Introduction
Breast cancer is the most frequent malignant disease
and the second cause of death from cancer in women in the United
States (1). In most of these
cases, relapse and consequent metastatic growth of cancer cells can
occur at distant sites, including bone, lung, liver and brain
(2–3). Metastasis to distant sites accounts
for >90% of breast cancer-related mortality (4). Despite its clinical importance,
metastasis remains the most insidious aspect of breast cancer, and
there are few successful treatments that directly target this
stage. Therefore, better understanding of the complexities of the
genetic and biochemical determinants of metastasis and developing
effective therapies are urgently needed to reduce the morbidity and
mortality from metastatic disease.
Natural products are still a major resource and have
become increasingly important for new drug discoveries (5). Chloranthaceae, a basal
angiosperm taxon, is composed of approximately 65–75 extant species
(6) and over 10 medical herbs,
most of which have antitumor associated usage (7). Sarcandra glabra Nakai,
colloquially known as Caoshanhu, is a medical herb in
Chloranthaceae that grows in the southern part of China
(8). It is widely used as a kind
of folk remedy for rheumatic arthralgia, bone fracture and other
ailments (7). Tablets of S.
glabra Nakai are clinically used as a well-known adjunctive
therapy agent for leukemia, pancreatic cancer, and liver cancer,
and can be found in the Pharmacopoeia of China (9). Unfortunately, the bioactive
ingredients and their mechanisms of action are largely unknown.
Interest has also been focused on the medical herb Chloranthus
japonicas (10,11), and the cytotoxicity of several
sesquiterpenes isolated from the whole herb was evaluated. In
addition, Kwon et al (12)
reported that dimeric sesquiterpenoids isolated from this herb
prevented monocyte adhesion to HUVECs via inhibiting the expression
of cell adhesion molecules.
Led by these concepts, we hypothesized that
antimetastatic activities may contribute to the antitumor related
usage of plants in Chloranthaceae. Thus, we focused our
interests on antimetastatic effects of four medical herbs in this
genius, Sarcandra glabra Nakai, Chloranthus henryi Hemsl,
Chloranthus fortune and Chloranthus multistachys Pei,
which have close relationships and grow in Jiangxi Province, China
(7). By invasion-inhibition-guide
fractionation, we screened 263 constituents from these plants,
including sesquiterpenes, diterpenes, chalcones, and sesquiterpene
lactones (data not shown). Codonolactone (CLT), one of the
sesquiterpene lactones isolated from C. henryi Hemsl,
exhibited the strongest anti-invasive properties in vitro.
In the present study, we report that CLT exhibited antimetastatic
effects in metastatic breast cancer cells, and these effects may be
mediated by inhibition of matrix metalloproteinases (MMPs) via
downregulation of the transcriptional activity of Runx2.
Materials and methods
Materials
CLT was purchased from Shanghai PureOne
Biotechnology (Shanghai, China; P0110) with a purity of ≥98.0%, as
determined by high-performance liquid chromatography. CLT was
dissolved in 100% dimethyl sulfoxide (DMSO) as a stock solution and
stored at −20°C. The final DMSO concentration did not exceed 0.1%
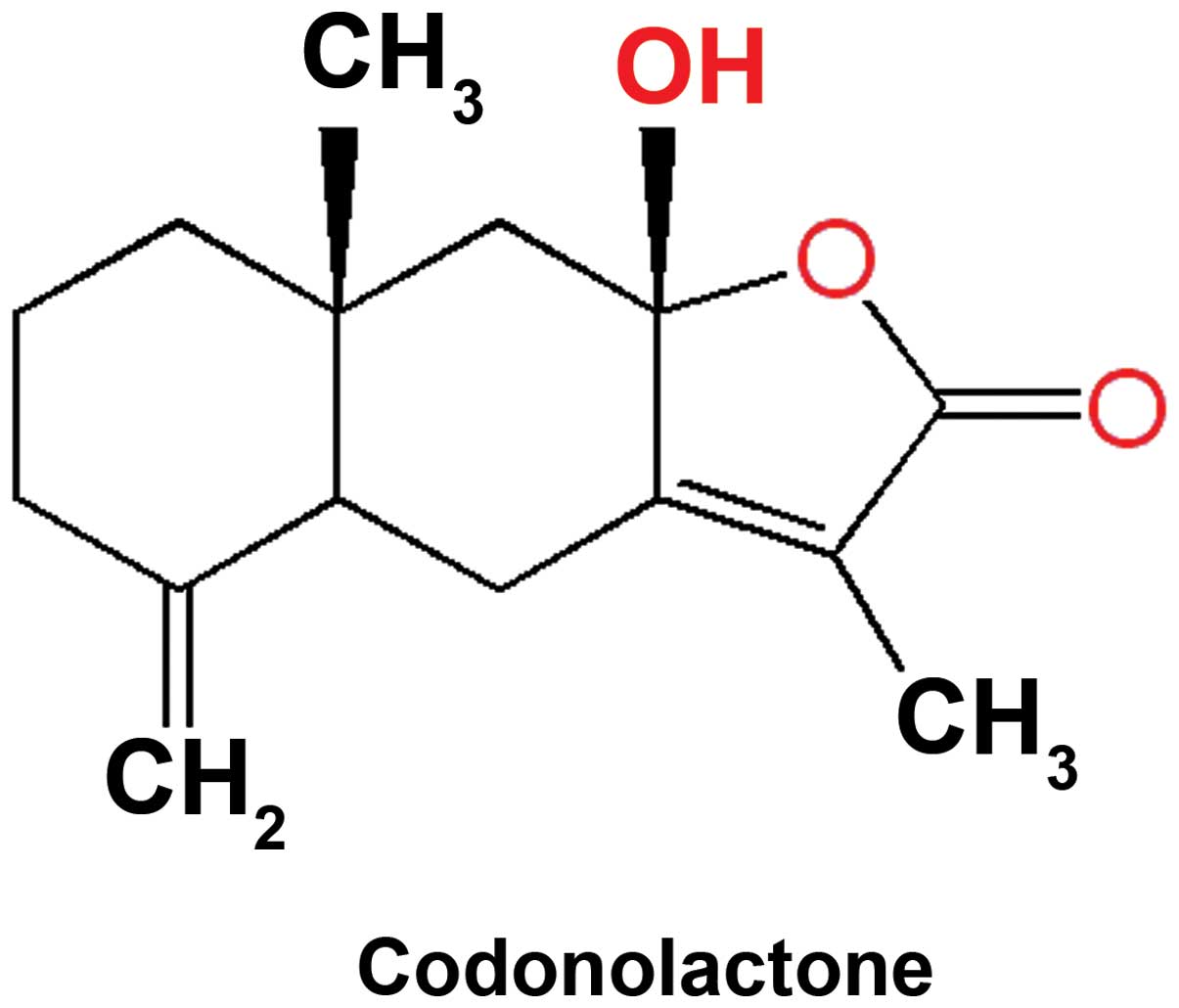
throughout the study. The molecular structure of this sesquiterpene
lactone is presented in Fig. 1.
Batimastat (Bat), a broad spectrum MMP inhibitor, was purchased
from Santa Cruz Biotechnology Inc. (SC-203833, Santa Cruz, CA,
USA), and served as a positive control.
Cell culture
MDA-MB-231 human breast cancer cells, purchased from
Cell Resource Center, Institute of Basic Medical Sciences, Chinese
Academy of Medical Sciences, were maintained in Dulbecco’s modified
Eagle’s medium (DMEM) supplemented with 10% fetal bovine serum
(FBS; Gibco, Inc.), 100 IU/ml penicillin, and 100 μg/ml
streptomycin (Invitrogen, Inc.) in a humidified incubator
containing 5% CO2 at 37°C. Another breast cancer line,
MDA-MB-157 was purchased from Sigma-Aldrich China (Beijing, China).
MDA-MB-157 cells were maintained in L15 supplemented with 10% fetal
bovine serum (FBS; Gibco, Inc.), 2 mM glutamine.
Animals and ethics statement
Five- to 6-week-old female NOD/SCID mice were
purchased from Vital River Laboratories (Beijing, China) to
establish an orthotopic xenograft tumor model and an experimental
lung metastasis xenograft model. All the animal experiments were
conducted in strict accordance with full ethical approval of the
Animal Ethics Committees of Jiangxi University of Traditional
Chinese Medicines. Surgery was performed under sodium pentobarbital
anesthesia, and all efforts were made to minimize suffering.
In vitro invasive assay
In vitro invasive capacity of cancer cells
was measured using the 48-well microchemotaxis system (AP 48, Neuro
Probe, Gaithersburg, MD, USA). Briefly, a polycarbonate membrane
with 8-μM pore size was pre-coated with 5 μg of fibronectin in a
volume of 50 μl on the rough (lower) surface. The Matrigel was
diluted to 100 μg/ml with cold phosphate-buffered saline (PBS) and
applied to the smooth (upper) surface of the filters (5 μg/filter),
and the filters were dried at room temperature. The lower
compartments of the plates were filled with 30 μl DMEM containing
0.1% bovine serum albumin (BSA). Log-phase cells were harvested,
washed three times with serum-free medium, and resuspended into a
final concentration of 2×106/ml in DMEM with 0.1% BSA.
Cell suspensions (100 μl) containing 2×105 cells with or
without CLT were added to the upper compartment and plates were
incubated for 14 h in an incubator containing 5% CO2 at
37°C. The filters were fixed with methanol for 10 min, stained with
0.5% crystal violet for 60 min, and then washed with distilled
water. The cells on the upper surface of the filters were removed
by wiping with cotton swabs. The cells invading the lower surface
of the filter through the Matrigel and filter were manually counted
under light microscopy (Leica DM 4000), and images were captured by
the Leica Application Suite (LAS V3.8.0).
In vitro migratory assay
In vitro migration of breast cancer cells was
measured using Oris™ cell migration assay (Platypus Technologies,
Madison, WI, USA) following the manufacturer’s instructions.
Log-phase cells were harvested, washed three times with serum-free
medium, and resuspended into a final concentration of
4×106/ml in culture medium. Suspended cells (100 μl)
were pipetted into each test well through one of the side ports of
the Oris™ Cell Seeding Stopper. The seeded plate containing the
Oris Cell Seeding Stoppers in a humidified chamber (37°C, 5%
CO2) was incubated for 24 h to permit cell attachment.
Using the Stopper Tool, all the stoppers were removed. Then, media
was gently removed and the wells washed with PBS to remove the
unattached cells. After that, 100 μl of FBS-free media containing
Calcein AM (final concentration 0.5 μg/ml) was added, and the cells
were incubated in a humidified chamber (37°C, 0.5% CO2)
for 40 min. The images were captured under a fluorescence
microscope (Leica DM 3000B), and these data served as the initial
control. After fluorescence intensity examination, media were
removed gently, and washed with PBS for 2 times. Then, 100 μl of
media containing vehicle, Bat or CLT was added, and the cells were
incubated in a humidified chamber (37°C, 0.5% CO2) for
24 h. At the end of treatment, data were obtained with the methods
mentioned above.
Cell proliferation assay
Cell proliferation was assayed by MTT assay.
Briefly, log phase cells were seeded in 96-well plates 24 h before
initiation of treatment with CLT. The vehicle-treated cells served
as controls. Three duplicate wells were set up in each sample.
After incubation with CLT, Bat or vehicle for 48 h, cells were
incubated with 3-(4, 5-dimethylthiazol-2-yl)-2,
5-diphenyltetrazolium bromide (MTT, Sigma-Aldrich) (final
concentration 0.5 mg/ml) for 4 h at 37°C. The media was carefully
removed from each well and 200 μl of DMSO was added. The plates
were gently agitated and optical absorbance at OD570 nm and OD450
nm was determined using an EL×800 microplate reader (BioTek,
Winooski, VT, USA). Vehicle only-treated cells served as the
indicator of 100% cell viability. Each experiment was repeated
three times.
In vivo growth assay
For the growth assay, MDA-MB-231 cell xenografts
were established by injection of 1×107 cells to the
mammary fat pad in NOD/SCID mice. After 3 weeks of growth, the
tumors were removed and chopped into 1×1×1-mm pieces, and the
pieces were then implanted at the mammary fat pad of mice. The mice
bearing tumor chunks were randomly divided into three groups:
control, CLT (25 mg/kg per day) treatment group, and CLT (75 mg/kg
per day) treatment group. Forty-eight hours later, CLT was
administered by gavage on a regimen of 6-day dosing per week for 5
weeks. Tumor growth was assessed by measuring the length and width
of tumors with electronic calipers every 3–4 days continuously.
Volumes were calculated using the formula: (length) ×
(width)2/2.
In vivo experimental metastasis assay and
survival assay
To establish experimental lung metastasis xenograft
model, 2×106 MDA-MB-231 cells in 200 μl normal saline
were injected through tail vein into NOD/SCID mice. The mice were
also divided into control, CLT (25 mg/kg per day) treatment group,
and CLT (75 mg/kg per day) treatment group. CLT treatment began 24
h after tumor injection. After 8-week treatment, the formation of
metastatic foci in lung tissues was measured by nest
reverse-transcription polymerase chain reaction (RT-PCR). The mice
were euthanized and dissected, and lungs were snap-frozen in liquid
nitrogen. Total RNA was isolated from each lung using the
QIAshredder and RNeasy Protect Mini kit (Qiagen) for detection of
human cytokeratin 19 (ck19) by nested PCR.
For the survival assay, the experimental lung
metastasis xenograft model was established using the same method as
in vivo experimental metastasis assay, but the observation
period was 120 days.
RNA isolation and nested RT-PCR
Nested RT-PCR was used to detect expression of the
human ck19 gene in lung tissues of tumor-bearing mice. Total RNA
was isolated from each lung using the QIAshredder and RNeasy
Protect Mini kit (Qiagen). Ck19 was reverse transcribed in the
presence of 2 μl enzyme mix, 10 μl RNA and outer primer for 30 min
at 50°C, according to the manufacturer’s instructions (Qiagen
OneStep RT-PCR kit, Qiagen). After Taq polymerase activation for 15
min at 95°C, samples were amplified for 37 cycles at 94°C for 45
sec, 58°C for 45 sec and 72°C for 90 sec. Target primers for
amplifying ck19 (outer primer) were designed using Primer Designer
(Scientific & Educational Software Version 2.0). The forward
primer for ck19 (GenBank no. BC010409) was 5′-cca cgt cgt cct tcg
gag gcc-3′ (64–84 bp) and reverse primer was 5′-gttc cgt ctc aaa
ctt ggt tc-3′ (529–549 bp). After a final extension for 10 min at
72°C, the RT-PCR product was further subjected to nested PCR using
the Qiagen Multiplex PCR kit (Qiagen). The inner primers were
forward 5′-tac agc cac tac tac acg acc atc c-3′ (432–456 bp) and
reverse 5′-gga caa tcc tgg agt tct caa tg-3′ (488–510 bp). The
nested PCR profile was as follows: 3 min at 94°C, followed by 35
three-step cycles of 45 sec at 94°C, 45 sec at 60°C and 1 min at
72°C. PCR reactions were subjected to final extension at 72°C for
10 min. Nested RT-PCR analysis was performed using the Mastercycler
gradient (Eppendorf). The β-actin gene was used as an internal
control for standardization, and the primers, Tm, and cycles are
the same as reported (13). The
nested PCR product was separated by 4% agarose (UltraPure™ Agarose,
Invitrogen) gel electrophoresis, and the gels were viewed by UV
transillumination and photographed by the UVP EC3 gel imaging
system.
FRET-based MMP activity assay
Activity of MMPs was also measured by the
SensoLyte® 570 Generic MMP assay kit (AnaSpec, Fremont,
CA, USA). This kit provides a FRET-based method to detect the
activities of a variety of MMPs including MMP-1, 2, 3, 7, 8, 9, 10,
11, 12, 13 and 14. It uses 5-FAM (fluorophore) and QXL520™
(quencher) labeled FRET peptide substrates for continuous
measurement of MMP activity. In an intact FRET peptide, the
fluorescence of 5-FAM is quenched by SensoLyte. Upon the cleavage
of FRET peptide by MMPs, the fluorescence of 5-FAM is recovered.
Analyses were performed according to the manufacturer’s
instructions. Briefly, supernatants of breast cancer cells were
collected after incubation with or without CLT for 12 h. The MMPs
in supernatants were activated by incubation with
4-aminophenylmercuric acetate for 1 h at 37°C. MMPs containing 50
μl of samples and 50 μl MMP substrate solution were added into a
96-well plate. The reagents were mixed by shaking the plate gently
for 30 sec. After a 50-min incubation period at 37°C, the reaction
was stopped by adding stop solution, and fluorescence intensity was
measured by a multilabel counter (Victor3™,
Perkin-Elmer, Waltham, MA, USA) at Ex/Em=540/575 nm.
Western blot analysis
Cells were rinsed twice with PBS, and total proteins
were extracted in 500 μl lysis buffer. Aliquots of whole cell
lysates were added to 10% SDS-PAGE and then transferred to Hybond
nitro-blotting membranes. The membranes were blocked with 3% BSA in
Tris-buffered saline containing 0.5 ml/l Tween-20 and then
incubated with primary antibodies against MMP-9 (Santa Cruz,
SC-6840), MMP-13 (Santa Cruz, SC-30073), Runx2 (Santa Cruz,
SC-10758) and phospho-Runx2 (Abgent, AP3559a), followed by
incubation with horseradish peroxidase (HRP)-conjugated secondary
antibodies. Immunoreactive proteins were detected using an enhanced
chemiluminescence kit (Millipore). β-actin (Santa Cruz, SC-130301)
served as an internal control.
Immunohistochemistry
MMP-13 in tumor tissues was determined by
immunostaining. Formalin-fixed, paraffin-embedded MDA-MB-231 tumor
tissues were cut into 3-μm thick sections and mounted on
poly-lysine coated slides. After dewaxing and hydration, antigen
retrieval was performed by incubation with Proteinase K at 37°C for
20 min. After three 5-min rinses in PBS (0.01 mol/l), the sections
were treated with 3% H2O2 in methanol for 10
min to block endogenous peroxidase activity per the manufacturer’s
instructions (Zhongshan Golden-bridge Biotech., Beijing, China).
Sections were then blocked with 10% normal goat serum (Nichirei,
Tokyo, Japan) for 30 min at room temperature. The tissues were
incubated with primary antibodies against MMP-13 (Santa Cruz,
SC-30073; 1:50) or PBS at 4°C overnight. After three rinses in PBS,
the slides were incubated with HRP-labeled IgG (SC-2004, Santa
Cruz) for 60 min, and then peroxidase activity was detected by the
AEC Substrate System (ab64252, Abcam). All the slides were examined
by light microscopy (Leica DM 4000), and images were captured using
the Leica Application Suite (LAS V3.8.0).
Runx2 transcription factor assay
Runx2 activity of nuclear extracts was detected
using the TransAM™ AML-3/Runx2 kit (Active Motif North America,
Carlsbad, CA, USA) following the manufacturer’s instructions. Cell
extracts were prepared using the Nuclear Extract kit (Active Motif)
with breast cancer cells that were treated with either CLT or DMSO
for 12 h. Then, 20 μl of extracts diluted in complete lysis buffer
and containing 15 μg nuclear extract were added into a 96-well
plate. This plate immobilizes oligonucleotides containing Runx2
consensus binding sites. Saos-2 nuclear extract served as a
positive control for Runx2 activation, and 20 μl complete lysis
buffer served as the blank. The wild-type consensus oligonucleotide
was provided as a competitor for Runx2 binding to monitor the
specificity of the assay. After 1-h incubation at room temperature,
the plate was washed three times with washing buffer. Diluted
primary antibody (100 μl) was added into wells and incubated for 1
h at room temperature without agitation. After three washes,
HRP-labeled secondary antibody was added and incubated for 1 h at
room temperature. Then, 100 μl developing solution was added to
initiate the color reaction. After 100 μl stop solution was added,
the absorbance was measured within 5 min at 450 nm with a reference
wavelength of 655 nm using an EL×800 microplate reader
(BioTek).
Chromatin immunoprecipitation assay
Chromatin immunoprecipitation assay (ChIP) was
performed using the Agarose ChIP kit as described in the
manufacturer’s instructions (Pierce™ Agarose ChIP kit, Thermo
Scientific). Briefly, cross-linking was performed for 10 min using
formaldehyde (final concentration 1%). Cross-linked cells then were
lysed by lysis buffer containing protease inhibitors. To obtain DNA
fragments with an average size of 0.3 kb, micrococcal nuclease was
added and incubated for 5 min. Protein-DNA complexes were
immunoprecipitated using anti-Runx2 antibody (Santa Cruz,
SC-10758). Normal rabbit IgG served as a negative control and
anti-RNA polymerase II antibody served as positive control. After
DNA recovery, purified DNA was subjected to PCR amplification on
the Mastercycler gradient (Eppendorf) using Phusion®
High-Fidelity PCR kit. The primers designed to amplify one of the
Runx2 binding regions (−1531 bp: acaccaa) in human mmp-13 promoter
(GeneBank no. NM_002427) were forward, 5′-aac ttg gta gct ttt atg
gtg g-3′ and reverse, 5′-gcc tct tca tca gat aat aag gg-3′. The PCR
profile was as follows: 15 min at 95°C, followed by 30 three-step
cycles of 15 sec at 95°C, 30 sec at 57°C and 30 sec at 72°C. PCR
reactions were subjected to final extension at 72°C for 10 min. PCR
analysis was performed using the Mastercycler gradient (Eppendorf).
Aliquots of total DNA before immunoprecipitation were saved as
input, and these input lysates were also processed as above. All
ChIP assays were performed at least three times, and the most
representative results are illustrated in the figures.
Statistical analysis
The data are presented as mean ± SD and were
analyzed with SPSS for Windows (13.0) software program (Chicago,
IL, USA). Comparison among different groups was carried out by
one-way analysis of variance (the one-way ANOVA). Analysis of
number of mice bearing metastatic foci in lung was performed by
χ2 test. The difference between the means was considered
statistically significant at p<0.05.
Results
CLT inhibits invasion and migration of
breast cancer cells in vitro
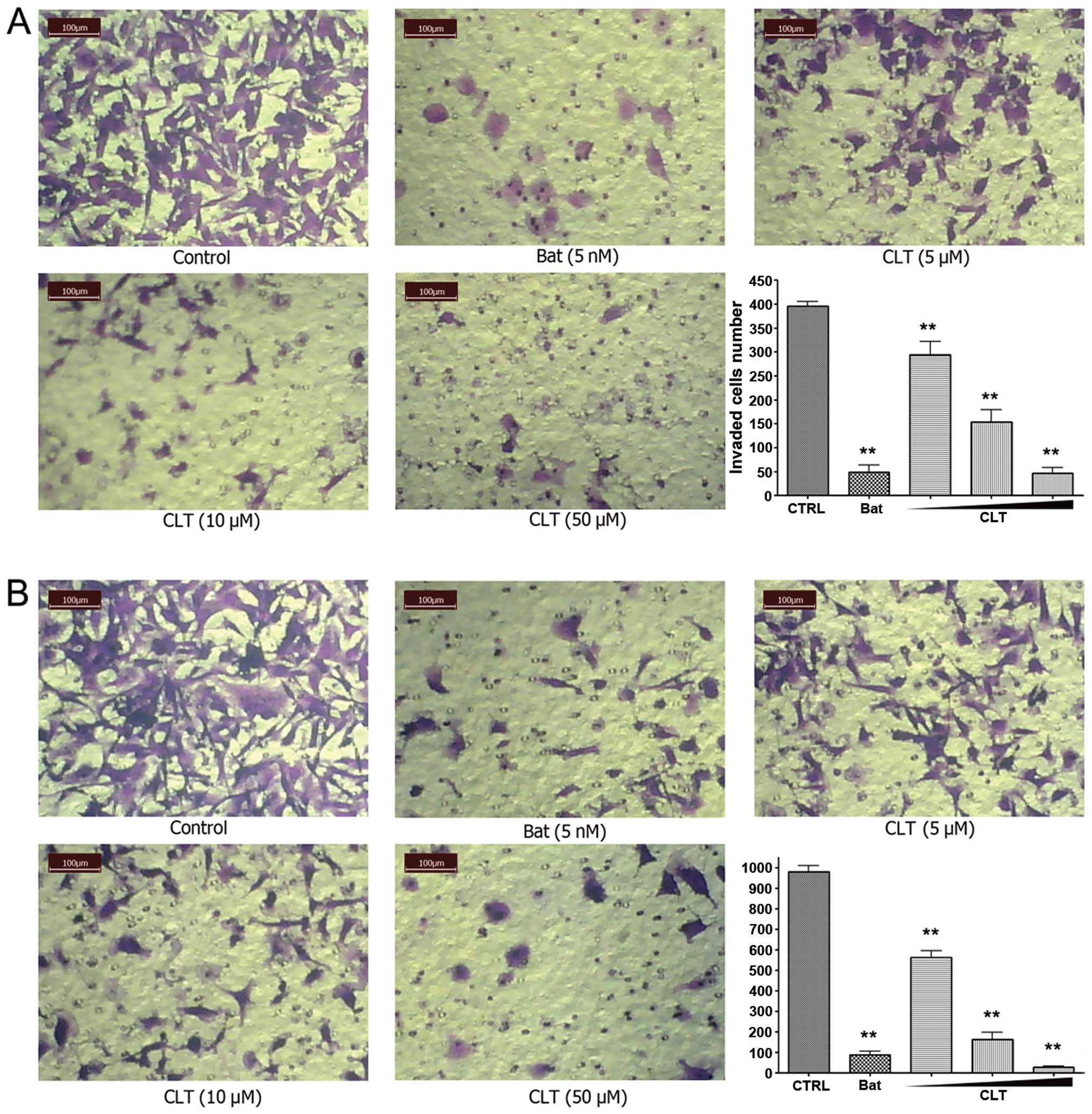
First, we evaluated the effects of CLT on invasion
using a reconstituted basement membrane system. We used fibronectin
as a chemo-attractant on the lower surface of a polycarbonate
membrane. Matrigel was plated on the upper surface of the
polycarbonate membrane to mimic the basement membrane. After 14-h
incubation, CLT significantly blocked the trans-membrane invasion
of MDA-MB-231 cells, with an 88.37% inhibition rate at 50 μM
(Fig. 2A). Invasion of MDA-MB-157
cells was also significantly blocked by CLT (Fig. 2B), with inhibitions of 42.58, 83.32
and 97.15% at concentrations of 5, 10 and 50 μM, respectively.
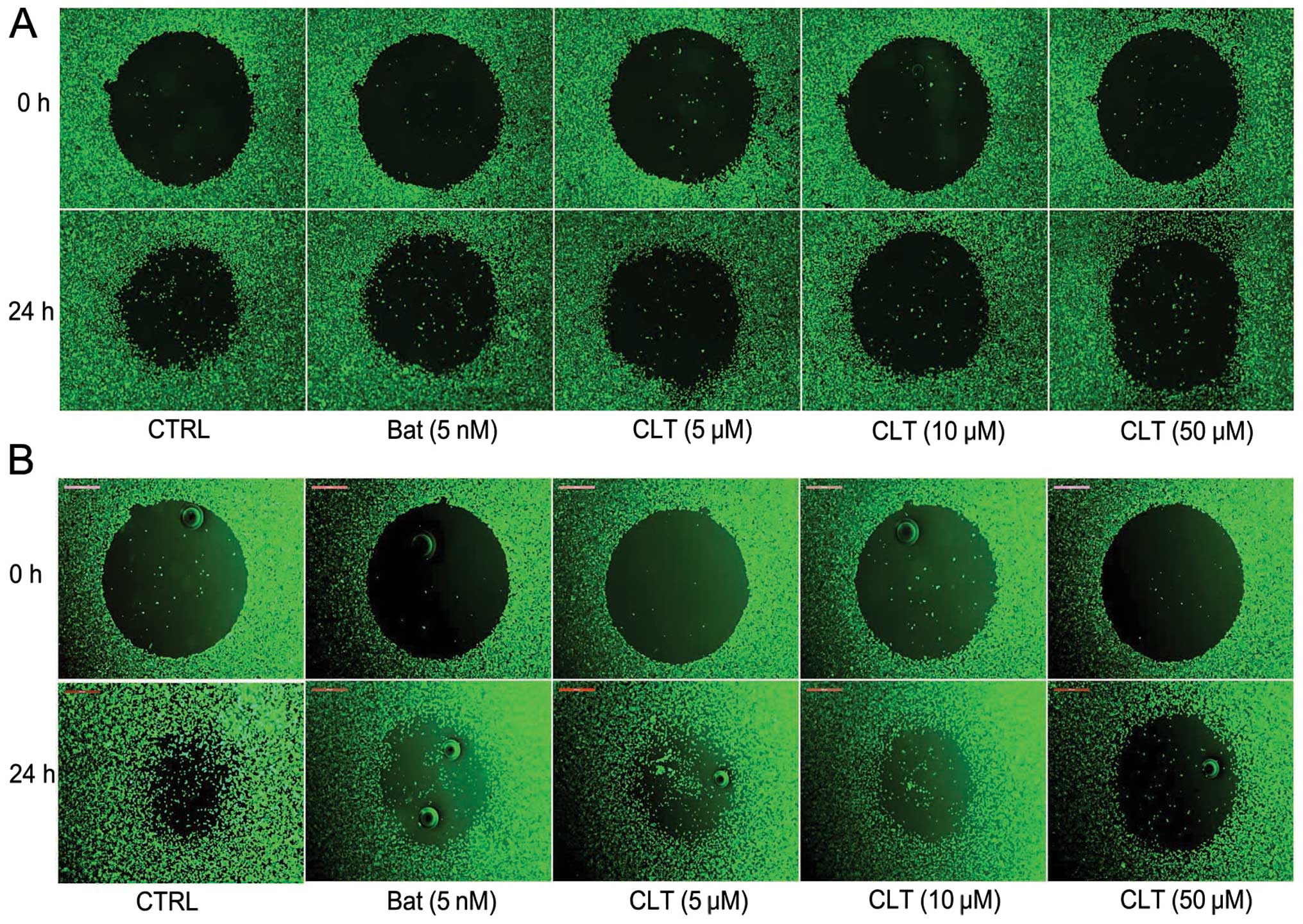
Next, the effect of CLT on cell migration in vitro was
tested using Oris cell migration system. As shown in Fig. 3, after 24-h incubation, CLT
inhibited the migration of breast cancer cells in a dose-dependent
manner. These results indicate that CLT could inhibit the invasion,
migration of MDA-MB-231 and MDA-MB-157 cells
concentration-dependently.
CLT slightly suppresses tumor growth in
vitro and in vivo, but induces a significant increase of overall
survival and a considerable inhibitory effect on formation of lung
metastatic foci in NOD/SCID mice
To assess the effect of CLT on the proliferation of
MDA-MB-231 and MDA-MB-157 cells, we first observed the growth
status of these cells treated by CLT or vehicle using the MTT
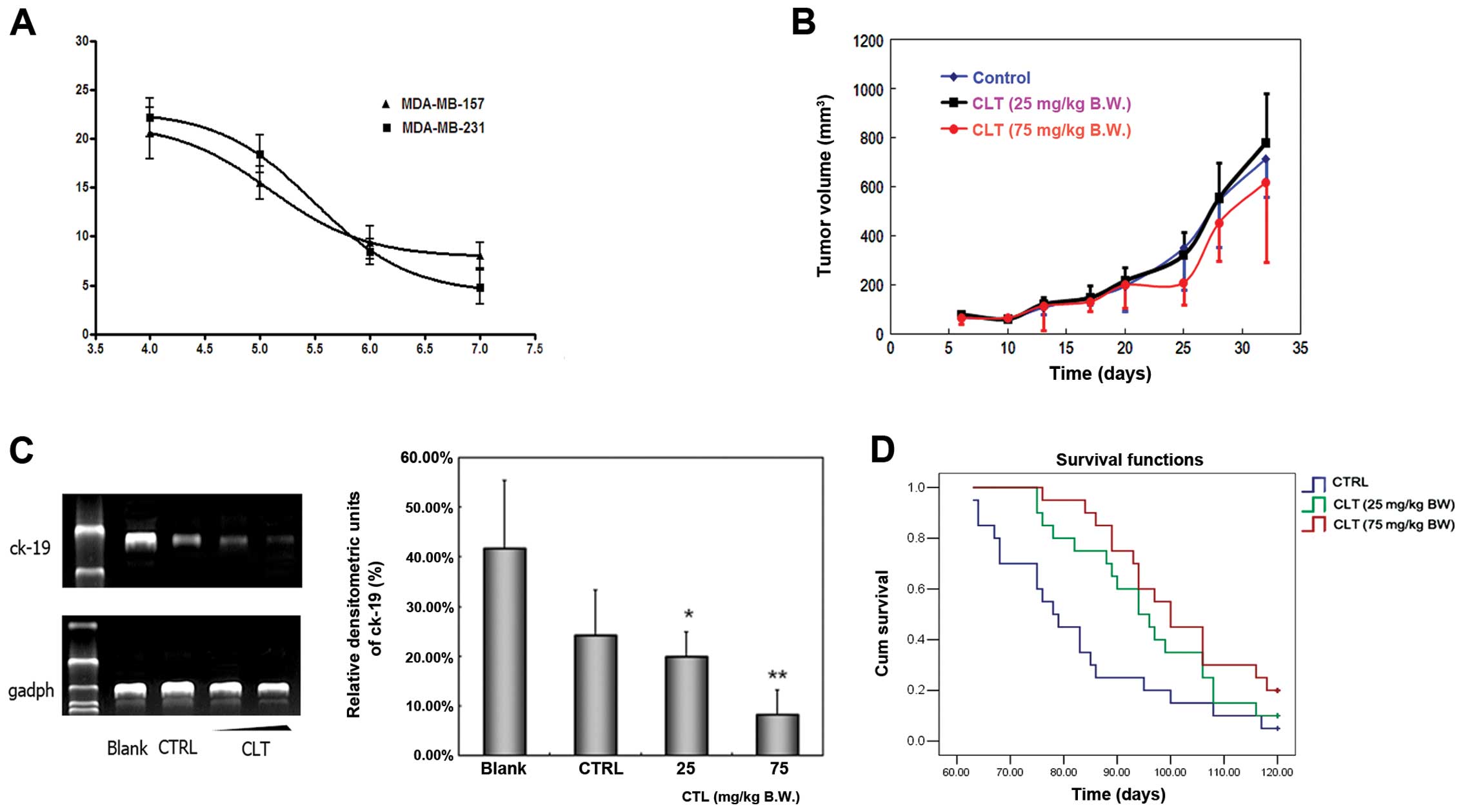
assay. The growth curve is shown in Fig. 4A. The proliferation of MDA-MB-231
and MDA-MB-157 cells was inhibited slightly after CLT exposure for
48 h. The inhibitory rate on MDA-MB-231 cell growth was 22.19% when
the cells were treated with 100 μM CLT for 48 h, and the inhibition
rate was 20.60% on MDA-MB-157 cells. Next, we examined the
inhibitory effects of CLT in vivo. An orthotopic xenograft
model was established by injection of MDA-MB-231 cells into mammary
fat pads of NOD/SCID mice. The tumor volume curve during this study
is shown in Fig. 4B. Compared with
vehicle-treated animals, CLT administered at 75 mg/kg/day caused
very slight inhibition on primary tumor growth.
Next, the antimetastatic effects of CLT were
evaluated using experimental metastasis model. MDA-MB-231 cells
spreading to the lung of cancer cell injected NOD/SCID mice were
assessed by detecting the expression of human ck19 using nested
PCR. The most typical bands and the statistical results of relative
densitometric units of lung tissues in each group are illustrated
in Fig. 4C. CLT treatment showed
overt inhibition of human ck-19 expression in lung tissues of mice
in a dose-dependent manner. Compared to control group, the average
inhibition rate of CLT treatment group (25 and 75 mg/kg/day) was
17.67 and 65.56%, respectively (Fig.
4C). These results suggest that the primary effects of CLT were
not in the growth inhibition of tumor cells, but in metastatic
suppression.
To better understand the overall benefits of CLT, we
evaluated the effect of CLT on the overall survival of tumor
cell-injected mice after treatment for 120 days. As shown in
Fig. 4D, both CLT-treated groups
showed a prolonged survival and an increasing survival time
dose-dependently. Animals in the control group died 63 days and
onwards after tumor inoculation, and the median survival period is
78 days. There is only 1 of the total 20 surviving mice at the
endpoint under this condition. However, for groups treated with CLT
25 and 75 mg/kg/day, the median survival days ranged from 94 to 100
days, and there were still two and four survivals at the endpoint,
respectively. These data indicated that tumor bearing mice may
benefit from CLT treatment, which may be associated with its
antimetastatic effects.
CLT inhibits MMP activity and expression
in MDA-MB-231 and MDA-MB-157 cells
Extensive work on the mechanisms of tumor invasion
and metastasis has identified MMPs as key players in these events
(14). Therefore, we next examined
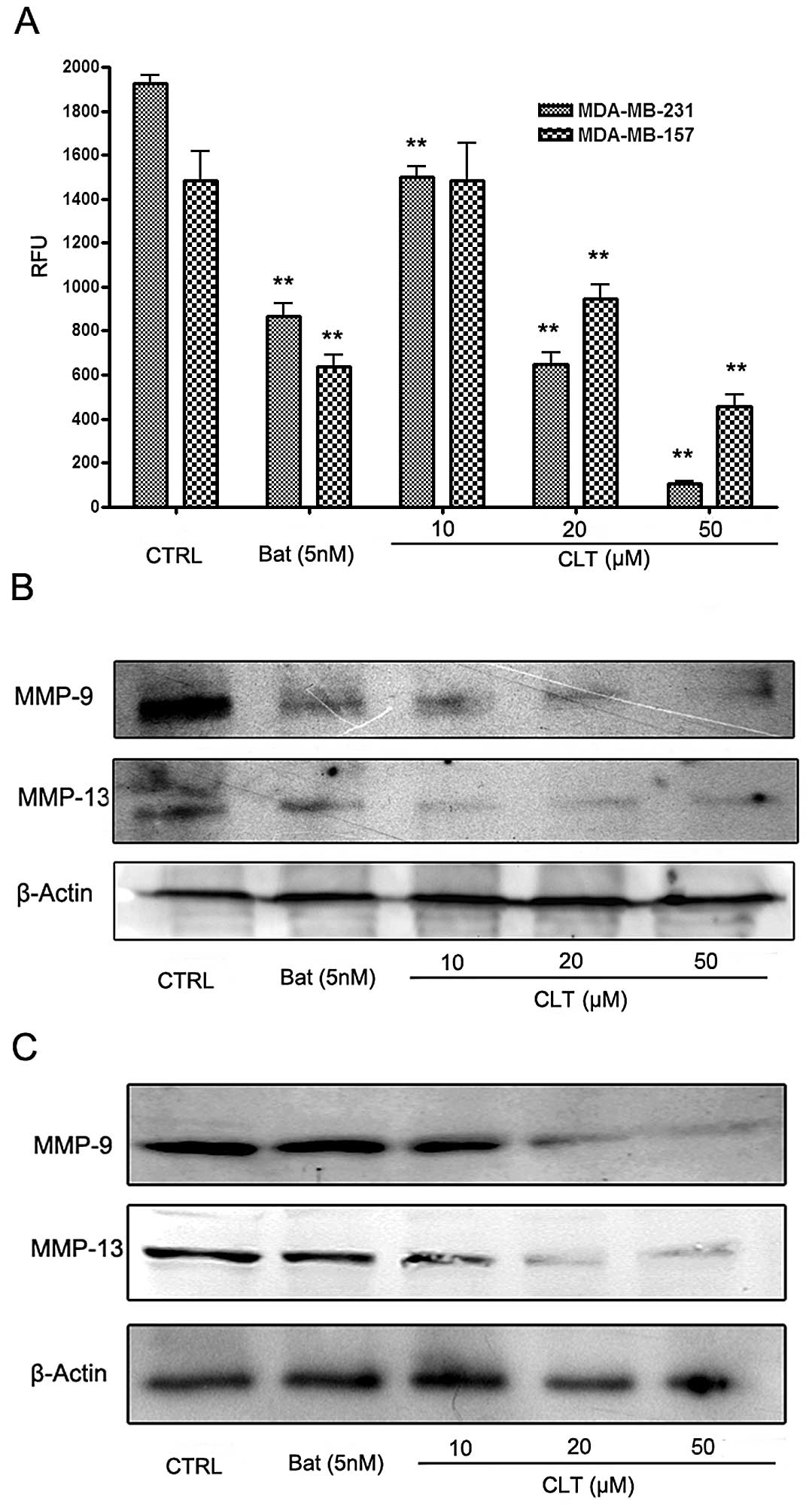
the effects of CLT on MMPs in breast cancer cells. Fig. 5 shows the effects of CLT on MMPs in
the metastatic breast cancer cells. Using a FRET-based analysis, we
found that CLT exhibited significant suppression of FRET substrate
cleavage of MMPs in a dose-dependent manner (Fig. 5A). To further identify whether CLT
inhibited the functional activity or expression of MMPs in the
breast cancer cells, we analyzed the expression of MMP-9 and MMP-13
in both culture cells and tumor tissues treated with CLT. As shown
in Fig. 4B and C, CLT
significantly decreased MMP-13 expression in MDA-MB-231 and
MDA-MB-157 breast cancer cells in a dose-dependent manner. Then,
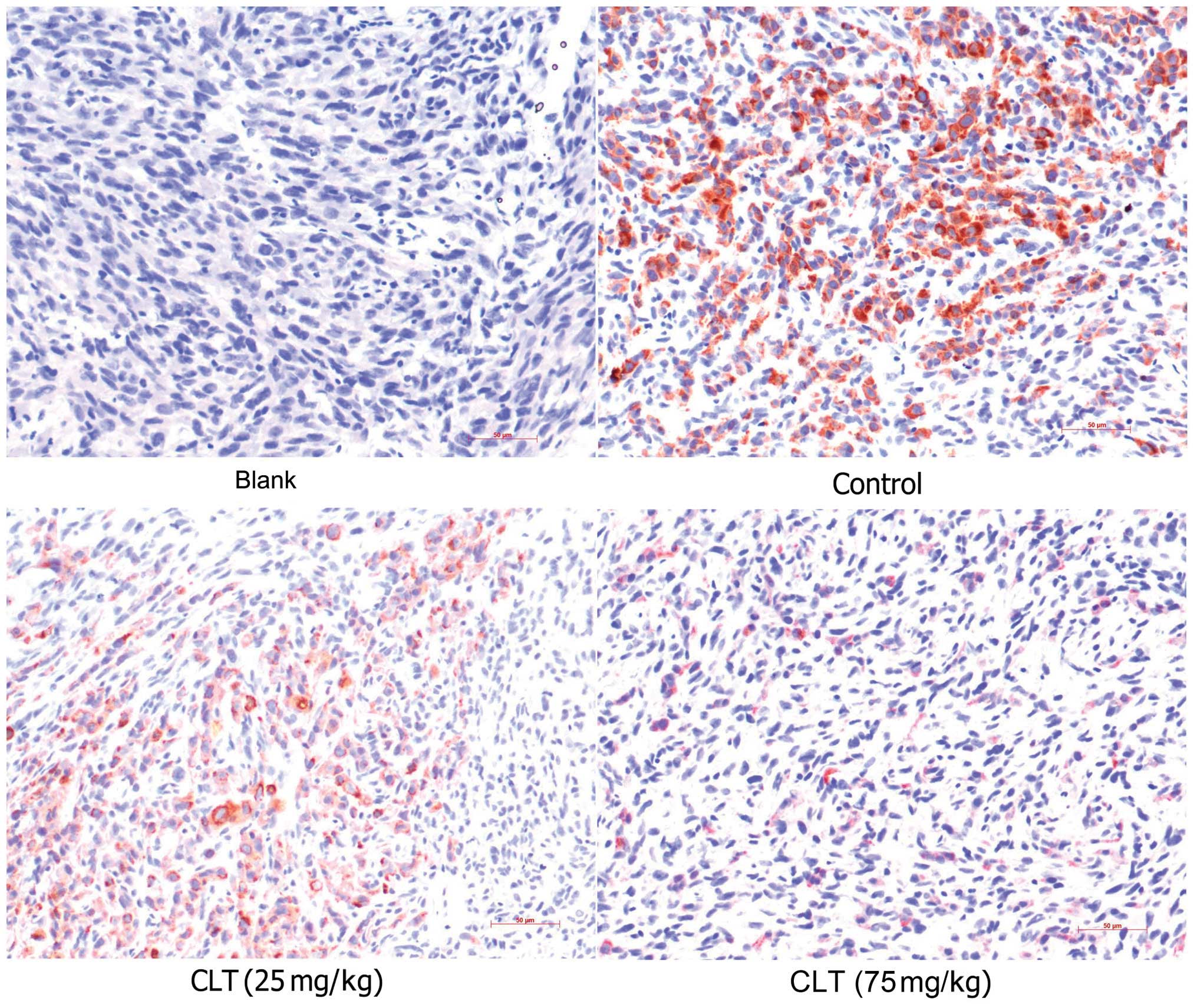
IHC assay was involved in present study to confirm the effects of
CLT on MMP-13 expression. It was also observed that CLT inhibited
MMP-13 expression in tumor tissues (Fig. 6 and Table I). These data suggested that the
antimetastatic properties of CLT may be due to downregulation of
expression of MMP-9 and MMP-13 in breast cancer.
 | Table ISemiquantitative analysis of MMP-13
in MDA-MB-231 tumor tissues. |
Table I
Semiquantitative analysis of MMP-13
in MDA-MB-231 tumor tissues.
| Group | Dose (mg/kg) | Relative area of
MMP-13 immunostaining (%)a | Inhibition rate
(%) |
|---|
| Blank | - | 1.62±0.69 | - |
| CTRL | - | 34.89±10.40 | - |
| CLT | 25 | 25.61±5.72 | 26.60 |
| 75 | 8.66±2.12b | 75.18 |
CLT suppresses activation of Runx2 and
impedes its binding to sequences in the mmp-13 promoter in
MDA-MB-231 cells
From current concepts, we understood that Runx2 is a
‘master’ transcriptional factor of metastatic growth of breast
cancer cells. Several genes required for the formation of
metastatic foci, including MMP-9, MMP-13, BSP, OPN, VEGF, are
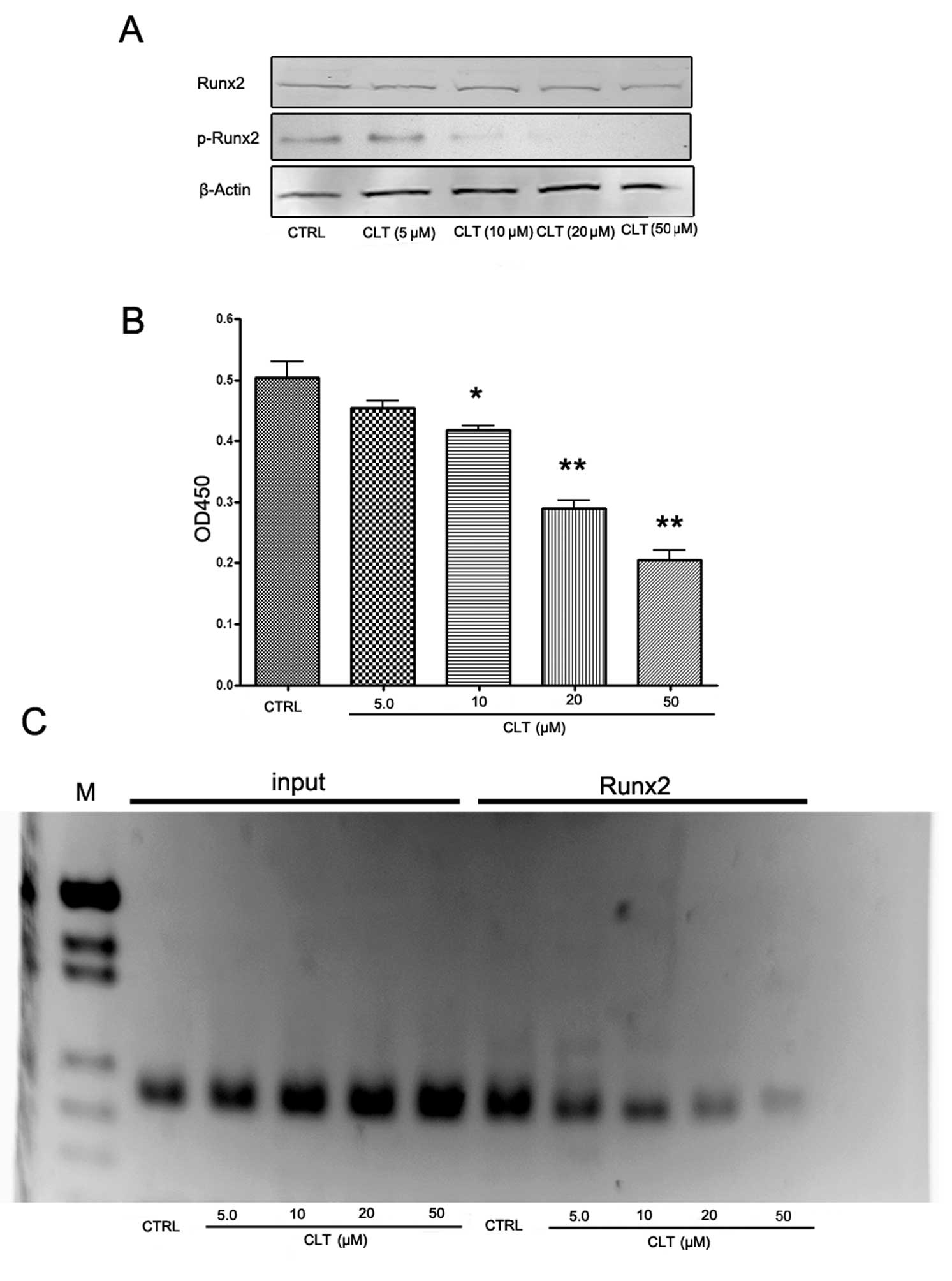
targets of this transcriptional factor (15). So we examined the levels of total
Runx2 and phospho-Runx2 in MDA-MB-231 cells. As shown in Fig. 7A, CLT had no effect on total Runx2
expression, but caused a significant decrease of phospho-Runx2
expression. These results indicate that CLT-induced downregulation
of MMP-13 may be associated with the suppression of Runx2
activation.
To confirm the western blot results, Runx2
transcription factor assay and ChIP assay were designed to evaluate
the effects of CLT on Runx2 activities in MDA-MB-231 cells. An
enzyme-linked immunoabsorbent assay (ELISA)-based kit for the Runx2
transcription factor was used to analyze the effects of CLT on
Runx2. MDA-MB-231 nuclear extracts incubated with CLT or vehicle
were prepared, and binding activity between Runx2 with its target
sequence was determined. The results showed that CLT downregulated
the binding activity of Runx2 in a dose-dependent manner (Fig. 7B). Using the ChIP assay, we found
that binding of Runx2 to one of the Runx2 binding domains (−1,531
bp: aca cca a) in the mmp-13 promoter region was inhibited by CLT
(Fig. 7C). These findings
suggested that the inhibitory effect of CLT on MMP-13 was caused by
CLT downregulation of Runx2 binding to the mmp-13 promoter, which
may be associated with suppression of Runx2 phosphorylation.
Discussion
Breast cancer is the leading cause of death in women
with cancer, and most of the deaths are due to the distant spread
of cancer cells (1), and growth in
distant organs of cancer cells (metastasis), is the most insidious
aspect of this malignancy. Better understanding of metastasis and
developing effective therapies are urgently required. On the other
hand, natural products are still the main resource of new drug
development (5), and therefore our
interests have been focused on identifying new potential
therapeutic agents from Chinese herbal medicine. Here we report
that CLT, a sesquiterpene lactone which can be found in several
Chinese herbal medicines, suppressed invasion, migration of
MDA-MB-231 and MDA-MB-157 cells. Furthermore, these in vitro
effects were confirmed by in vivo data. CLT exhibited
significant suppression of formation of lung metastatic foci and
improvement on total survival of tumor-bearing mice.
MMPs are a family of structurally and functionally
related zinc-dependent endopeptidases. To date, 23 human MMPs,
including 17 soluble, secreted enzymes and six membrane-associated
enzymes have been identified (14). These enzymes are involved in a wide
range of physiological and pathological processes, such as
embryonic development, wound healing, tumor growth, invasion and
metastasis (16). During the past
30 years, the role of MMPs in human cancer has been widely
investigated. These enzymes participate in the proteolysis of the
extracellular matrix, modulation of cell adhesion, migration
(17), and epithelial to
mesenchymal transition (EMT) (18), processing of growth factors, and
tumor-induced angiogenesis. In breast cancer, intensive data
suggested critical roles for MMPs in both breast cancer initiation
and progression (17). Based on
this evidence, we examined the effects of CLT on MMP-9 and MMP-13.
Our results showed that CLT induced a significant suppression of
the activity and expression of MMP-9 and MMP-13. These data
suggested that CLT attenuated the metastatic properties of breast
cancer cells by MMPs inhibition.
Runx2, also named PEBP2αA/AML3/Cbfa1, is a critical
transcription factor for osteoblastic differentiation and skeletal
morphogenesis (19–21). This protein belongs to the Runx
family encoding proteins homologous to Drosophila Runt and
has a conserved Runt DNA-binding domain. Originally, Runx2 was
found to act as a master regulatory factor in skeletal development
(22). This transcriptional factor
acts as a master regulatory factor involved in skeletal gene
expression by binding DNA as a monomer or, with more affinity, as a
subunit of a heterodimeric complex (19–22).
To date, extensive evidence shows a close association between Runx2
and breast cancer metastasis, and this transcriptional factor is
becoming a potential target of novel antimetastatic agents and
diagnostic approaches to breast cancer control (15,23–25).
Runx2 is highly expressed in both breast cancer cell lines which
have the tendency to form metastatic foci and breast primary
tumors. In breast cancer cells, several genes that correlated with
the occurrence of skeletal metastasis have been shown to be
regulated by Runx2, such as bsp and opn (26,27).
However, the functions of Runx2 on tumor metastasis are far beyond
colonizing bone. Recently, it was reported that Runx2 is not only a
novel transcriptional regulator of tumor-induced angiogenesis
(28) but also plays a critical
role in cancer cell induced EMT (29).
Runx2 is also involved in the regulation of MMP in
metastatic breast cancer cells. Jimenez et al first found
that MMP-13, also named Collagenase-3, was highly expressed in
MDA-MB-231 cells, and that it was one of target genes of Runx2
(30). These observations were
demonstrated by Selvamurugan and colleagues (31,32).
Studies showed that the runt domain (RD) binding site and Runx2
were required for maximal constitutive and basal expression of
MMP-13 in MDA-MB-231 cells. Furthermore, the ChIP assay confirmed
two Runx2 binding sites in the mmp-13 promoter, and these sites are
occupied by Runx2. Pratap et al investigated the role of
Runx2 in the regulation of the promoter of MMP-9, in MDA-MB-231 and
MCF-7 cells. MMP-9 was found to be another direct target of Runx2
in metastatic breast cancer cells, and the modulation of Runx2
activity by either forced expression or RNA interference directly
affected MMP-9 expression and the invasive properties of metastatic
cancer cells (33). Thus, Runx2
acts a master transcription factor of MMPs expression. Therefore,
we wondered whether the inhibitory effects of CLT on MMP-9 and
MMP-13 were associated with the downregulation of Runx2 activities.
Our findings from western blotting demonstrated that CLT
significantly inhibited the expression of p-Runx2, indicating that
the transcriptional ability of Runx2 is impaired by CLT. To confirm
these findings, we used an ELISA-based Runx2 transcription factor
assay in this study, and the results revealed that the interaction
between Runx2 and its target sequences was significantly inhibited
by CLT. On the other hand, the interaction of Runx2 and mmp-13
promoter was evaluated by ChIP. It was found that binding between
Runx2 and mmp-13 promoter was impaired significantly by CLT.
In conclusion, here we report that CLT, a
sesquiterpene lactone, impairs the metastatic potential of breast
cancer, and the inhibitory effects are due to its ability to reduce
the expression of MMP-9 and MMP-13, targets of the Runx2
transcription factor. These effects may be associated with
inhibition of transcriptional activity of Runx2.
Acknowledgements
This study was supported by Grants from the National
Natural Science Foundation of China (grant nos. 30860375, 81160530
and 81260656), Key Research Project from the Ministry of Education
of China (Grant Number: 211091), and the Natural Science Foundation
of Jiangxi Province (grant no. 2010GQY0147).
References
|
1
|
Siegel R, Naishadham D and Jemal A: Cancer
Statistics, 2013. CA Cancer J Clin. 63:11–30. 2013. View Article : Google Scholar
|
|
2
|
Leong SP, Cady B, Jablons DM, et al:
Clinical patterns of metastasis. Cancer Metastasis Rev. 25:221–232.
2006. View Article : Google Scholar
|
|
3
|
Stevanovic A, Lee P and Wilcken N:
Metastatic breast cancer. Aust Fam Physician. 35:309–312. 2006.
|
|
4
|
Liu S, Goldstein RH, Scepansky EM, et al:
Inhibition of rho-associated kinase signaling prevents breast
cancer metastasis to human bone. Cancer Res. 69:8742–8751. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Newman DJ: Natural products as leads to
potential drugs: an old process or the new hope for drug discovery?
J Med Chem. 51:2589–2599. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Hristova K, Lam M, Feild T, et al:
Transmitting tissue ECM distribution and composition, and pollen
germinability in Sarcandra glabra and Chloranthus
japonicus (Chloranthaceae). Ann Bot. 96:779–791. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Editorial Committee of the Administration
Bureau of Traditional Chinese Medicine: genus
Chloranthaceae. Chinese Materia Medica (zhonghua
Bencao). Shanghai Science & Technology Press; Shanghai: pp.
2051–2061. 1998
|
|
8
|
Pan C, Xu H, Peng H, et al: Resource
investigation and exploitable foreground of Sarcandra
glabra. Zhong Yao Cai. 27:556–557. 2004.PubMed/NCBI
|
|
9
|
National Committee of Pharmacopoeia.
Pharmacopoeia of People’s Republic of China. 2010 ed. China Medical
Science Press; Beijing: pp. 8422010
|
|
10
|
Wang QH, Kuang HX, Yang BY, et al:
Sesquiterpenes from Chloranthus japonicus. J Nat Prod.
74:16–20. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Zhang M, Wang JS, Wang PR, et al:
Sesquiterpenes from the aerial part of Chloranthus japonicus
and their cytotoxicities. Fitoterapia. 83:1604–1609. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Kwon OE, Lee HS, Lee SW, et al: Dimeric
sesquiterpenoids isolated from Chloranthus japonicus
inhibited the expression of cell adhesion molecules. J
Ethnoparmacol. 104:270–277. 2006.PubMed/NCBI
|
|
13
|
Fu J, Ding Y, Huang D, et al: The retinoid
X receptor-selective ligand, LGD1069, inhibits tumor-induced
angiogenesis via suppression of VEGF in human non-small cell lung
cancer. Cancer Lett. 248:153–163. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Chambers AF and Matrisian LM: Changing
views of the role of matrix metalloproteinases in metastasis. J
Natl Cancer Inst. 89:1260–1270. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Inman CK and Shore P: The osteoblast
transcription factor Runx2 is expressed in mammary epithelial cells
and mediates osteopontin expression. J Biol Chem. 278:48684–48689.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Roy R, Yang J and Moses MA: Matrix
metalloproteinases as novel biomarkers and potential therapeutic
targets in human cancer. J Clin Oncol. 27:5287–5297. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Duffy MJ, Maguire TM, Hill A, et al:
Metalloproteinases: role in breast carcinogenesis, invasion and
metastasis. Breast Cancer Res. 2:252–257. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Radisky ES and Radisky DC: Matrix
metalloproteinase-induced epithelial-mesenchymal transition in
breast cancer. J Mammary Gland Biol Neoplasia. 15:201–212. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Lian JB, Stein JL, Stein GS, et al:
Runx2/Cbfa1 functions: diverse regulation of gene transcription by
chromatin remodeling and co-regulatory protein interactions.
Connect Tissue Res. 44:141–148. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Karsenty G: Role of Cbfa1 in osteoblast
differentiation and function. Semin Cell Dev Biol. 11:343–346.
2000. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Komori T: Runx2, a multifunctional
transcription factor in skeletal development. J Cell Biochem.
87:1–8. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Stein GS, Lian JB, van Wijnen AJ, et al:
Runx2 control of organization, assembly and activity of the
regulatory machinery for skeletal gene expression. Oncogene.
23:4315–4329. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Inman CK, Li N and Shore P: Oct-1
counteracts autoinhibition of Runx2 DNA binding to form a novel
Runx2/Oct-1 complex on the promoter of the mammary gland-specific
gene beta-casein. Mol Cell Biol. 25:3182–3193. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Barnes GL, Javed A, Waller SM, et al:
Osteoblast-related transcription factors Runx2 (Cbfa1/AML3) and
MSX2 mediate the expression of bone sialoprotein in human
metastatic breast cancer cells. Cancer Res. 63:2631–2637. 2003.
|
|
25
|
Shore P: A role for Runx2 in normal
mammary gland and breast cancer bone metastasis. J Cell Biochem.
96:484–489. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Ganss B, Kim RH and Sodek J: Bone
sialoprotein. Crit Rev Oral Biol Med. 10:79–98. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Bellahcène A and Castronovo V: Expression
of bone matrix proteins in human breast cancer: potential roles in
microcalcification formation and in the genesis of bone metastases.
Bull Cancer. 84:17–24. 1997.PubMed/NCBI
|
|
28
|
Sun L, Vitolo M and Passaniti A:
Runt-related gene 2 in endothelial cells: inducible expression and
specific regulation of cell migration and invasion. Cancer Res.
61:4994–5001. 2001.PubMed/NCBI
|
|
29
|
Chimge NO, Baniwal SK, Little GH, et al:
Regulation of breast cancer metastasis by Runx2 and estrogen
signaling: the role of SNAI2. Breast Cancer Res. 13:R1272011.
View Article : Google Scholar
|
|
30
|
Jiménez MJ, Balbín M, López JM, et al:
Collagenase 3 is a target of Cbfa1, a transcription factor of the
runt gene family involved in bone formation. Mol Cell Biol.
19:4431–4442. 1999.PubMed/NCBI
|
|
31
|
Selvamurugan N and Partridge NC:
Constitutive expression and regulation of collagenase-3 in human
breast cancer cells. Mol Cell Biol Res Commun. 3:218–223. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Selvamurugan N, Kwok S and Partridge NC:
Smad3 interacts with JunB and Cbfa1/Runx2 for transforming growth
factor-beta1-stimulated collagenase-3 expression in human breast
cancer cells. J Biol Chem. 279:27764–27773. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Pratap J, Javed A, Languino LR, et al: The
Runx2 osteogenic transcription factor regulates matrix
metalloproteinase 9 in bone metastatic cancer cells and controls
cell invasion. Mol Cell Biol. 25:8581–8591. 2005. View Article : Google Scholar
|