Cytoskeleton. 65:116–129. 2008. View
Article : Google Scholar : PubMed/NCBI
|
|
33
|
Hall A: Rho GTPases and the control of
cell behaviour. Biochem Soc Trans. 33:891–895. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Jeggo P: The role of the DNA damage
response mechanisms after low-dose radiation exposure and a
consideration of potentially sensitive individuals. Radiat Res.
174:825–832. 2010. View
Article : Google Scholar : PubMed/NCBI
|
|
35
|
Jeggo P and Lavin MF: Cellular
radiosensitivity: how much better do we understand it? Int J Radiat
Biol. 85:1061–1081. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Grimm D, Wise P, Lebert M, Richter P and
Baatout S: How and why does the proteome respond to microgravity?
Expert Rev Proteomics. 8:13–27. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Gabryś, Greco O, Patel G, Prise KM, Tozer
GM and Kanthou C: Radiation effects on the cytoskeleton of
endothelial cells and endothelial monolayer permeability. Int J
Radiat Oncol Biol Phys. 69:1553–1562. 2007.PubMed/NCBI
|
|
38
|
Rousseau M, Gaugler MH, Rodallec A,
Bonnaud S, Paris F and Corre I: Rhoa GTPase regulates
radiation-induced alterations in endothelial cell adhesion and
migration. Biochem Biophys Res Commun. 414:750–755. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Etienne-Manneville S and Hall A: Rho
GTPases in cell biology. Nature. 420:629–635. 2002. View Article : Google Scholar
|