Introduction
High-mobility-group-box chromosomal protein 1
(HMGB1), a protein of 215 amino acids, is a ubiquitous and abundant
nuclear protein in eukaryotic cells. As a nuclear protein, HMGB1
stabilizes nucleosomes and facilitates DNA replication,
recombination, repair and gene transcription (1–4).
Nuclear HMGB1 serves an important role in maintaining nuclear
stability under stress (5,6). However, an increasing number of
previous studies revealed that extracellular HMGB1 exerts actions,
which are distinctly different compared with its intracellular
functions (7–12). HMGB1 is able to be rapidly
mobilized into the extracellular space, or it can be released
passively as a cytokine by cells undergoing unprogrammed cell death
or necrosis (6,13). HMGB1, when released
extracellularly, is an extremely potent innate signal, which
initiates host defense mechanisms or tissue regeneration (8), and it has been identified as a
macrophage-stimulating factor and a pro-inflammatory mediator
(10).
Serum HMGB1 levels are reported to be raised in
patients with acute organ injuries, including trauma, stroke, acute
myocardial infarction, acute respiratory distress, acute
pancreatitis and ischemia-reperfusion injury (6). Additionally, the increases observed
in circulating levels of the HMGB1 protein were positively
correlated with the severity of the disease in human and animal
models. Complete inhibition or neutralization of HMGB1 may
attenuate the severity of these diseases, decrease the incidence of
organ dysfunction and improve survival, and consequently there is a
burgeoning interest in HMGB1 as a therapeutic target. Among the
strategies investigated, agents which bind to HMGB1 and neutralize
its activity, including antibodies raised against HMGB1 and
molecules which inhibit HMGB1 activation or its interaction with
specific receptors, are gathering attention.
HMGB1 has two separate and characteristic
DNA-binding domains, termed the HMG A and B boxes. Each domain
contains ~80 amino acid residues with a of molecular mass of ~10
kDa. The B box domain contains the pro-inflammatory cytokine
functionality of the molecule, whereas the A box region exerts an
antagonistic, anti-inflammatory effect, and is a putative
therapeutic target (7). The
purified A box has been identified as an antagonist of the
pro-inflammatory actions of the HMGB1 B box, since it competes with
HMGB1 for receptor activation and attenuates the HMGB1-induced
release of pro-inflammatory cytokines (6). The HMGB1 A box markedly ameliorated
damage status. For example, the administration of the HMGB1 A box
in a mouse model of transient coronary vessel occlusion was
associated with a marked attenuation of tissue damage, whereas
systemically administered recombinant HMGB1 protein increased the
severity of the damage by an appreciable extent (14). The administration of the A box
markedly enhanced cardiac allograft survival, and it was associated
with reduced levels of expression of tumor necrosis factor (TNF),
interferon (IFN)-γ and HMGB1 in allografts (13). The administration of the HMGB1 A
box protein to wild-type mice, into which HMGB1 had been injected
directly into the hippo-campus, led to a marked attenuation in
HMGB1-induced seizure severity (15). In a model which emulated
collagen-induced arthritis inflammation, the damage was markedly
attenuated by inhibitors of HMGB1, including anti-HMGB1 antibodies
and HMGB1 A box protein (11).
Additionally, the administration of the A box afforded a high level
of protection against sepsis lethality, reduced the mean arthritis
score, and circumvented disease-induced weight loss and the
histological severity of arthritis (16). Consequently, counteracting
extracellular HMGB1 with its specific antagonist, the HMGB1 A box,
may offer a novel method for therapeutic intervention.
Rapid, efficient and cost-effective protein
expression and purification strategies are required for the
production of therapeutic proteins (17). The small ubiquitin-like modifier
(SUMO) fusion expression system is able to meet these requirements.
SUMO family proteins function as post-translational modifiers by
making covalent and reversible connections with other proteins
(18). SUMO and its associated
enzymes are present in all eukaryotes, and are highly conserved
from yeast to humans, although they are absent in prokaryotes
(19–21). SUMO, fused at the N-terminus with
heterologous proteins, was revealed to improve the folding of the
protein of interest, to enhance its level of expression and to
protect the protein from degradation via its chaperoning properties
(22,23). SUMO-fusion proteins may be cleaved
by SUMO proteases, which recognizes the three-dimensional structure
of the SUMO protein rather than a specific peptide sequence, and
the target protein is obtainable with its native N-terminus intact
(17). The aim of the present
study was to develop a SUMO-fusion expression system in order to
express and purify high levels of the HMGB1 A box protein.
Materials and methods
Reagents
Primers were synthesized by Shanghai Generay Biotech
Co., Ltd (Shanghai, China). The restriction enzymes StuI and
HindIII were purchased from New England Biolabs, Ltd.
(Beijing, China). The Taq DNA polymerase was obtained from
Takara Biotechnology (Dalian) Co., Ltd. (Dalian, China). The
reverse transcription-polymerase chain reaction (RT-PCR)
purification, gel extraction and plasmid miniprep kits were
purchased from Axygen (Corning Inc., Corning, NY, USA). The
expression vector, pSumo-Mut, was modified and obtained from
Novobio Scientific (Shanghai, China). Escherichia coli and
ArcticExpress™ competent cells (DH5α and DE3 cells, respectively)
were also obtained from Novobio Scientific. The
Ni2+-IDA-Sepharose CL-6B affinity column was from
Novagen®, Merck KGaA (Darmstadt, Germany). SUMO protease
was purchased from Novobio Scientific, and the cell counting kit 8
(CCK8) was purchased from Dojindo Molecular Technologies, Inc.
(Rockville, MD, USA). Lipopolysaccharide (LPS) was from
Sigma-Aldrich (St. Louis, MO, USA). Enzyme-linked immunosorbent
assay (ELISA) kits for TNF-α and IL-1β were obtained from R&D
Systems China Co., Ltd. (Shanghai, China).
Construction of the SUMO-HMGB1-A-Box
fusion protein expression vector
The full-length HMGB1 cDNA (NM_002128.4), with
certain synonymous mutations incorporated to render it more
appropriate for prokaryotic expression, was used as a template.
Primers for the HMGB1-A-Box were synthesized, according to the
sequence of the modified HMGB1 cDNA. The sequences of the primers
were as follows: Forward, 5′-GGAGGTATGGGCAAAGGAGATCCTAAGAAG-3′ and
reverse: 5′-AAGCTTTGTTTCCCCTTTAGGAGGGATATAG-3′. Thermal cycling
conditions were as follows: 5 min at 94°C, followed by 30 cycles at
94°C for 30 sec, 51°C for 30 sec and 72°C for 30 sec using a T100
Thermal Cycler (Bio-Rad Laboratories, Inc., Hercules, CA, USA).
Briefly, each PCR reaction mixture (100 µl) contained 10
µl 10X buffer, 20 µM dNTP, 2.5 µl Taq
DNA polymerase, 1.5 mM Mg2+, 1 µl sense and
antisense primers (2.5 µM), and 2 µg cDNA. The PCR
products were digested with the restriction enzymes, StuI
and HindIII, and were ligated into pre-digested vector,
pSumo-Mut, to make the SUMO-HMGB1-A-BOX fusion protein expression
vector, pSumo-Mut-HMGB1-A-BOX. The accuracy of the inserted DNA
segment was confirmed by DNA sequencing at Genewix (Suxhou,
China).
Induction and expression of the
SUMO-HMGB1-A-Box fusion protein expression vector
Aliquots of 1 µl (~5 µg) recombinant
pSumo-Mut-HMGB1-A-BOX plasmid harboring the accurate sequence of
the SUMO-HMGB1-A-BOX fusion gene were transferred into
ArcticExpress™ (DE3) cells (Agilent Technologies, Inc., Santa
Clara, CA, USA). Single transformed colonies were inoculated into 3
ml lysogeny broth (LB), containing 50 µg/ml kanamycin
(Sangon Biotech Co., Ltd., Shanghai, China), and agitated in a
shaker (ZQZY-70Bl ZhiCu, Shanghai, China) at 5 × g overnight at
37°C. A total of 300 µl culture was transferred into 30 ml
LB medium the following day, and this was cultured with agitation
at 5 × g at 37°C until the absorbance at 600 nm reached 0.4. Isopro
pyl-β-d-thiogalactopyranoside (IPTG;
Sangon Biotech Co., Ltd.) was added into the culture at a final
concentration of 0.2 mM. Following induction with agitation at 5 ×
g at 11°C for 4 h, the samples were prepared for subsequent
expression analysis by SDS-PAGE (Bio-Rad Laboratories, Inc.) using
18% SDS-PAGE gels (Amresco LLC, Solon, OH, USA).
Purification of the SUMO-HMGB1-A-Box
fusion protein expression vector
The cells were collected by centrifugation at 95 × g
for 5 min at room temperature following IPTG induction. Cell
pellets of 15 l bacterium solution were resuspended in 600 ml
Ni2+-IDA binding buffer and lysed by sonication using an
ultrasonic cell disruption system (FB705; Thermo Fisher Scientific,
Waltham, MA, USA). The parameters of the sonicator were adjusted to
45% amplitude, 20 min sonication and 3 sec sonication with 3 sec
between pulses. The lysate was subsequently centrifuged at 12,000 g
for 20 min at 4°C. The supernatant was applied onto an
Ni2+-IDA-Sepharose CL-6B affinity column
pre-equilibrated with Ni2+-IDA binding buffer. Following
washing of the column with Ni2+-IDA washing buffer [160
mM Tris-HCl (pH 7.9), 20 mM imidazole and 0.5 M NaCl], bound
proteins were eluted with Ni2+-IDA elution buffer [160
mM Tris-HCl (pH 7.9), 250 mM imidazole and 0.5 M NaCl]. The
fractions were collected prior to SDS-PAGE analysis.
Cleavage of the SUMO-HMGB1-A-BOX and
subsequent purification of HMGB1-A-BOX
The purified fusion protein was dialyzed overnight
with 20 mM Tris-HCl (pH 8.0) buffer. A total of 2 units SUMO
protease/50 µg fusion protein was added and the mixture was
incubated at 30°C for 30 min. The cleaved sample was applied onto
the Ni2+-IDA-Sepharose CL-6B affinity column to separate
the target HMGB1-A-BOX protein from the His-tagged
SUMO-HMGB1-A-BOX, SUMO and SUMO protease. The purified protein,
HMGB1-A-BOX, was dialyzed overnight with phosphate-buffered saline
(PBS; pH 7.4). The concentration of purified HMGB1-A-BOX was
evaluated using a Nanodrop 2000 UV-vis spectrophotometer (Thermo
Fisher Scientific, Wilmington, DE, USA).
Cell culture
Murine macrophage-like RAW 264.7 cells were
purchased from the cell bank of the Shanghai Institutes for
Biological Sciences of the Chinese Academy of Sciences (Shanghai,
China). Macrophage-like RAW 264.7 cells were cultured in Dulbecco's
modified Eagle's medium, supplemented with 15% fetal bovine serum,
100 U/ml penicillin and 100 U/ml streptomycin (all from Invitrogen
Life Technologies, Carlsbad, CA, USA) at 37°C with 5%
CO2. The supernatant was replaced with fresh medium
every 48 h, and cells, which were in a healthy condition with a
viability >98%, were used for subsequent experiments.
Cell viability measurements and
ELISA
A total of 100 µl macrophage-like RAW 264.7
cell suspension was seeded into 96-well plates at a density of
1×105/ml. LPS was added at a final concentration of 20
µg/ml. Following 1 h incubation with LPS, HMGB1-A-BOX
protein at final concentrations of 100 or 200 µg/ml were
added into the experimental group (Exp). An identical volume of PBS
was added into the control group (Ctrl). Blank controls (Blank)
were established with PBS and lacking LPS stimulation. Following 2,
6, 12, 24 or 48 h incubation, 10 µl CCK8 medium was added to
corresponding wells, and the plates were incubated at 37°C for an
additional 1 h. The absorbance at a wavelength of 450 nm (A) was
determined using an ELISA reader (BioRad M450; Bio-Rad
Laboratories, Inc.), and the cell viability was calculated
according to the following equation: Cell viability =
(AExp − ABlank)/(ACtrl −
ABlank) × 100%. The data were acquired from experiments
performed in triplicate. The concentration of TNF-α and IL-1β in
the cell culture supernatants was determined by ELISA, according to
the manufacturer's instructions.
Statistical analysis
The results are expressed as the mean ± standard
deviation. Statistical analyses were performed using the paired
Student's t-test, with paired comparisons being made where relevant
using GraphPad Prism 5 (GraphPad Software, Inc., La Jolla, CA,
USA). P<0.05 was considered to indicate a statistically
significant difference.
Results
Construction of the SUMO-HMGB1-A-BOX
expression strain
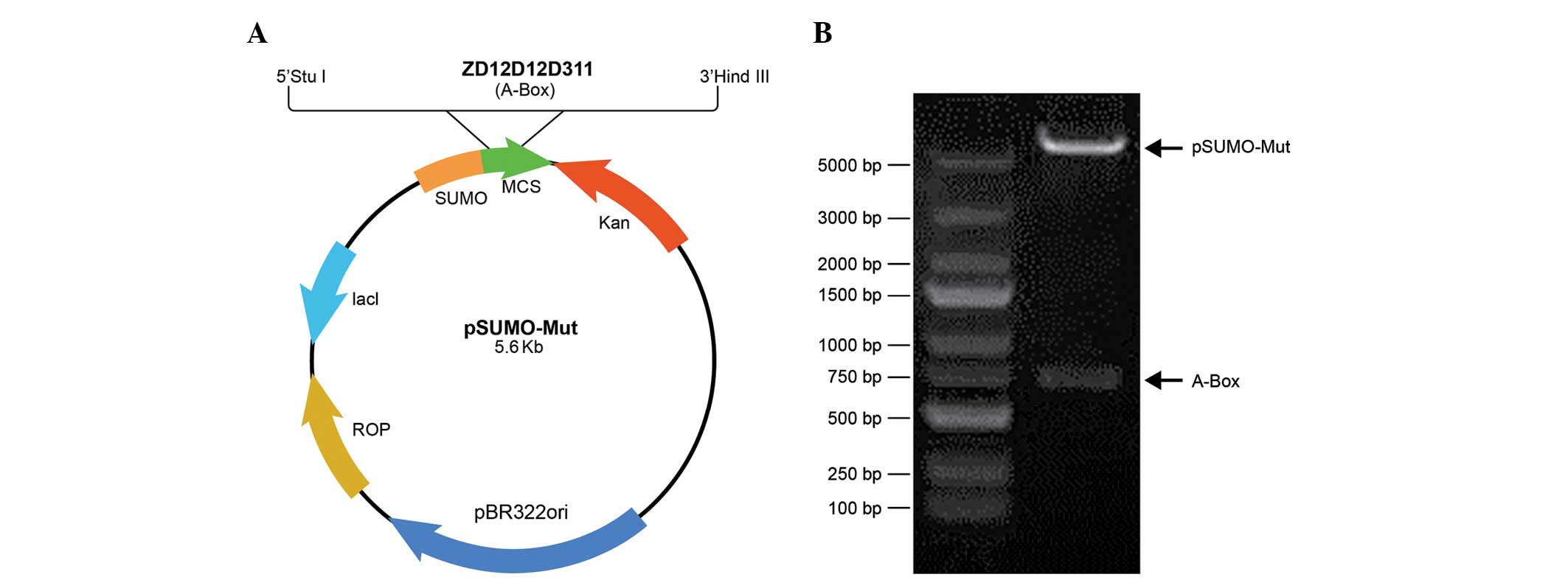
The PCR products of HMGB1-A-BOX were digested with
the restriction enzymes, StuI and HindIII, and were
ligated into pre-digested pSumo-Mut vector to create the
SUMO-HMGB1-A-BOX fusion protein expression vector,
pSumo-Mut-HMGB1-A-BOX (Fig. 1A).
Two restriction enzyme sites, Xbal I and Xhol I, were
located at the ends of the sequence of the fusion protein, and
restriction enzyme analysis of the pSumo-Mut-HMGB1-A-BOX plasmid by
Xbal I and Xhol I was performed to confirm that the
construct of pSumo-Mut-HMGB1-A-BOX was obtained (Fig. 1B).
Expression of the SUMO-HMGB1-A-BOX fusion
protein
The recombinant plasmid, pSumo-Mut-HMGB1-A-BOX,
harboring the accurate sequence of the SUMO-HMGB1-A-BOX fusion gene
was transferred into ArcticExpress™ DE3 cells. A total of two
single transformed colonies were inoculated and cultured on a large
scale, prior to IPTG induction. Each colony expressed high levels
of the SUMO-HMGB1-A-BOX fusion protein following 0.2 mM IPTG
induction at 11°C for 4 h (Fig.
2A; lanes 2 and 3).
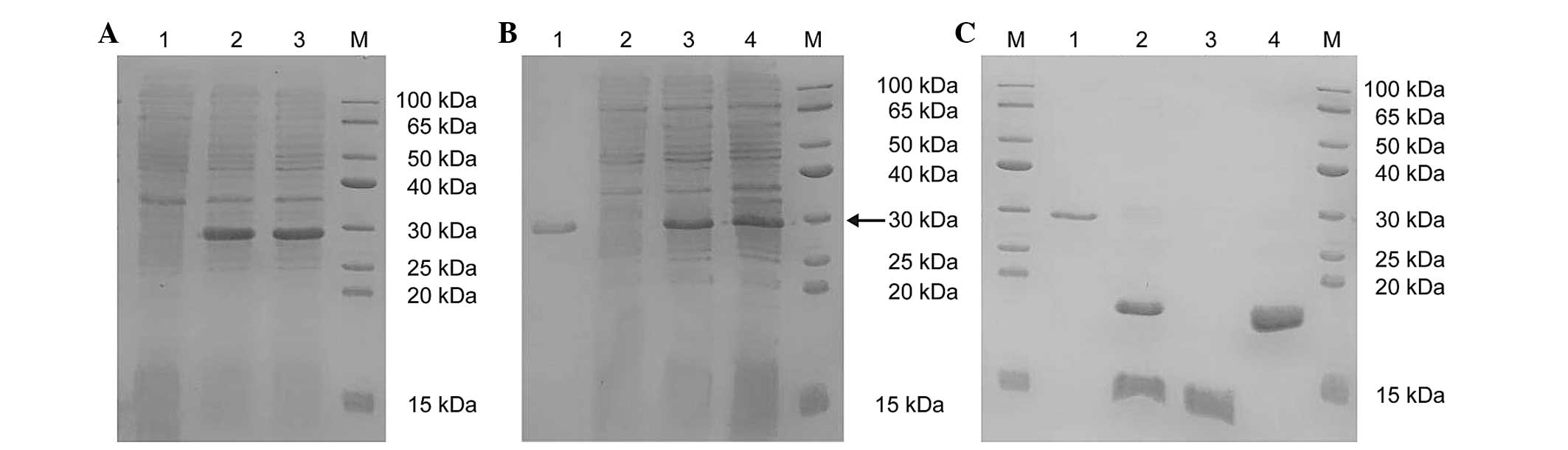 | Figure 2SDS-PAGE analysis of protein
expression levels. (A) The SDS-PAGE analysis of the protein
expression of SUMO-HMGB1-A-BOX (M, protein marker; lane 1,
non-induced ArcticExpress™ (DE3 cells)/Sumo-Mut-HMGB1-A-BOX; lanes
2 and 3, supernatant of ArcticExpress™ (DE3)/pSumo-Mut-HMGB1-A-BOX
induced at 11°C with 0.2 mM IPTG for 4 h). (B) The SDS-PAGE
analysis of the purified SUMO-HMGB1-A-BOX fusion protein (M,
protein marker; lane 1, precipitate of ArcticExpress™ (DE3
cells)/pSumo-Mut-HMGB1-A-BOX induced at 11°C with 0.2 mM IPTG for 4
h; lane 2, supernatant of ArcticExpress™
(DE3)/pSumo-Mut-HMGB1-A-BOX induced at 11°C with 0.2 mM IPTG for 4
h; lane 3, flow through; lane 4, purified SUMO-HMGB1-A-BOX eluted
with Ni2+-IDA elution buffer containing 250 mM
imidazole). The fusion protein is indicated by the black arrow. (C)
The SDS-PAGE analysis of purified HMGB1-A-BOX protein (M, Protein
marker; lane 1, purified SUMO-HMGB1-A-BOX fusion protein; lane 2,
digested mixture of the purified fusion protein using SUMO protease
at 30°C for 30 min; lane 3, purified target protein of HMGB1-A-BOX
in the flow through; lane 4, remaining mixture eluted with the
Ni2+-IDA elution buffer, containing 250 mM imidazole).
HMGB1, high-mobility-group box chromosomal protein 1; IPTG,
isopropyl-β-D-thiogalactopyranoside; SUMO, small ubiquitin-like
modifier. |
Purification of the SUMO-HMGB1-A-BOX
fusion protein
Following IPTG induction, the cells were collected
by centrifugation, and the cell pellets were resuspended in
Ni2+-IDA binding buffer and lysed by sonication. The
results of the SDS-PAGE experiment revealed that the
SUMO-HMGB1-A-BOX fusion protein was expressed in the precipitate
and in the supernatant of ArcticExpress™ DE3
cells/pSumo-Mut-HMGB1-A-BOX induced at 11°C with 0.2 mM IPTG for 4
h (Fig. 2B, lanes 1 and 2). The
supernatant was loaded onto an Ni2+-IDA-Sepharose CL-6B
affinity column pre-equilibrated with Ni2+-IDA binding
buffer. Proteins with a His-tag were retained on the column,
whereas proteins lacking the His-tag were removed by the
Ni2+-IDA washing buffer. The bound fusion proteins were
eluted with Ni2+-IDA elution buffer, containing 250 mM
imidazole, and the purity was revealed to be >90% (Fig. 2B; lane 4).
Cleavage of the SUMO-HMGB1-A-BOX fusion
protein and the subsequent purification of HMGB1-A-BOX
The HMGB1-A-BOX fusion protein was released by
cleaving the purified SUMO-HMGB1-A-BOX with SUMO protease. The
results of the SDS-PAGE experiment revealed that almost all the
fusion protein was cleaved within 30 min at 30°C (Fig. 2C; lane 2). The digested mixture was
subsequently reloaded onto the Ni2+-IDA-Sepharose CL-6B
affinity column for further purification. Since all the non-cleaved
fusion protein, SUMO fragments and SUMO protease possessed a
His-tag and were retained on the column, the released HMGB1-A-BOX
target protein was obtained in the flow-through, and the purity of
HMGB1-A-BOX was revealed to be >90% (Fig. 2C; lane 3). The purified HMGB1-A-BOX
protein was dialyzed overnight with PBS (pH 7.4), and the final
concentration of purified HMGB1-A-BOX protein, and the total yield
of this target protein, were determined to be 0.72 mg/ml and 10.8
mg, respectively.
HMGB1-A-BOX ameliorates LPS-impaired cell
viability
Previous results have indicated that the HMGB1
protein may be released by LPS-activated RAW 264.7 cells, and this
may contribute to the LPS-regulated cell viability (24). To assess the anti-inflammatory
function of the HMGB1 A box following its expression and
purification, the cell viability was evaluated following LPS
treatment with or without HMGB1-A-Box incubation. After treatment
with 20 µg/ml LPS for 1 h, the HMGB1-A-BOX protein at final
concentrations of 100 and 200 µg/ml were added into the
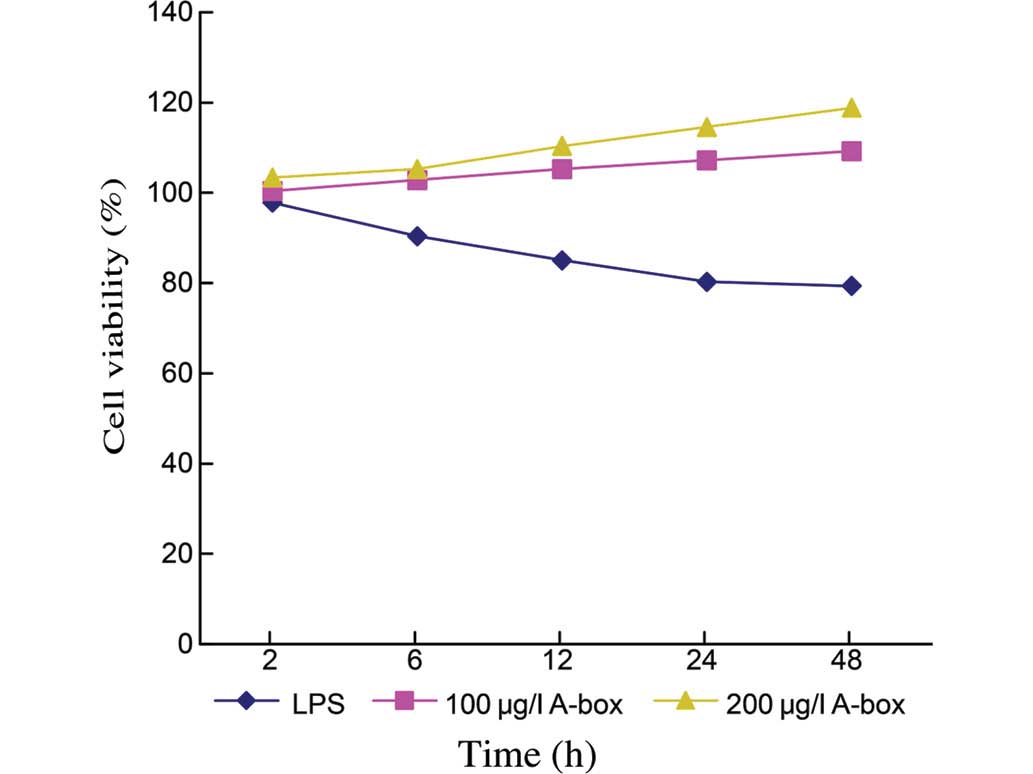
experimental groups and incubated for 2, 6, 12, 24 or 48 h. The
control groups, with LPS stimulation only, demonstrated an impaired
cell viability throughout the duration of the experiment. By
contrast, the cell viability was gradually increased over time on
addition of the HMGB1-A-BOX protein, and this increase was
dose-dependent (Fig. 3). The cell
viabilities at 48 h were 79.33±0.49% for the Control group (n=3),
109.26±0.63% for the group treated with 100 µg/ml
HMGB1-A-BOX protein (n=3, P<0.05) and 118.81±0.64% for the group
treated with 200 µg/ml HMGB1-A-BOX protein (n=3;
P<0.05).
HMGB1-A-BOX protein attenuates the levels
of TNF-α and IL-1β
TNF-α and IL-1β are released at an early stage
during the onset of systemic inflammatory responses (16). Furthermore, HMGB1 protein released
by LPS-activated RAW 264.7 cells has been demonstrated to stimulate
macrophages to release TNFα and IL-1β (12). Therefore, an objective of the
present study was to observe the effect of the purified HMGB1-A-BOX
protein on the levels of TNF-α and IL-1β stimulated by LPS/HMGB1 in
the supernatant. The level of TNF-α in the control group reached a
maximum at 48 h (Table I), whereas
the level of IL-1β peaked at 12 h (Table II). The level of TNF-α was reduced
immediately following a 2 h incubation with HMGB1-A-BOX, compared
with the control group (P<0.05), and the inhibitory effect was
time-dependent. The maximum inhibition of TNF-α was achieved
following 48 h incubation with HMGB1-A-BOX and reached ~30.20%
(P<0.05; Table I). The levels
of IL-1β observed upon incubation with HMGB1-A-BOX were inhibited,
reaching the maximum at 24 h, and this was postponed compared with
the control group (Table II).
 | Table IEffect of the HMGB1-A-BOX protein on
the levels of TNFα in the supernatant (pg/ml). |
Table I
Effect of the HMGB1-A-BOX protein on
the levels of TNFα in the supernatant (pg/ml).
| Group | N | Levels of TNFα at
various time points (h)
|
|---|
| 2 | 6 | 12 | 24 | 48 |
|---|
| Ctrl | 20 | 3,339.5±56.3 | 3,569.7±69.3 | 3,673.4±82.5 | 4,102.7±66.3 | 6,189.8±71.4 |
| Exp | 20 | 2,167.2±76.6a | 2,348.2±49.8a | 3,015.4±59.1a | 3,265.8±58.9a | 4,320.5±63.2a |
 | Table IIEffect of the HMGB1-A-BOX protein on
the levels of IL-1β in the supernatant (pg/ml). |
Table II
Effect of the HMGB1-A-BOX protein on
the levels of IL-1β in the supernatant (pg/ml).
| Group | N | Levels of IL-1β at
various time points (h)
|
|---|
| 2 | 6 | 12 | 24 | 48 |
|---|
| Ctrl | 20 | 3,216.1±44.2 | 3,389.6±49.6 | 3,521.9±51.7 | 3,306.8±62.7 | 3,209.3±47.6 |
| Exp | 20 |
2,757.2±70.1a |
2,978.5±39.4a |
3,068.8±60.5a | 3,187.4±54.2 |
2,685.5±81.7a |
Discussion
Theoretically, the size of the SUMO-fusion protein
is ~24 kDa. However, the size of the SUMO-fused HMGB1-A-BOX
protein, as determined by SDS-PAGE analysis, appeared to be
marginally larger than the predicted size. The possible explanation
for this phenomenon may be attributable to the SUMO protein itself:
SUMO is a ubiquitin-like protein, which is able to form tertiary
structures with itself (17),
which consequently leads to increases in molecular weight. This was
commonly observed when other SUMO-fusion proteins were being
expressed. The sought-after target protein, HMGB1-A-BOX, is a
low-molecular-weight protein of ~10 kDa. SUMO proteases have been
observed to successfully cleave a broad range of sizes (6–110 kDa)
of partner proteins fused to SUMO (17). The achievement in obtaining
purified HMGB1-A-BOX protein with the SUMO-fusion expression system
used in the present study also supported the capability of this
expression system to accommodate SUMO fusion partner proteins.
Since the A box region of HMGB1 exerts an
antagonistic, anti-inflammatory effect with therapeutic potential,
the ultimate goal for expressing and purifying HMGB1-A-BOX is for
therapeutic interventions. The production of therapeutic proteins
is required to be rapid, efficient and cost-effective. Furthermore,
tags should be removed in order that the protein activity is
neither attenuated nor modified. Different methods used in
preliminary investigations to express the HMGB1-A-BOX protein
failed to yield satisfactory results (data not shown). The yields
of the HMGB1-A-BOX fusion protein obtained with a glutathione
S-transferase (GST) tag were acceptable, although removal of the
GST tag affected the activity of the target protein. Expressing the
target protein without a GST tag led to low solubility. One
advantage of the SUMO-fusion technology used in the present study
is that 10.8 mg soluble HMGB1-A-BOX protein of a purity >90% was
obtained from 15 l bacterium solution. SUMO fusion was reported to
enhance protein expression, solubility and purification in
prokaryotes (22). This fusion
technology is able to improve the expression of the recombinant
proteins by protecting them from degradation. In addition, SUMO has
an external hydrophilic surface and inner hydrophobic core, which
may exert a detergent-like effect on otherwise-insoluble proteins
(17,22). Furthermore, the presence of the
His-tag on SUMO and the SUMO protease provides a simplified means
of purification to obtain high levels of the target proteins.
Notably, the lack of an endogenous SUMO protease in prokaryotes
facilitated the use of SUMO as a purification tag in E.
coli. SUMO proteases are accurate and efficient agents at
cleaving the SUMO tag and in allowing for retention of the desired
N-terminus, without the extraneous residues, which are usually
produced by other proteases, since SUMO proteases recognize the
tertiary structure of the SUMO tag instead of peptide sequences.
The use of the SUMO-fusion expression system in the present study
has overcome several problems, including low protein yield,
precipitation of the target protein and a failure to recover
active, structurally intact protein.
The recombinant A box of HMGB1 is antagonistic to
the B box and full-length HMGB1 protein, and is considered to
compete with HMGB1 for receptor activation. A previous study has
reported that the recombinant HMGB1-A-BOX protein may attenuate the
development of the inflammatory disease, thromboangiitis
obliterans, in rat models, which provided further evidence that
HMGB1-A-BOX may be a putative therapeutic protein for the treatment
of certain inflammatory diseases (25). Furthermore, the truncation of HMGB1
into individual structural domains revealed that HMGB1-A-box, a
DNA-binding motif, specifically antagonizes the activity of HMGB1
and rescues mice from lethal sepsis (9). Accordingly, strategies that target
HMGB1 with specific antibodies or antagonists are potentially
useful as therapies for lethal systemic inflammatory disease. The
recombinant A box of HMGB1 obtained with SUMO fusion technology in
the present study was demonstrated to attenuate the levels of TNF-α
and IL-1β in the supernatant, and to ameliorate LPS-impaired cell
viability. In addition, this technology has allowed the production
of active HMGB1-A-BOX protein with a high yield and purity, which
is valuable for producing high levels of this target protein for
subsequent therapeutic studies.
Acknowledgments
This study was supported by the National Natural
Science Foundation of China (no. 81200279), the Medical Engineering
(Science) Cross-Research Fund of Shanghai Jiao Tong University (no.
YG2011MS29) and the LI Jieshou Bowel Function Barrier Medical
Foundation (no. LJS-201111).
References
|
1
|
Bustin M, Lehn DA and Landsman D:
Structural features of the HMG chromosomal proteins and their
genes. Biochim Biophys Acta. 1049:231–243. 1990. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Boonyaratanakornkit V, Melvin V,
Prendergast P, Altmann M, Ronfani L, Bianchi ME, Taraseviciene L,
Nordeen SK, Allegretto EA and Edwards DP: High-mobility group
chromatin proteins 1 and 2 functionally interact with steroid
hormone receptors to enhance their DNA binding in vitro and
transcriptional activity in mammalian cells. Mol Cell Biol.
18:4471–4487. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Melvin VS and Edwards DP: Coregulatory
proteins in steroid hormone receptor action: The role of chromatin
high mobility group proteins HMG-1 and -2. Steroids. 64:576–586.
1999. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Vaccari T, Beltrame M, Ferrari S and
Bianchi ME: Hmg4, a new member of the Hmg1/2 gene family. Genomics.
49:247–252. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Funayama A, Shishido T, Netsu S, Narumi T,
Kadowaki S, Takahashi H, Miyamoto T, Watanabe T, Woo CH, Abe J, et
al: Cardiac nuclear high mobility group box 1 prevents the
development of cardiac hypertrophy and heart failure. Cardiovasc
Res. 99:657–664. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Tsung A, Tohme S and Billiar TR: High
mobility group box-1 in sterile inflammation. J Intern Med.
276:425–443. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Dumitriu IE, Baruah P, Manfredi AA,
Bianchi ME and Rovere-Querini P: HMGB1: Guiding immunity from
within. Trends Immunol. 26:381–387. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Yang D, Chen Q, Yang H, Tracey KJ, Bustin
M and Oppenheim JJ: High mobility group box-1 protein induces the
migration and activation of human dendritic cells and acts as an
alarmin. J Leukoc Biol. 81:59–66. 2007. View Article : Google Scholar
|
|
9
|
Yang H, Wang H, Czura CJ and Tracey KJ:
HMGB1 as a cytokine and therapeutic target. J Endotoxin Res.
8:469–472. 2002. View Article : Google Scholar
|
|
10
|
Andersson U and Tracey KJ: HMGB1 in
sepsis. Scand J Infect Dis. 35:577–584. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Kokkola R, Li J, Sundberg E, Aveberger AC,
Palmblad K, Yang H, Tracey KJ, Andersson U and Harris HE:
Successful treatment of collagen-induced arthritis in mice and rats
by targeting extracellular high mobility group box chromosomal
protein 1 activity. Arthritis Rheum. 48:2052–2058. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Wang H, Bloom O, Zhang M, Vishnubhakat JM,
Ombrellino M, Che J, Frazier A, Yang H, Ivanova S, Borovikova L, et
al: HMG-1 as a late mediator of endotoxin lethality in mice.
Science. 285:248–251. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Huang Y, Yin H, Han J, Huang B, Xu J,
Zheng F, Tan Z, Fang M, Rui L, Chen D, et al: Extracellular hmgb1
functions as an innate immune-mediator implicated in murine cardiac
allograft acute rejection. Am J Transplant. 7:799–808. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Andrassy M, Volz HC, Igwe JC, Funke B,
Eichberger SN, Kaya Z, Buss S, Autschbach F, Pleger ST, Lukic IK,
et al: High-mobility group box-1 in ischemia-reperfusion injury of
the heart. Circulation. 117:3216–3226. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Andersson U and Tracey KJ: HMGB1 is a
therapeutic target for sterile inflammation and infection. Annu Rev
Immunol. 29:139–162. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Yang H, Ochani M, Li J, Qiang X, Tanovic
M, Harris HE, Susarla SM, Ulloa L, Wang H, DiRaimo R, et al:
Reversing established sepsis with antagonists of endogenous
high-mobility group box 1. Proc Natl Acad Sci USA. 101:296–301.
2004. View Article : Google Scholar :
|
|
17
|
Butt TR, Edavettal SC, Hall JP and Mattern
MR: SUMO fusion technology for difficult-to-express proteins.
Protein Expr Purif. 43:1–9. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Johnson ES: Protein modification by SUMO.
Annu Rev Biochem. 73:355–382. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Hochstrasser M: Evolution and function of
ubiquitin-like protein-conjugation systems. Nat Cell Biol.
2:E153–E157. 2000. View
Article : Google Scholar : PubMed/NCBI
|
|
20
|
Jentsch S and Pyrowolakis G: Ubiquitin and
its kin: How close are the family ties? Trends Cell Biol.
10:335–342. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Müller S, Hoege C, Pyrowolakis G and
Jentsch S: SUMO, ubiquitin's mysterious cousin. Nat Rev Mol Cell
Biol. 2:202–210. 2001. View
Article : Google Scholar : PubMed/NCBI
|
|
22
|
Malakhov MP, Mattern MR, Malakhova OA,
Drinker M, Weeks SD and Butt TR: SUMO fusions and SUMO-specific
protease for efficient expression and purification of proteins. J
Struct Funct Genomics. 5:75–86. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Zuo X, Mattern MR, Tan R, Li S, Hall J,
Sterner DE, Shoo J, Tran H, Lim P, Sarafianos SG, et al: Expression
and purification of SARS coronavirus proteins using SUMO-fusions.
Protein Expr Purif. 42:100–110. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Zhang Z, Zhang L, Zhou C and Wu H:
Ketamine inhibits LPS-induced HGMB1 release in vitro and in vivo.
Int Immunopharmacol. 23:14–26. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Kong X, Yuan H, Wu X, Zhang J, Zhou H,
Wang M, Liu Y and Jin X: High-mobility-group box protein 1A box
reduces development of sodium laurate-induced thromboangiitis
obliterans in rats. J Vasc Surg. 57:194–204. 2013. View Article : Google Scholar
|