Introduction
Rhabdomyosarcoma (RMS) is the most prevalent soft
tissue sarcoma in children and adolescents (1,2).
Chemotherapy intensification has not improved the survival outcomes
for metastatic or relapsed RMS (3,4).
Moreover, the long-term side effects of chemotherapy are
significant, including infertility and secondary malignancy. The
heterogeneous genomic landscape of different types of RMS has
challenged the introduction of targeted molecular therapies.
Therefore, the search for molecular targets in RMS could allow for
superior antitumor effects and fewer off-targeted side effects.
Forkhead box protein M1 (FOXM1) is a member of the
forkhead box (FOX) transcription factor family that shares homology
in the Forkhead DNA-binding domain (5,6).
FOXM1 is a known oncogene that is highly expressed in various
cancers with little to no expression in terminally differentiated
cells (7-12). FOXM1 overexpression has been
associated with advanced tumor stage and worse overall survival in
many solid tumors, including rhabdomyosarcoma (13,14). Moreover, FOXM1 is notably
upregulated in chemotherapy-resistant solid tumors, partly due to
the role of FOXM1 in enhancing DNA repair response and protecting
tumor cells from apoptosis (15,16). Collectively, FOXM1 contributes to
multiple hallmarks of cancer (6,17).
Considering the essential role of FOXM1 in
tumorigenesis, extended efforts have been directed towards
identifying FOXM1 inhibitors (16). Using high throughput screening, a
non-toxic, small molecule named Robert Costa Memorial Drug-1 or
RCM-1 was developed (18). The
anti-tumor efficacy and safety of RCM-1 were demonstrated in mouse
models of different cancers. RCM-1 inhibited FOXM1 nuclear
localization (18,19) and suppressed RMS tumor cell
proliferation, colony formation and migration (19,20).
The B-cell lymphoma 2 (Bcl2) protein family
regulates the intrinsic apoptotic pathway, balancing the pro- and
anti-apoptotic proteins to maintain cellular homeostasis.
Disturbing this balance to evade apoptosis is a hallmark of cancer
(21). The anti-apoptotic protein
Bcl2 is commonly expressed in rhabdomyosarcoma and has been
associated with poor survival outcomes (22,23). Venetoclax, an FDA-approved
Bcl2-specific inhibitor, is effective in hematologic malignancies
but shows limited activity in solid tumors, necessitating rational
combination therapies to enhance anti-tumor activity (24).
Previous preclinical RMS studies have shown that
inhibiting Bcl2 has synergistic effects with chemotherapy (25-27). However, combining venetoclax with
cytotoxic chemotherapy increases toxicity, notably prolonged
neutropenia and thrombocytopenia (28). Therefore, the combination of
venetoclax with targeted therapies such as RCM-1 may offer a more
favorable safety profile.
The object of the present study was to determine
whether the combination of RCM-1 and venetocalx exerted synergistic
anti-tumor effects in RMS, while potentially offering fewer toxic
side effects via targeting the apoptotic pathway.
Materials and methods
To evaluate the anti-tumor effects of RCM-1 and
venetocalx in RMS, the present study performed a series of in
vitro and in vivo experiments using mouse and human RMS
cell lines. Cell viability, cytotoxicity and apoptosis were
measured using CCK-8 and caspase 3/7 assay. To investigate the
mechanism of synergy between RCM-1 and venetocalx, the present
study performed RNA sequencing, followed by gene knockdown and
overexpression, which were utilized for a series of in vitro
and in vivo experiments, including immunofluorescence,
reverse transcription-quantitative (RT-q) PCR and growth and
migration assays.
Cell lines and reagents
RD and RH30 (ATCC) are fusion-negative and
fusion-positive human RMS cells, respectively (29). 76-9 is a murine-derived RMS cell
line isolated from a methylcholanthrene-induced mouse RMS tumor in
a female C57BL/6 mouse and was provided by Dr Tim Cripe (Nationwide
Children's Hospital, Columbus, OH, USA) (30,31). Human RMS cells were authenticated
using STR profiling (Arizona Genetic Core). Cells were cultured in
DMEM (76-9 and RD) or RPMI (RH30) complete media (Gibco; Thermo
Fisher Scientific, Inc.) and kept at 37°C and a 5% CO2
incubator. The small molecule compound RCM1
(2-[2-oxo-2-(thiophen-2-yl) ethyl]sulfanyl-4,6-di(thiophen-2-yl)
pyridine-3-carbonitrile) was synthesized by Vitas-M Laboratory (95%
purity) and dissolved in DMSO for in vitro studies.
Venetoclax (ABT-199) purchased from APeXBIO Technology LLC (cat.
no. A8194) was dissolved in DMSO for in vitro studies and
reconstituted in the solution that contained 10% ethanol, 30%
polyethylene glycol 400 (PEG 400) and 60% lecithin dissolved in
propylene glycol for in vivo studies.
Cell proliferation assays
76-9 (10×104) and RD cells
(8×104) per well were seeded in 6-well plates and
allowed to grow for 24 h. RCM-1 was added at 24 h and venetoclax
was added at 48 h in the sequential treatment regimen. Trypan blue
staining was performed by incubating the cell suspension with 0.4%
trypan blue at room temperature for 1 min. to exclude dead cells
and viable cells were counted using a hemocytometer. To measure
cell viability, cells were incubated with the cell counting kit-8
(CCK8) solution (GLPBIO Technology LLC) at 37°C for 1 h, followed
by detecting the absorbance at 450 nm using a microplate reader.
Experiments were performed in triplicate.
Caspase 3/7 activation assay
Apoptosis in RMS cells following treatment with
RCM-1 and venetoclax was assessed using the Caspase-Glo®
3/7 Assay (Promega Corporation) according to the manufacturer's
protocol. After 72 h of drug incubation, as aforementioned, the
Caspase-Glo 3/7 reagent was added to each sample at a 1:1 ratio
with the culture volume. The plate was then incubated at 37°C for
30 min and luminescence was subsequently measured using a
microplate reader.
Nanoparticle synthesis
All chemicals for nanoparticle synthesis were used
as received without any further purification and were obtained from
MilliporeSigma.
The Poly(β-Amino Ester) (PBAE) polymer backbone was
synthesized via a modified Michael Addition, as described in our
previous studies (20,32). Briefly, Bisphenol A glycerolate
diacrylate was initially mixed with 6-amino-1-hexanol in DMSO at
90°C for 24 h. Following this, 4,4'-Trimethylenedipiperidine in
DMSO was added to the mixture as the temperature was reduced to
50°C and maintained for another 24 h. The PBAE backbone was then
capped with methoxypolyethylene glycol amine and folic
acid-modified polyethylenimine in DMSO at 40°C. The folic acid
modification was performed via EDC/NHS coupling, as previously
described (33). To encapsulate
RCM-1, the PBAE polymers were mixed with RCM-1 in DMSO at a mass
ratio of 10:1, followed by transitioning the mixture to an aqueous
environment to facilitate DMSO diffusion and nanoparticle
self-assembly. The resulting nanoparticles were then dialyzed for
48 h to remove DMSO, excess drug and extra polymers. The
concentration of RCM-1 encapsulated in the nanoparticles was
determined by UV/Vis spectroscopy as described (34).
Mouse model
A total of 55 C57Bl/6J mice (8-12 weeks old, 1:1
male and female) were purchased from the Jackson Laboratory. The
mean male mouse weight was 25 g and the average female mouse weight
was 20 g. All mice were kept under SPF (specific-pathogen-free)
conditions in 12-h light/dark cycle, 18-23°C and 40-60% humidity.
To generate the subcutaneous syngeneic murine model,
1×106 76-9 rhabdomyosarcoma cells were re-suspended in
equal volumes of PBS: Matrigel (Corning, Inc.) and were injected
subcutaneously into the flanks of mice (35). On day 7 after tumor cell
inoculation, tumor-bearing mice were randomly assigned into control
(n=19), single-agent RCM-1 (n=12), venetocalx (n=12), or
combination therapy (n=12) groups. RCM-1 encapsulated into
nanoparticles was prepared as previously described (RCM-1
NPFA) (20) and
administered via tail vein injection every other day for a total of
seven injections, using the half-maximal inhibitory concentrations
(IC50) dose of 8 μg. Venetoclax (100 mg/kg/dose)
was administered via oral gavage 5 days a week for 2 weeks. Control
mice were injected and orally given a vehicle control. Tumors were
harvested on day 21. Mice were monitored for signs of distress and
weighed every other day until day 21. Mice were euthanized using IP
pentobarbital (100 mg/kg/dose) followed by cervical dislocation to
ensure irreversible death (36,37). Tumors were measured using calipers
and volumes were calculated in cubic millimeters using V=
0.5xLxW2, where L was the tumor length and W was the
tumor width.
Generating gene knockdown and
overexpression in RMS cells
To knockdown ATP2B4 in vitro, RMS cells were
transfected with SMARTpool siRNA (Horizon Discovery; cat. no.
L-066791-00-0010) and non-targeting siRNA pool as a control
(Horizon Discovery; cat. no. D-001810-10) by using Dharmafect
transfection reagent (Dharmacon, Inc.; Revvity, Inc.) as previously
described (38). For stable
knockdown of ATPase Plasma Membrane Ca2+ Transporting 4
(Atp2b4) in 76-9 cells, pre-packaged lentiviral particles
containing ATP2B4 short hairpin (sh)RNA were purchased directly
from Origene (cat. no. TL508592V; titer 4.8×107 TU/ml).
Wild-type (WT) 76-9 cells were transduced with two mouse lentiviral
particles (C, D) at a multiplicity of infection (MOI=50) in the
presence of 8 μg/ml polybrene, and incubated at 37°C
overnight. Medium was replaced the following day, and cells were
cultured for 48-72 h. GFP+ transfected cells were
isolated by FACS. Briefly, cells were dissociated into a
single-cell suspension, washed with PBS, resuspended in culture
medium and filtered through a 40-μm cell strainer prior to
sorting. GFP+ cells were gated based on unstained
control cells and collected using a Sony SH800 cell sorter. FACS
analysis was performed as previously prescribed (18). To knockdown FOXM1, 76-9 and
RH-30 were transfected with pLKO-shFOXM1 plasmid DNA (clone
ID: TRCN0000015546; MilliporeSigma) against FOXM1using
TransIT-X2 (Mirus Bio, LLC; cat. no. MIR 6004). To overexpress
ATP2B4a and ATP2B4b, 76-9 cells were transfected with CMV-ATP2B4a
(Origene; cat. no. MC223809) and CMV-ATP2B4b (Origene; cat. no.
MR215322) or an empty overexpression (OE) plasmid as a control. For
both FOXM1 knockdown and ATP2B4 overexpression transfections
performed using TransIT-X2, cells were plated at 8×104
cells per well in a 24-well plate and transfected with 0.5
μg plasmid DNA premixed with 1.5 μl TransIT-X2,
followed by incubation at 37°C for 24 h before subsequent selection
or analysis. Sequences are listed in Table SI.
Immunofluorescence staining
Control, siATP2B4 and pATP2B4 76-9 cells were seeded
at a density of 1×105 cells per 24 mm square coverslip
in 6-well plates and allowed to grow for 48 h. Cells were then
fixed in 4% paraformaldehyde at room temperature for 15 min, washed
three times with PBS, and stained as previously described (20). Blocking was performed using 4%
Normal Goat Serum (Jackson ImmunoResearch Labs; cat. no.
005-000-121) at room temperature for 3 h. Primary antibodies were
incubated overnight at 4°C, followed by secondary antibody
incubation at room temperature for 1 h, during which Hoechst 33342
(Invitrogen; Thermo Fisher Scientific, Inc.; cat. no. H3570) was
included as a nuclear counterstain. For quantification, five random
fields per sample were acquired at 20× magnification using the EVOS
FL Auto 2 Cell Imaging System and EVOS imaging software (Thermo
Fisher Scientific, Inc.). To perform immunostaining of tumor
tissue, paraffin-embedded 76-9 subcutaneous tumor sections were
stained as described previously (39). Briefly, tissues were fixed in 4%
paraformaldehyde at 4°C overnight, dehydrated through graded
ethanol and xylene, and embedded in paraffin at 60°C. Sections (5
μm) were deparaffinized, rehydrated, and subjected to
antigen retrieval in citrate buffer (pH 6.0) at 95°C for 20 min.
Slides were blocked with 4% Normal Goat Serum at room temperature
for 1 h, incubated with primary antibodies overnight at 4°C, and
then incubated with secondary antibodies at room temperature for 1
h with Hoechst 33342 included as a nuclear counterstain. Five
random fields per sample were acquired and quantified using ImageJ,
with imaging performed on the same EVOS FL Auto 2 platform.
Antibodies used for immunostaining were anti-Ki-67 (1:250;
Invitrogen, MA5-14520), anti-Cleaved-Caspase 3 (1:200; R&D,
MAB835), anti-BAX (1:150; Santa Cruz, sc-7480), anti-ATP2B4 (1:200;
Thermo Fisher Scientific, Inc.; cat. no. PA5-87634).
RT-qPCR
RNA was isolated using the RNeasy kit (Qiagen; cat.
no.74104) and was subsequently reversed to cDNA using the iScript
cDNA synthesis kit (Bio-Rad Laboratories, Inc.; cat. no.1708891).
qPCR was carried out using TaqMan Gene Expression Assays (Applied
Biosystems) according to the manufacturer's instructions. The
thermal cycling conditions were: An initial hold at 50°C for 2 min,
followed by 95°C for 10 min, and 45 cycles of 95°C for 15 sec and
60°C for 1 min (annealing/extension with data acquisition).
Experiments were performed in triplicates, and relative gene
expression was analyzed using the 2−ΔΔCq method
(40). Assay catalog numbers are
provided in Table SII.
Clonogenic growth assay
Control and small interfering (si)ATP2B4 knockdown
76-9 cells were seeded at 2,000 cells per well in 6-well culture
plates and cultured in a 37°C, 5% CO2 incubator. The
culture medium was replaced every 2 days. After 6 and 9 days in
culture, the cells were washed with Dulbecco's phosphate-buffered
saline (DPBS), fixed with 4% paraformaldehyde (PFA) for 10 min at
room temperature and then treated with 100% ice-cold methanol for 5
min to permeabilize the cells. Subsequently, the cells were stained
with 0.5% crystal violet (dissolved in 25% methanol) for 10 min at
room temperature. After staining, the crystal violet solution was
removed and the cells were washed with DPBS at least twice to
achieve a clear background. Representative images were captured and
the crystal violet staining was quantified using ImageJ software
(version 1.53e; National Institutes of Health).
Wound healing assay
Ibidi two-well culture inserts (Ibidi GmbH; cat.
no.80209) were used for the experiment. A total of 4×104
control or siATP2B4 knockdown 76-9 cells were seeded into the
culture inserts and allowed to adhere overnight. Cells were
maintained in complete growth medium. Afterward, the culture
inserts were removed, creating open areas with clear edges. Images
of these open areas were captured using the EVOS FL Auto 2 Cell
Imaging System (Thermo Fisher Scientific, Inc.) at 0, 6 and 24 h
using a light (phase-contrast) microscope, the EVOS FL Auto 2 Cell
Imaging System (Thermo Fisher Scientific, Inc.). The images were
analyzed using TScratch software (CSElab; ETH Zurich; version
1.0).
Fura-2 AM staining
76-9 cells were transfected with siAtp2b4 or a
non-targeting control (siNC) for 24 h, then stained with 5
μM Fura-2 AM (AAT Bioquest; cat. no. 21020) in a 37°C, 5%
CO2 incubator. After 30 min of incubation, the cells
were rinsed with DPBS and images were captured using the EVOS FL
Auto 2 Cell Imaging System (Thermo Fisher Scientific, Inc.).
Apoptosis measurement and Caspase 3
activation assay
Control, siATP2B4 and ATP2B4 overexpressed 76-9
cells were seeded in a 24-well plate at a density of
8×104 cells/ml overnight and treated with venetoclax (5
and 8 μM) for 24 h. Then, apoptotic cells were assessed via
Annexin V-iFluor 488 staining and Caspase 3 activity was measured
using TF3-DEVD-FMK staining with the Cell Meter Live Cell Caspase
3/7 and Phosphatidylserine Detection Kit (AAT Bioquest; cat. no.
22850). Cells were incubated with Annexin V–iFluor 488 and
TF3-DEVD-FMK at 37°C in a 5% CO2 incubator for 1 h,
followed by Hoechst staining at room temperature for 10 min.
Fluorescence images were captured and analyzed using the EVOS FL
Auto 2 Cell Imaging System (Thermo Fisher Scientific, Inc.). The
apoptotic rate was calculated as the percentage of Annexin
V-positive cells, including both early and late apoptotic
populations, relative to the total number of Hoechst-positive
cells.
RNA-sequencing (RNA-seq) and data
analysis
RNA extracted from control and treated 76-9 cell
line were sent to the CCHMC Genomics Sequencing Facility,
Cincinnati, Ohio (USA) for sequencing. The quality of RNA was
determined using a Fragment Analyzer with an average RNA Quality
Number for all samples of 9.87. RNA libraries were prepared for all
samples using Illumina Stranded total RNA Prep, Ligation with
Ribo-Zero (Illumina, Inc.) to generate non-stranded RNA libraries.
Sequencing was performed using NoveSeq6000 (Illumina, Inc.) with an
estimated 30 million read per sample. Reads were aligned to the
GRCm38 mouse genome and quantified using an index transcriptome
version of GRCm38 using Kallisto and standard settings. This
was performed by the CCHMC Genomics Sequencing Facility,
Cincinnati, Ohio (USA). Raw counts were normalized using DESeq2
(41). Differential gene
expression between conditions was performed using DEseq2, which
uses a negative binomial model for each gene. The Wald test was
used for hypothesis testing when comparing the two groups. All
P-values attained were corrected for multiple testing using the
Benjamini and Hochberg method, which is the default method in
DESeq2. In the standard DESeq2 algorithm, the α for the
false-discovery rate is set to 0.1 by default. Heatmap was
generated using the pheatmap R package (version 1.0.12;
https://cran.r-project.org/package=pheatmap) and the
volcano plot was generated using the EnhancedVolcano R
package (https://github.com/kevinblighe/EnhancedVolcano). Venn
diagrams for differentially expressed genes were created using
AltAnalyze (42). Gene list
functional enrichment was created using TopGene Suite and graphed
using sRplot (43,44).
Single-cell RNA-seq data analysis
The datasets GSE143704 (45) and GSE195709 (46) were obtained from the GEO database
(https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE143704
and https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE195709).
The GSE143704 dataset contains muscle tissue cell populations from
10 healthy human donors, while the GSE195709 dataset includes
rhabdomyosarcoma samples from 4 primary rhabdomyosarcoma patients.
The datasets were both generated using the NextSeq 500 platform
(Illumina, Inc.). Data analysis was performed using the Seurat R
package (version 4.3.1; https://satijalab.org/seurat/). Cells with >100
genes detected or with a mitochondrial gene percentage >20% were
excluded. The present study used the FindClusters and
FindAllMarkers functions in Seurat to identify clusters for Smooth
Muscle Cells (ACTA2), Skeletal Muscle (ACTA1) and Myoblasts (PAX7).
These three clusters were then integrated with the scRNA-seq
dataset of the 4 rhabdomyosarcoma samples using the SCTintegration
function of the Seurat package. The cells were clustered using the
FindClusters function and cluster visualization was performed using
Uniform Manifold Approximation and Projection (UMAP). ATP2B4
expression levels were analyzed to generate a violin plot.
Cloning of the human ATP2B4 promoter
region and the luciferase reporter assay
The human ATP2B4 promoter region spanning-373
to 0 bp was obtained from 293 genomic DNA through PCR amplification
using the following primers (5'-3'): Forward:
TGAGCAAGAGTCTGGCCCGGGGTACCCC; reverse:
GGGGTACCCCGGGCCAGACTCTTGCTCA. This promoter region was then cloned
into the KpnI site of the pGL4.23[luc2/minP] luciferase
reporter plasmid (Promega Corporation). A Dual-Glo luciferase
reporter assay was performed on 76-9 cells, which were
co-transfected with the luciferase reporter, Renilla and
either a CMV-empty or CMV-FOXM1 overexpression plasmid.
Statistical analysis
Data were expressed as mean ± SD. Statistical
significance was determined using an unpaired Student's t-test, or
one-way or two-way analysis of variance (ANOVA) followed by Tukey's
multiple comparisons post hoc test, where appropriate. The in
vivo experiment was a pilot study with no formal power
calculation required and was analyzed in a blinded manner. All
statistical analyses were obtained using GraphPad Prism (version
9.5.1; Dotmatics). P<0.05 was considered to indicate a
statistically significant difference.
Results
The sequential treatment of RCM-1 and
venetoclax decreases RMS cell viability and enhances apoptosis in
vitro
To establish the IC50 of RCM-1 and
venetoclax, mouse 76-9 and human RD rhabdomyosarcoma cell lines
were treated with increasing doses of RCM-1 and venetoclax to
generate a dose-response curve. The IC50 of RCM-1 was
2.5 μM in RD cells and 1.37 μM in 76-9 cells, whereas
venetoclax had higher IC50 values of 9.4 μM in RD
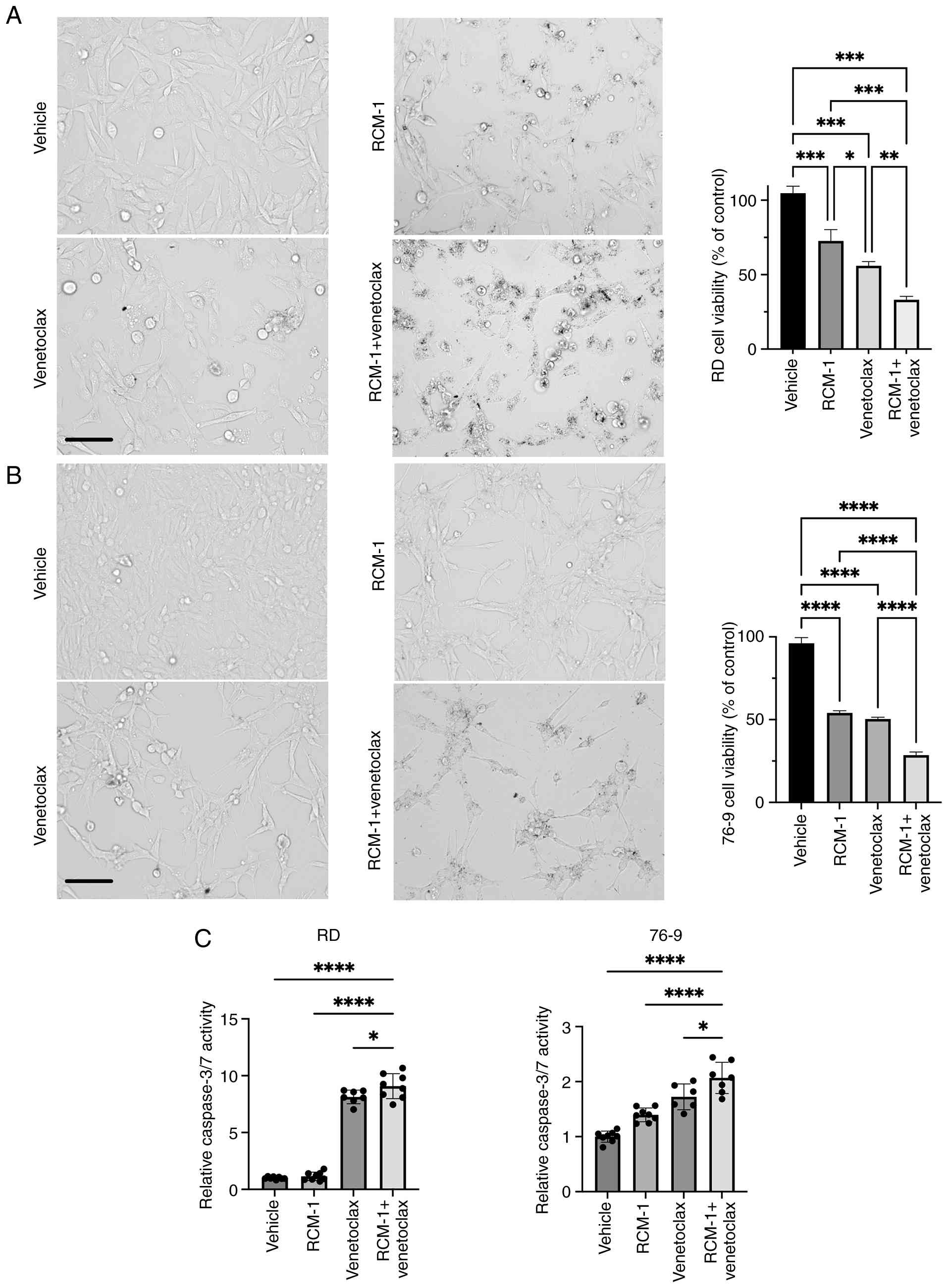
cells and 6.3 μM in 76-9 cells (Fig. S1A). Next, RMS cells were treated
with either RCM-1 alone, venetoclax alone, or a combination of both
agents sequentially. The combination therapy of RCM-1 and
venetocalx reduced the number of viable tumor cells by 70% compared
with 40-50% with single agents (Fig.
1A and B). Also, the CCK8 assay showed a significant reduction
in RD, 76-9 and RH30 cell viability after RCM-1 and venetocalx
treatment compared with a single agent or vehicle (Figs. S1B and C). Next, caspase 3/7
activity was assessed using Caspase-Glo® 3/7 Assay kit.
The combination therapy significantly increased apoptosis in RD,
76-9 and RH30 cells compared with single agents (Figs. 1C and S1C). Thus, the combination of RCM-1 and
venetoclax inhibits RMS tumor cell growth and induces apoptosis
more efficiently than either drug alone.
Combination therapy with
RCM-1-NPFA and venetoclax inhibits tumor growth and
promotes tumor cell apoptosis in the mouse model of RMS
To verify that the efficacy of combination therapy
is not limited to in vitro conditions, a mouse model of RMS
was used. 76-9 RMS tumor cells were inoculated subcutaneously into
the flanks of C57Bl/6J mice. The tumor-bearing mice were randomly
assigned to 4 groups and were treated with either vehicle
(control), venetoclax alone, IC50 dose of
nanoparticle-encapsulated RCM-1 alone, or a combination of both
drugs (20) (Fig. 2A). Monotherapy with venetoclax did
not have anti-tumor efficacy in the RMS mouse model (Fig. 2B), which is consistent with
previously published studies (26,47) (Fig.
2B). Monotherapy with the IC50 dose of
RCM-1-NPFA had only limited anti-tumor efficacy
(Fig. 2B). However, the
combination therapy efficiently reduced RMS tumor growth in a mouse
model (Fig. 2B). The combination
therapy effectively suppressed cell proliferation, as indicated by
the decreased number of Ki-67 positive cells, and significantly
promoted apoptosis, evidenced by the increased percentage of BAX
and cleaved caspase-3 positive cells compared with single agents
(Fig. 2C-E). The combination
therapy was well tolerated in mice, with no observed weight loss
throughout the experiment (Fig.
S2A). Venetoclax-treated mice exhibited reduced white blood
cell counts compared with controls (Fig. S2B), consistent with known
myelosuppressive effects of venetoclax in humans. Importantly, the
combination therapy did not induce liver dysfunction (Fig. SC). Thus, the combination of
RCM-1-NPFA and venetoclax efficiently inhibits tumor
growth and is not toxic in the animal model of RMS.
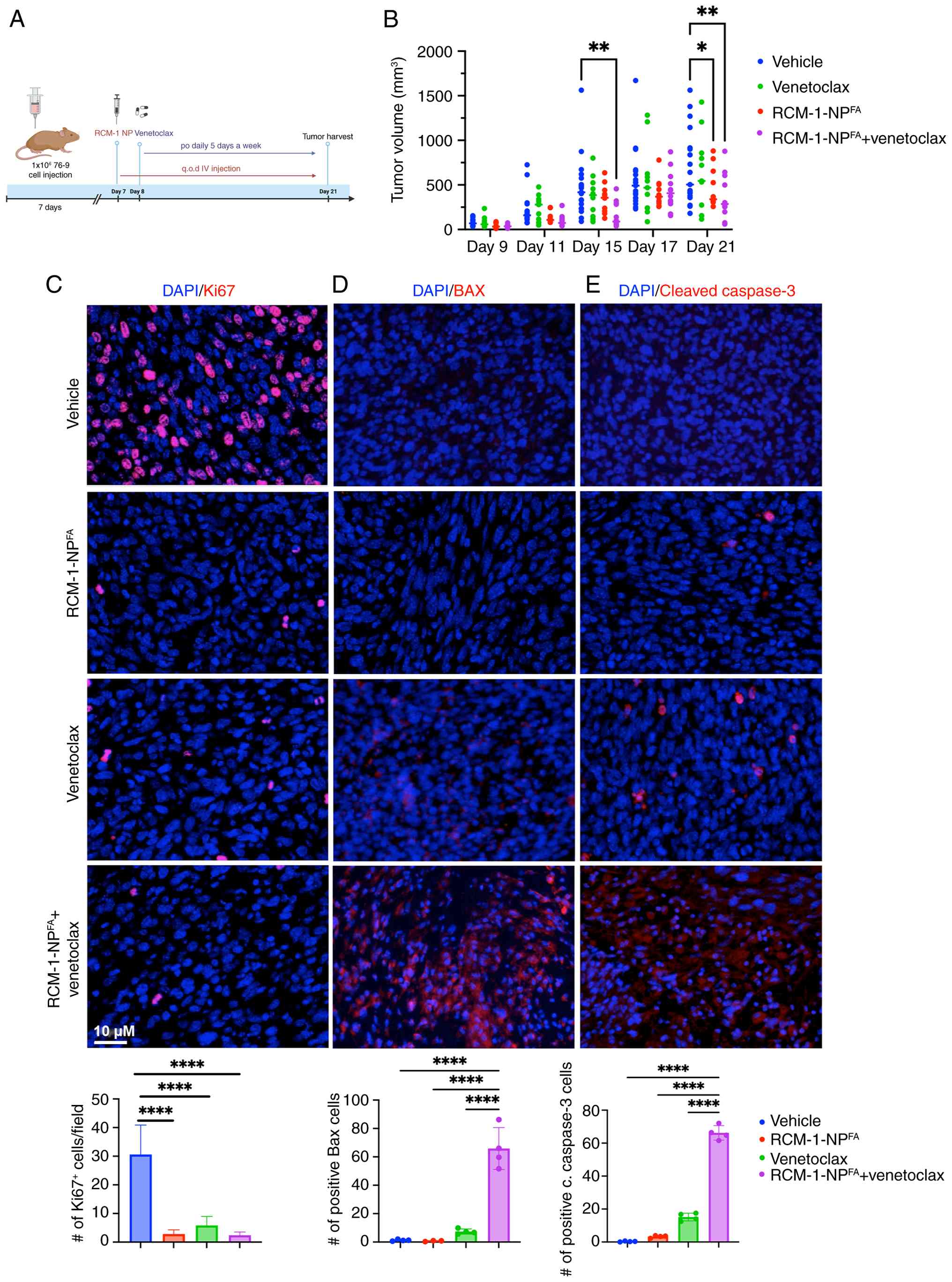 | Figure 2The combination of
RCM1-NPFA and venetoclax decreases tumor growth and
enhances caspase-mediated apoptosis in a murine RMS model. (A)
Schematic diagram of tumor cells inoculation and treatment. The
figure is created in BioRender. Merjaneh, N. (2024) https://BioRender.com/z67t342. (B) Combination
therapy significantly reduced tumor burden compared with vehicle.
The mean vehicle tumor volume on day 21 was 685 mm3,
compared with the average tumor volume of the combination therapy
of 361 mm3. The maximum tumor diameter was 17.3×11.5 mm
and the corresponding maximum tumor volume was 1,144
mm3. Tumor volume was measured at different time points
during the experiment (P≤0.01; n=12). (C) Combination therapy
inhibited proliferation, as indicated by the decreased number of
Ki67-positive cells. Combination therapy increased apoptosis, shown
by the increased number of (D) BAX-positive cells and (E) the
number of caspase3-positive cells compared with single agents
and/or vehicle. A total of five random fields per sample were used
to quantify the number of Ki67, BAX and cleaved caspase 3 positive
cells per group. Values are shown as mean ± SD. Scale bar, 10
μm. *P≤0.05, **P≤0.01,
****P≤0.0001. RMS, rhabdomyosarcoma. |
RNA-seq analysis identifies a signature
of cellular stress in the combination therapy
To determine the molecular mechanism of increased
apoptosis in the combination therapy, bulk RNA sequencing was
performed to compare the transcriptome of untreated control and
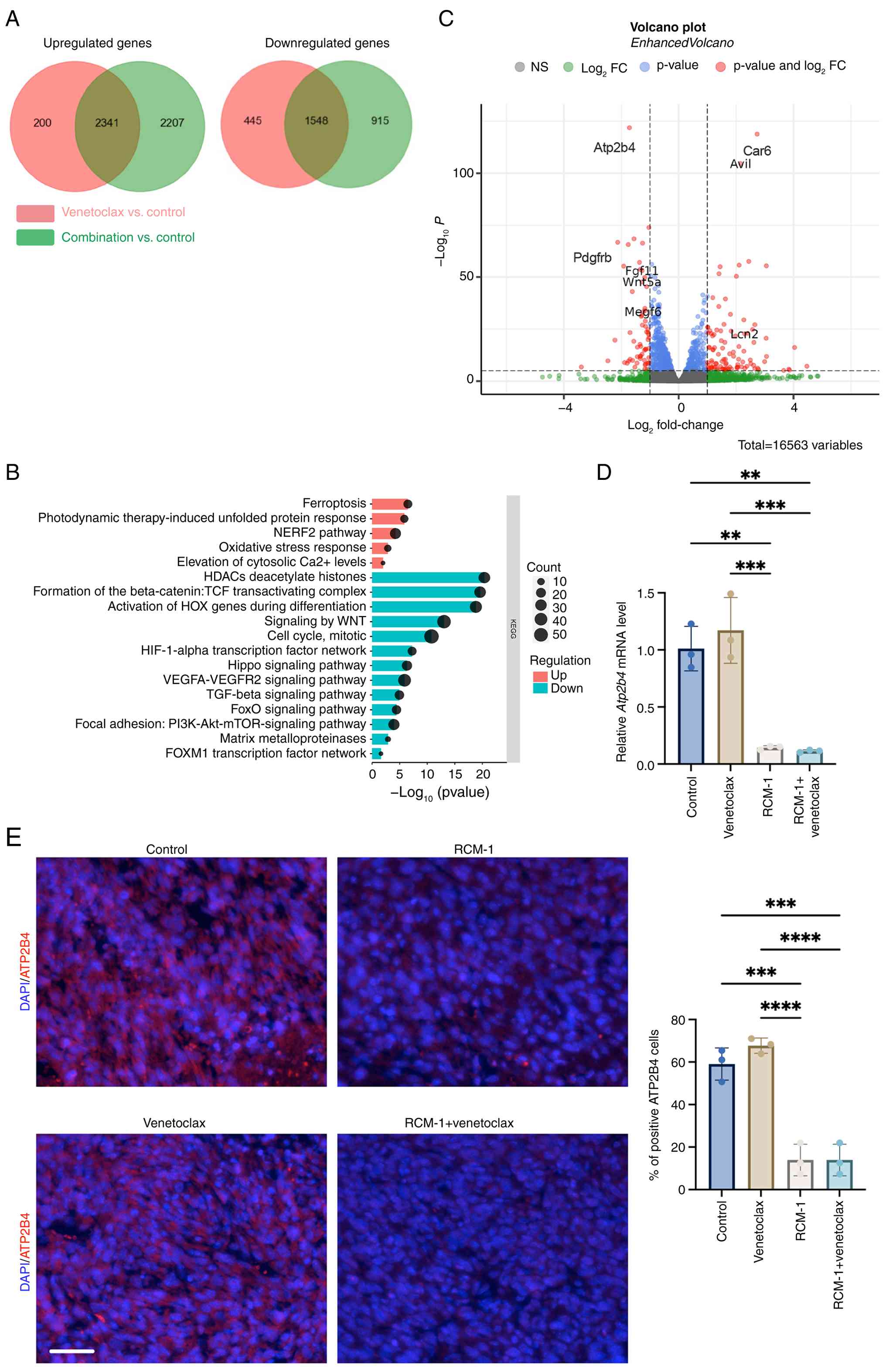
treated 76-9 RMS cells (Fig. 3A).
The gene enrichment analysis of RMS cells treated with the
combination therapy compared with venetoclax revealed an
upregulation in multiple biologic pathways that are activated in
cells under stress, such as ferroptosis, unfolded protein response
and oxidative stress response. The cytosolic Ca2+ level
pathway was significantly elevated in the combination therapy
compared with venetoclax as well. The downregulated pathways
included WNT signaling, mitotic cell cycle, hypoxia-inducible
factor-1α signaling, Hippo signaling, TGF-β signaling, AKT-mTOR and
FOXM1 network pathways, reflecting the role of RCM-1 in FOXM1
inhibition (Fig. 3B). A volcano
plot was generated to visualize the most differentially expressed
genes in combination therapy compared with venetoclax treatment
(Fig. 3C). Atp2b4 was one
of the most downregulated genes in the combination treatment. The
decreased expression of Atp2b4 mRNA was confirmed using
RT-qPCR, demonstrating that both RCM-1 treatment and combination
therapy were sufficient to decrease the Atp2b4 mRNA level
(Fig. 3D). Furthermore, using
immunofluorescence staining, the present study showed that the
protein level of ATP2B4 was also decreased in RMS tumors after
RCM-1 or combination treatment (Fig.
3E).
ATP2B4 is differentially expressed in
rhabdomyosarcoma cells and is essential for RMS cell proliferation
and migration
Since it has been shown that ATP2B4, a calcium
channel located on the plasma membrane, plays an important role in
different cancers (48,49), the present study analyzed the
publicly available sc-RNA genomic databases of human normal muscle
tissue and RMS tumors (45,46) (Figs. S3A and 4A). Consistent with previously published
data, Bcl2 was overexpressed in RMS (22,23) and CDKN1A (a cell cycle
inhibitor) was overexpressed in normal muscle cells (Fig. S3B). Based on the sc-RNA seq
analysis, the mRNA level of ATP2B4 was markedly higher in
fusion-negative (RMS 1 and 3), fusion-positive (RMS 2) and spindle
(RMS 4) rhabdomyosarcoma compared with normal human muscle cells,
including skeletal muscle, smooth muscle cells and myoblasts
(Fig. 4A). Also, ATP2B4 mRNA
level was 3-4 fold higher in human RMS cell lines (RD and RH30)
compared with normal skeletal muscle cells (HSkMC) (Fig. 4B). To determine the role of
ATP2B4, 76-9 RMS cells were transfected with siRNA against
Atp2b4, resulting in ~45% knockdown of Atp2b4 mRNA
after 24 h and ~87% after 48 h (Fig.
S4A; left panel). Immunostaining also demonstrated that the
protein level of ATP2B4 was decreased 24 h after siATP2B4
transfection (Fig. S4A; right
panel). Depletion of ATP2B4 increased the basal cytosolic
Ca2+ level in RMS tumor cells (Fig. 4C), which is consistent with the
upregulated signaling pathway related to cytosolic Ca2+
levels shown in RNA seq pathway analysis (Fig. 3B). Furthermore, depletion of
ATP2B4 in rhabdomyosarcoma cells led to a time-dependent decrease
in tumor cell proliferation, with a 40% reduction observed at 72 h
as well as reduced colony formation and cell migration (Fig. 4D-F). Thus, ATP2B4 is
differentially expressed in rhabdomyosarcoma cells and is essential
for RMS cell proliferation and migration.
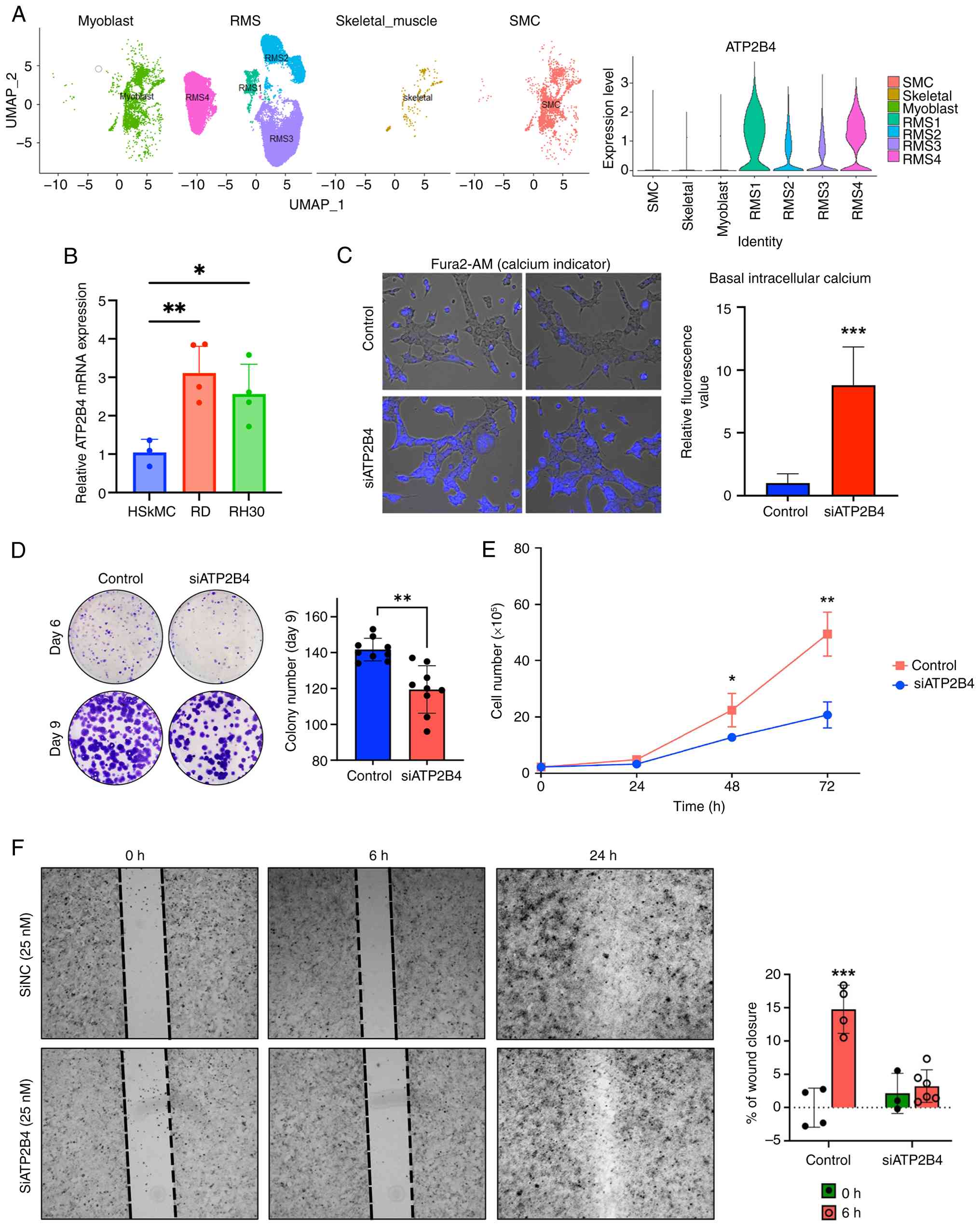 | Figure 4ATP2B4 is differentially
overexpressed in RMS cells vs. normal muscle cells and its
knockdown decreases tumor cell proliferation, migration and colony
formation. (A) Left panel, Human myoblast (muscle progenitor
cells), RMS, normal skeletal muscle and smooth muscle scRNA
sequencing datasets were visualized using UMAP. Data were extracted
from GSE143704 for normal muscle tissue, GSE 195709 for RMS (RMS1
and 3 are fusion-negative RMS, RMS2 is fusion-positive and RMS4 is
spindle (fusion-negative). Right panel, ATP2B4 mRNA
expression was higher in RMS cells compared with normal muscle
cells. (B) Reverse transcription-quantitative PCR showed the
upregulation of ATP2B4 mRNA level in RMS cells (RD and RH30)
compared with HSkMC. (C) Knockdown of ATP2B4 increased the
intracellular calcium level. Calcium level was assessed using the
calcium-sensitive fluorescent dye fura-2 AM. Scale bar, 10
μm. (D) Knockdown of ATP2B4 inhibited colony
formation in 76-9 RMS cells. Data presented as mean ± SD. (E)
Knockdown of ATP2B4 suppressed 76-9 cell proliferation in
culture. Data presented as mean ± SD. (F) ATP2B4 knockdown
decreased RMS cell migration. The percentage of wound closure
presented as mean ± SD (magnification, ×4). *P≤0.05,
**P≤0.01, ***P≤0.001. RMS, rhabdomyosarcoma;
UMAP, Uniform Manifold Approximation and Projection; HSkMC, human
skeletal muscle cells; SMC, smooth muscle cells. |
Knockdown of ATP2B4 enhances
venetoclax-mediated apoptosis in 76-9 RMS cells
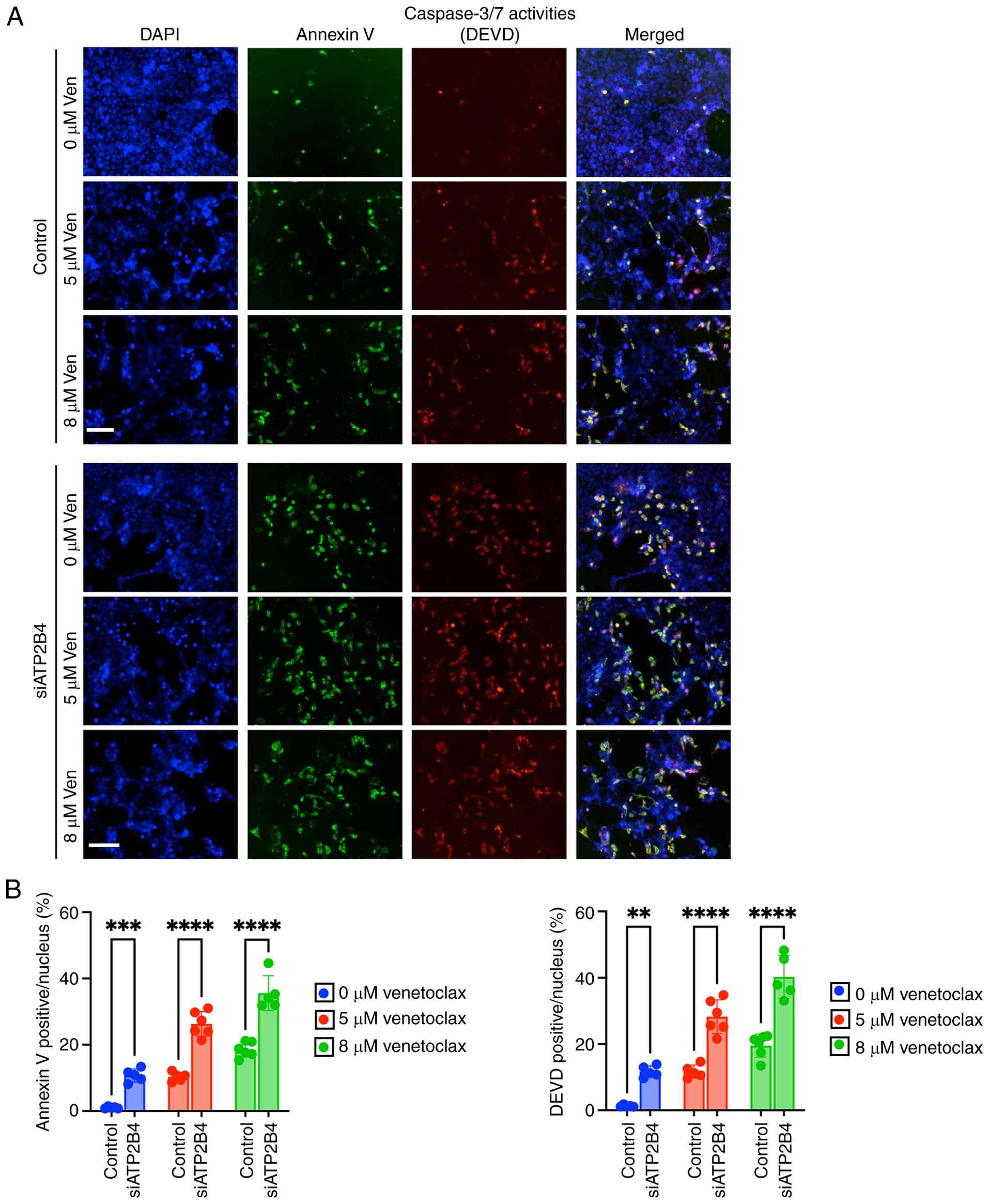
To determine whether ATP2B4 is important for
venetoclax-induced apoptosis in rhabdomyosarcoma, the present study
treated control and ATP2B4-depleted tumor cells with increasing
doses of venetoclax. Using immunostaining with antibodies against
caspase 3/7 (DEVD) and annexin V, it was shown that venetoclax
induced a dose-dependent increase in apoptosis of control tumor
cells (Fig. 5A and B). Moreover,
the depletion of ATP2B4 in the siATP2B4-KD cells increased the
efficacy of the same doses of venetoclax, which was demonstrated by
a higher percentage of Annexin V-positive cells (10-35% in
siATP2B4-KD vs. 1-18% in controls, P£0.0001) and DEVD-positive
cells (10-40% vs. 1-20%, P<0.0001; Fig. 5A and B). Thus, depletion of ATP2B4
increases venetoclax-mediated apoptosis in 76-9 RMS cells.
ATP2B4 overexpression induces resistance
to apoptosis and decreases the therapeutic effect of venetoclax in
76-9 RMS cells
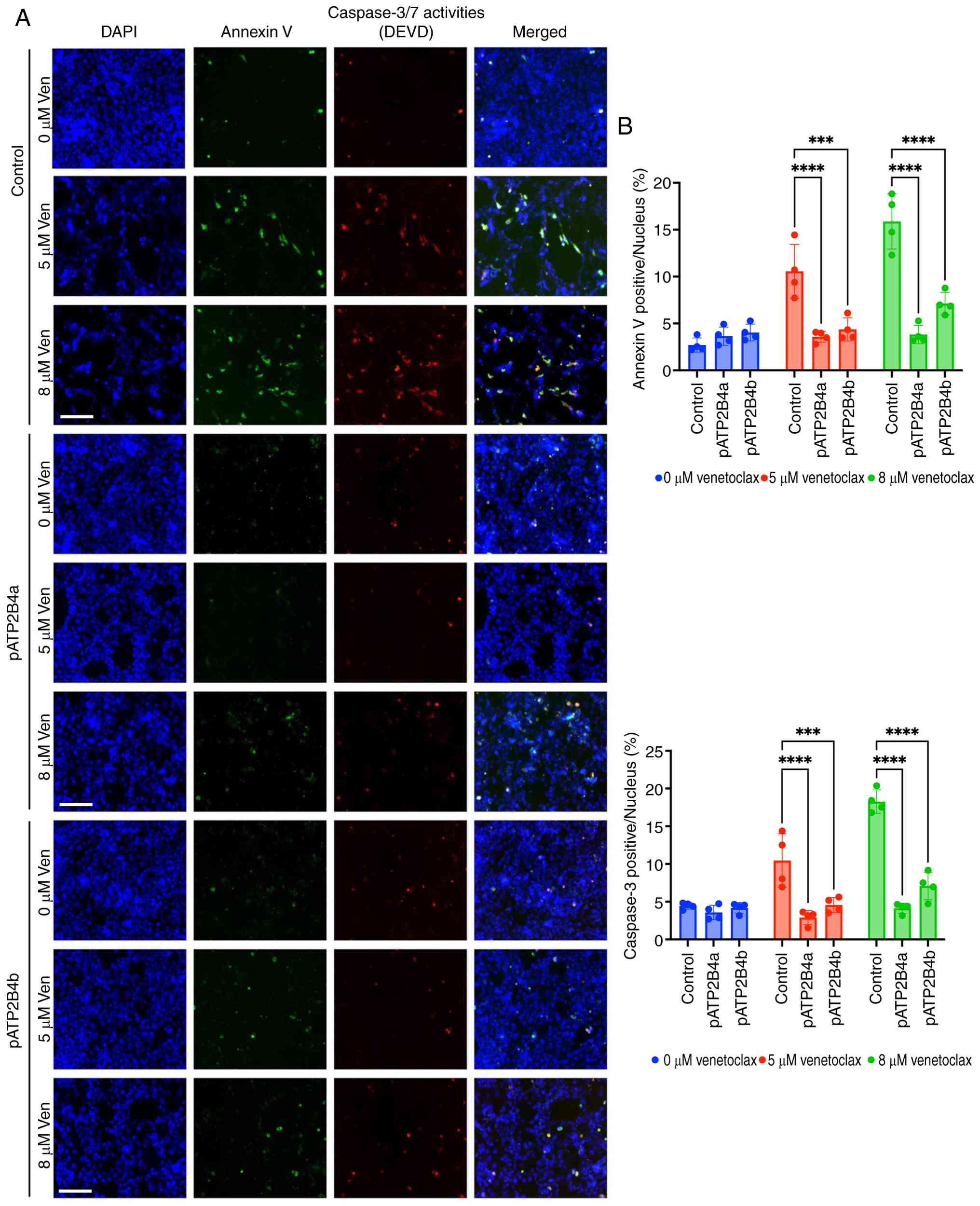
As the depletion of ATP2B4 sensitized RMS cells to
venetoclax-mediated apoptosis, the present study next determined
whether overexpression of Atp2b4 would inhibit
venetoclax-mediated apoptosis. ATP2B4 has 2 main isoforms, 4a and
4b, with isoform 4b being more predominant and active in different
cancers (50). Therefore, the
present study overexpressed Atp2b4a and Atp2b4b in
76-9 cells. The transfection efficiency was examined using RT-qPCR
and immunofluorescence (Fig.
S4B). The WT and pATP2B4 cells were treated with increasing
doses of venetoclax. Overexpression of Atp2b4 markedly
decreased cellular apoptosis after venetoclax treatment, measured
by decreased caspase 3/7 (DEVD) positive cells (3-8% in pATP2B4 vs.
4-18% in controls, P≤0.0001) and decreased cellular death, measured
by decreased annexin positive cells (3-8% in pATP2B4 vs. 2-18% in
controls, P≤0.0001; Fig. 6A and
B). Thus, increasing ATP2B4 levels induces RMS resistance to
apoptosis and decreases the therapeutic effect of venetoclax in RMS
cells.
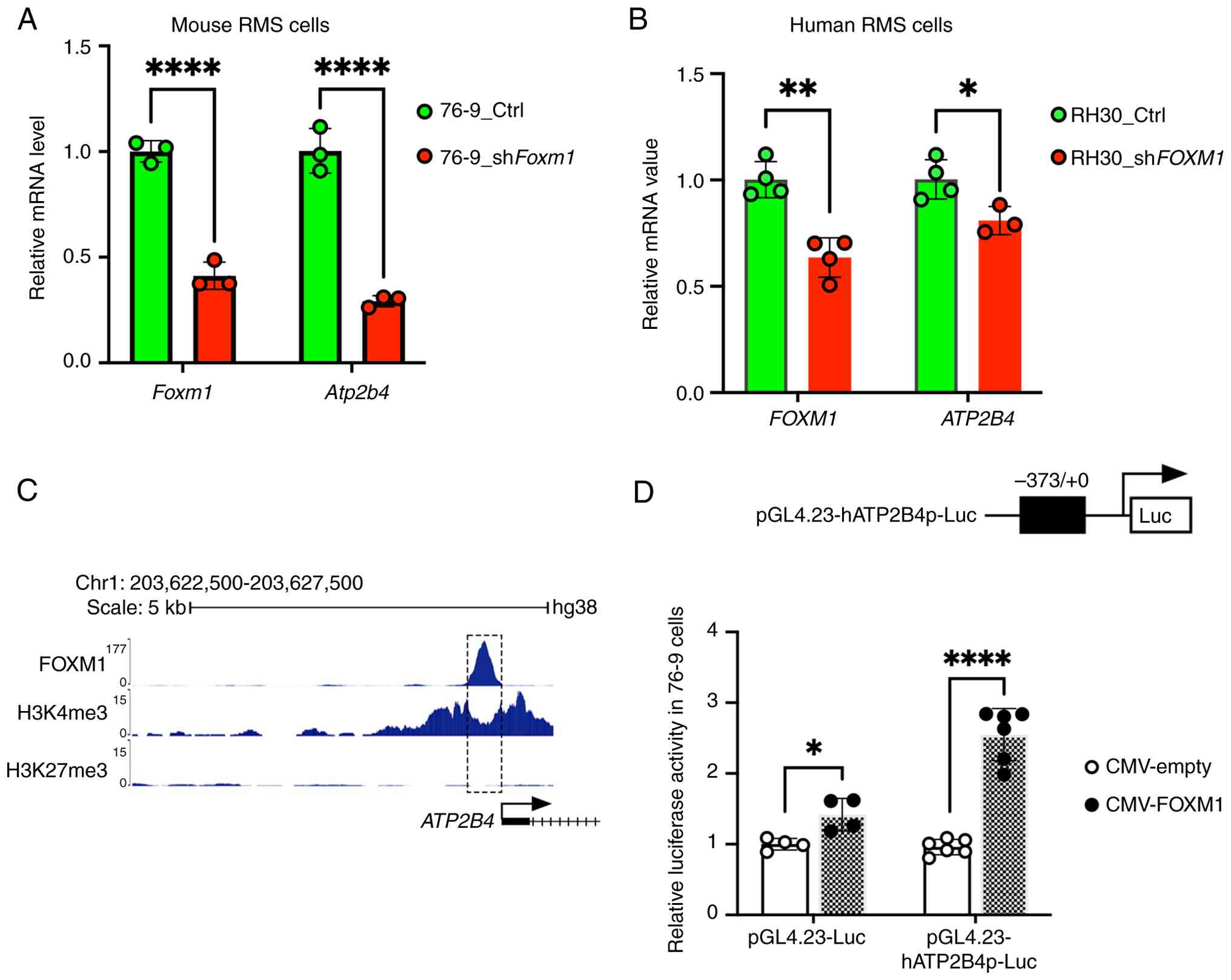
FOXM1 transcriptionally regulates ATP2B4
expression and RCM-1 downregulates ATP2B4 expression via FOXM1
inhibition
As it is known that RCM-1 specifically inhibits
FOXM1 and we have shown that RCM-1 decreases ATP2B4 in RMS
(Fig. 3D-E), the present study
next examined whether FOXM1 directly regulated ATP2B4 in RMS. Mouse
RMS cells were transfected with control and shFoxm1. The
shRNA-mediated knockdown of Foxm1 in 76-9 decreased
expression of Atp2b4 mRNA (Fig. 7A). Similarly, shRNA-mediated
knockdown of FOXM1 in human RH30 rhabdomyosarcoma cells also
decreased the expression of ATP2B4 (Fig. 7B). Next, a publicly available
ChIP-seq dataset from the ENCODE portal (51,52) was used and it was demonstrated
that FOXM1 directly binds to the ATP2B4 promoter region in
cancer cells (Fig. 7C). The
FOXM1-binding region in the ATP2B4 promoter had H3K4me3 but
not H3K27me3 marks, suggesting that FOXM1 activates the
ATP2B4 gene promoter (Fig.
7C). To verify that FOXM1 activates ATP2B4 gene
expression, the-373/+0 bp ATP2B4 promoter region, containing
the FOXM1-binding site identified by ChIP-seq (Fig. 7C), was cloned into the pGL4.23
luciferase reporter plasmid (Fig.
7D). In co-transfection experiments using mouse RMS cells, the
CMV-Foxm1 expression vector increased transcriptional activity of
the-373/+0 bp ATP2B4 promoter region compared with CMV-empty
vector (Fig. 7D). CMV-Foxm1
overexpression efficiency is shown in Fig. S5A. Thus, ATP2B4 is a direct
transcriptional target of FOXM1 in rhabdomyosarcoma cells.
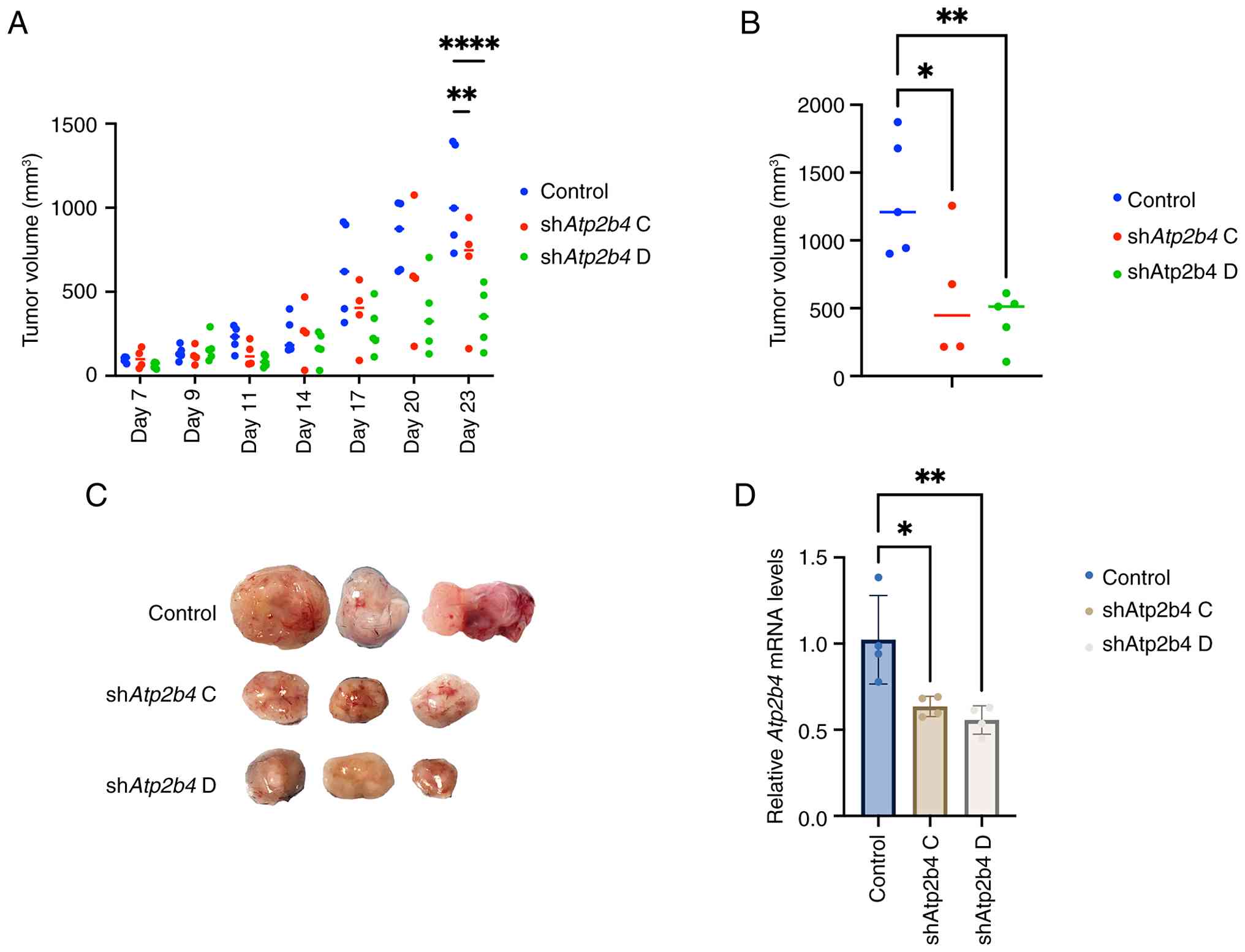
Deletion of ATP2B4 inhibits RMS tumor
growth in an animal model
As the depletion of ATP2B4 sensitized RMS cells to
venetoclax-mediated apoptosis in vitro, the present study
next determined the role of ATP2B4 knockdown in rhabdomyosarcoma
growth in an animal model. 76-9 cells with stable deletion of
Atp2b4 were generated using shAtp2b4. GFP-positive
cells were isolated by FACS, followed by clonal expansion. Knockout
efficiency was evaluated by RT-qPCR, revealing that shAtp2b4
C and shAtp2b4 D achieved the most substantial reduction in
ATP2B4 expression, with knockdown levels of approximately 70-80%.
(Fig. S6A). shAtp2b4 and
control cells were then both subcutaneously injected into C57BL/6J
mice. Deletion of Atp2b4 decreased tumor growth and reduced
final tumor volume compared with controls (Figs. 8A-C and S5B). RT-qPCR analysis of tumor-derived
mRNA confirmed a knockdown efficiency of approximately 50% in the
shAtp2b4 group (Fig. 8D).
Immunostaining demonstrated that the protein level of ATP2B4
decreased in shAtp2b4 harboring tumors (Fig. S6B). Thus, efficient deletion of
ATP2B4 inhibits RMS tumor growth in an animal model of RMS.
Discussion
Cytotoxic chemotherapy has been the cornerstone for
the treatment of localized and metastatic rhabdomyosarcoma. Despite
the heterogeneous genomic landscape of rhabdomyosarcoma, all types
of childhood rhabdomyosarcomas are treated with similar cytotoxic
agents (53). Multiple molecular
targets have been identified as potential therapeutic hits in
rhabdomyosarcoma, but the lack of efficacy or significant toxicity
has halted the progress of these agents in clinical trials
(54,55). Targeted therapies with less
toxicity and better efficacy are an unmet need in
rhabdomyosarcoma.
The present study evaluated a novel combination of
RCM-1 and venetoclax in mouse and human rhabdomyosarcoma cell
lines, as well as a mouse model of RMS. FOXM1 is an oncogene
that induces the transcription of many genes involved in cell cycle
and DNA repair (56,57) However, the present study, to the
best of the authors' knowledge, was the first to investigate the
role of FOXM1 in regulating the expression of the ATP-dependent
plasma membrane calcium channel, ATP2B4. ATP2B is a family
of calcium channels located on the plasma membrane and has a
crucial role in extruding calcium and maintaining intracellular
calcium hemostasis (58). ATP2B
downregulation has been connected to intracellular calcium
overload, cellular stress and death in different cancers (48,58). ATP2B is the predominant pathway in
non-excitable cells and has 4 isoforms (ATP2B 1-4) (58). While ATP2B1 and ATP2B4 are
ubiquitously expressed in tissues, ATP2B2 and ATP2B3 are
predominantly expressed in excitable cells such as neurons
(58). Knockdown of ATP2B1
is lethal in embryos, indicating its crucial role in early embryo
development. However, despite the extensive distribution in
tissues, ATP2B4 ablation is not lethal in embryos. This
suggests that ATP2B4 may have specialized roles in disease
pathogenesis and could serve as a potential therapeutic target in
RMS.
FOXM1 binds the ATP2B4 promoter and
regulates its expression. RCM-1 via FOXM1 inhibition downregulates
ATP2B4 expression and sensitizes RMS cells to venetoclax treatment.
The present study showed that ATP2B4 is overexpressed in
rhabdomyosarcoma of different types compared with normal muscle and
progenitor cells. Moreover, the present study showed that the
knockdown of ATP2B4 in rhabdomyosarcoma inhibits cell
proliferation, colony formation and cell migration as well as tumor
growth in a mouse model. Notably, the knockdown of ATP2B4
enhanced the caspase-mediated apoptosis of venetoclax in
rhabdomyosarcoma. On the other hand, ATP2B4 overexpression
resulted in increased resistance to apoptosis. The synergistic
effect of RCM-1 and venetoclax may, in part, result from
RCM-1-mediated downregulation of ATP2B4, although additional
mechanisms are likely involved (19). The present study demonstrated that
the decrease in ATP2B4 expression resulted in elevated
intracellular calcium levels. Increased intracellular calcium has
been linked to mitochondrial dysfunction, inhibiting Bcl2 and
Bcl-XL while promoting BAK and BAX oligomerization, ultimately
driving cells toward apoptosis (58). The enhanced apoptotic effect of
venetoclax by the downregulation of ATP2B4 is also reported in
breast cancer cells (48).
Moreover, ATP2B4 overexpression confers worse survival outcomes in
pancreatic adenocarcinoma and its inhibition sensitizes tumor cells
to apoptosis (49,59). Notably, the role of ATP2B4 in
cancer pathogenesis differs across types of cancer. For example,
ATP2B4 overexpression suppresses melanoma cell migration and
induces colon and gastric cell differentiation (60-62). Nevertheless, these findings
underscore the role of ATP2B4 in RMS pathobiology, highlighting its
potential as a therapeutic target.
The present study was limited by the short
observation period in the animal model, which may not fully capture
long-term treatment effects. Additionally, patient-derived
xenograft models of different types of RMS may give an improved
reflection of tumor heterogeneity seen in patients compared with
the murine xenografts used in the present study. Finally, future
studies should evaluate the combination of RCM-1 and venetocalx
across different types of rhabdomyosarcoma and in patient-derived
xenograft models to facilitate its transition into early-phase
clinical trials. Additional studies are warranted to identify
biomarkers that may guide precision treatment and help identify
patients who are most likely to benefit from this combination.
In conclusion, RCM-1 sensitizes rhabdomyosarcoma to
venetoclax-induced apoptosis by inhibiting FOXM1 and downregulating
ATP2B4 expression. These findings highlight the potential
therapeutic benefits of combining RCM-1 with venetoclax through the
modulation of the calcium signaling pathway. This underscores the
need for further investigation into this novel approach in
rhabdomyosarcoma and supports exploring the calcium signaling
pathway as a therapeutic strategy in rhabdomyosarcoma.
Supplementary Data
Availability of data and materials
The datasets generated during and/or analyzed
during the current study are available in the NCBI Sequence Read
Archive repository (SRA) under accession number: PRJNA1204196
(https://dataview.ncbi.nlm.nih.gov/object/PRJNA1204196?reviewer=mfdqbt192mum79u708ccn1r6od).
Authors' contributions
NM and TVK conceived and designed the study. NM and
YWL performed the in vivo experiments. ZD designed and
synthesized the nanoparticle. NM, YWL, XX, TJ and JDon participated
in vitro and culture experiments. YWL and GW designed and
performed the RNA-seq bioinformatics analysis. YWL and JDo designed
and performed the dual luciferase analysis. NM and YWL analyzed
data. NM, VVK and TVK interpreted the data. TVK and VVK provided
critical reagents and intellectual discussions. NM and TVK wrote
the paper. All authors discussed the data. NM and YWL confirmed the
authenticity of all the raw data. TVK approved the submission of
the manuscript. All authors read and approved the final
manuscript.
Ethics approval and consent to
participate
All animal studies were approved by the University
of Arizona Institutional Animal Care and Use Committee and covered
under animal protocol approval no. IACUC2023-1128).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Acknowledgements
The call sets were downloaded from the ENCODE
portal (63) (https://www.encodeproject.org/) with the
following identifiers: ENCFF162QJJ, ENCFF660WUG and
ENCFF139KZL.
Funding
The present study was supported by a CTI Research Grant,
CancerFree KIDS to NM, Hyundai Hope on Wheels, Young Investigator
Award to NM, NHLBI grant no. R01HL158659 to TVK and NHLBI grant no.
R01HL141174 to VVK.
References
|
1
|
Martin-Giacalone BA, Weinstein PA, Plon SE
and Lupo PJ: Pediatric rhabdomyosarcoma: Epidemiology and genetic
susceptibility. J Clin Med. 10:20282021. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
McEvoy MT, Siegel DA, Dai S, Okcu MF,
Zobeck M, Venkatramani R and Lupo PJ: Pediatric rhabdomyosarcoma
incidence and survival in the United States: An assessment of 5656
cases, 2001-2017. Cancer Med. 12:3644–3656. 2023. View Article : Google Scholar :
|
|
3
|
Oberlin O, Rey A, Lyden E, Bisogno G,
Stevens MCG, Meyer WH, Carli M and Anderson JR: Prognostic factors
in metastatic rhabdomyosarcomas: Results of a pooled analysis from
United States and European cooperative groups. J Clin Oncol.
26:2384–2389. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Weigel BJ, Lyden E, Anderson JR, Meyer WH,
Parham DM, Rodeberg DA, Michalski JM, Hawkins DS and Arndt CA:
Intensive multiagent therapy, including dose-compressed cycles of
ifosfamide/etoposide and vincristine/doxorubicin/cyclophosphamide,
irinotecan, and radiation, in patients with high-risk
rhabdomyosarcoma: A report from the children's oncology group. J
Clin Oncol. 34:117–122. 2016. View Article : Google Scholar :
|
|
5
|
Clark KL, Halay ED, Lai E and Burley SK:
Co-crystal structure of the HNF-3/fork head DNA-recognition motif
resembles histone H5. Nature. 364:412–420. 1993. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Kalin TV, Ustiyan V and Kalinichenko VV:
Multiple faces of FoxM1 transcription factor: Lessons from
transgenic mouse models. Cell Cycle. 10:396–405. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Milewski D, Balli D, Ustiyan V, Le T,
Dienemann H, Warth A, Breuhahn K, Whitsett JA, Kalinichenko VV and
Kalin TV: FOXM1 activates AGR2 and causes progression of lung
adenomas into invasive mucinous adenocarcinomas. PLoS Genet.
13:e10070972017. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Weiler SME, Pinna F, Wolf T, Lutz T,
Geldiyev A, Sticht C, Knaub M, Thomann S, Bissinger M, Wan S, et
al: Induction of chromosome instability by activation of
yes-associated protein and forkhead box M1 in liver cancer.
Gastroenterology. 152:2037–2051.e22. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Cheng XH, Black M, Ustiyan V, Le T,
Fulford L, Sridharan A, Medvedovic M, Kalinichenko VV, Whitsett JA
and Kalin TV: SPDEF inhibits prostate carcinogenesis by disrupting
a positive feedback loop in regulation of the Foxm1 oncogene. PLoS
Genet. 10:e10046562014. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Wang IC, Ustiyan V, Zhang Y, Cai Y, Kalin
TV and Kalinichenko VV: Foxm1 transcription factor is required for
the initiation of lung tumorigenesis by oncogenic Kras(G12D.).
Oncogene. 33:5391–5396. 2014. View Article : Google Scholar
|
|
11
|
Cai Y, Balli D, Ustiyan V, Fulford L,
Hiller A, Misetic V, Zhang Y, Paluch AM, Waltz SE, Kasper S and
Kalin TV: Foxm1 expression in prostate epithelial cells is
essential for prostate carcinogenesis. J Biol Chem.
288:22527–22541. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Balli D, Ren X, Chou FS, Cross E, Zhang Y,
Kalinichenko VV and Kalin TV: Foxm1 transcription factor is
required for macrophage migration during lung inflammation and
tumor formation. Oncogene. 31:3875–3888. 2012. View Article : Google Scholar
|
|
13
|
Li L, Wu D, Yu Q, Li L and Wu P:
Prognostic value of FOXM1 in solid tumors: A systematic review and
meta-analysis. Oncotarget. 8:32298–32308. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Kuda M, Kohashi K, Yamada Y, Maekawa A,
Kinoshita Y, Nakatsura T, Iwamoto Y, Taguchi T and Oda Y: FOXM1
expression in rhabdomyosarcoma: A novel prognostic factor and
therapeutic target. Tumour Biol. 37:5213–5223. 2016. View Article : Google Scholar
|
|
15
|
Nestal de Moraes G, Bella L, Zona S,
Burton MJ and Lam EWF: Insights into a critical role of the
FOXO3a-FOXM1 axis in DNA damage response and genotoxic drug
resistance. Curr Drug Targets. 17:164–177. 2016. View Article : Google Scholar
|
|
16
|
Merjaneh N, Hajjar M, Lan YW, Kalinichenko
VV and Kalin TV: The promise of combination therapies with FOXM1
inhibitors for cancer treatment. Cancers (Basel). 16:7562024.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Ustiyan V, Zhang Y, Perl AKT, Whitsett JA,
Kalin TV and Kalinichenko VV: β-catenin and Kras/Foxm1 signaling
pathway are critical to restrict Sox9 in basal cells during
pulmonary branching morphogenesis. Dev Dyn. 245:590–604. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Sun L, Ren X, Wang IC, Pradhan A, Zhang Y,
Flood HM, Han B, Whitsett JA, Kalin TV and Kalinichenko VV: The
FOXM1 inhibitor RCM-1 suppresses goblet cell metaplasia and
prevents IL-13 and STAT6 signaling in allergen-exposed mice. Sci
Signal. 10:eaai85832017. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Shukla S, Milewski D, Pradhan A, Rama N,
Rice K, Le T, Flick MJ, Vaz S, Zhao X, Setchell KD, et al: The
FOXM1 inhibitor RCM-1 decreases carcinogenesis and nuclear
β-catenin. Mol Cancer Ther. 18:1217–1229. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Donovan J, Deng Z, Bian F, Shukla S,
Gomez-Arroyo J, Shi D, Kalinichenko VV and Kalin TV: Improving
anti-tumor efficacy of low-dose vincristine in rhabdomyosarcoma via
the combination therapy with FOXM1 inhibitor RCM1. Front Oncol.
13:11128592023. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Fairlie WD and Lee EF: Targeting the
BCL-2-regulated apoptotic pathway for the treatment of solid
cancers. Biochem Soc Trans. 49:2397–2410. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Steinert DM, Salganick J, Ballo M, Zhang
W, Munsell M, Raney B, Jaffe N, Koh J, El-Naggar A and Trent J:
Expression of Bax and Bcl-2 in human rhabdomyosarcoma: Correlation
with survival in 64 patients. J Clin Oncol. 23(Suppl 16):
S90412005. View Article : Google Scholar
|
|
23
|
Armistead PM, Salganick J, Roh JS,
Steinert DM, Patel S, Munsell M, El-Naggar AK, Benjamin RS, Zhang W
and Trent JC: Expression of receptor tyrosine kinases and apoptotic
molecules in rhabdomyosarcoma: Correlation with overall survival in
105 patients. Cancer. 110:2293–2303. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Ploumaki I, Triantafyllou E,
Koumprentziotis IA, Karampinos K, Drougkas K, Karavolias I,
Trontzas I and Kotteas EA: Bcl-2 pathway inhibition in solid
tumors: A review of clinical trials. Clin Transl Oncol.
25:1554–1578. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Heinicke U, Haydn T, Kehr S, Vogler M and
Fulda S: BCL-2 selective inhibitor ABT-199 primes rhabdomyosarcoma
cells to histone deacetylase inhibitor-induced apoptosis. Oncogene.
37:5325–5339. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Alcon C, Manzano-Muñoz A, Prada E, Mora J,
Soriano A, Guillén G, Gallego S, Roma J, Samitier J, Villanueva A
and Montero J: Sequential combinations of chemotherapeutic agents
with BH3 mimetics to treat rhabdomyosarcoma and avoid resistance.
Cell Death Dis. 11:6342020. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Alcon C, Martín F, Prada E, Mora J,
Soriano A, Guillén G, Gallego S, Roma J, Samitier J, Villanueva A
and Montero J: MEK and MCL-1 sequential inhibition synergize to
enhance rhabdomyosarcoma treatment. Cell Death Discov. 8:1722022.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Goldsmith KC, Verschuur A, Morgenstern DA,
van Eijkelenburg N, Federico SM, Fraser C, Forlenza CJ, Ziegler DS,
Gerber NU, Khaw AL, et al: The first report of pediatric patients
with solid tumors treated with venetoclax. J Clin Oncol. 38(15
Suppl): S105242020. View Article : Google Scholar
|
|
29
|
Hinson ARP, Jones R, Crose LES, Belyea BC,
Barr FG and Linardic CM: Human rhabdomyosarcoma cell lines for
rhabdomyosarcoma research: Utility and pitfalls. Front Oncol.
3:1832013. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Leddon JL, Chen CY, Currier MA, Wang PY,
Jung FA, Denton NL, Cripe KM, Haworth KB, Arnold MA, Gross AC, et
al: Oncolytic HSV virotherapy in murine sarcomas differentially
triggers an antitumor T-cell response in the absence of virus
permissivity. Mol Ther Oncolytics. 1:140102015. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Milewski D, Pradhan A, Wang X, Cai Y, Le
T, Turpin B, Kalinichenko VV and Kalin TV: FoxF1 and FoxF2
transcription factors synergistically promote rhabdomyosarcoma
carcinogenesis by repressing transcription of p21Cip1
CDK inhibitor. Oncogene. 36:850–862. 2017. View Article : Google Scholar :
|
|
32
|
Sun F, Wang G, Pradhan A, Xu K,
Gomez-Arroyo J, Zhang Y, Kalin GT, Deng Z, Vagnozzi RJ, He H, et
al: Nanoparticle delivery of STAT3 alleviates pulmonary
hypertension in a mouse model of alveolar capillary dysplasia.
Circulation. 144:539–555. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Deng Z, Lin J, Bud'ko SL, Webster B, Kalin
TV, Kalinichenko VV and Shi D: Dual targeting with cell surface
electrical charge and folic acid via superparamagnetic
Fe3O4@Cu2-xS for photothermal
cancer cell killing. Cancers (Basel). 13:52752021. View Article : Google Scholar
|
|
34
|
Deng Z, Gao W, Kohram F, Li E, Kalin TV,
Shi D and Kalinichenko VV: Fluorinated amphiphilic Poly(β-Amino
ester) nanoparticle for highly efficient and specific delivery of
nucleic acids to the Lung capillary endothelium. Bioact Mater.
31:1–17. 2023.
|
|
35
|
Milewski D, Shukla S, Gryder BE, Pradhan
A, Donovan J, Sudha P, Vallabh S, Pyros A, Xu Y, Barski A, et al:
FOXF1 is required for the oncogenic properties of PAX3-FOXO1 in
rhabdomyosarcoma. Oncogene. 40:2182–2199. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Singh S, Uppuluri P, Mamouei Z, Alqarihi
A, Elhassan H, French S, Lockhart SR, Chiller T, Edwards JE Jr and
Ibrahim AS: The NDV-3A vaccine protects mice from multidrug
resistant Candida auris infection. PLoS Pathog. 15:e10074602019.
View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Li Z, Wang H, Li S, Deng B, Zhang W, Wu P,
Li W, Xin P, Zhao L and Gao S: Anti-pseudomonas aeruginosa activity
of the scorpion-derived peptide GK8. Probiotics Antimicrob
Proteins. Jul 3–2025.Epub ahead of print.
|
|
38
|
Acharya A, Bian F, Gomez-Arroyo J, Wagner
KA, Kalinichenko VV and Kalin TV: Hypoxia represses FOXF1 in lung
endothelial cells through HIF-1α. Front Physiol. 14:13091552024.
View Article : Google Scholar
|
|
39
|
Shukla S, Saha T, Rama N, Acharya A, Le T,
Bian F, Donovan J, Tan LA, Vatner R, Kalinichenko V, et al:
Ultra-high dose-rate proton FLASH improves tumor control. Radiother
Oncol. 186:1097412023. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
41
|
Love MI, Huber W and Anders S: Moderated
estimation of fold change and dispersion for RNA-seq data with
DESeq2. Genome Biol. 15:5502014. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Emig D, Salomonis N, Baumbach J, Lengauer
T, Conklin BR and Albrecht M: AltAnalyze and DomainGraph: Analyzing
and visualizing exon expression data. Nucleic Acids Res.
38:W755–W762. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Chen J, Bardes EE, Aronow BJ and Jegga AG:
ToppGene Suite for gene list enrichment analysis and candidate gene
prioritization. Nucleic Acids Res. 37:W305–W311. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Tang D, Chen M, Huang X, Zhang G, Zeng L,
Zhang G, Wu S and Wang Y: SRplot: A free online platform for data
visualization and graphing. PLoS One. 18:e02942362023. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
De Micheli AJ, Spector JA, Elemento O and
Cosgrove BD: A reference single-cell transcriptomic atlas of human
skeletal muscle tissue reveals bifurcated muscle stem cell
populations. Skelet Muscle. 10:192020. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Wei Y, Qin Q, Yan C, Hayes MN, Garcia SP,
Xi H, Do D, Jin AH, Eng TC, McCarthy KM, et al: Single-cell
analysis and functional characterization uncover the stem cell
hierarchies and developmental origins of rhabdomyosarcoma. Nat
Cancer. 3:961–975. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Dalton KM, Krytska K, Lochmann TL, Sano R,
Casey C, D'Aulerio A, Khan QA, Crowther GS, Coon C, Cai J, et al:
Venetoclax-based rational combinations are effective in models of
MYCN-amplified neuroblastoma. Mol Cancer Ther. 20:1400–1411. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Curry MC, Luk NA, Kenny PA,
Roberts-Thomson SJ and Monteith GR: Distinct regulation of
cytoplasmic calcium signals and cell death pathways by different
plasma membrane calcium ATPase isoforms in MDA-MB-231 breast cancer
cells. J Biol Chem. 287:28598–28608. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Sritangos P, Pena Alarcon E, James AD,
Sultan A, Richardson DA and Bruce JIE: Plasma membrane
Ca2+ ATPase isoform 4 (PMCA4) has an important role in
numerous hallmarks of pancreatic cancer. Cancers (Basel).
12:2182020. View Article : Google Scholar
|
|
50
|
Naffa R, Hegedűs L, Hegedűs T, Tóth S,
Papp B, Tordai A and Enyedi Á: Plasma membrane Ca2+ pump
isoform 4 function in cell migration and cancer metastasis. J
Physiol. 602:1551–1564. 2024. View Article : Google Scholar
|
|
51
|
ENCODE Project Consortium: An integrated
encyclopedia of DNA elements in the human genome. Nature.
489:57–74. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Zhang J, Liu J, Lee D, Lou S, Chen Z,
Gürsoy G and Gerstein M: DiNeR: A differential graphical model for
analysis of co-regulation network rewiring. BMC Bioinformatics.
21:2812020. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Ruymann FB: The development of VAC
chemotherapy in rhabdomyosarcoma: What does one do for an encore?
Curr Oncol Rep. 5:505–509. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Nguyen TH and Barr FG: Therapeutic
approaches targeting PAX3-FOXO1 and its regulatory and
transcriptional pathways in rhabdomyosarcoma. Molecules.
23:27982018. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Chen C, Dorado Garcia H, Scheer M and
Henssen AG: Current and future treatment strategies for
rhabdomyosarcoma. Front Oncol. 9:14582019. View Article : Google Scholar
|
|
56
|
Kalinichenko VV and Kalin TV: Is there
potential to target FOXM1 for 'undruggable' lung cancers? Expert
Opin Ther Targets. 19:865–867. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Black M, Arumugam P, Shukla S, Pradhan A,
Ustiyan V, Milewski D, Kalinichenko VV and Kalin TV: FOXM1 nuclear
transcription factor translocates into mitochondria and inhibits
oxidative phosphorylation. Mol Biol Cell. 31:1411–1424. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Stafford N, Wilson C, Oceandy D, Neyses L
and Cartwright EJ: The plasma membrane calcium ATPases and their
role as major new players in human disease. Physiol Rev.
97:1089–1125. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
James AD, Chan A, Erice O, Siriwardena AK
and Bruce JIE: Glycolytic ATP fuels the plasma membrane calcium
pump critical for pancreatic cancer cell survival. J Biol Chem.
288:36007–36019. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Hegedũs L, Garay T, Molnár E, Varga K,
Bilecz Á, Török S, Padányi R, Pászty K, Wolf M, Grusch M, et al:
The plasma membrane Ca2+ pump PMCA4b inhibits the
migratory and metastatic activity of BRAF mutant melanoma cells.
Int J Cancer. 140:2758–2770. 2017. View Article : Google Scholar
|
|
61
|
Ribiczey P, Tordai A, Andrikovics H,
Filoteo AG, Penniston JT, Enouf J, Enyedi A, Papp B and Kovács T:
Isoform-specific up-regulation of plasma membrane Ca2+ATPase
expression during colon and gastric cancer cell differentiation.
Cell Calcium. 42:590–605. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Aung C, Kruger W, Poronnik P,
Roberts-Thomson S and Monteith G: Plasma membrane Ca2+-ATPase
expression during colon cancer cell line differentiation. Biochem
Biophys Res Commun. 355:932–936. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Sloan CA, Chan ET, Davidson JM, Malladi
VS, Strattan JS, Hitz BC, Gabdank I, Narayanan AK, Ho M, Lee BT, et
al: ENCODE data at the ENCODE portal. Nucleic Acids Res.
44:D726–D732. 2016. View Article : Google Scholar :
|