Introduction
Glioma is the most common malignant tumor in the
central nervous system, with incidence rate of 5–7 per 100,000 and
5-year survival rate of approximately 20–30% (1). Gliomas originate from astrocytes,
oligodendrocytes and neural stem cells or their precursors.
According to WHO classification, gliomas are classified into four
different malignant grades ranging from grade I to grade IV based
on histopathological features and related molecular aberrations, of
which the glioblastoma multiforme (GBM) is classified as grade IV,
the most aggressive group and one of the hardest tumors to treat
(2). The therapy method of GBM
mainly include surgery, radiation and chemotherapy, but the
survival of patients is still quite short, especially when invasion
and metastasis occur (3).
To improve the prognosis of GBM, great efforts have
been taken to identify molecular markers and therapeutic targets.
In which, several genes, such as RINT1, mTORC2 and WT1 have been
reported to participate in the carcinogenesis and progression of
GBM (4). However, there is no
biomarker with enough sensitivity and specificity for assessment of
response to therapy and prognosis in GBM. Thus, a better
understanding of the mechanisms involved in regulating tumor growth
requires the identification of novel genes is associated with
GBM.
MicroRNAs (miRNAs) are small 19–24 nucleotide,
non-coding RNAs that can inhibit the post-transcription of target
mRNA by completely or incompletely matching with the target mRNA,
and then participate in the regulation of many biological
processes, such as cell proliferation, metabolism, differentiation
and apoptosis (5). Therefore,
aberrant miRNA expression associated with the development of
different kinds of cancer.
Accumulated evidence indicates that several mRNAs
and genes, respectively participate in the progression of GBM.
Currently, global analyses have revealed that several miRNAs are
clinically implicated with GBM, such as miR-103a, miR-107 and
miR-182 (6). Previous reports
showed that miR-372 regulates GBM cell proliferation and invasion
by directly targeting PHLPP2 (7).
miR-210 upregulation inhibits proliferation and induces apoptosis
in GBM cells by targeting SIN3A (8) and miR-429 inhibits GBM invasion
through BMK1 suppression (9).
However, no previous report exists on investigation
of the correlation between the expression of miR-181a and target
genes in GBM. We profiled miRNAs and genes expression by microarray
to identify their differentially expression in GBM and normal brain
tissues, and then explored the correlation between miR-181a and its
target genes in GBM to provide further insight into the
pathogenesis of GBM.
Materials and methods
Ethics statement
The study protocol and acquisition of tissue
specimens were approved by the Ethics Committee of the Ganzhou City
People's Hospital (2015-RES-10). Each participant provided written
informed consent before participating in the present study.
Acquisition of clinical specimens
GBM samples were collected from patients undergoing
surgical resection and classified according to the last WHO
classification of central nervous tumors and clinical histories
were recorded, confirmed by two experienced pathologists
independently.
Collection of clinical information and
follow up
Demographic and clinicopathological characteristics
were recorded including the patient's characteristics (e.g., age
and gender), tumor characteristics (size, necrosis, boundary and
cystic degeneration), overall survival time. All previous data were
collected by reviewing the clinical history. For analysis, patients
were stratified according to age, ≥40 or <40 years. The tumor
size was described by mean tumor diameter (MTD, defined as the
geometric mean of 3 diameters on MRI scan), and tumors were grouped
according to size, ≥5 cm and <5 cm. The follow-up was conducted
by telephone or direct correspondence. The time of tumor relapse or
death was verified by the patient or their relatives, by medical
recording, or by the social security record. Overall survival (OS)
was calculated in months from the date of diagnosis to the time of
death, regardless of cause.
RNA extraction
According to the manufacturer's guideline, total RNA
was isolated using TRIzol reagent (Invitrogen, Carlsbad, CA, USA).
RNA quantity was determined using NanoDrop ND-1000
spectrophotometer and the integrity of RNA was measured by gel
electrophoresis.
Gene microarray analysis
For gene expression microarray analysis, tumor
tissue from 22 GBM patients was assessed on an Affymetrix array
platform (Affymetrix, Inc., Santa Clara, CA, USA). The sample
preparation and microarray hybridization were performed based on
the manufacturer's standard protocols and published studies
(10–13).
Cell lines and transfection
Human glioblastoma cell line U251 were purchased
from Shanghai Institute for Biological Sciences (Shanghai, China)
and cultured in Dulbecco's modified Eagle's medium (DMEM; Gibco,
Carlsbad, CA, USA) supplemented with 10% fetal bovine serum (FBS;
HyClone Laboratories, Inc., Logan, UT, USA), 100 units/ml
penicillin and 100 mg/ml streptomycin at 37°C in a humidified
chamber supplemented with 5% CO2.
miR-181a mimic and corresponding negative control
miRNA (miR-NC) were purchased from Shanghai GenePharma, Co., Ltd.
(Shanghai, China). These molecular productions were transiently
transfected into U251 cells, respectively, using Oligofectamine™
transfection reagent (Invitrogen) according to the manufacturer's
instructions.
Reporter assays
The human ANGPT2, ARHGAP18 or LAMC1 3′-UTR
oligonucleotides containing the wild-type (Wt) or mutant (Mut)
miR-181a binding site were subcloned into the XhoI and
NotI sites of the psiCHECK2 vector (Promega, Madison, WI,
USA). For luciferase assay, U251 cells were inoculated into 24-well
plates and cultured for 24 h; then, cells were co-transfected with
the Wt/Mut reporter plasmid (100 ng) and miR-181a mimic/miR-NC (100
nM). Forty-eight hours after the transfection, luciferase assay was
determined using the Dual-Luciferase kit (Promega).
Quantitative RT-PCR (qRT-PCR)
For qRT-PCR of ANGPT2, ARHGAP18 and LAMC1, cDNA was
synthesized from total RNA (10 ng), and quantitative PCR reactions
were performed with the TaqMan™ Universal PCR kit (Life
Technologies, Grand Island, NY, USA). GAPDH was used as the
internal control. Quantitative qRT-PCR for miR-181a was performed
using the TaqMan Universal PCR kit (Life Technologies). U6 small
nuclear RNA was used as internal control, and the 2−ΔΔCT
method was used to analyze the expression levels of miR-181a and 3
gene levels.
Bioinformatic analysis
The expression levels of miRNAs were investigated in
GBM and normal tissue samples in the GEO datasets using the NCBI
Platform (http://www.ncbi.nlm.nih.gov/). Hierarchical clustering
was performed using the multiple experiment viewer (MeV) 4.7.1
software (http://www.tm4.org/mev/). Each row
represents a miRNA or gene, and each column represents a sample in
the heat map diagram. The clustering tree of miRNAs or genes are
shown on the left, and the sample clustering tree appears at the
top.
We used the target gene prediction software
DIANA-TarBase (14), TargetScan
(http://www.targetscan.org) and PicTar
(http://pictar.mdc-berlin.de/) to
forecast several potential target genes of miR-181a.
To elucidate the relationship between gene
differential expression patterns, we used GO classifications
(http://www.geneontology.org/), which
mainly analyzed three aspects including biological process,
molecular function and cellular components, which reflect the
biological function of differential expression genes in GBM.
Subsequent bioinformatic analysis of these genes was performed by
the Kyoto Encyclopedia of Genes and Genomes Pathway analysis (KEGG
Pathway analysis) (http://www.genome.jp/kegg/). Pathways were selected
with a P-value <0.05 and gene count >2.
Statistical analysis
The results were expressed as mean ± SD (standard
deviation). The statistical significance between the groups was
assessed by using one-way analysis of variance (ANOVA). Univariate
survival analysis and multivariate analyses were carried out using
the Kaplan-Meier method. All calculations were performed with the
SPSS 20.0 software program (SPSS, Inc., Chicago, IL, USA). The
level of significance was chosen as P<0.05.
Results
miRNAs microarray analysis
The expression profiles of human miRNAs were
analyzed by microarrays in three paired GBM tissues download from
the Gene Expression Omnibus (GEO) database (GSE65626-GPL19117).
After hierarchical clustering of miRNAs, the GBM samples and normal
samples were completely separated. Of all the miRNAs represented on
the array, 39 miRNAs displayed at least a 2-fold increased or
0.5-fold decreased in expression at the P<0.01 level with a
false discovery rate. Of which, 30 miRNAs were upregulated in human
GBM tissues, while 9 miRNAs were downregulated in human GBM tissues
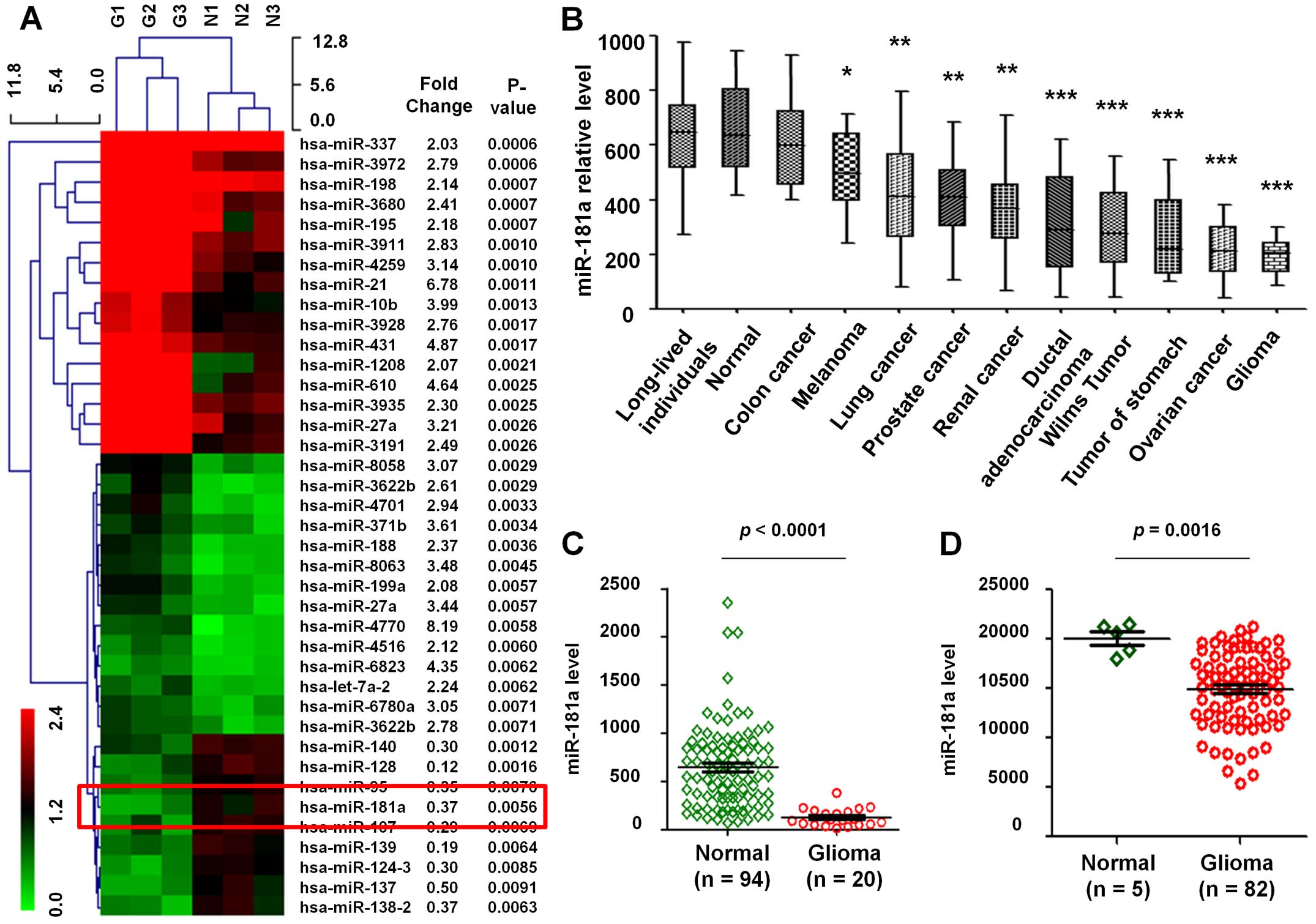
(Fig. 1A). Several common miRNAs,
such as miR-21, miR-10b and miR-27a, linked to the risk of GBM.
However, no previous report exists on investigation of the
correlation between the expression of miR-181a and the target genes
in GBM. Moreover, we found miR-181a that is rarely referred to
presented downregulation in GBM.
Subsequently, we analyzed the expression level of
the above mentioned differential miRNAs in 94 normal controls and
448 various cancers included 20 GBM from the GEO datasets
(GSE61741). This project analyzed peripheral blood miRNA profiles
and each miRNA was measured in at least seven replicates, the
median of the replica had been computed. The results showed that
the expression level of miR-181a in various tumors was lower than
in normal samples (Fig. 1B). Then,
94 normal and 20 GBM samples were used to evaluate the expression
profile of miR-181a. Results show that the expression level of
miR-181a in GBM was remarkably lower than in normal tissues
(P<0.0001, fold change, 0.21; Fig.
1C).
In addition, data of 5 normal and 82 GBM samples
from the GEO database (GSE25632-GPL8179) were also analyzed, which
similarly reflected that miR-181a was downregulated in GBM compared
to normal tissues (P=0.0016, fold change, 0.66; Fig. 1D), thus miR-181a was downregulated
in tumors especially in GBM.
Gene microarray analysis
We further analyzed gene expression profile of the
above three paired GBM tissue samples. Of all the genes
represented, 46 genes displayed at least a 1.5-fold increased or
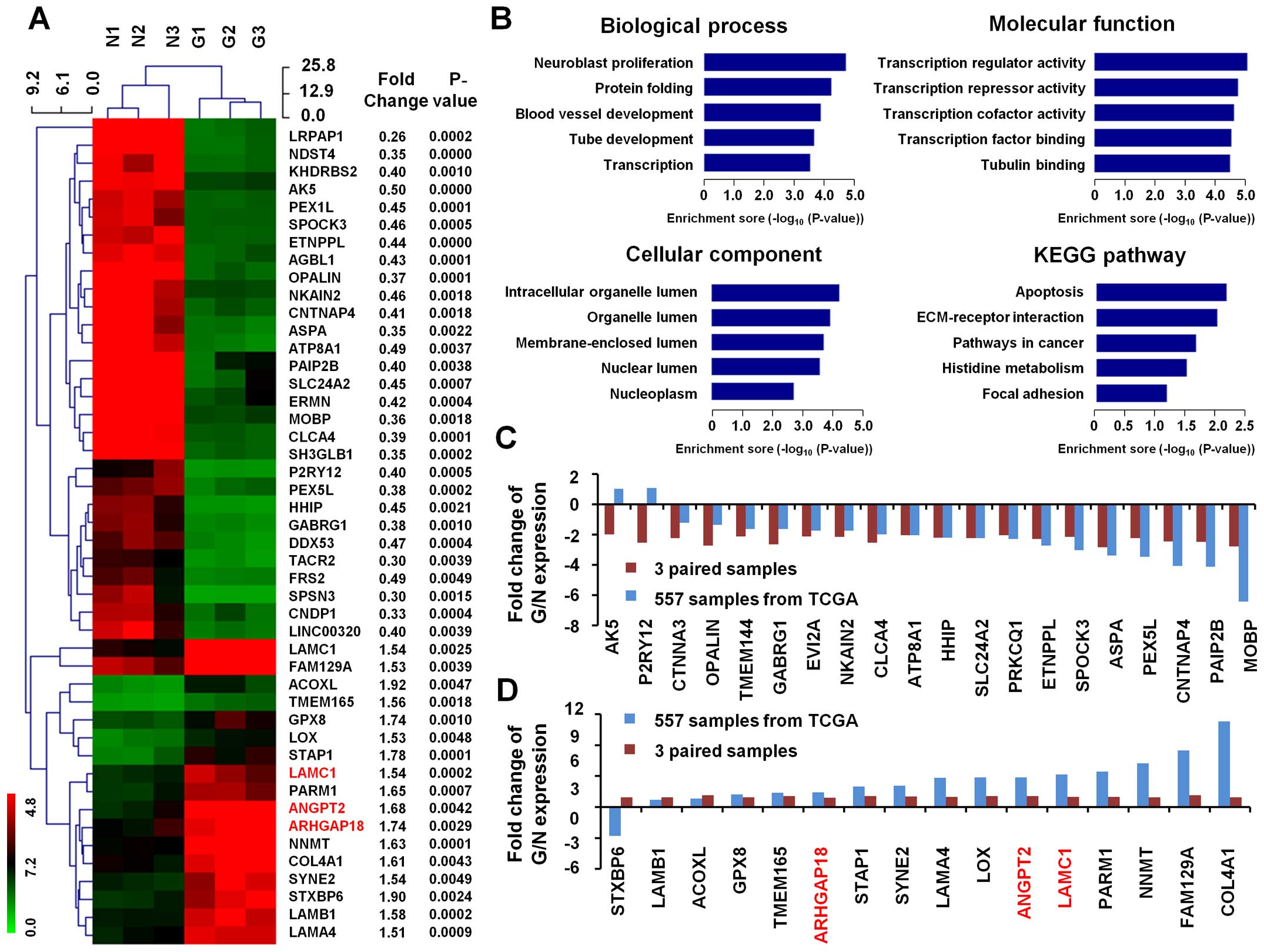
0.5-fold decreased in expression at the P<0.01 level. Of which,
17 genes represent upregulated function, while 29 genes were
downregulated in human GBM tissues (Fig. 2A).
GO and KEGG pathway analysis
In biological process, the top 5 GO terms of these
46 differential genes are neuroblast proliferation, protein
folding, blood vessel development, tube development and
transcription. In molecular function, the top 5 GO terms are
transcription regulator activity, transcription repressor activity,
transcription cofactor activity, transcription factor binding and
tubulin binding. The top 5 GO terms in cellular component are
intracellular organelle lumen, organelle lumen, membrane-enclosed
lumen, nuclear lumen and nucleoplasm (Fig. 2B).
KEGG pathways analysis of these differentially
expressed gene patterns revealed several enrichment-related
pathways, including pathways in apoptosis, ECM-receptor
interaction, pathways in cancer, histidine metabolism and focal
adhesion (Fig. 2B).
TCGA dataset comparative analysis
We obtained a total of 557 GMB cases with level 3
gene expression data based on microarray by search of the TCGA
dataset. In contrast to standard TCGA dataset on gene expression,
we found that upregulated genes (in our 3 paired samples) are
largely in keeping with TCGA dataset in fold change of GBM
(Fig. 2C). In addition,
downregulated genes are also mostly in accordance with TCGA dataset
in fold change of expression in GBM (Fig. 2D).
Computational prediction of associations
between miRNAs and target genes
To elucidate whether there were paired miRNAs and
target mRNAs profiles aberrantly expressed in GBM, we used target
gene prediction software including DIANA-TarBase, TargetScan and
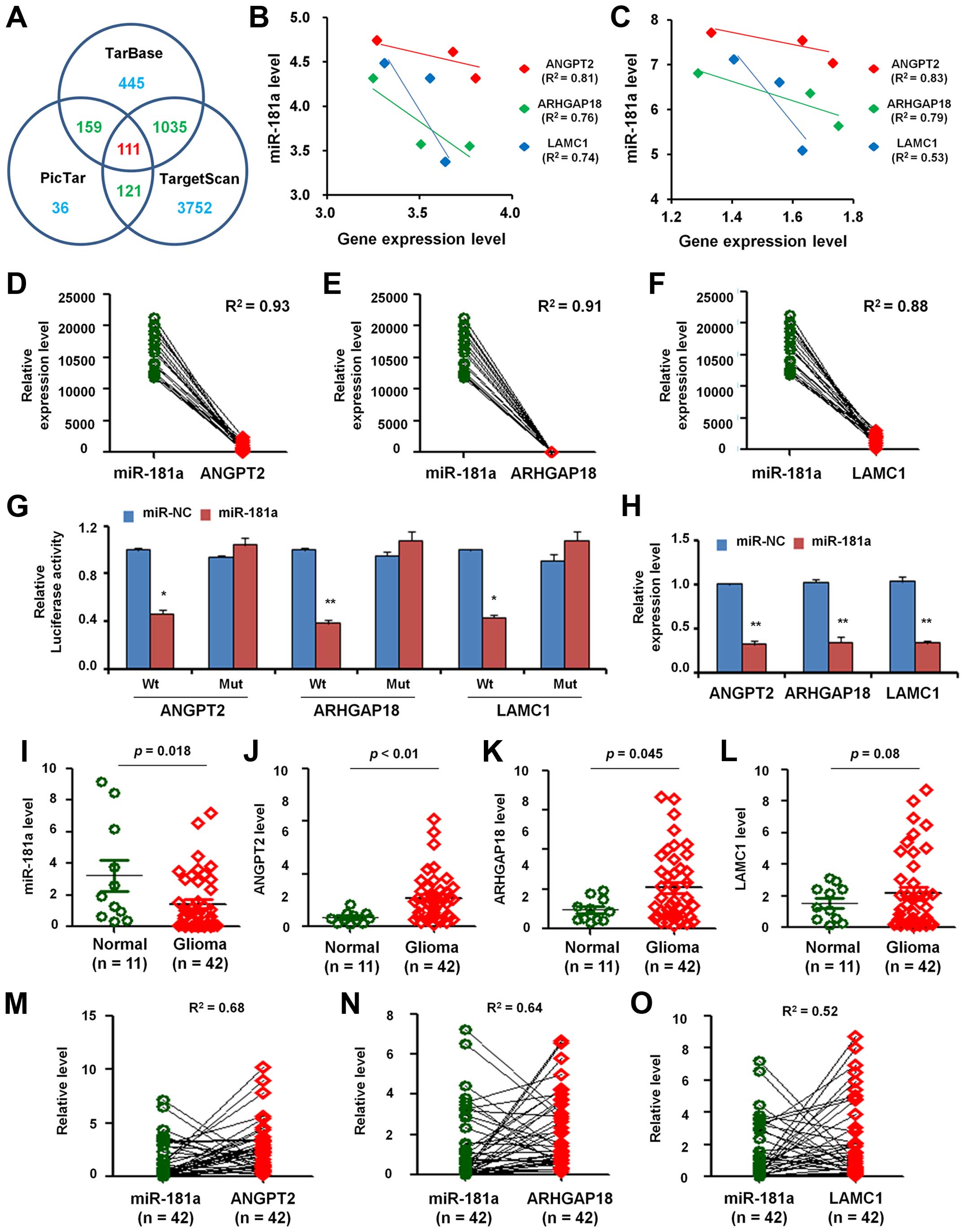
PicTar to search several potential target genes of miR-181a. There
were 1750, 5019 and 427 predicted target genes found correlated
with miR-181a in DIANA-TarBase, TargetScan and PicTar, respectively
(Fig. 3A). Based on the analysis
of the three databases, we found 111 common target genes.
Then, we analyzed the expression of these common
target genes in 3 paired GBM and normal tissues mentioned before
and found the negative relationships between the expression level
of miR-181a and the predicted target genes (ANGPT2, ARHGAP18 and
LAMC1) either in normal brain (Fig.
3B) or in GBM tissues (Fig.
3C).
Besides, we analyzed the correlation between the
expression level of miR-181a and 3 predicted target genes in 21
paired GBM samples from GEO dataset (GPL8179). As shown in Fig. 3D, the relationship between miR-181a
expression level and ANGPT2 expression level presented inverse
correlation (R2=0.93). Similarly, the inverse
correlation also emerged in miR-181a expression with mRNA levels of
ARHGAP18 (R2=0.91; Fig.
3E) and LAMC1 (R2=0.88; Fig.
3F).
Identification of ANGPT2, ARHGAP18 and
LAMC1 as the miR-181a direct targets in U251 cells
To confirm the predicted target genes as direct
targets of miR-181a, we performed luciferase reporter assays in
U251 cells. As shown in Fig. 3G,
transfection of miR-181a mimic caused a significant decrease in
luciferase activity in cells transfected with the reporter plasmid
with wild-type targeting sequence of mRNA, but not reporter plasmid
with mutant sequence of the genes.
To confirm miR-181a regulation of the expression of
ANGPT2, ARHGAP18 and LAMC1, we detected their expression in U251
cells after transfection with miR-181a or miR-NC. It was found that
miR-181a overexpression significantly inhibited mRNA expression
level (Fig. 3H).
Validation of miR-181a/mRNAs expression
and correlation using qRT-PCR
To further verify the correlation between miR-181a
and the target genes, we extracted RNA from 11 normal brain tissues
and 42 GBM tissues and carried out qRT-PCR. Fig. 3I shows that the expression level of
miR-181a in GBM was lower than in normal tissues. However, ANGPT2
(Fig. 3J), ARHGAP18 (Fig. 3K) and LAMC1 (Fig. 3L) were overexpressed in GBM when
compare with normal tissues, which imply that the expression level
of miR-181a was negatively correlated with ANGPT2, ARHGAP18 and
LAMC1 in GBM.
Furthermore, we analyzed the correlation of miR-181a
and the three genes in 42 paired GBM samples and found that the
relative expression level between miR-181a and ANGPT2 presented
inverse correlation (R2=0.68; Fig. 3M). In the same samples, the
relative expression level between miR-181a and ARHGAP18 showed
negative correlation (R2=0.64; Fig. 3N). Similarly, the relative
expression level between miR-181a and LAMC1 also indicated negative
relation (R2=0.52; Fig.
3O).
Clinical significance of 3 genes in
GBM
We analyzed the microarray data from 22 cases of GBM
which included 12 GBM with good prognosis and 10 GBM with poor
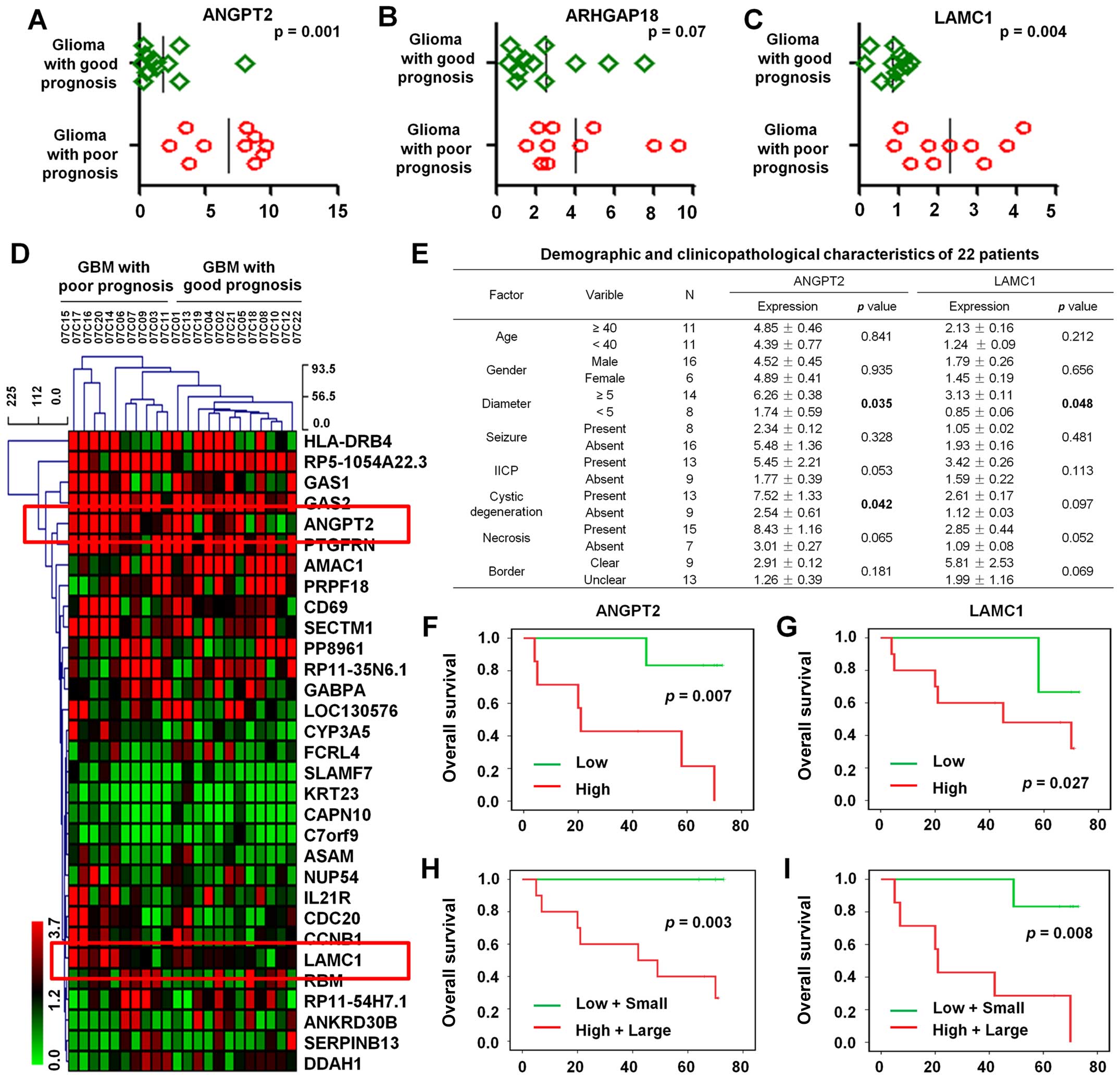
prognosis. The results showed that the expression of ANGPT2 and
LAMC1 in GBM patients who had a good prognosis was significantly
lower than who had a poor prognosis (P=0.001 and 0.004,
respectively) (Fig. 4A and B).
However, there is no significant difference in ARHGAP18 expression
(P=0.07; Fig. 4C). Hierarchical
clustering analysis was further used to detect the expression
profile of 31 differential genes (fold change ≥2 or ≤0.5;
P<0.01) in these 22 GBM patients with different prognosis, of
which ANGPT2 and LAMC1 were shown to upregulate (Fig. 4D).
Next, we investigated the relationship between
ANGPT2 and LAMC1 expression level and clinical characteristics.
Univariate analysis of 22 cases of GBM demonstrated that ANGPT2
expression levels were significantly correlated with tumor diameter
(P=0.035) and cystic degeneration (P=0.042) and the expression
levels of LAMC1 was significantly associated with tumor diameter
(P=0.048; Fig. 4E).
Kaplan-Meier survival analysis was then used to
explore the prognostic significance of ANGPT2 and LAMC1. The
results indicated high expression of ANGPT2 was negatively
correlated with higher OS (P=0.007; Fig. 4F). Likewise, high expression of
LAMC1 was negatively associated with higher OS (P=0.027; Fig. 4G). Therefore, high expression of
ANGPT2 and LAMC1 correlated with poor prognosis in GBM
patients.
Multivariate analysis of ANGPT2 expression with
diameter using Kaplan-Meier estimation suggested that high
expression of ANGPT2 with large diameter tumor was significantly
associated with poor OS (P=0.003; Fig.
4H). In addition, high expression of LAMC1 with large diameter
tumor was also related with poor OS (P=0.008; Fig. 4I). Take together, ANGPT2 and LAMC1
might be predictors of prognosis in GBM patients.
Discussion
GBM is one of the most deadly human malignancies
worldwide and has increasing incidence with short survival rate and
high mortality (15). Thus, the
novel molecular mechanisms involved in the aggressive growth of
GBM, and further new targeted therapies are required for prolonging
the survival of glioma patients. Useful biomarkers are needed as we
rely on their potential significance as a molecular targeted
therapy in the treatment of GBM patients (16). Enormous efforts have been made to
explore a molecular signature that would assist to cancer diagnose
and therapy, in order to improve the current standard of clinical
care (17). A large number of
signal pathways were identified related to the development of GBM,
which are meaningful characteristic biomarkers in individual
patients (18).
More than 100 miRNAs were investigated that
correlated with the progression of cell proliferation, metastasis
and apoptosis among GBM (6).
miR-181a was proven to be related to certain type of cancers
(19). However, the clinical
relevance and function of miR-181a in GBM remains elusive.
In the present study, via the hierarchical
clustering analysis of microarray data, we found 30 upregulated and
9 downregulated miRNAs in 3 pairs of GBM, of these, miR-181a was
included in the downregulated miRNAs. By systematical analysis of
differential miRNAs, we discovered that miR-181a was downregulated
in many cancers, especially lower expressed in GBM. Thus, miR-181a
may be a tumor suppressor in GBM.
The hierarchical clustering analysis of microarray
data indicated 46 upregulated and 17 downregulated genes in 3 pairs
of GBM. We selected 3 upregulated genes (ANGPT2, ARHGAP18 and
LAMC1), which were validated to be the target genes of miR-181a by
luciferase reporter assays. Furthermore, both GO analysis and KEGG
pathway analysis indicated that ANGPT2, ARHGAP18 and LAMC1
significantly associated with GBM formation. Our results showed
that expression of miR-181a was negatively correlated with mRNA
levels of the target genes. Moreover, high expression of ANGPT2 and
LAMC1 is correlated with a shorter median OS in GBM, which
suggesting that ANGPT2 and LAMC1 are underlying diagnostic
biomarkers and prognostic factors for GBM.
GBM is characterized by exuberant angiogenesis, a
key event in tumor growth and progression. Tumor vessel density,
which represents angiogenesis, has prognostic value in various
malignant tumors, including glioma (20). Angiogenesis is controlled by the
interplay between numerous positive and negative factors (21). ANGPT2 is a ligand of the tyrosine
kinase receptor Tie2 and integrin receptors (22). It functions as an autocrine
controller of endothelial cells in a context-dependent manner,
promoting either blood vessel growth or regression depending on the
levels of other growth factors such as vascular endothelial growth
factor (VEGF) (23). ANGPT2 level
has been associated with tumor angiogenesis in various cancers
(24). However, the diagnostic
value of angiopoietin-2 in glioma was not investigated. Considering
that the use of angiogenesis inhibitors may offer novel strategies
in brain tumor therapy, our results that ANGPT2 upregulation
correlated with a shorter median OS in glioma may provide new
insight for diagnosis and treatment of glioma.
Tumor development and progression depend not only on
the perturbed genes that govern cell proliferation and apoptosis,
but is also highly determined by tumor microenvironment (TME)
(25,26). Extracellular matrix proteins
constitute an integral part of TME and play critical roles in
regulating tumor cell proliferation, survival, autophagy, migration
and invasion. Although the role and molecular mechanisms in
upregulation of LAMC1, an extracellular matrix glycoprotein in
cancer are unclear, a previous report suggested that LAMC1 is an
oncogene in breast, prostate and ovarian cancer and may contribute
to the development and progression of uterine carcinoma through
enhancing tumor cell motility and invasion (27), which warrants further investigation
regarding its role as a biomarker and therapeutic target in
cancer.
In conclusion, we have demonstrated that miR-181a is
downregulated, but the target genes ANGPT2, ARHGAP18 and LAMC1 are
upregulated in GBM. Our results indicated that high expression of
ANGPT2 or LAMC1 in GBM is associated with shorter overall survival,
which suggest that ANGPT2 and LAMC1 might be a predictor of
prognosis in GBM patients.
Acknowledgements
The present study was supported in part by grants
from the Key Science and Technology Project of Hainan Province
(ZDXM2015070), the Social Science Development Foundation of Hainan
Province (SF201414), the Natural Science Foundation of Hainan
Province (813201) and the Hainan Health Institution Project
(2012PT-06).
References
|
1
|
Wang H, Xu T, Jiang Y, Xu H, Yan Y, Fu D
and Chen J: The challenges and the promise of molecular targeted
therapy in malignant gliomas. Neoplasia. 17:239–255. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Demuth T and Berens ME: Molecular
mechanisms of glioma cell migration and invasion. J Neurooncol.
70:217–228. 2004. View Article : Google Scholar
|
|
3
|
See SJ and Gilbert MR: Anaplastic
astrocytoma: Diagnosis, prognosis, and management. Semin Oncol.
31:618–634. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Wang CL, Wang JY, Liu ZY, Ma XM, Wang XW,
Jin H, Zhang XP, Fu D, Hou LJ and Lu YC: Ubiquitin-specific
protease 2a stabilizes MDM4 and facilitates the p53-mediated
intrinsic apoptotic pathway in glioblastoma. Carcinogenesis.
35:1500–1509. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Wang X, Zhu Y, Ma Y, Wang J, Zhang F, Xia
Q and Fu D: The role of cancer stem cells in cancer metastasis: New
perspective and progress. Cancer Epidemiol. 37:60–63. 2013.
View Article : Google Scholar
|
|
6
|
Srinivasan S, Patric IRP and Somasundaram
K: A ten-microRNA expression signature predicts survival in
glioblastoma. PLoS One. 6:e174382011. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Chen X, Hao B, Han G, Liu Y, Dai D, Li Y,
Wu X, Zhou X, Yue Z, Wang L, et al: miR-372 regulates glioma cell
proliferation and invasion by directly targeting PHLPP2. J Cell
Biochem. 116:225–232. 2015. View Article : Google Scholar
|
|
8
|
Shang C, Hong Y, Guo Y, Liu YH and Xue YX:
MiR-210 up-regulation inhibits proliferation and induces apoptosis
in glioma cells by targeting SIN3A. Med Sci Monit. 20:2571–2577.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Chen W, Zhang B, Guo W, Gao L, Shi L, Li
H, Lu S, Liu Y and Li X: miR-429 inhibits glioma invasion through
BMK1 suppression. J Neurooncol. 125:43–54. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Yan Y, Zhang L, Xu T, Zhou J, Qin R, Chen
C, Zou Y, Fu D, Hu G, Chen J, et al: SAMSN1 is highly expressed and
associated with a poor survival in glioblastoma multiforme. PLoS
One. 8:e819052013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Xu T, Jiang Y, Yan Y, Wang H, Lu C, Xu H,
Li W, Fu D, Lu Y and Chen J: VSIG4 is highly expressed and
correlated with poor prognosis of high-grade glioma patients. Am J
Transl Res. 7:1172–1180. 2015.PubMed/NCBI
|
|
12
|
Qin R, Zhou J, Chen C, Xu T, Yan Y, Ma Y,
Zheng Z, Shen Y, Lu Y, Fu D, et al: LIN28 is involved in glioma
carcinogenesis and predicts outcomes of glioblastoma multiforme
patients. PLoS One. 9:e864462014. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Wang Q, Wang JY, Zhang XP, Lv ZW, Fu D, Lu
YC, Hu GH, Luo C and Chen JX: RLIP76 is overexpressed in human
glioblastomas and is required for proliferation, tumorigenesis and
suppression of apoptosis. Carcinogenesis. 34:916–926. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Vlachos IS, Paraskevopoulou MD, Karagkouni
D, Georgakilas G, Vergoulis T, Kanellos I, Anastasopoulos IL,
Maniou S, Karathanou K, Kalfakakou D, et al: DIANA-TarBase v7.0:
indexing more than half a million experimentally supported
miRNA:mRNA interactions. Nucleic Acids Res. 43(D1): D153–D159.
2015. View Article : Google Scholar :
|
|
15
|
Manterola L, Guruceaga E, Gállego
Pérez-Larraya J, González-Huarriz M, Jauregui P, Tejada S,
Diez-Valle R, Segura V, Samprón N, Barrena C, et al: A small
noncoding RNA signature found in exosomes of GBM patient serum as a
diagnostic tool. Neuro Oncol. 16:520–527. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Tumilson CA, Lea RW, Alder JE and Shaw L:
Circulating microRNA biomarkers for glioma and predicting response
to therapy. Mol Neurobiol. 50:545–558. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Zemp FJ, McKenzie BA, Lun X, Reilly KM,
McFadden G, Yong VW and Forsyth PA: Cellular factors promoting
resistance to effective treatment of glioma with oncolytic myxoma
virus. Cancer Res. 74:7260–7273. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Grant R, Kolb L and Moliterno J: Molecular
and genetic pathways in gliomas: The future of personalized
therapeutics. CNS Oncol. 3:123–136. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Shi L, Cheng Z, Zhang J, Li R, Zhao P, Fu
Z and You Y: hsa-miR-181a and hsa-mir-181b function as tumor
suppressors in human glioma cells. Brain Res. 1236:185–193. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Wu SD, Ma YS, Fang Y, Liu LL, Fu D and
Shen XZ: Role of the microenvironment in hepatocellular carcinoma
development and progression. Cancer Treat Rev. 38:218–225. 2012.
View Article : Google Scholar
|
|
21
|
Wang Y, Ma Y, Fang Y, Wu S, Liu L, Fu D
and Shen X: Regulatory T cell: A protection for tumour cells. J
Cell Mol Med. 16:425–436. 2012. View Article : Google Scholar
|
|
22
|
Janelidze S, Lindqvist D, Francardo V,
Hall S, Zetterberg H, Blennow K, Adler CH, Beach TG, Serrano GE,
van Westen D, et al: Increased CSF biomarkers of angiogenesis in
Parkinson disease. Neurology. 85:1834–1842. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Holash J, Wiegand SJ and Yancopoulos GD:
New model of tumor angiogenesis: Dynamic balance between vessel
regression and growth mediated by angiopoietins and VEGF. Oncogene.
18:5356–5362. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Tsai YC, Lee CS, Chiu YW, Kuo HT, Lee SC,
Hwang SJ, Kuo MC and Chen HC: Angiopoietin-2 as a prognostic
biomarker of major adverse cardiovascular events and all-cause
mortality in chronic kidney disease. PLoS One. 10:e01351812015.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Stewart TA, Yapa KT and Monteith GR:
Altered calcium signaling in cancer cells. Biochim Biophys Acta.
1848(10 Pt B): 2502–2511. 2015. View Article : Google Scholar
|
|
26
|
Han DY, Fu D, Xi H, Li QY, Feng LJ, Zhang
W, Ji G, Xiao JC and Wei Q: Genomic expression profiling and
bioinformatics analysis of pancreatic cancer. Mol Med Rep.
12:4133–4140. 2015.PubMed/NCBI
|
|
27
|
Kashima H, Wu RC, Wang Y, Sinno AK,
Miyamoto T, Shiozawa T, Wang TL, Fader AN and Shih IeM: Laminin C1
expression by uterine carcinoma cells is associated with tumor
progression. Gynecol Oncol. 139:338–344. 2015. View Article : Google Scholar : PubMed/NCBI
|