Introduction
Hepatocellular carcinoma (HCC) has become the third
most fatal type of neoplasm worldwide (1). An improved understanding of its
molecular mechanisms may assist in identifying novel targets for
therapeutics.
Ubiquitin specific peptidase 7 (UPS7), also known as
the herpes simplex virus associated ubiquitin-specific protease,
was initially isolated as a binding partner of the herpes simplex
virus protein Vmw110/infected cell polypeptide 0 (2). Subsequent studies demonstrated that
USP7 has critical roles in tumor development and progression
(3). Downregulation of its
expression usually contributes to oncogenic transformation and is
crucial for tumor cell proliferation (4,5). At
the molecular level, USP7 has been identified as a key regulator of
the p53 signaling pathway, through stabilizing p53 protein and
preventing its degradation (6). In
this regard, USP7 may act as a tumor suppressor and a deficiency in
USP7 may result in cell proliferation and an increase in genomic
instability leading to mutagenesis (2,7).
However, the molecular determinants of USP7 expression in human
cancer remain to be elucidated.
MicroRNAs (miRNAs) are a class of small non-coding
RNA molecules, which repress gene expression through translational
repression or degradation (8,9). In
human cancer, increasing lines of evidence have indicated that
miRNAs have important roles in tumor cell proliferation, metastasis
and angiogenesis, through regulation of multiple signaling
pathways, including the p53, Wnt/β-Catenin and nuclear factor
(NF)-κB signaling pathways (10–12).
Therefore, the aim of the present study was to investigate whether
USP7 expression could be controlled by miRNAs in HCC.
Materials and methods
Human tissue samples
HCC tissues and adjacent non-tumor normal tissues
were collected from patients undergoing routine therapeutic surgery
at the Department of Interventional Radiology, Zhongshan Hospital
(Shanghai, China), between May 2009 and July 2011. All samples were
obtained with informed consent and approved by the Zhongshan
Hospital Institutional Review Board.
Cell culture
HepG2 cells derived from HCC were obtained from the
Cell Bank of Type Culture Collection of the Chinese Academy of
Sciences (Shanghai, China). Cells were grown in Dulbecco's modified
Eagle's medium (DMEM, Gibco-BRL, Shanghai, China) supplemented with
10% fetal bovine serum (Gibco-BRL) and maintained at 37°C in a
humidified atmosphere with 5% CO2.
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
Total RNA from tissues and cells was extracted using
the miRNA isolation kit (Ambion, Austin, TX, USA) according to the
manufacturer's instructions. cDNA synthesis was performed for each
RNA sample using a Reverse Transcription system (Promega
Corporation, Madison, WI, USA), and oligo dT from the system was
used to prime cDNA synthesis. Expression levels of mature miRNAs
were assayed using a Taqman MicroRNA assay (Applied Biosystems,
Foster City, CA, USA). RT-qPCR was performed using an Applied
Biosystems 7300 Real-time PCR system and a TaqMan Universal PCR
master mix (Applied Biosystems). PCR conditions included an initial
hold period at 95°C for 5 min, followed by a two-step PCR program
consisting of 95°C for 5 sec and 60°C fo 30 sec for 40 cycles. The
primer sequences were as follows: p21 forward,
5′-TGTCCGTCAGAACCCATGC-3′, reverse, 5′-CCAGCCCATGATGGTTCTGAT-3′;
Bax forward, 5′-CCCGAGAGGTCTTTTTCCGAG-3′, reverse,
5′-CCAGCCCATGATGGTTCTGAT-3′; and GADD45 forward,
5′-GAGAGCAGAAGACCGAAAGGA-3′ and reverse,
5′-CACAACACCACGTTATCGGG-3′. All the primers were obtained from
Bioyare Company (Shanghai, China). Data were collected and
quantitatively analyzed accoring to the 2−ΔΔCt method.
Expression of the miRNAs was normalized to that of the U6
spliceosomal RNA.
Western blot analysis
Cells were harvested and lysed with ice-cold lysis
buffer (50 mM Tris-HCl, pH 6.8; 100 mM 2-mercaptoethanol, 2% w/v
SDS and 10% glycerol). Following centrifugation at 10,000 × g for
15 min at 4°C, proteins in the supernatants were quantified and
separated using 10% SDS-PAGE. Western blot analysis was performed
using the following rabbit antibodies: Anti-USP7 (1:1,000; cat. no.
3277) and p53 (1:1,000; cat. no. 9282) (polyclonal), and p21
(1:2,000; cat. no. 2947), Bax (1:1,000; cat. no. 5023) p27
(1:2,000; cat. no. 3688) and growth arrest and DNA damage 45
(GADD45; 1:1,000; cat. no. 4632) (monoclonal) (Cell Signaling
Technology, Inc., Danvers, MA, USA). The secondary antibodies
(1:5000, goat anti-rabbit IgG-HRP; sc-2004) were purchased from
Santa Cruz Biotechnology, Inc. (Santa Cruz, CA, USA). Protein
levels were normal-ized to total GAPDH, using a rabbit anti-GAPDH
antibody (1:5,000; cat. no. H-83) (Santa Cruz Biotechnology, Inc.).
The blots were visualized using chemiluminescence reagents
(Amersham Pharmacia, Little Chalfont, UK).
Transfection and luciferase reporter
assay
miR-205 mimics and antisense oligonucleotides were
purchased from Ambion (Invitrogen Life Technologies, Carlsbad, CA,
USA). The human USP7 3′UTR was cloned into the pMir-Report miRNA
expression reporter vector system (Ambion), yielding
pMir-Report-USP7. Mutations were introduced in potential miR-205
binding sites using the QuikChange site-directed mutagenesis kit
(Stratagene, Santa Clara, CA, USA). Transfection was performed with
Lipofectamine 2000 (Invitrogen Life Technologies) according to the
manufacturer's instructions. The Renilla luciferase control
reporter vector pRL-TK (Promega Corporation) carrying the
Renilla luciferase gene was used to normalize the
transfection efficiency. Luciferase values were measured using the
Dual-Luciferase reporter assay system (Promega Corporation).
Bromodeoxyuridine (BrdU) assays
A cell proliferation enzyme-linked immunosorbent
assay (BrdU kit; Beyotime Institute of Biotechnology, Haimen,
China) was used to analyze the incorporation of BrdU during DNA
synthesis following the manufacturer's instructions. Absorbance was
measured at 450 nm in the Spectra Max 190 ELISA reader (Molecular
Devices, Sunnyvale, CA, USA). All experiments were performed in
triplicate.
Statistical analysis
The data shown are presented as the mean ± standard
error of the mean of three independent experiments. Statistical
analyses were conducted using GraphPad version 5.0 (GraphPad
Software, Inc., La Jolla, CA, USA). Significance was analyzed using
Student's t-test. P<0.05, P<0.01 and P<0.001 were
considered to indicate a statistically significant difference.
Results
miR-205 reduces USP7 protein expression
in HepG2 cells
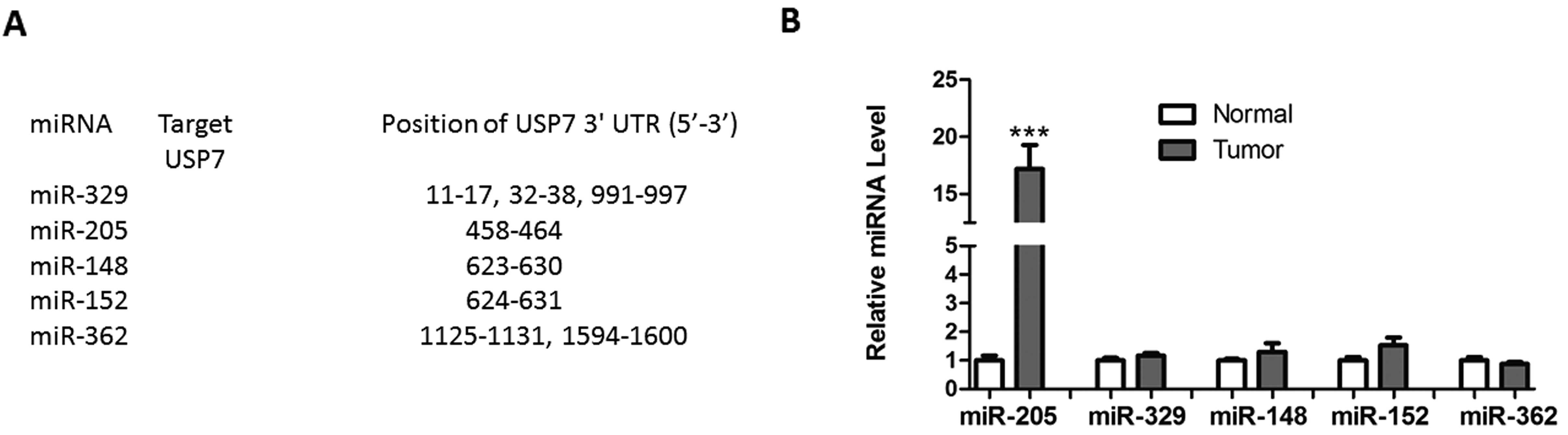
Using the TargetScan algorithm based on seed
recognition, several miRNAs were identified, which may potentially
interact with the USP7 transcript (Fig. 1A). However, only miR-205 was
significantly upregulated in HCC tissues, compared with normal
tissues (Fig. 1B). To assess
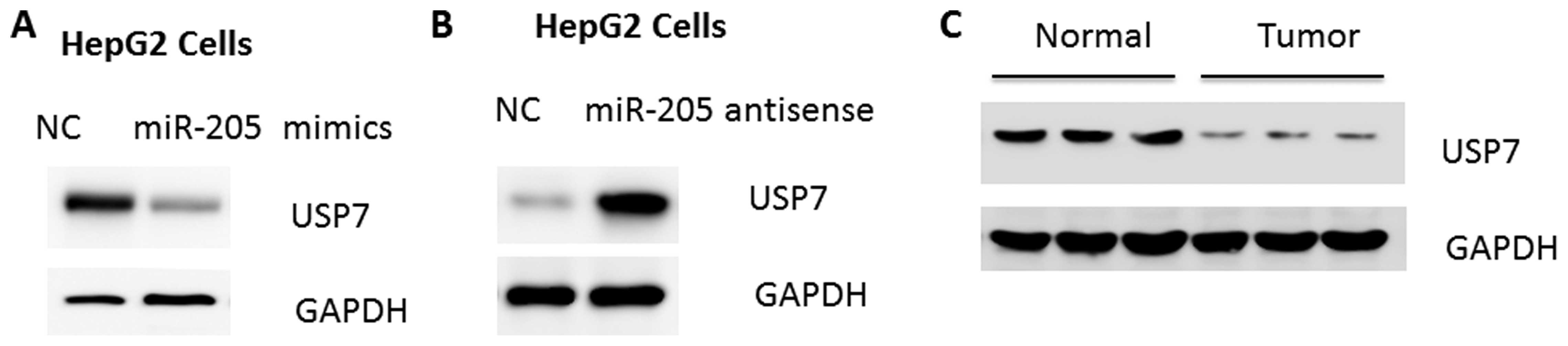
whether miR-205 regulates USP7, miR-205 mimic or antisense were
introduced into HepG2 human hepatoma cells. As a result, expression
of miR-205 and anti-miR-205 reduced and increased the quantity of
USP7 protein, respectively (Fig. 2A
and B). Concurrent with the upregulation of miR-205, a marked
reduction of USP7 protein levels was observed in HCC tissues
(Fig. 2C), further suggesting that
USP7 may be a target of miR-205 in HCC development.
miR-205 interacts with the 3′UTR of
USP7
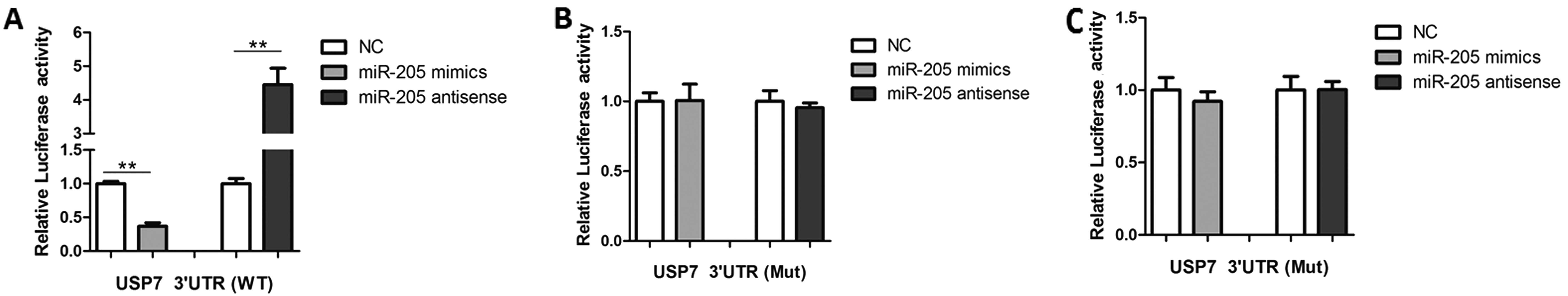
To understand how miR-205 regulates USP7 expression,
the luciferase reporter plasmids containing the 3′UTR of USP7 were
co-transfected with miR-205 mimics or antisense oligonucleotides.
As shown in Fig. 3A, miR-205
mimics led to a reduction and antisense led to an increase in
luciferase activity. Furthermore, mutagen-esis of the seed sequence
eliminated the effects of miR-205 mimics or antisense
oligonucleotides on USP7 activity (Fig. 3B). When co-expressed with a plasmid
encoding green fluorescent protein-tagged human USP7 lacking the
3′UTR, neither miR-205 nor anti-miR-205 had an effect on USP7
activity (Fig. 3C). These results
suggest that miR-205 binds to the 3′UTR of USP7 and reduces its
protein contents.
miR-205 regulates the p53 signaling
pathway in HepG2 cells
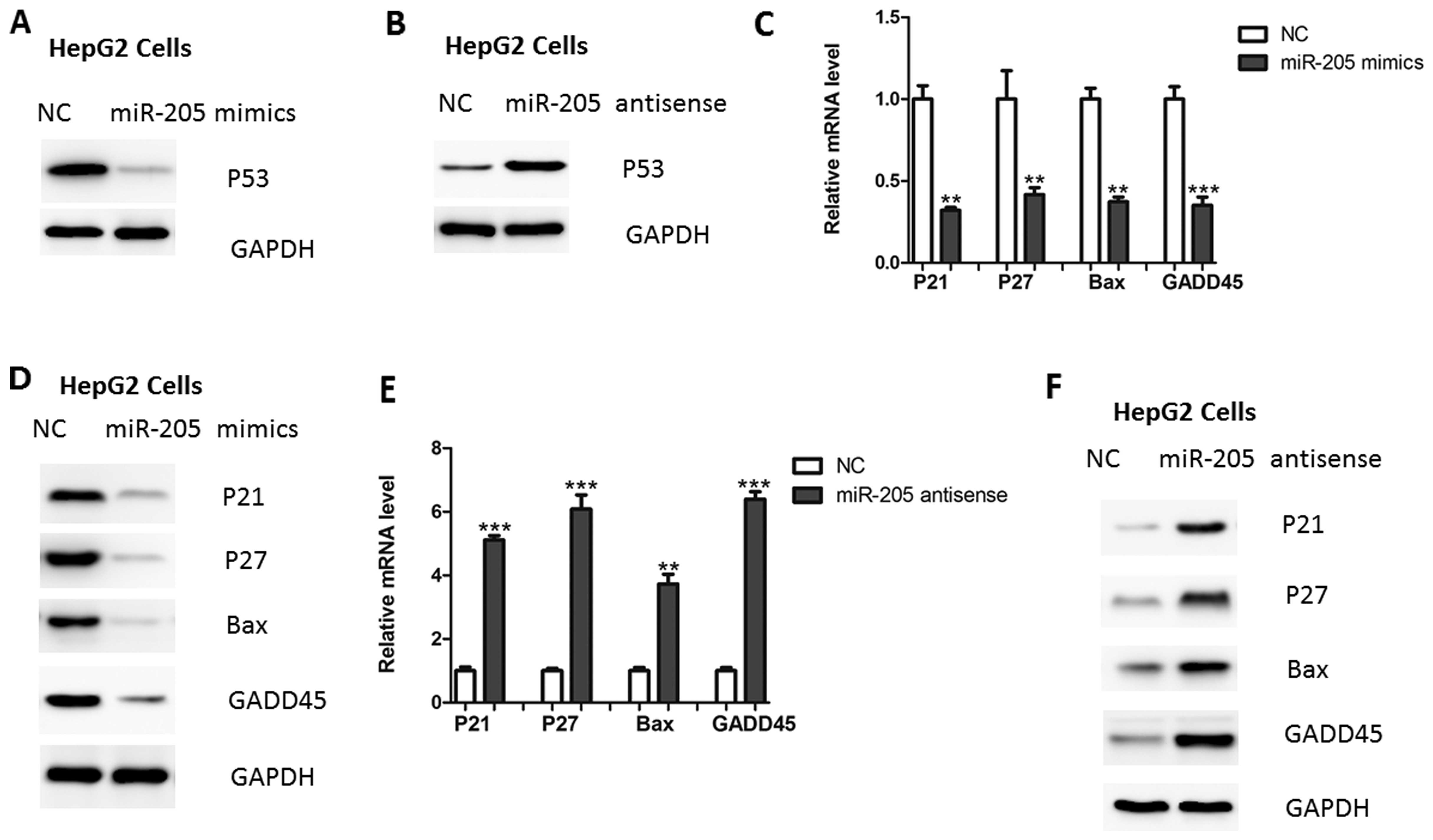
Due to the protective role of USP7 in p53 protein
stability (6), it was determined
whether miR-205 may affect the p53 stability. As shown in Fig. 4A, overexpression of miR-205 led to
reduced p53 protein expression in HepG2 cells. Consistently,
inhibition of miR-205 led to an increased expression of p53
(Fig. 4B). In addition, the
downstream targets of p53 signaling, including p21, p27 and GADD45,
were also regulated by miR-205 over-expression or inhibition
(Fig. 4C–F).
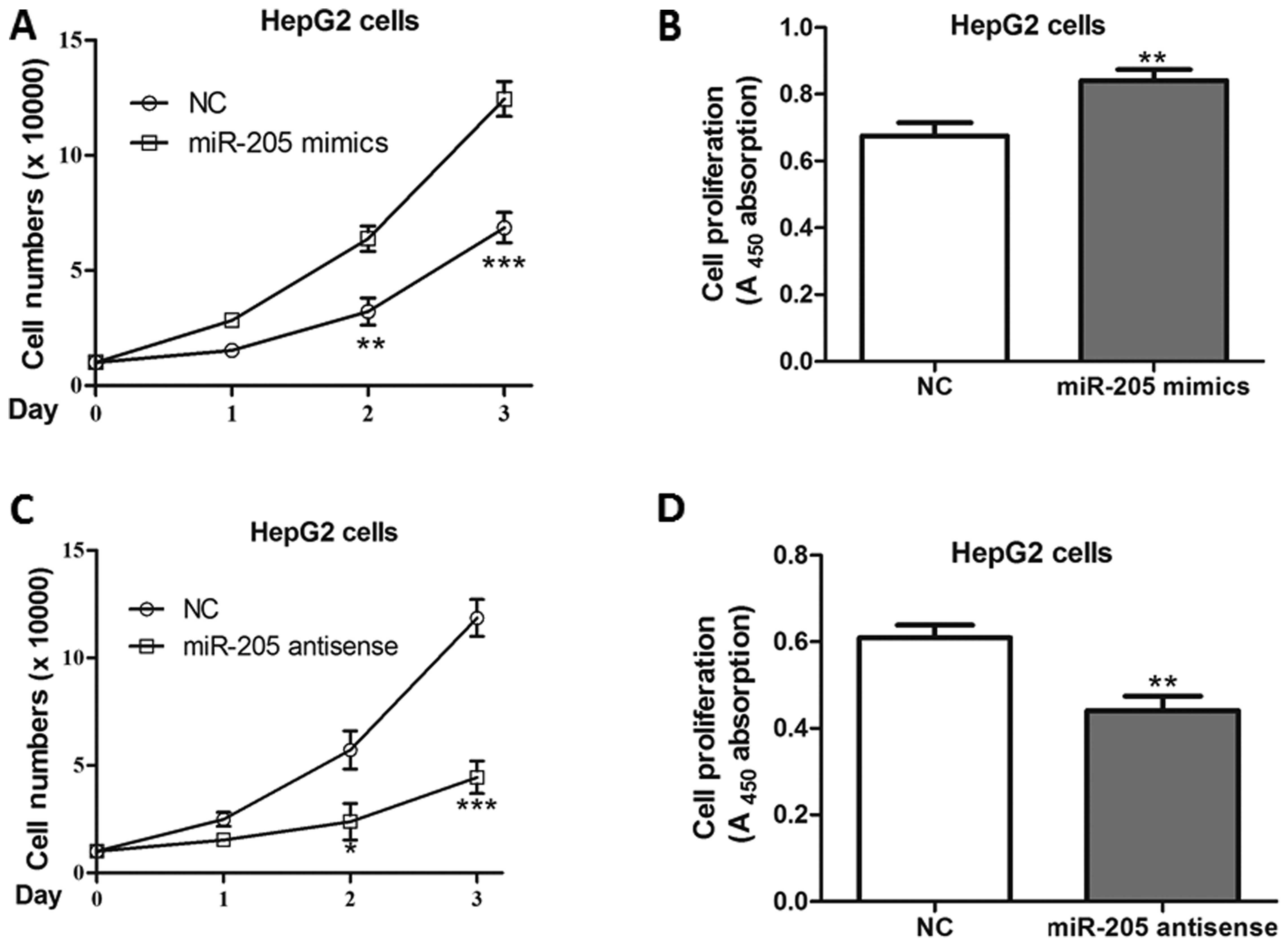
miR-205 positively promotes cell
proliferation
As the critical roles of the USP7-p53 regulatory
axis is involved in cell proliferation, the effects of miR-205 on
HepG2 cell growth were assessed. As a result, miR-205 mimics
significantly increased cell numbers and promoted proliferation in
cells post-transfection (Fig. 5A and
B). Consistently, its antisense inhibited the growth of HepG2,
compared with negative control-transfected cells (Fig. 5C and D).
Discussion
In the present study, it was demonstrated that
miR-205 is upregulated in HCC tissues. In addition, miR-205 is able
to inhibit cell proliferation in HepG2 cells, through regulation of
USP7 protein levels. Therefore, for the first time, to the best of
our knowledge, the present study identified that miR-205 may be an
onco-microRNA involved in the progression of HCC. However, further
studies are required to investigate its role in vivo.
It has been demonstrated that several miRNAs were
dysregulated in HCC tissues or cell lines (13,14).
For instance, an in-depth analysis of miRNomes in human HCC and
normal liver has been performed, which identified that
miR-199a/b-3p, the third most highly expressed miRNA in the liver,
was consistently decreased in HCC (15). In addition, miR-199a/b-3p may
target tumor-promoting p21 protein (Cdc42/Rac)-activated kinase 4
(PAK4) to suppress HCC growth through inhibiting the
PAK4/Raf/mitogen-activated protein kinase kinase/extracellular
signal-regulated kinase pathway in vitro and in vivo
(15). Therefore, identification
of more dysregulated miRNAs may aid in improving understanding of
the important deregulated miRNAs in HCC.
The roles of miR-205 have been elucidated in other
types of human cancer. miR-205 promotes tumor proliferation and
invasion through targeting estrogen-related receptor-γ in
endometrial carcinoma (16). In
addition, miR-205 targets phosphatase and tensin homolog and PH
domain and leucine rich repeat protein phosphatase 2 to augment
protein kinase B signaling and drive malignant phenotypes in
non-small cell lung cancer (17).
However, miR-205 is frequently down-regulated in prostate cancer
and acts as a tumor suppressor by inhibiting tumor growth (18). Although the reason for the
inconsistency remains to be elucidated at present, the effects of
miR-205 may be cell- or tissue-specific.
Previous studies have demonstrated that USP7 is
regulated by several signals, such as interleukin-6-mediated signal
transducer and activator of transcription 3 activation (19,20).
However, whether its expression is controlled by miRNAs remains to
be examined. Therefore, the present data on the functional
interaction of miR-205 and USP7/p53 signaling in HCC may assist in
elucidating mechanisms underlying tumorigenesis in HCC.
References
|
1
|
Villanueva A, Hernandez-Gea V and Llovet
JM: Medical therapies for hepatocellular carcinoma: a critical view
of the evidence. Nat Rev Gastroenterol Hepatol. 10:34–42. 2013.
View Article : Google Scholar
|
|
2
|
Qing P, Han L, Bin L, Yan L and Ping WX:
USP7 regulates the stability and function of HLTF through
deubiquitination. J Cell Biochem. 112:3856–3862. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Nicholson B and Suresh Kumar KG: The
multifaceted roles of USP7: new therapeutic opportunities. Cell
Biochem Biophys. 60:61–68. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Lee JT and Gu W: The multiple levels of
regulation by p53 ubiquitination. Cell Death Differ. 17:86–92.
2010. View Article : Google Scholar
|
|
5
|
Colland F: The therapeutic potential of
deubiquitinating enzyme inhibitors. Biochem Soc Trans. 38:137–143.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Lee MH and Lozano G: Regulation of the
p53-MDM2 pathway by 14-3-3 sigma and other proteins. Semin Cancer
Biol. 16:225–234. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Meulmeester E, Maurice MM, Boutell C, et
al: Loss of HAUSP-mediated deubiquitination contributes to DNA
damage-induced destabilization of Hdmx and Hdm2. Mol Cell.
18:565–576. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Sun K and Lai EC: Adult-specific functions
of animal microRNAs. Nat Rev Genet. 14:535–548. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Ameres SL and Zamore PD: Diversifying
microRNA sequence and function. Nat Rev Mol Cell Biol. 14:475–488.
2013. View
Article : Google Scholar : PubMed/NCBI
|
|
10
|
Liu Y, Xing R, Zhang X, et al: miR-375
targets the p53 gene to regulate cellular response to ionizing
radiation and etoposide in gastric cancer cells. DNA Repair.
12:741–750. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Nagano H, Tomimaru Y, Eguchi H, et al:
MicroRNA-29a induces resistance to gemcitabine through the
Wnt/beta-catenin signaling pathway in pancreatic cancer cells. Int
J Oncol. 43:1066–1072. 2013.PubMed/NCBI
|
|
12
|
Zhang S, Shan C, Kong G, Du Y, Ye L and
Zhang X: MicroRNA-520e suppresses growth of hepatoma cells by
targeting the NF-kappaB-inducing kinase (NIK). Oncogene.
31:3607–3620. 2012. View Article : Google Scholar
|
|
13
|
Giordano S and Columbano A: MicroRNAs: new
tools for diagnosis, prognosis and therapy in hepatocellular
carcinoma? Hepatology. 57:840–847. 2013. View Article : Google Scholar
|
|
14
|
Wong CM, Kai AK, Tsang FH and Ng IO:
Regulation of hepatocar-cinogenesis by microRNAs. Front Biosci
(Elite edi). 5:49–60. 2013.
|
|
15
|
Hou J, Lin L, Zhou W, et al:
Identification of miRNomes in human liver and hepatocellular
carcinoma reveals miR-199a/b-3p as therapeutic target for
hepatocellular carcinoma. Cancer Cell. 19:232–243. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Su N, Qiu H, Chen Y, Yang T, Yan Q and Wan
X: miR-205 promotes tumor proliferation and invasion through
targeting ESRRG in endometrial carcinoma. Oncology Reports.
29:2297–2302. 2013.PubMed/NCBI
|
|
17
|
Cai J, Fang L, Huang Y, et al: miR-205
targets PTEN and PHLPP2 to augment AKT signaling and drive
malignant phenotypes in non-small cell lung cancer. Cancer Res.
73:5402–5415. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Wang N, Li Q, Feng NH, et al: miR-205 is
frequently down-regulated in prostate cancer and acts as a tumor
suppressor by inhibiting tumor growth. Asian J Androl. 15:735–741.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Yang Z, Huo S, Shan Y, et al: STAT3
repressed USP7 expression is crucial for colon cancer development.
FEBS Lett. 586:3013–3017. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
van Loosdregt J, Fleskens V, Fu J, et al:
Stabilization of the transcription factor Foxp3 by the
deubiquitinase USP7 increases Treg-cell-suppressive capacity.
Immunity. 39:259–271. 2013. View Article : Google Scholar : PubMed/NCBI
|