|
1
|
Klein AP: Identifying people at a high
risk of developing pancreatic cancer. Nat Rev Cancer. 13:66–74.
2013. View
Article : Google Scholar : PubMed/NCBI
|
|
2
|
Torre LA, Bray F, Siegel RL, Ferlay J,
Lortet-Tieulent J and Jemal A: Global cancer statistics, 2012. CA
Cancer J Clin. 65:87–108. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Jemal A, Siegel R, Ward E, Hao Y, Xu J and
Thun MJ: Cancer statistics, 2009. CA Cancer J Clin. 59:225–249.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Lin QJ, Yang F, Jin C and Fu DL: Current
status and progress of pancreatic cancer in China. World J
Gastroenterol. 21:7988–8003. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Heinemann V, Reni M, Ychou M, Richel DJ,
Macarulla T and Ducreux M: Tumour-stroma interactions in pancreatic
ductal adenocarcinoma: Rationale and current evidence for new
therapeutic strategies. Cancer Treat Rev. 40:118–128. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Yachida S and Iacobuzio-Donahue CA: The
pathology and genetics of metastatic pancreatic cancer. Arch Pathol
Lab Med. 133:413–422. 2009.PubMed/NCBI
|
|
7
|
Simard EP, Ward EM, Siegel R and Jemal A:
Cancers with increasing incidence trends in the United States: 1999
through 2008. CA Cancer J Clin. 62:118–128. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Siegel R, Naishadham D and Jemal A: Cancer
statistics, 2013. CA Cancer J Clin. 63:11–30. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Brower JV, Clark PA, Lyon W and Kuo JS:
MicroRNAs in cancer: Glioblastoma and glioblastoma cancer stem
cells. Neurochem Int. 77:68–77. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Shukla GC, Singh J and Barik S: MicroRNAs:
Processing, maturation, target recognition and regulatory
functions. Mol Cell Pharmacol. 3:83–92. 2011.PubMed/NCBI
|
|
11
|
Kalla R, Ventham NT, Kennedy NA, Quintana
JF, Nimmo ER, Buck AH and Satsangi J: MicroRNAs: New players in
IBD. Gut. 64:504–517. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Zhang W, Dahlberg JE and Tam W: MicroRNAs
in tumorigenesis. Am J Pathol. 171:728–738. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Zhou N and Mo YY: Roles of microRNAs in
cancer stem cells. Front biosci (School Ed). 4:810–818. 2012.
|
|
14
|
Tutar Y: miRNA and cancer; computational
and experimental approaches. Curr Pharm Biotechnol. 15:4292014.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Vetter G, Saumet A, Moes M, Vallar L, Le
Béchec A, Laurini C, Sabbah M, Arar K, Theillet C, Lecellier CH and
Friederich E: miR-661 expression in SNAI1-induced epithelial to
mesenchymal transition contributes to breast cancer cell invasion
by targeting Nectin-1 and StarD10 messengers. Oncogene.
29:4436–4448. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Li C, Duan P, Wang J, Lu X and Cheng J:
miR-320 inhibited ovarian cancer oncogenicity via targeting TWIST1
expression. Am J Transl Res. 9:3705–3713. 2017.PubMed/NCBI
|
|
17
|
Jensen KP and Covault J: Human miR-1271 is
a miR-96 paralog with distinct non-conserved brain expression
pattern. Nucleic Acids Res. 39:701–711. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Maurel M, Jalvy S, Ladeiro Y, Combe C,
Vachet L, Sagliocco F, Bioulac-Sage P, Pitard V, Jacquemin-Sablon
H, Zucman-Rossi J, et al: A functional screening identifies five
microRNAs controlling glypican-3: Role of miR-1271 down-regulation
in hepatocellular carcinoma. Hepatology. 57:195–204. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Zhou Z, Niu X, Li C, Sheng S and Lu S:
Inhibition of the growth of non-small cell lung cancer by
miRNA-1271. Am J Transl Res. 7:1917–1924. 2015.PubMed/NCBI
|
|
20
|
Yu T, Yu HR, Sun JY, Zhao Z, Li S, Zhang
XF, Liao ZX, Cui MK, Li J, Li C and Zhang Q: miR-1271 inhibits ERα
expression and confers letrozole resistance in breast cancer.
Oncotarget. 8:107134–107148. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
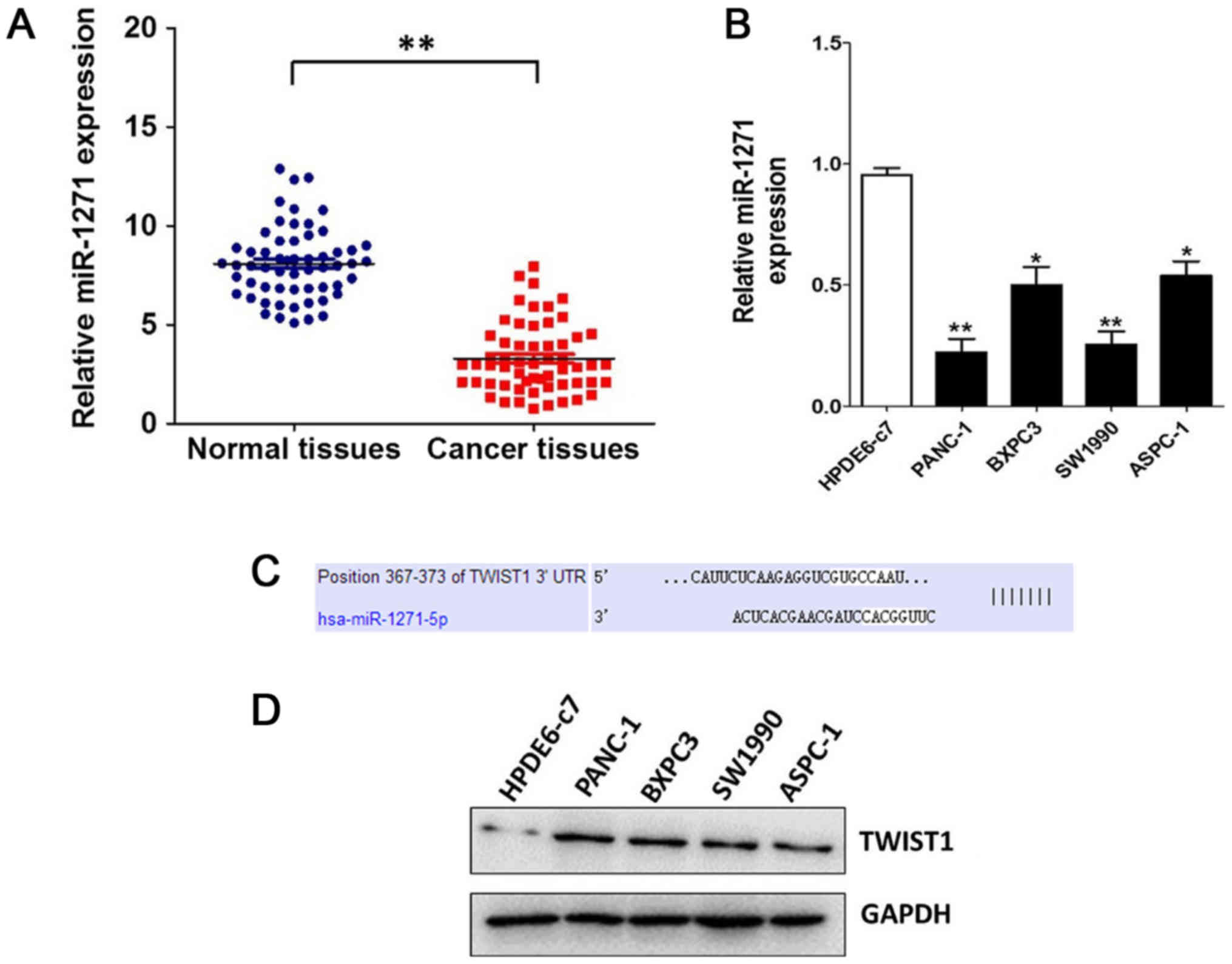
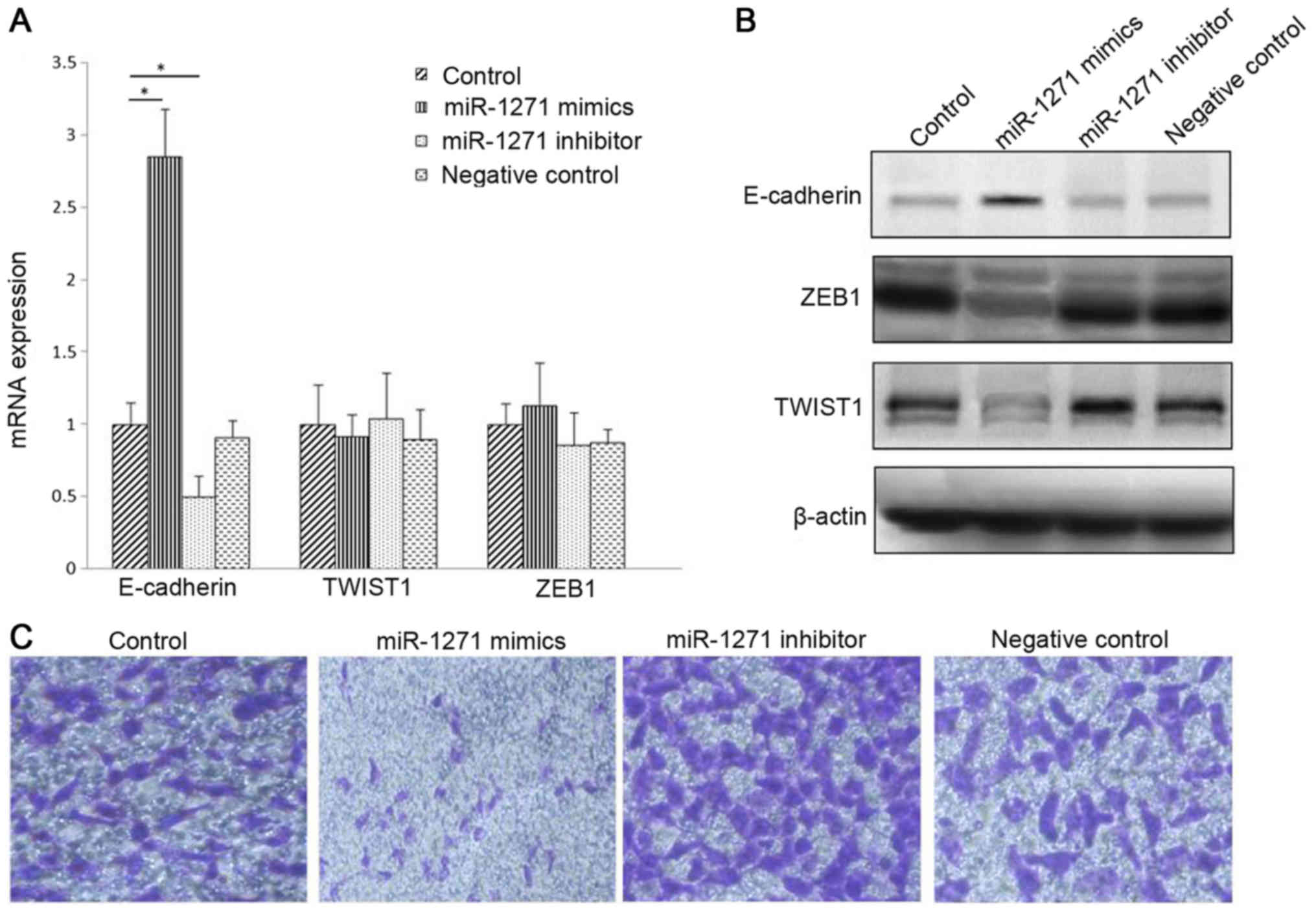
Liu H, Wang H, Liu X and Yu T: miR-1271
inhibits migration, invasion and epithelial-mesenchymal transition
by targeting ZEB1 and TWIST1 in pancreatic cancer cells. Biochem
Biophys Res Commun. 472:346–352. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Yang M, Shan X, Zhou X, Qiu T, Zhu W, Ding
Y, Shu Y and Liu P: miR-1271 regulates cisplatin resistance of
human gastric cancer cell lines by targeting IGF1R, IRS1, mTOR and
BCL2. Anticancer Agents Med Chem. 14:884–891. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Galván JA, Zlobec I, Wartenberg M, Lugli
A, Gloor B, Perren A and Karamitopoulou E: Expression of E-cadherin
repressors SNAIL, ZEB1 and ZEB2 by tumour and stromal cells
influences tumour-budding phenotype and suggests heterogeneity of
stromal cells in pancreatic cancer. Br J Cancer. 112:1944–1950.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Chen X, Peng H, Xiao J, Guan A, Xie B, He
B and Chen Q: Benzo(a)pyrene enhances the EMT-associated migration
of lung adenocarcinoma A549 cells by upregulating Twist1. Oncol
Rep. 38:2141–2147. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Ma L, Liu L, Ma Y, Xie H, Yu X, Wang X,
Fan A, Ge D, Xu Y, Zhang Q and Song C: The role of
E-Cadherin/β-catenin in hydroxysafflor yellow a inhibiting
adhesion, invasion, migration and lung metastasis of hepatoma
cells. Biol Pharm Bull. 40:1706–1715. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Du L and Wang-Gillam A: Trends in
neoadjuvant approaches in pancreatic cancer. J Natl Compr Canc
Netw. 15:1070–1077. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Srinivas G, Babykutty S, Sathiadevan PP
and Srinivas P: Molecular mechanism of emodin action: Transition
from laxative ingredient to an antitumor agent. Med Res Rev.
27:591–608. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Dey D, Ray R and Hazra B: Antitubercular
and antibacterial activity of quinonoid natural products against
multi-drug resistant clinical isolates. Phytother Res.
28:1014–1021. 2014. View
Article : Google Scholar : PubMed/NCBI
|
|
30
|
Shrimali D, Shanmugam MK, Kumar AP, Zhang
J, Tan BK, Ahn KS and Sethi G: Targeted abrogation of diverse
signal transduction cascades by emodin for the treatment of
inflammatory disorders and cancer. Cancer Lett. 341:139–149. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Gu J, Cui CF, Yang L, Wang L and Jiang XH:
Emodin inhibits colon cancer cell invasion and migration by
suppressing epithelialmesenchymal transition via the Wnt/β-catenin
pathway. Oncol Res. 2018.doi: 10.3727/096504018X15150662230295.
View Article : Google Scholar
|
|
32
|
Min H, Niu M, Zhang W, Yan J, Li J, Tan X,
Li B, Su M, Di B and Yan F: Emodin reverses leukemia multidrug
resistance by competitive inhibition and downregulation of
P-glycoprotein. PLoS One. 12:e01879712017. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Tseng HS, Wang YF, Tzeng YM, Chen DR, Liao
YF, Chiu HY and Hsieh WT: Aloe-emodin enhances tamoxifen
cytotoxicity by suppressing ras/ERK and PI3K/mTOR in breast cancer
cells. Am J Chin Med. 45:337–350. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Wang X, Li L, Guan R, Zhu D, Song N and
Shen L: Emodin inhibits ATP-induced proliferation and migration by
suppressing P2Y receptors in human lung adenocarcinoma cells. Cell
Physiol Biochem. 44:1337–1351. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Hu L, Cui R, Liu H and Wang F: Emodin and
rhein decrease levels of hypoxia-inducible factor-1α in human
pancreatic cancer cells and attenuate cancer cachexia in athymic
mice carrying these cells. Oncotarget. 8:88008–88020. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Pan FP, Zhou HK, Bu HQ, Chen ZQ, Zhang H,
Xu LP, Tang J, Yu QJ, Chu YQ, Pan J, et al: Emodin enhances the
demethylation by 5-Aza-CdR of pancreatic cancer cell
tumor-suppressor genes P16, RASSF1A and ppENK. Oncol Rep.
35:1941–1949. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Mooney SM, Talebian V, Jolly MK, Jia D,
Gromala M, Levine H and McConkey BJ: The GRHL2/ZEB feedback loop-a
key axis in the regulation of emt in breast cancer. J Cell Biochem.
118:2559–2570. 2017. View Article : Google Scholar : PubMed/NCBI
|