Introduction
Lung cancer is the leading cause of cancer deaths in
the world and cancer metastasis is the major cause of death in lung
cancer patients (1). Non-small
cell lung cancer (NSCLC) accounts for ~85% of all lung cancer
cases, and to date, its overall 5-year survival rate is ~15%
(2). Lung cancer is expected to
remain a major health hurdle for the next 40–50 years (3,4).
Therefore, development of effective therapies for NSCLC is urgently
needed.
miRNAs are small non-coding RNAs that can degrade
target mRNA and suppress protein translation (5,6).
miRNAs could post-transcriptionally regulate gene expression and
therefore, modulate cell function and homeostasis and act as tumor
suppressor genes or oncogenes (7).
On average, each miRNA is estimated to be able to regulate several
hundred protein-coding genes, and ~60% of proteins in cells is
thought to be regulated by miRNAs (8,9).
Thus, changes in miRNA expression pattern are linked to profound
effects on cell phenotypes, and miRNAs have an emerging role in
diverse physiological and pathological processes (10). In this regard, altered miRNA
expression will lead to human carcinogenesis and cancer
progression. For example, miR-143 targets K-Ras, Bcl-2, and other
genes to regulate tumor cell proliferation and invasion capacity
(11,12). Furthermore, transforming growth
factor-β (TGF-β) acts as a tumor suppressor in normal tissues, but
the suppressive effects are lost during early carcinogenesis of
advanced tumors.
TGF-β-induced epithelial-to-mesenchymal transition
(EMT) contributes to human carcinogenesis (13–15).
EMT usually occurs during critical phases of embryonic development
(such as mesoderm and neural tube formation), wound healing, or
initiation of metastasis during cancer progression. During EMT,
epithelial cells lose polarity and cell-cell adhesion, and acquire
migration and invasion properties of mesenchymal cells (13–15).
At the gene level, various genes participate in EMT; for example,
TGF-β, a protein that controls proliferation, cellular
differentiation, and other functions in most human cells, can
promote NSCLC cell EMT (16–18).
However, it is unknown how TGF-β regulates miRNA expression or EMT
to regulate cell apoptosis and tumor progression and whether there
is a relationship between miR-143 and TGF-β, previous studies
indeed show that miR-143 can regulate EMT and tumor cell growth and
apoptosis (19–21). For example, Davis et al
showed that TGF-β interacts with myocardin (Myocd) or Myocd-related
transcription factors to induce pri-miR-143/145 expression and in
turn, miR-143/145 downregulates KLF4 transcription to control
vascular smooth muscle cells contractile phenotypes and mediate
induction of contractile genes (22). Thus, in this study, we investigated
the effects of TGF-β induction on miR-143 expression and then
assessed the effects of miR-143 expression or knockdown on the
regulation of NSCLC cell viability and invasion capacity in
vitro, along with xenograft formation and growth in nude
mice.
Materials and methods
Cell line and culture
Human NSCLC A549 cell line was purchased from
American Type Culture Collection (ATCC, Manassas, VA, USA) and
cultured with RPMI-1640 medium supplemented with 10% (v/v)
heat-inactivated fetal bovine serum (Hyclone Laboratories, Inc.,
Logan, UT, USA) in an atmosphere of 95% air and 5% CO2
at 37°C. Recombinant human TGF-β was purchased from R&D Systems
(MN, USA) and used to treat the cultured cells (400 pM). Cells were
cultured under starvation conditions (0.2% FBS) overnight (~16 h)
prior to growth factor stimulation as described previously
(23).
miRNA microarray analysis
Total cellular RNA was isolated from the cultured
cells using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) and
purified using an RNA easy mini kit (Qiagen, Frankfurt, Germany)
according to the manufacturer’s instructions. The isolated RNA
samples were quantified using a Nanodrop instrument (Thermo Fisher,
Wilmington, DE, USA) and then labeled with the miRCURY™ Hy3™/Hy5™
Power labeling kit (Qiagen). After that, the labeled RNA samples
were used as probes to hybridize onto the miRCURY™ LNA Array
(v.2.0, Affymetrix) in a hybridization station (Capitalbio,
Beijing, China) according to a previous study (24). The array sections were scanned
using the Axon GenePix 4000B microarray scanner (Capitalbio) and
the data were read and analyzed using GenePix pro V6.0 software
(Capitalbio) for the raw intensity of the images.
The threshold value used to screen up- and
downregulated miRNAs was the degree of change (≥1.5- and
≤0.75-fold; p≤0.05). A heat map diagram was used to display data
for two-way hierarchical clustering of genes and samples.
Quantitative reverse
transcription-polymerase chain reaction (qRT-PCR)
Total cellular RNA was isolated using a
TRIzol® reagent (Invitrogen) from cultured cells. RT
reactions were performed using the iScript cDNA synthesis kit
(Bio-Rad, Hercules, CA, USA) and qPCR was performed in a Bio-Rad
IQTM5 Multicolor Real-Time PCR detection system (Bio-Rad). The qPCR
cycle was set an initial 98°C for 2 min, 40 cycles of 95°C for 15
sec, 60°C for 30 sec, and a final melt-curve analysis (60–95°C) was
included. The standard curve was produced with slopes at ~−3.32
(~100% efficiency); miRNA PCR quantification used the
2ΔΔct method against the U6 for normalization. mRNA PCR
quantification used the 2ΔΔct method against the β-actin
for normalization. The data are representative of the means of
three experiments. Primers for qPCR were designed to detect miRNA
based on the miRNA sequences provided by the Sanger Center miRNA
Registry, were and synthesized and purified by Shanghai Gene-Pharma
Co. (Shanghai, China). Mature miRNA primers and mRNA are described
in Table I. They were all
synthesized and purified by GenePharma Co.
 | Table IPrimer sequences. |
Table I
Primer sequences.
| Primer | Primer
sequences |
|---|
| miRNA primers |
| miR-143 |
5′-TGAGATGAAGCACTGTAGCTC-3′ |
| miR-145 |
5′-GTCCAGTTTTCCCAGGAATCCCT-3′ |
| miR-181a-2 |
5′-AACATTCAACGCTGTCGGTGAGT-3′ |
| miR-487b |
5′-GTGGTTATCCCTGTCCTGTTCG-3′ |
| miR-1972 |
5′-TCAGGCCAGGCACAGTGGCTCA-3′ |
| miR-138 |
5′-AGCTGGTGTTGTGAATCAGGCCG-3′ |
| miR-21 |
5′-TAGCTTATCAGACTGATGTTGA-3′ |
| miR-1246 |
5′-AATGGATTTTTGGAGCAGG-3′ |
| mRNA primers |
| Smad3 | F
5′-CCCCAGAGCAATATTCCAGA-3′
R 5′-GGCTCGCAGTAGGTAACTGG-3′ |
| CD44 | F
5′-AAGGTGGAGCAAACACAACC-3′
R 5′-AGCTTTTTCTTCTGCCCACA-3′ |
| K-Ras | F
5′-TGTGGTAGTTGGAGCTGGTG-3′
R 5′-TGACCTGCTGTGTCGAGAAT-3′ |
|
Pri-miR-143/145 | F
5′-AACTCCAGCTGGTCCTTAG-3′
R 5′-TCTTGAACCCTCATCCTGT-3′ |
| GAPDH | F
5′-AAGGTGAAGGTCGGAGTC-3′
R 5′-GATTTTGGAGGGATCTCG-3′ |
Immunohistochemistry and in situ
hybridization
Human NSCLC tissue specimens and nude mouse
xenografts were fixed in formalin and embedded in paraffin for
preparation of tissue sections for immunohistochemistry and in
situ hybridization. Tissue sections 4–6-μm thick were dewaxed
and re-hydrated. For immunohistochemistry, the sections were heated
in a microwave in an antigen retrieval buffer (0.01 M citrate
buffer, pH 6.0) for 5 min, incubated in 3%
H2O2 solution in phosphate-buffered saline
(PBS) at room temperature for 30 min to block potential endogenous
peroxidase activity, and then incubated in 20% normal goat serum to
block nonspecific binding of the secondary antibody. Then the
tissue sections were incubated with a primary antibody, Smad3 at a
dilution of 1:50; CD44 at a 1:50 dilution (both from Cell Signaling
Technology, Danvers, MA, USA, or K-Ras from Millipore, Billerica,
USA at a 1:50 dilution) at 4°C overnight in a humidified chamber.
The next day, after washing with PBS three times, the sections were
further incubated with a biotinylated goat anti-rabbit IgG antibody
(UltraSensitive S-P kit, Maixin Biotechnology Co., Fuzhou,
China).
For in situ hybridization, the rehydrated
tissue sections were transferred into a 3%
H2O2 and protease buffer for inactivation of
endogenous peroxidase. The sections were then treated with pepsin
solution diluted in 3% citric acid, and were hybridized overnight
at 59°C with a 50 nM DIG-labeled locked nucleic acid (LNA)-based
probe specific for miR-143 or LNA-src-miR negative control probe
(Exiqon, Vedbaek, Denmark). After rinsing in the washing buffer,
the sections were blocked in blocking buffer. An antibody specific
for DIG-AP Fab fragments antibody was applied to the sections.
For color reaction, both immunohistochemical and
hybridized sections were then washed and incubated in
3′-diaminobenzidine hydrochloride (DAB) solution and then briefly
counterstained with haematoxylin solution. The sections were then
dehydrated and mounted for review and scoring under an Olympus
microscope (Model BX-51, Tokyo, Japan). For negative and blank
control of immunohistochemistry, goat serum and PBS were used to
replace the primary antibody, respectively. A semi-quantitative
scoring criterion was used for these sections, in which two
pathologists independently in a blinded manner recorded staining
intensity and percentage of positively stained tumor cells. For the
staining intensity, we used 0 to represent no staining, 1 for weak
staining, 2 for positive staining, and 3 for strongly positive
staining. For percentage values, we scored the sections as 1 for
0–25% stained, 2 for 26–50% stained, 3 for 51–75% stained, and 4
for 76–100% stained tumor cells. The sum of both counts was used as
the final score for each case. A total score <8 was considered a
low level of expression, while a total score ≥8 was considered
high-level expression.
siRNA and transfection into NSCLC
cells
To knock down Smad3 expression in NSCLC cells, we
utilized Smad3 siRNA; for a negative control, we used non-targeting
scrambled siRNA (both from Qiagen). Smad3 siRNA and control siRNA
(40 nM) were then transfected into NSCLC cells using RNAi Max kit
(Qiagen) according to the manufacturer’s instructions. After 24 h,
the cells were used for phenotypic and gene expression
experiments.
Cell viability MTT assay
Cells were seeded at a density of 5×103
cells/well in 96-well plates 24 h after siRNA transfection and
incubated for 24 h. After that, 20 μl of MTT solution (5 mg/ml) was
added to each well, and the cell culture plates were further
incubated for additional 4 h at 37°C. The growth medium was removed
and replaced with 150 μl of dimethyl sulfoxide (DMSO) in each well
to dissolve MTT formaz crystals for 10 min; optical density (OD)
was measured using a scanning multi-well spectrophotometer at a
wavelength of 490 nm. Six parallel wells were measured each time,
and the experiment was repeated three times.
Wound healing assay
Cells were transfected with miR-143 mimics and grown
to 90% confluency in a 6-well dish. A wound was created using a
sterile 10-μl pipette tip, followed by a wash with 1X PBS to remove
detached cells. The cells were then further cultured in the growth
medium containing 5% FCS for up to 48 h, during which cell
migration at the corresponding wound site was documented using a
Nikon microscope (Nikon) at 0, 24 and 48 h. The wound distance was
then measured, and the mean and standard deviation of each
treatment were determined.
Tumor cell Transwell invasion assay
Corning Costar Transwell 24-well plate inserts (8-μm
pores; Corning) were coated with 20 μl of Matrigel (BD, New York,
USA) and placed in the cell culture hood for 1 h at 37°C. Cells
(2×104) were then seeded into the upper chambers of the
inserts after transfections, while normal growth medium was placed
in the bottom chambers. Cells were then allowed to grow for 24 or
48 h. The invaded cells were fixed in 100% methanol for 1 min and
allowed to air dry. Invasive cells on the lower surface of the
membrane were stained by dipping inserts in 1% crystal violet
staining solution for 2 min and were counted. Specifically, five
randomly selected 20x microscopic fields of cells were counted per
insert.
Nude mouse xenograft assay
Nude mice were obtained from Slaccas Co. (Shanghai,
China), and the A549 cells overexpressing LAN-miR-143 were injected
subcutaneously (1×106 cells per mouse with five mice per
group). After that, the mice were monitored for xenograft formation
and growth for 21 days. At the end of experiments, the mice were
sacrificed and xenografts were removed and weighed.
Luciferase report assay
We first performed bioinformatical analysis of
miR-143-targeted genes using web tools (such as TargetScan and
pictar) and found Smad3, CD44, and K-Ras as the putative
miR-143-targeted genes. We then constructed a PMIR vector carrying
Smad3, CD44, K-Ras-binding site sequences or their deletion
mutation sequences. Binding of miR-143 to these gene sequences was
assessed using a Luciferase reporter system from Sigma Chemical Co.
(St. Louis, MO, USA) according to the kit instructions.
Protein extraction and western
blotting
Protein extraction and western blotting were
performed as described previously (11). Protein samples (50 μg each) were
resolved on 10% SDS-polyacrylamide gels. The proteins were then
transferred onto PVDF membranes, incubated with 5% skim milk at
room temperature in TBS-T (20 mM Tris-HCl, pH 7.5, 500 mM NaCl
(13,14), and 0.1% Tween-20), and then further
incubated at 4°C overnight with a primary mouse monoclonal antibody
against Smad3, CD44 or K-Ras (Cell Signaling Technologies,
Cambridge, UK) at a dilution of 1:1,000, or a rabbit polyclonal
anti-β-actin antibody from Boshide (Wuhan, China). After washing
three times with TBS-T, the membranes were incubated with a
secondary antibody for 1 h at room temperature. Finally, peroxide
and luminol solutions of 40:1 were added to cover the whole
membrane surface for 5 min at room temperature, and positive
protein bands were visualized by placing the membrane in a
developing cassette against X-ray film.
Statistical analysis
All experimental data were summarized as mean ±
standard deviation, if possible, and statistically analyzed using
SPSS 17.0. (SPSS, Chicago, IL, USA). The Student’s t-test was used
to compare between two groups. Two-way ANOVA was used to compare
differences among three or more experimental groups. All p-values
were two-sided and values <0.05 were considered statistically
significant.
Results
TGF-β induces miR-143 expression in NSCLC
cells
To explore the molecular mechanisms by which TGF-β
induces cell contractile phenotype in NSCLC cells, we first grew
and treated A549 cells with either 400-pM TGF-β or PBS (as a
control) for 48 h, and then performed a miRNA chip array analysis.
We found 16 upregulated and 42 downregulated miRNAs in NSCLC cells
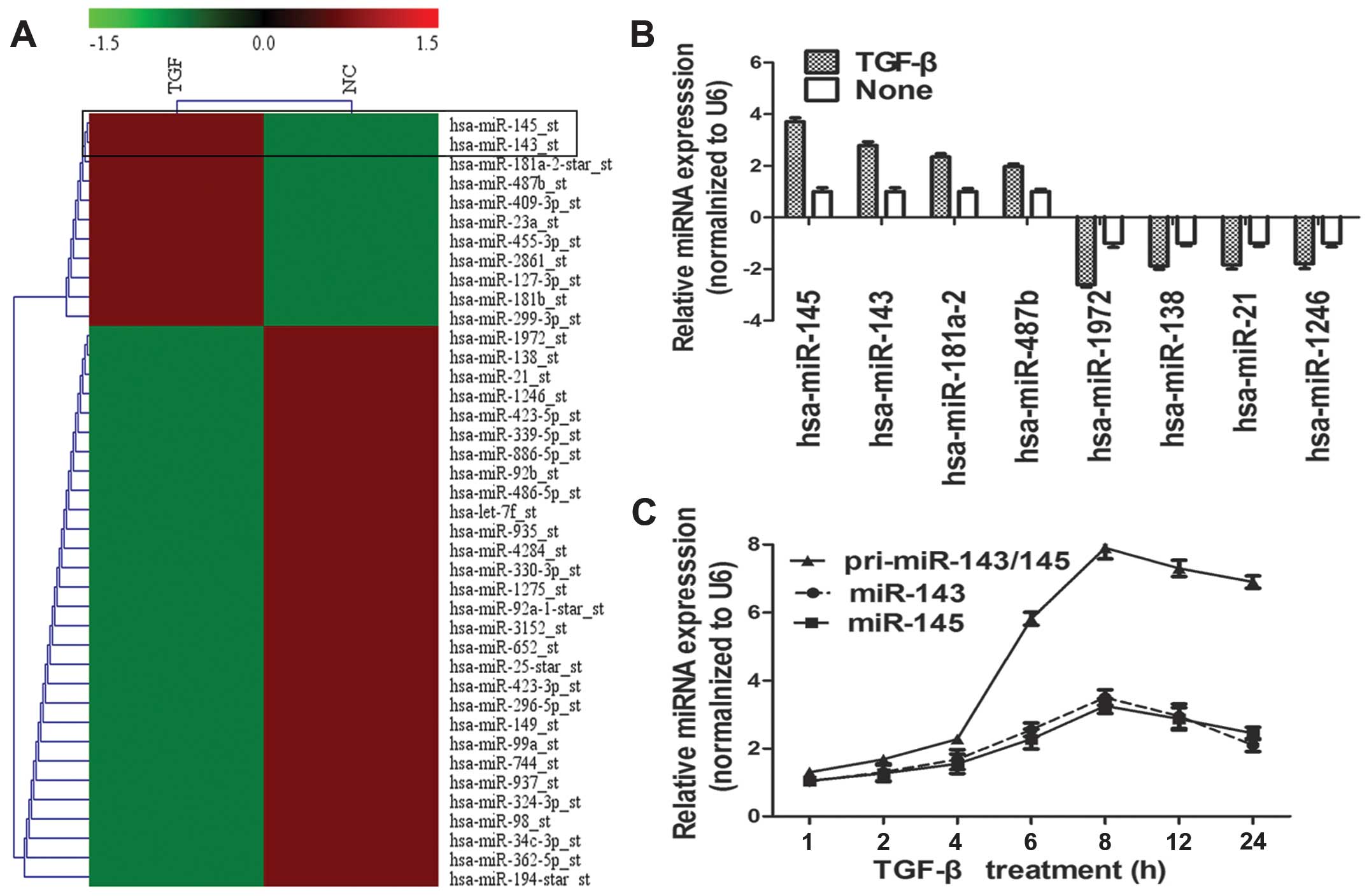
after TGF-β treatment. A heat map diagram analysis showed that the
maximally upregulated miRNA was in the miR-143/145 cluster, and the
maximally downregulated miRNA was miR-1972 (Fig. 1A). We then verified expression of
four upregulated and four downregulated miRNAs using qPCR (Fig. 1B). We then assessed expression of
the miR-143 cluster and pri-miR-143/145 in NSCLC cells after
treatment with TGF-β for different periods of time. Pri-miR-143/145
expression was dramatically upregulated after 4 h after TGF-β
treatment and reached a maximum at 8 h after treatment (a 7.9-fold
increase, Fig. 1C).
Downregulation of miR-143 expression in
NSCLC tissue specimens
To confirm the altered miR-143 expression ex
vivo, we performed qRT-PCR analysis of miR-143 in 24 cases of
matched NSCLC and the corresponding distant normal tissue specimens
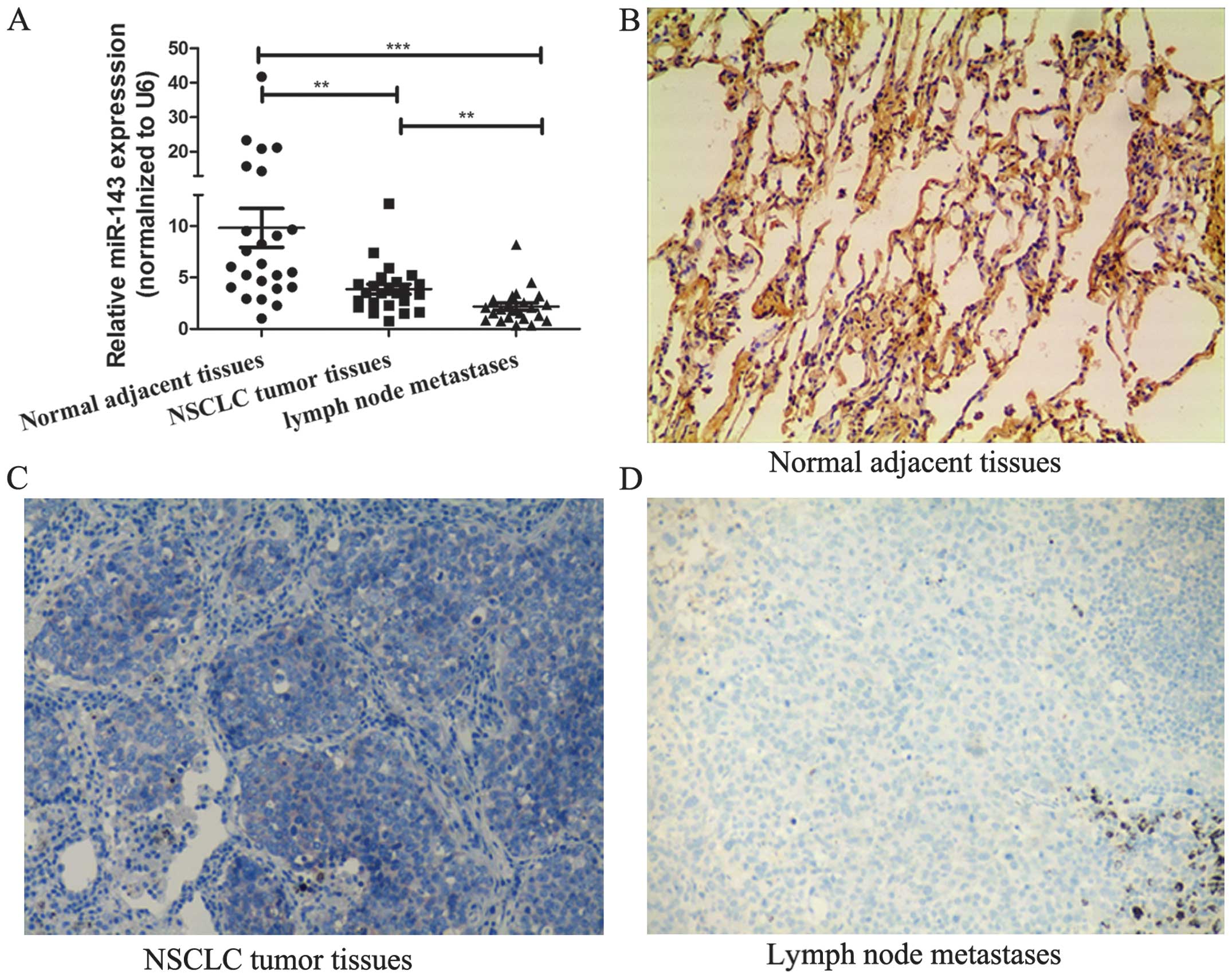
and in lymph node metastasized NSCLCs. As shown in Fig. 2A, levels of miR-143 expression were
significantly reduced by 1.2–9.8-fold in 87.5% (21 of 24) of NSCLC
tumor tissues compared to the corresponding distant normal lung
tissues; miR-143 expression was further reduced by 1.5–65.7-fold in
91.7% (22 in 24) of lymph node metastasized NSCLCs (Fig. 2A). Next, we assessed expression of
miR-143 in a panel of NSCLC tissue specimens using in situ
hybridization with similar findings (Fig. 2B).
Effects of miR-143 on regulation of A549
cell viability and invasion capacity
To explore the role of miR-143 in NSCLC cells, we
transfected miR-143 mimics (miR-143) and negative controls
(miR-NC), or miR-143 inhibitors (antagomiR-143) and inhibitor
controls (antagomiR NC) into A549 cells. Expression of miR-143
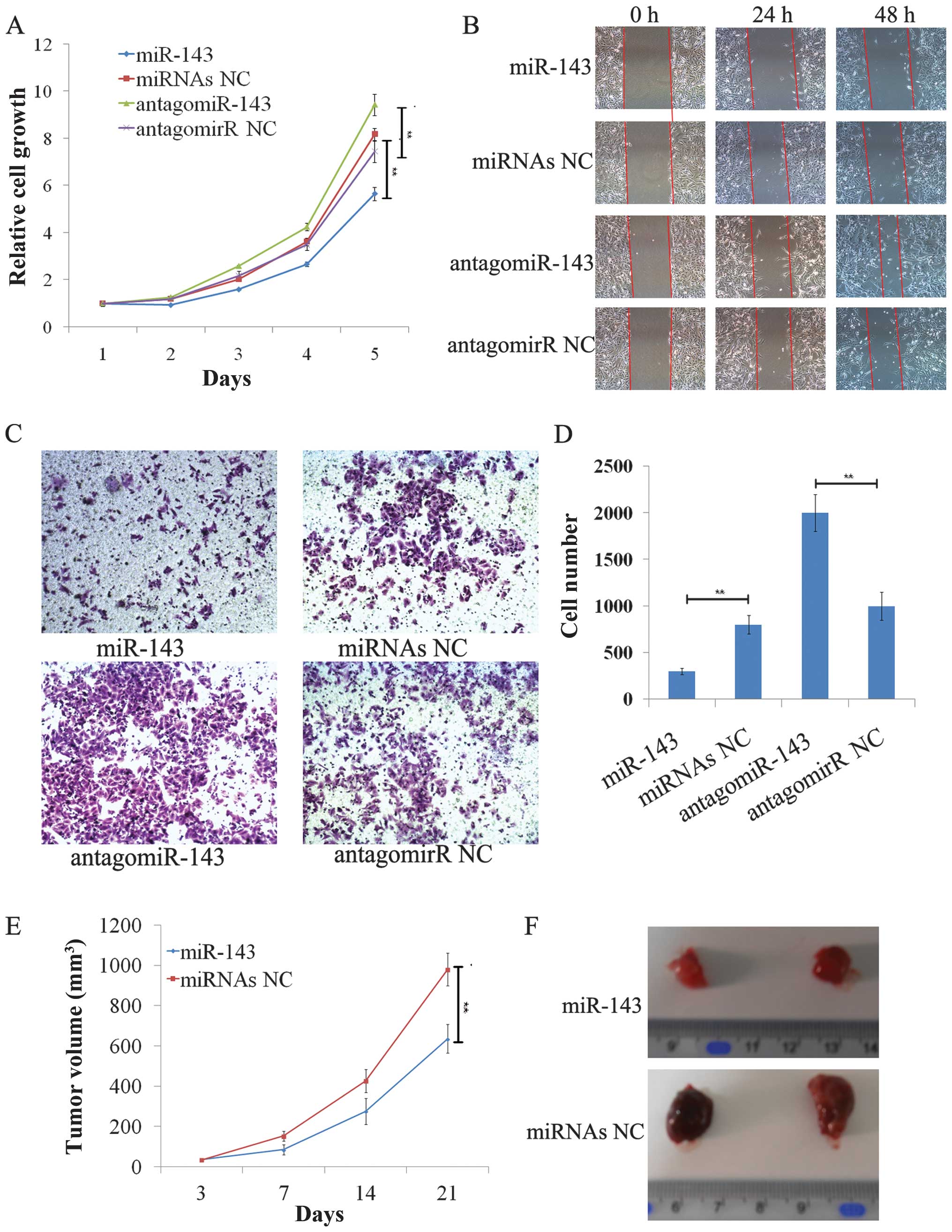
inhibited A549 cell viability, whereas miR-143 inhibitors increased
A549 cell viability (Fig. 3A).
Moreover, during tumor cell wound healing assays, over-expression
of miR-143 significantly delayed closure of the wound in a
time-dependent manner, whereas miR-143 inhibitor promoted the wound
healing (Fig. 3B). During a
Transwell tumor cell invasion assay, the number of cells
transfected with miR-143 mimics that passed through the 8-μm pore
insert underneath the well was significantly less than the number
of cells transfected with negative control. The miR-143 inhibitor
obviously promoted the invasion capacity of A549 cells (Fig. 3C and D).
The in vivo role of miR-143 in tumor cell
growth was also assessed using a nude mouse xenograft assay.
Following subcutaneous injection of LAN-miR-143-overexpressed A549
cells into nude mice, xenografts from miR-143-overexpressed-A549
cells were much smaller than those from control mice (n=4–5/group,
p<0.05; Fig. 3E). The tumor
volume of xenografts from miR-143-overexpressed A549 cells was also
smaller than from control cells (Fig.
3F).
miR-143 downregulation of Smad3, CD44,
and K-Ras expression in NSCLC cells
To elucidate the underlying molecular events of
miR-143 expression in NSCLC cells, putative target genes of miR-143
were searched using online algorithms for miRNA target prediction
(i.e., Pictar, RNAhybrid, and TargetScan). We found that Smad3,
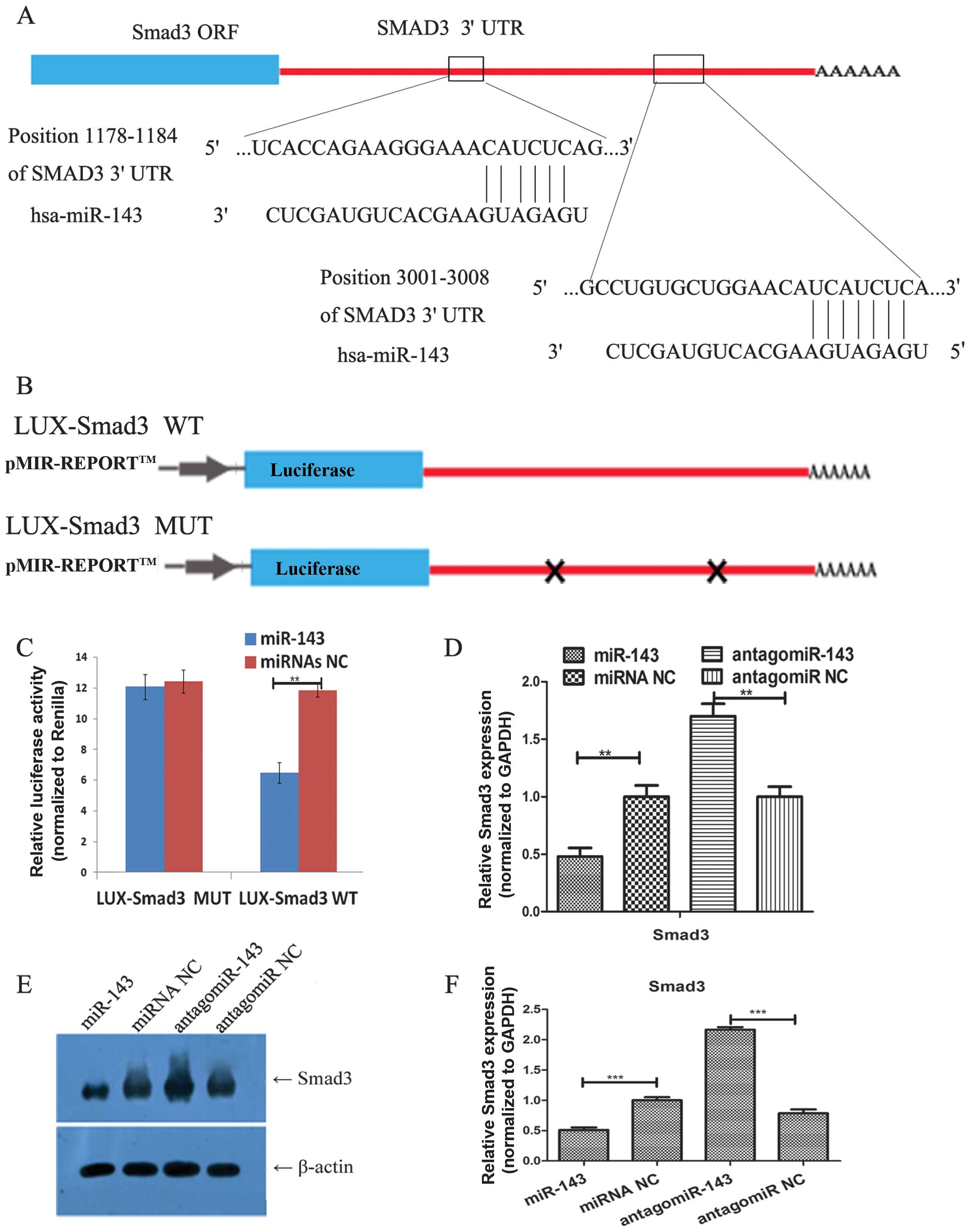
CD44, or K-Ras could be the target gene. The schematic diagram of
miR-143 binding sites in Smad3 3′-UTR is shown in Fig. 4A. To verify the diagram, we
constructed a luciferase reporter vector using cloned Smad3 3′-UTR,
both wild-type and mutant sequences (Fig. 4B), and performed gene transfection
and a luciferase assay. We observed a significant decrease in
luciferase activity in cells overexpressing miR-143. In contrast,
luciferase activity of mutant 3′-UTR remained unchanged in cells
overexpressing miR-143 (Fig. 4C),
indicating that miR-143 did bind to Smad3 3′-UTR. After that, we
performed qRT-PCR and western blot analyses to confirm that miR-143
overexpression inhibited expression of endogenous Smad3 mRNA and
protein, in NSCLC A549 cell lines. The knockdown of miR-143
expression resulted in upregulation of endogenous Smad3 expression
(Fig. 4D–F).
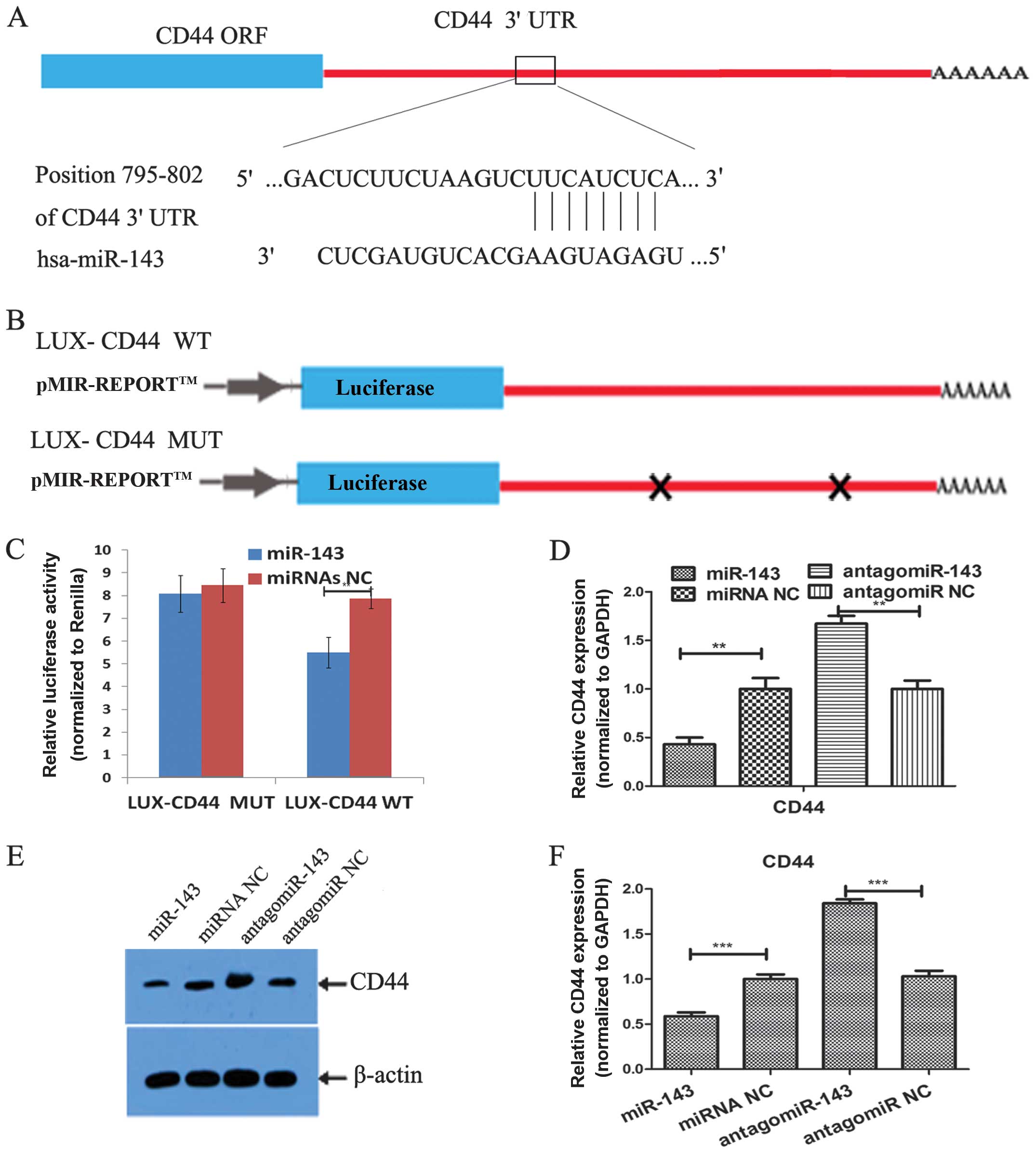
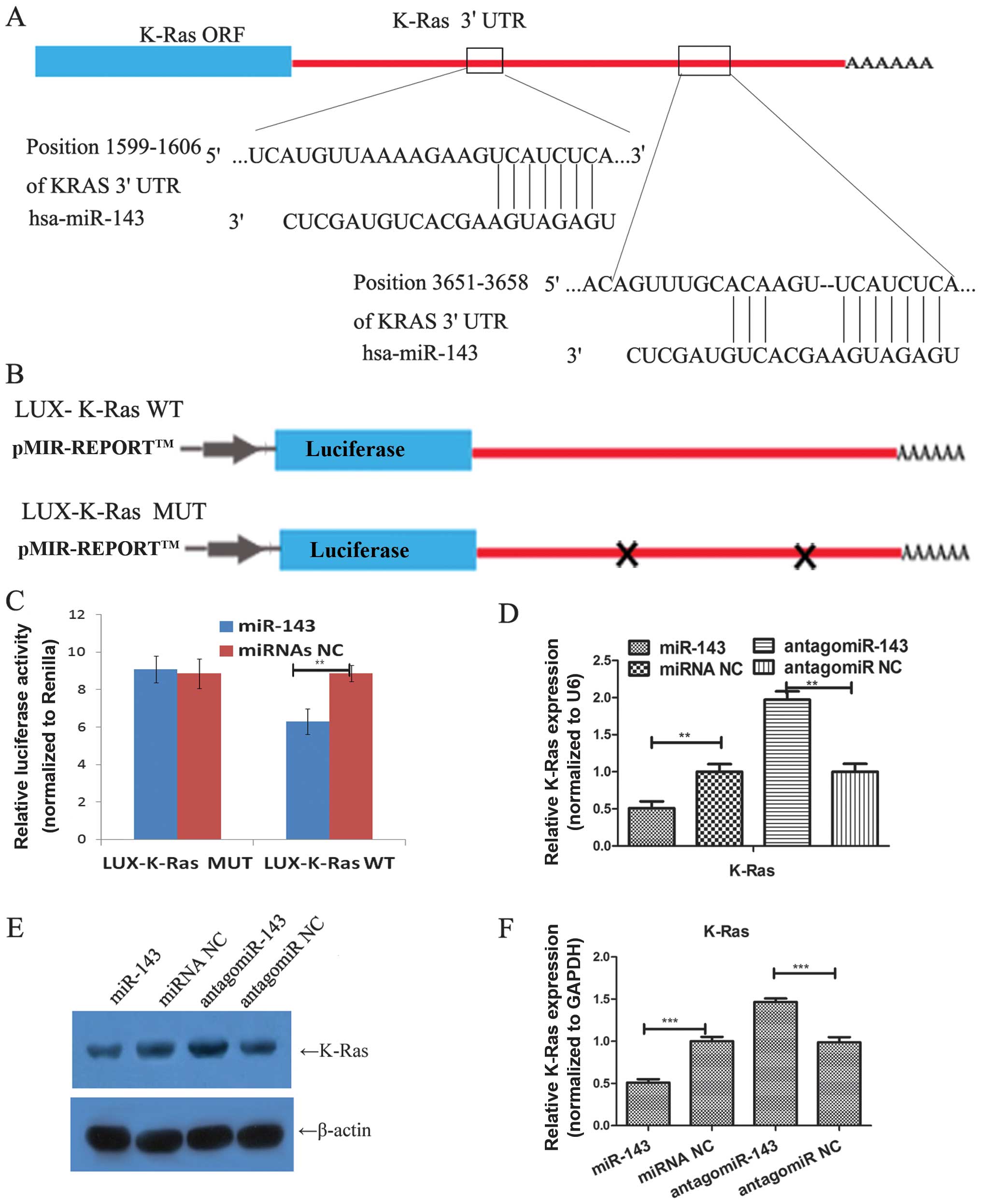
Furthermore, miR-143 binding sites in CD44 and K-Ras
3′-UTR were also analyzed (Figs. 5
and 6). The luciferase assay also
verified miR-143 binding to CD44 and K-Ras 3′-UTR. qRT-PCR and
western blotting confirmed that these two molecules were also
regulated by miR-143.
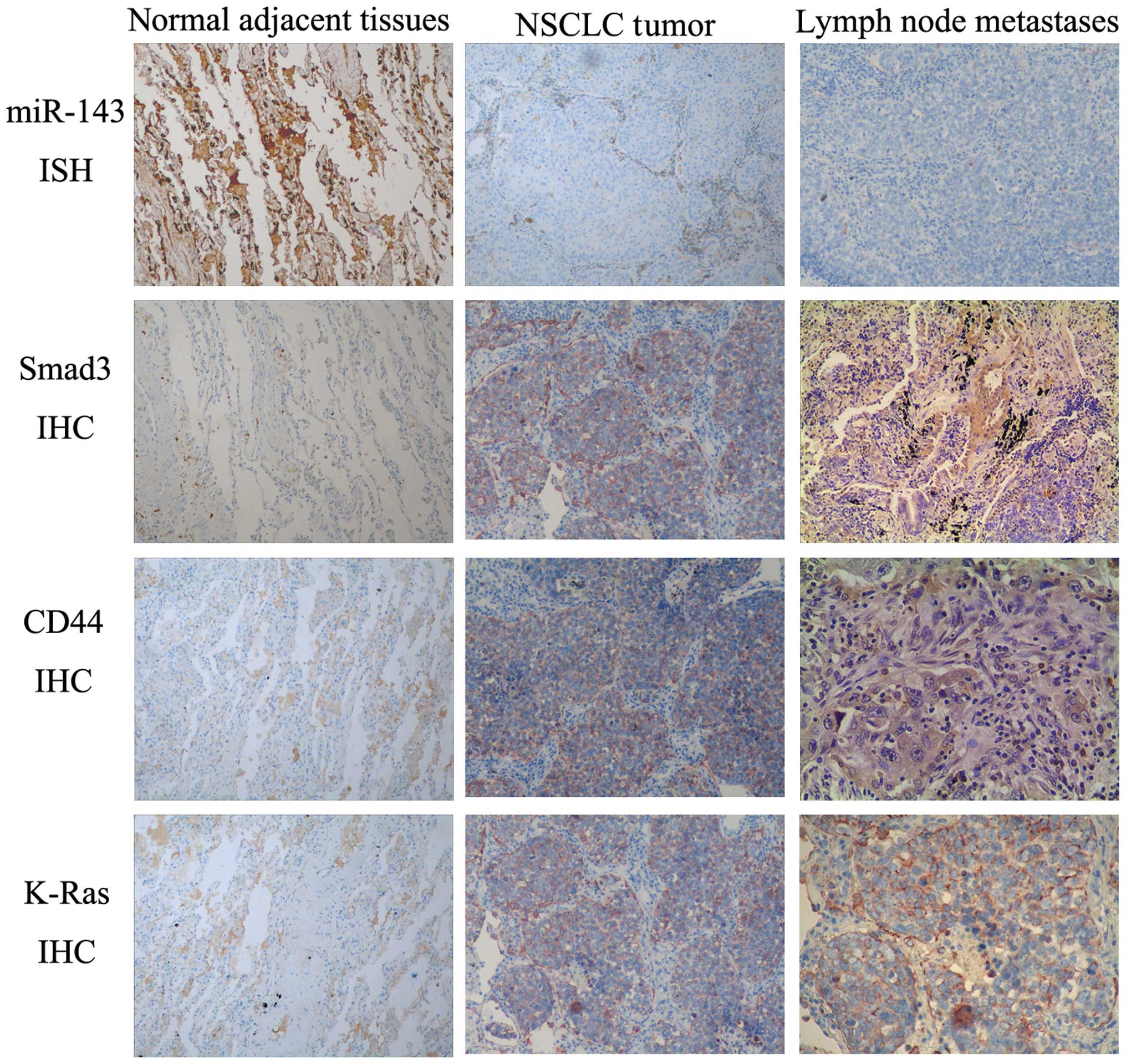
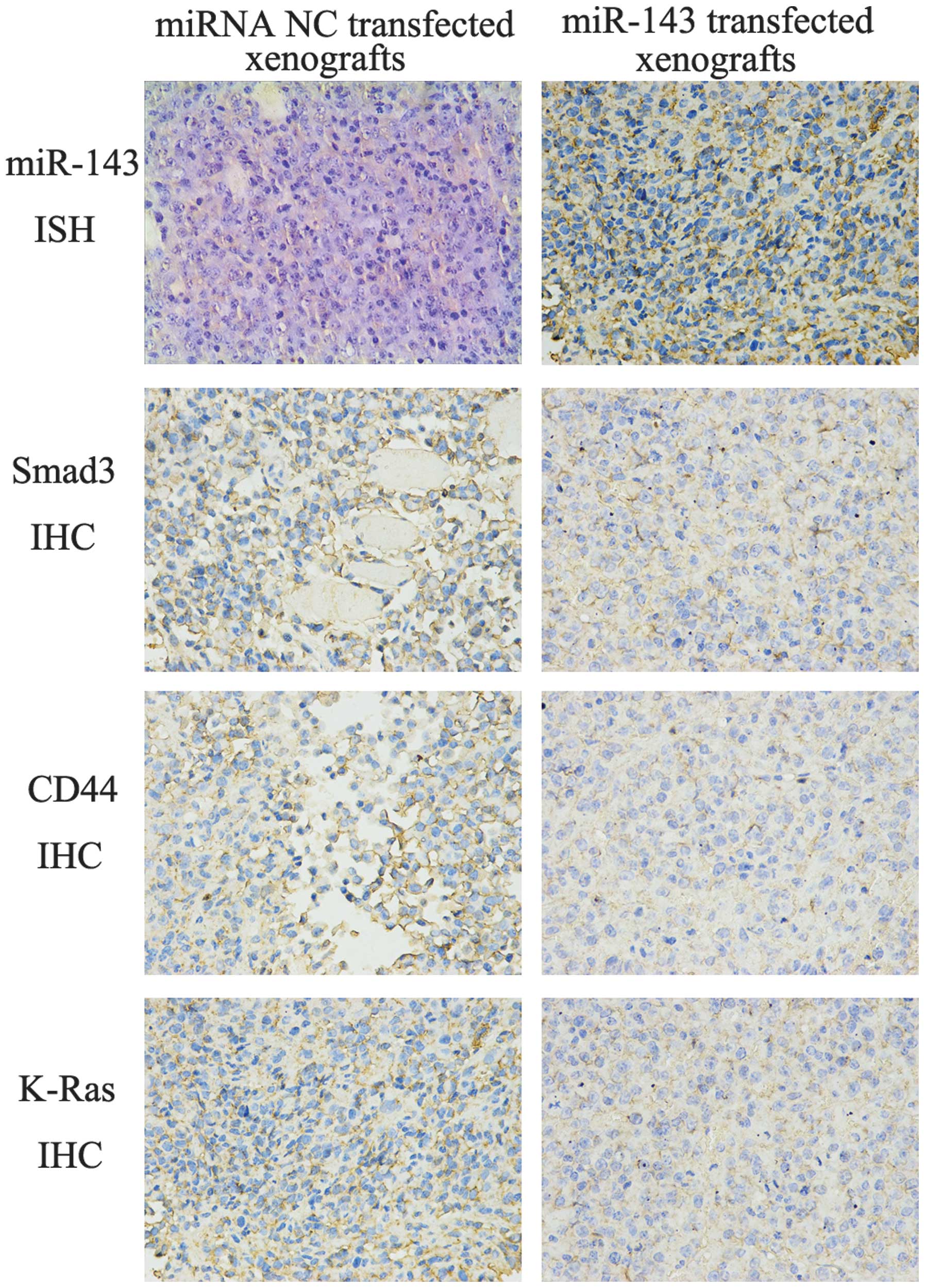
We also verified the regulation of Smad3, CD44 and
K-Ras in ex vivo tissue specimens and A549 cell xenografts
using in situ hybridization and immunohistochemistry. There
was an association between miR-143 expression and the target genes
(Smad3, CD44 and K-Ras) in the NSCLC and normal lung tissues and
lymph node metastatic tissues as well (Fig. 7). Subcutaneous A549 xenografts
displayed similar results (Fig.
8).
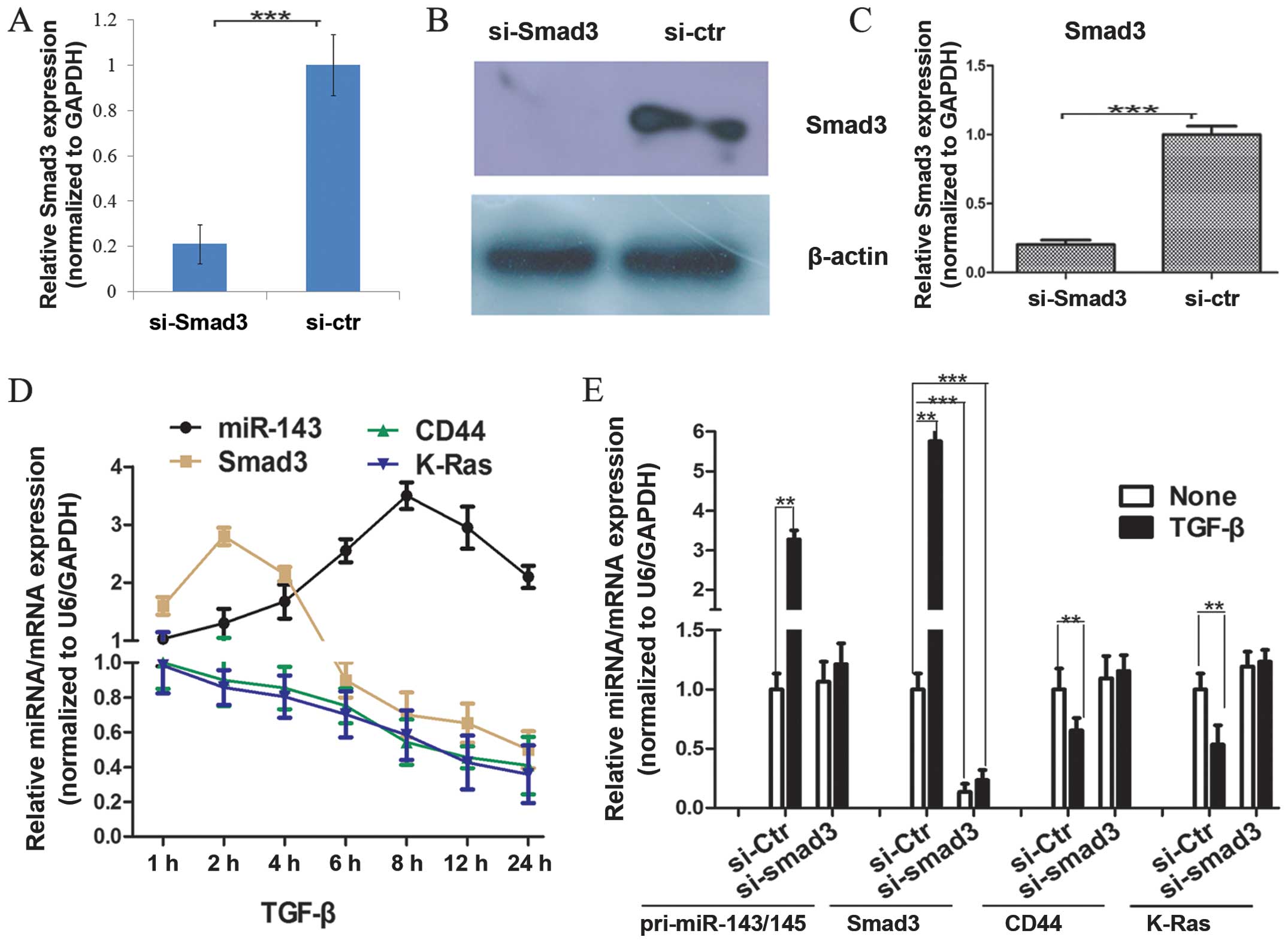
To assess the coordinated effects of TGF-β on
regulation of miR-143 and Smad3 expression, we first analyzed
expression of miR-143 and Smad3 in A549 cells after varying periods
of TGF-β treatment. Expression of miR-143 was slowly upregulated by
TGF-β from 1 to 8 h and reached a maximum after 8 h of treatment.
The opposite trend was observed for expression of CD44 and K-Ras
expression, which were downregulated at each time point from 1 h to
24 h (Fig. 9A). Smad3 expression,
however, was upregulated during the first 4 h, and then
downregulated from 6 to 24 h. Smad3, as a signal transducer that is
critical for transcriptional regulation by TGF-β, was first
stimulated by TGF-β, but later was downregulated by expression of
miR-143 that had been induced by TGF-β.
Furthermore, we knocked down Smad3 expression in
A549 cells using Smad3 siRNA (si-Smad3) followed by TGF-β treatment
which resulted in 85% suppression of Smad3 expression (Fig. 9A), induction of miR-143 expression
by TGF-β was not different than in TGF-treated cells; the TGF-β
treatment induced the pri-miR-143/145 expression without si-Smad3
in A549 cells. This would suggest that Smad3 plays a critical role
in transcriptional activation of miR-143 expression (Fig. 6D). TGF-β-induction of CD44 and
K-Ras mRNAs was abolished in si-Smad4-transfected cells (Fig. 9E).
Discussion
TGF-β is secreted by many types of normal cells, and
the TGF-β signaling pathway can inhibit cell cycle progression at
the G1 phase and therefore, suppress cell proliferation, but induce
cell differentiation, and promote apoptosis (13,14).
However, after transformation into cancer cells, the cells will
alter expression of TGF-β and its signaling pathway. Moreover,
TGF-β secretion is also induced in tumor-surrounding stoma cells
and TGF-β acts on tumor stoma cells, such as fibroblasts, immune
cells, endothelial and smooth-muscle cells causing tumor cell EMT,
immune suppression and angiogenesis for cancer progression
(25,26). In addition, previous studies showed
that miR-143 expression is downregulated in various cancers,
including lung cancer. For example, Yanaihara et al reported
that miR-143 expression was significantly decreased in 104 pairs of
lung cancer tissues (27). Gao
et al detected low expression of miR-143 in lung cancer
tissues using qRT-PCR (28). Ma
et al also found reduced levels of miR-143 expression NSCLC
tissues and cells (29). Thus, in
this study, we investigated the role of these molecules in NSCLC
progression. We first treated NSCLC A549 cells with TGF-β; miRNA
array analysis showed upregulation of miR-143, but our ex
vivo data showed and confirmed downregulation of miR-143 in
NSCLC tissues and lymph node-metastatic tissues. These data support
antitumor effects of TGF-β-induced miR-143 in NSCLC cells.
In order to verify antitumor effects of miR-143 in
NSCLC cells, we over-expressed miR-143 in A549 cells and observed
significantly reduced NSCLC cell viability and invasion. Wound
healing capacity in vitro and xenograft formation and growth
in nude mice further supported the notion of miR-143 as a tumor
suppressor gene in NSCLC. We then searched for the target gene of
miR-143 using Pictar, RNAhybrid, and TargetScan and identified
CD44, K-Ras and Smad3 as potential target genes. CD44 and K-Ras
were confirmed as the target genes of miR-143 by previous studies
(11,29–31).
CD44 is a cell-surface glycoprotein involved in cell-cell
interactions, adhesion, and migration (32,33).
It is a receptor for hyaluronic acid (HA) and can also interact
with other ligands, such as osteopontin, collagens, and matrix
metal lipoproteinases (MMPs). CD44 has a wide variety of cellular
functions, including lymphocyte activation, recirculation and
homing, hematopoiesis, and tumor metastasis (32–34).
K-Ras is a member of the small GTPase super family. The RAS (rat
sarcoma)/MAPK (mitogen-activated protein kinase) pathway plays a
key role in regulation of various physiological cellular processes,
including proliferation, differentiation, and cell death. Aberrant
mutational activation of K-Ras/MAPK signaling is closely linked to
cancer development in many organs in humans and laboratory animals
(23,35). We confirmed that expression of
miR-143-inhibited expression of CD44 and K-Ras in NSCLC cells.
Thus, NSCLC cell viability might be due to downregulation of K-Ras,
while miR-143-inhibited NSCLC cell invasion capacity might be due
to downregulation of CD44. In addition, one important element of
the TGF-β pathway, Smad3, is also a target gene of miR-143 that has
not been reported to date. In the present study, we verified that
miR-143 bound to Smad3 3′UTR, and expression of miR-143 reduced
Smad3 expression in NSCLC cells. There was an inverse association
between miR-143 and Smad3, CD44, and K-Ras in NSCLC tissue
specimens and A549 xenografts.
TGF-β plays different roles in normal cells vs.
tumors. For example, TGF-β acts as a tumor suppressor in normal
tissues or in early carcinogenesis (14,36,37),
but the tumor suppressive effects are lost in advanced stages of
cancer. In the present study, we hypothesized that TGF-β would
first induce miR-143 expression and upregulation of miR-143 would
in turn have a tumor suppressing function by inhibiting target
genes (such as CD44, K-Ras, and Smad3). Reduced Smad3 expression
could affect the TGF-β pathway by downregulated expression of the
Smad2/Smad3 complex. This complex will lose the effects on the
transcription factor binding to the DNA promoter, including the
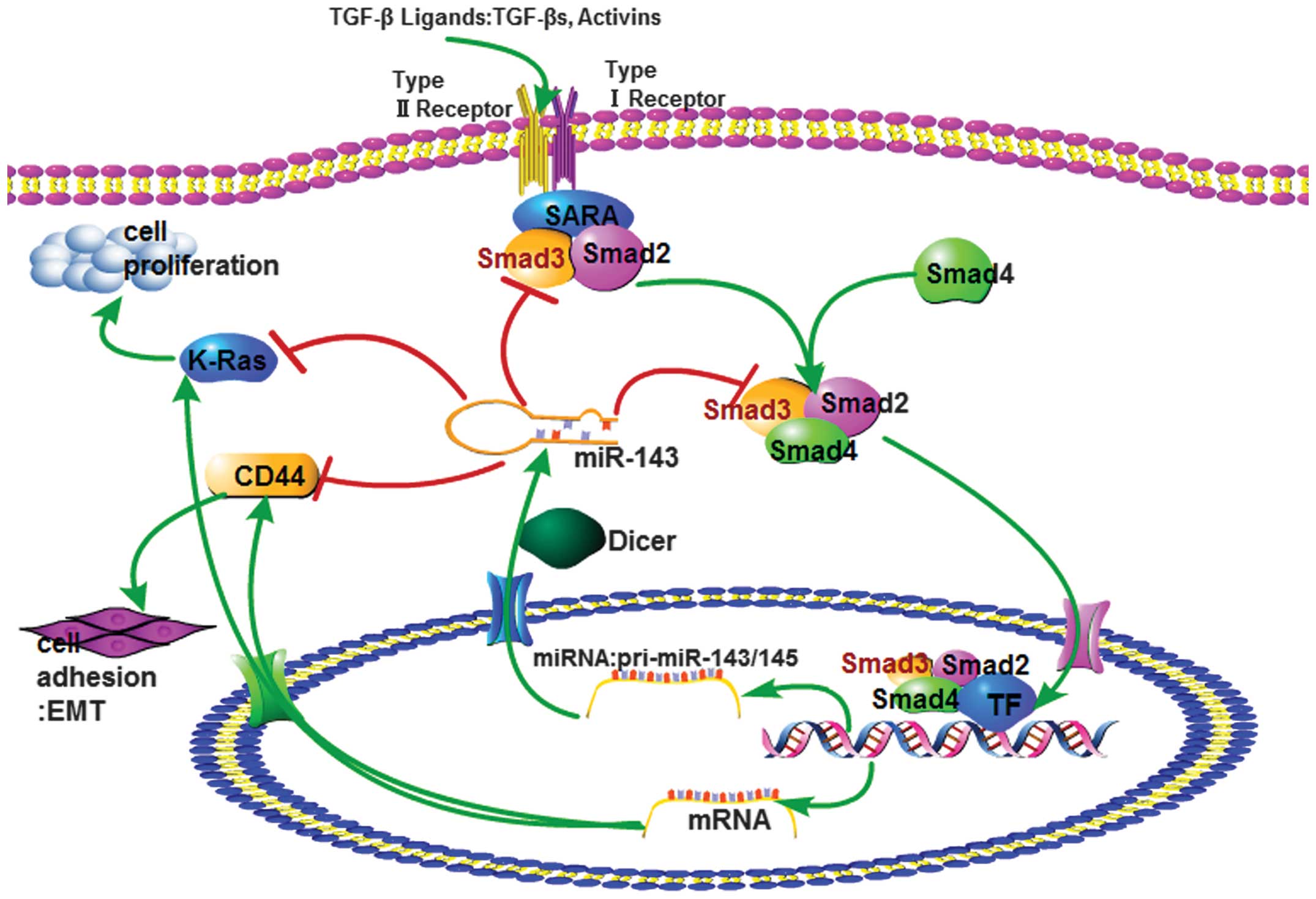
pri-miR-143 promoter. TGF-β binds to its receptor, which is
activated and then recruits the Smad2 and Smad3 to form a complex,
and in turn together with Smad4, translocates into the cell nucleus
to promote gene transcription, including expression of the
pri-miR-143/145 (Fig. 10). The
pri-miR-143/145 is then cleaved to the mature miR-143. miR-143
downregulated its target gene CD44 and K-Ras function as a tumor
suppressor gene. miR-143 also downregulates Smad3 expression, and
the downregulated Smad3 restrains TGF-β. This is the
miR-143-Smad3-TGF-β pathway loop. The loop might explain why TGF-β
acts as a tumor suppressor in normal tissues, and in early
carcinogenesis functions as an oncogene. TGF-β induces the
expression of miR-143 at first, and later the downregulated Smad3
represses miR-143 expression, enriching the miR-143 and TGF-β
pathway.
Our studies confirm that TGF-β could induce
expression of pri-miR-143/145, as miR-143 was frequently
downregulated in NSCLC and lymph node metastases. Our studies
demonstrate the biological functions of miR-143 in significantly
inhibiting the migration and invasion of NSCLC cells. We also
identified Smad3, CD44 and K-Ras as target genes possibly involved
in miR-143-mediated migration and invasion suppression for lung
cancer. Most importantly, we confirmed that TGF-β induced the
expression of miR-143, and later was repressed by the Smad3 target.
In this way, miR-143 and Smad3 composed a circular loop for cancer.
The miR-143 and TGF-β pathway might provide a potential therapeutic
strategy for NSCLC.
Acknowledgements
This study was supported in part by grants from
Hunan Provincial Innovation Foundation For Postgraduates
(CX2013B110), National Key Scientific & Technology Support
Program: Collaborative innovation of Clinical Research for chronic
obstructive pulmonary disease and lung cancer (2013BAI09B09) and
Science and Technology Department of Hunan Province Key Project
(2014FJ2009). Most of the financial support was provided by
C.H.M.
References
|
1
|
Field JK, Oudkerk M, Pedersen JH and Duffy
SW: Prospects for population screening and diagnosis of lung
cancer. Lancet. 382:732–741. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Curioni-Fontecedro A, Husmann L, Soldini D
and Stahel RA: Primary non-small cell lung cancer response upon
treatment with denosumab. Lung Cancer. 82:506–508. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Navarro A, Diaz T, Gallardo E, et al:
Prognostic implications of miR-16 expression levels in resected
non-small-cell lung cancer. J Surg Oncol. 103:411–415. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Vosa U, Vooder T, Kolde R, Vilo J,
Metspalu A and Annilo T: Meta-analysis of microRNA expression in
lung cancer. Int J Cancer. 132:2884–2893. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Yates LA, Norbury CJ and Gilbert RJ: The
long and short of microRNA. Cell. 153:516–519. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Chitwood DH and Timmermans MC: Small RNAs
are on the move. Nature. 467:415–419. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Lujambio A and Lowe SW: The microcosmos of
cancer. Nature. 482:347–355. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Ebert MS and Sharp PA: Roles for microRNAs
in conferring robustness to biological processes. Cell.
149:515–524. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Bartel DP: MicroRNAs: target recognition
and regulatory functions. Cell. 136:215–233. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Bowen T, Jenkins RH and Fraser DJ:
MicroRNAs, transforming growth factor beta-1, and tissue fibrosis.
J Pathol. 229:274–285. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Luo Z, Dai Y, Zhang L, et al: miR-18a
promotes malignant progression by impairing microRNA biogenesis in
nasopharyngeal carcinoma. Carcinogenesis. 34:415–425. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Liu L, Yu X, Guo X, et al: miR-143 is
downregulated in cervical cancer and promotes apoptosis and
inhibits tumor formation by targeting Bcl-2. Mol Med Rep.
5:753–760. 2012.PubMed/NCBI
|
|
13
|
Yoshida K, Saito T, Kamida A, et al:
Transforming growth factor-beta transiently induces vimentin
expression and invasive capacity in a canine mammary gland tumor
cell line. Res Vet Sci. 94:539–541. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Dong L, Ge XY, Wang YX, et al:
Transforming growth factor-beta and epithelial-mesenchymal
transition are associated with pulmonary metastasis in adenoid
cystic carcinoma. Oral Oncol. 49:1051–1058. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Tian M, Neil JR and Schiemann WP:
Transforming growth factor-beta and the hallmarks of cancer. Cell
Signal. 23:951–962. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Choe C, Shin YS, Kim SH, et al:
Tumor-stromal interactions with direct cell contacts enhance
motility of non-small cell lung cancer cells through the hedgehog
signaling pathway. Anticancer Res. 33:3715–3723. 2013.PubMed/NCBI
|
|
17
|
Huang S, Holzel M, Knijnenburg T, et al:
MED12 controls the response to multiple cancer drugs through
regulation of TGF-beta receptor signaling. Cell. 151:937–950. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Argast GM, Krueger JS, Thomson S, et al:
Inducible expression of TGFbeta, snail and Zeb1 recapitulates EMT
in vitro and in vivo in a NSCLC model. Clin Exp Metastasis.
28:593–614. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Tai J, Xiao X, Huang ZG, et al: MicroRNAs
regulate epithelial-mesenchymal transition of supraglottic
laryngeal cancer. Zhonghua Er Bi Yan Hou Tou Jing Wai Ke Za Zhi.
48:499–503. 2013.(In Chinese).
|
|
20
|
Peng X, Guo W, Liu T, et al:
Identification of miRs-143 and -145 that is associated with bone
metastasis of prostate cancer and involved in the regulation of
EMT. PLoS One. 6:e203412011. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Boominathan L: The tumor suppressors p53,
p63, and p73 are regulators of microRNA processing complex. PLoS
One. 5:e106152010. View Article : Google Scholar
|
|
22
|
Davis-Dusenbery BN, Chan MC, Reno KE, et
al: Down-regulation of Kruppel-like factor-4 (KLF4) by
microRNA-143/145 is critical for modulation of vascular smooth
muscle cell phenotype by transforming growth factor-beta and bone
morphogenetic protein 4. J Biol Chem. 286:28097–28110. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Wang YY, Ren T, Cai YY and He XY: MicroRNA
let-7a inhibits the proliferation and invasion of nonsmall cell
lung cancer cell line 95D by regulating K-Ras and HMGA2 gene
expression. Cancer Biother Radiopharm. 28:131–137. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Shi L, Campbell G, Jones WD, et al: The
MicroArray Quality Control (MAQC)-II study of common practices for
the development and validation of microarray-based predictive
models. Nat Biotechnol. 28:827–838. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Gordon KJ and Blobe GC: Role of
transforming growth factor-beta superfamily signaling pathways in
human disease. Biochim Biophys Acta. 1782:197–228. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Blobe GC, Schiemann WP and Lodish HF: Role
of transforming growth factor beta in human disease. N Engl J Med.
342:1350–1358. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Yanaihara N, Caplen N, Bowman E, et al:
Unique microRNA molecular profiles in lung cancer diagnosis and
prognosis. Cancer Cell. 9:189–198. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Gao W, Yu Y, Cao H, et al: Deregulated
expression of miR-21, miR-143 and miR-181a in non small cell lung
cancer is related to clinicopathologic characteristics or patient
prognosis. Biomed Pharmacother. 64:399–408. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Ma Q, Jiang Q, Pu Q, et al: MicroRNA-143
inhibits migration and invasion of human non-small-cell lung cancer
and its relative mechanism. Int J Biol Sci. 9:680–692. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Hu Y, Ou Y, Wu K, Chen Y and Sun W:
miR-143 inhibits the metastasis of pancreatic cancer and an
associated signaling pathway. Tumour Biol. 33:1863–1870. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Liu H, Zhang SZ, Cai SR, Peng JP and Zheng
S: Effect of microRNA143 expression on cell proliferation in
colonic carcinoma. Zhonghua Zhong Liu Za Zhi. 30:498–501. 2008.(In
Chinese).
|
|
32
|
Tabyaoui I, Tahiri-Jouti N, Serhier Z, et
al: Immunohistochemical expression of CD44s in human neuroblastic
tumors: Moroccan experience and highlights on current data. Diagn
Pathol. 8:392013. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Hiraga T, Ito S and Nakamura H: Cancer
stem-like cell marker CD44 promotes bone metastases by enhancing
tumorigenicity, cell motility, and hyaluronan production. Cancer
Res. 73:4112–4122. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Raso-Barnett L, Banky B, Barbai T, Becsagh
P, Timar J and Raso E: Demonstration of a melanoma-specific CD44
alternative splicing pattern that remains qualitatively stable, but
shows quantitative changes during tumour progression. PLoS One.
8:e538832013. View Article : Google Scholar
|
|
35
|
Zeller E, Hammer K, Kirschnick M and
Braeuning A: Mechanisms of RAS/beta-catenin interactions. Arch
Toxicol. 87:611–632. 2013. View Article : Google Scholar
|
|
36
|
Sawaki D and Suzuki T: Targeting
transforming growth factor-beta signaling in aortopathies in Marfan
syndrome. Circ J. 77:898–899. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Jakobsson L and van Meeteren LA:
Transforming growth factor beta family members in regulation of
vascular function: in the light of vascular conditional knockouts.
Exp Cell Res. 319:1264–1270. 2013. View Article : Google Scholar : PubMed/NCBI
|