Introduction
Gastric cancer remains a major health problem and a
leading cause of cancer-related death, although its incidence has
decreased worldwide (1). Numerous
patients are identified for gastric cancers at the advanced stage,
which is associated with increased recurrence and low overall
survival (OS) following potentially curative resection (2). Radical surgery, including lymph node
(LN) dissection, has been the standard treatment for early gastric
cancer; however, 50% of gastric cancer patients suffer from tumor
relapses even after radical surgery (3,4), and
their overall prognosis is suboptimal despite aggressive treatment.
In recent years, much evidence has clearly demonstrated that
multiple genetic changes are responsible for the development and
progression of gastric cancer. Thus, it is imperative to
investigate the molecular mechanism to improve the outcome of
patients with gastric cancer.
Studies have demonstrated that the aggressive nature
of gastric cancer is related to mutations of various oncogenes and
tumor suppressor genes, as well as abnormalities in certain growth
factors and their receptors (5).
Recent studies have shown that runt-related transcription factor 3
(Runx3) gene mutation is significantly associated with
primary gastric cancer progression. However, the detailed mechanism
for this relationship has yet to be completely clarified (6). The Runx3 gene encodes a protein
that belongs to the runt domain family of transcription factors
involved in mammalian development pathways (7). Runx3 protein mediates the
growth suppression effects of TGF-β in association with SMAD, a
downstream protein in the signaling pathway (8,9).
Previous studies demonstrated that Runx3 is markedly
down-regulated in gastric cancers compared to the surrounding
mucosa, and that lack of Runx3 is causally related to the
growth and progression of human gastric cancer (10), indicating that Runx3 is a
novel tumor suppressor (11–14).
It is known that the down-regulation of Runx3
in primary gastric cancer tissues is associated with poor
prognosis. However, less is known about its expression in LNs,
particularly the relationship between the expression of
Runx3 and the prognosis of patients. Lymph node metastasis
(LNM) is a significant factor for determining the prognosis of
patients with gastric cancer and is a frequent target for
chemotherapy. Therefore, it is essential to analyze Runx3
gene expression in LNs from gastric cancer. In this study, reverse
transcriptase-polymerase chain reaction (RT-PCR) and Western
blotting were used to assess the expression level of Runx3
in 101 LNs of stomach carcinoma, and to describe for the first time
the association between the expression of Runx3 gene in LNs
and the outcome of patients with gastric cancer.
Materials and methods
Tissue samples
LN specimens were obtained from 101 patients who
were diagnosed with primary gastric cancer and underwent
gastrectomy and LN dissection in the Department of Surgery, the
Tenth People's Hospital of Shanghai, Tongji University, China,
between October 2000 and October 2002. The samples were rapidly
frozen in liquid nitrogen and stored at −80°C until being used for
the extraction of RNA and protein. The gastric cancer patients had
well-documented clinical histories and follow-up information.
Clinicopathological data were obtained from a retrospectively
constructed medical database, which had been reviewed and confirmed
by two pathologists. Patients who had been preoperatively treated
with radiation and/or chemotherapy were excluded. Informed consent
was obtained from each of the patients and the study protocol was
approved by the Ethics Committee of the Tenth People's Hospital of
Shanghai, China. Details of the patient characteristics and
Runx3 expression are provided in Table I. Following radical gastrectomy,
patients were followed up until death or October 31, 2009, as
appropriate. The median follow-up duration was 35.6 months. At the
last follow-up examination, 22 (21.8%) patients were still alive,
whereas 79 (78.2%) patients had succumbed.
 | Table IAssociation between the expression of
Runx3 and clinicopathological characteristics of patients
with gastric cancer. |
Table I
Association between the expression of
Runx3 and clinicopathological characteristics of patients
with gastric cancer.
| Clinicopathological
parameters | No. | Runx3
mRNA | P-value |
|---|
|
|---|
| ≤0.362 | >0.362 |
|---|
| Gender | | | | >0.050 |
| Male | 69 | 38 | 31 | |
| Female | 32 | 12 | 20 | |
| Age (years) | | | | >0.050 |
| <60 | 46 | 21 | 25 | |
| ≥60 | 55 | 29 | 29 | |
| Growth pattern | | | | <0.050 |
| Expansive | 39 | 24 | 15 | |
| Infiltrative | 62 | 41 | 21 | |
| Histological
grade | | | | >0.050 |
| WD and MD | 55 | 31 | 24 | |
| PD | 46 | 20 | 26 | |
| Infiltrative
depth | | | | <0.050 |
| T1+T2 | 44 | 29 | 15 | |
| T3+T4 | 56 | 32 | 24 | |
| LN metastasis | | | | <0.001 |
| Absence | 27 | 18 | 9 | |
| Presence | 74 | 53 | 21 | |
| Distant
metastasis | | | | <0.001 |
| Absence | 73 | 42 | 31 | |
| Presence | 28 | 17 | 11 | |
| TNM stage | | | | <0.001 |
| I and II | 24 | 19 | 5 | |
| III and IV | 77 | 55 | 22 | |
RNA extraction and RT-PCR
LN tissues were homogenized with an ultrasound
homogenizer. Total RNA was extracted using TRIzol reagent
(Invitrogen, Carlsbad, CA, USA) according to the manufacturer's
instructions. Total RNA (1 μg) was reversely transcribed into cDNA
using dNTPs (1 mM), 5X reverse transcription buffer (500 mM
Tris-HCL pH 8.3, 250 mM KCL, 50 mM MgCl2 and 50 mM DTT),
16 units RNasin and 2.5 units AMV reverse transcriptase (Gibco BRL,
Life Technologies, Carlsbad, CA, USA). PCR conditions were:
pre-heating at 95°C for 5 min followed by 35 cycles of denaturation
for 30 sec at 95°C, annealing for 1 min at 55°C and extension for 1
min at 72°C, with a final extension for 5 min at 72°C. PCR products
were separated on 1.5% agarose gel and saved as digital images
(Perkin-Elmer, Wellesley, MA, USA). These experiments were
performed in triplicate and the mean value was calculated. The
value was normalized as the target gene divided by β-actin. The
primers used were: Runx3 gene, forward
5′-ATGACGAGAACTACTCCGCT-3′ and reverse 5′-GGTCGGAGAATGGGTTCAGT-3′
(PCR product, 396 bp).
Western blotting
The LN homogenates were heated in boiling water for
5 min. Protein concentrations were measured using Bradford's method
(Bio-Rad, Hercules, CA, USA). Protein (50 μg) was loaded, separated
by 10% SDS-polyacrylamide gel electrophoresis under reducing
conditions, and transferred onto equilibrated polyvinylidene
difluoride membrane (PVDF; Amersham) by electronic transfer.
Membranes were blocked by 5% non-fat dried milk and then incubated
with an antibody against Runx3 (dilution 1:200) overnight at
4°C; incubation with the secondary antibody was 1 h at room
temperature, with three washes after each incubation. ECL reagents
were used to show the positive bands on the membrane.
Statistical analysis
Statistical analysis was performed using SPSS 17.0
software. Continuous variables were determined as the means ± SD
and compared using the two-tailed version of Fisher's exact test.
The correlation was evaluated using regression analysis. Survival
was analyzed by the Kaplan-Meier method, and differences in the
distribution were evaluated using the log-rank test. P<0.05 was
considered to be statistically significant.
Results
Expression of Runx3 in LN specimens
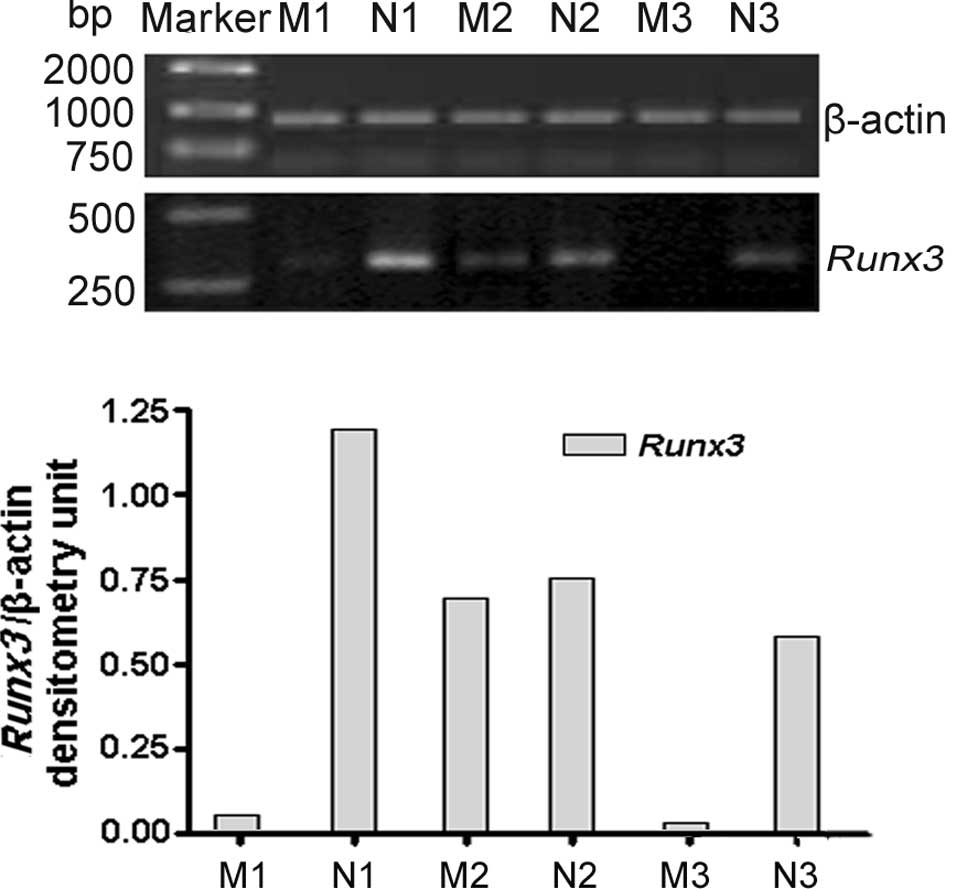
As shown in Fig. 1,
Runx3 mRNA was examined in 74 LNs with metastasis and 27 LNs
without metastasis. RT-PCR results revealed that Runx3 was
positively expressed in 28.4% (21 out of 74) LNs with metastasis
and 33.3% (9 out of 27) without metastasis (P<0.001). The
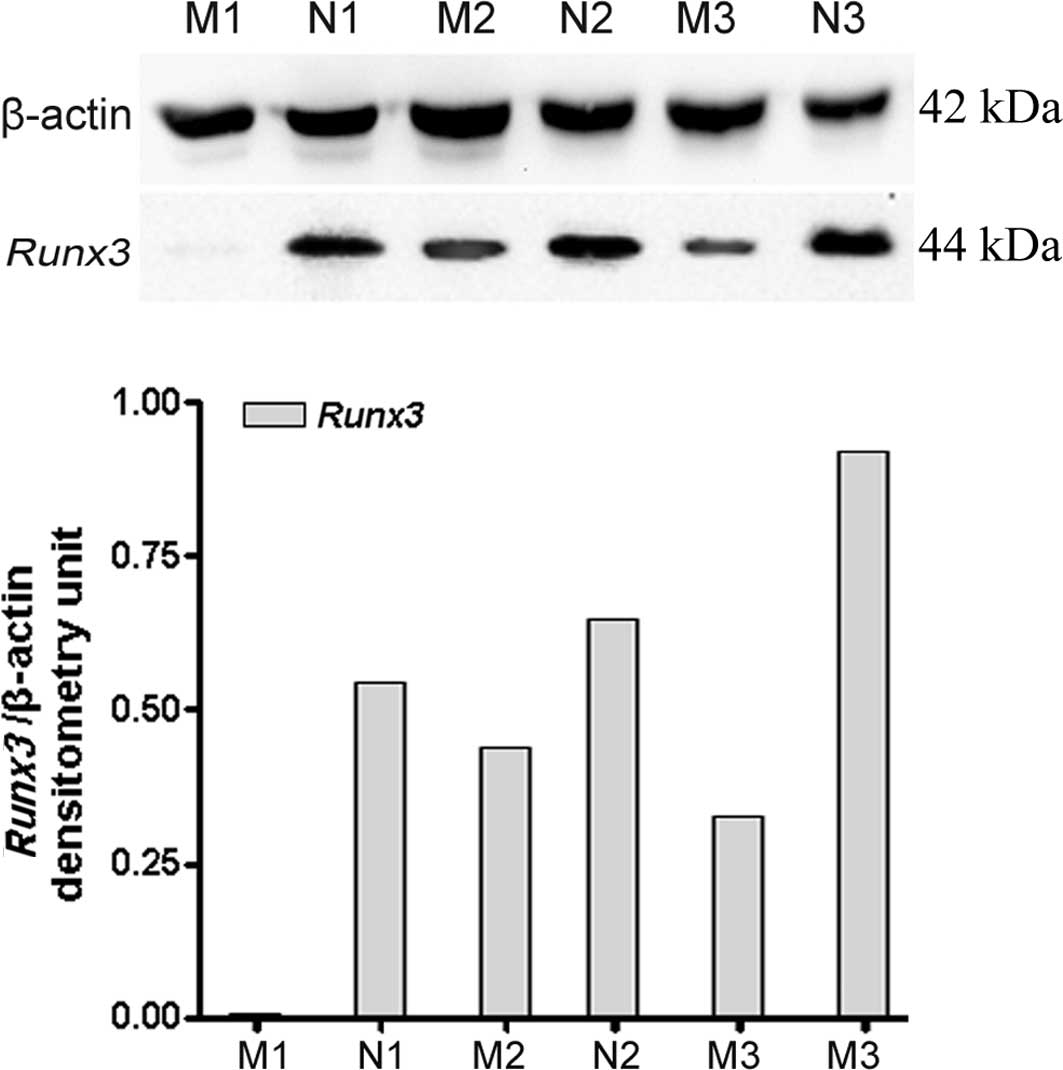
expression of Runx3 protein in LNs was further confirmed by
immunoblotting, which revealed a 44-kDa protein band (Fig. 2). Similarly, the results also
revealed that expression of the Runx3 protein was
significantly lower in LNM tissues than in those without
metastasis. A consistent correlation was found between the protein
and mRNA expression of Runx3.
Relationship between the expression
levels of Runx3 and clinicopathological parameters
The clinicopathological characteristics of the
patients and associations with Runx3 mRNA expression in LNs
are shown in Table I. The gastric
cancer patients (69 males and 32 females with a median age of 68.6
years; range 45–88) had undergone gastrectomy and lymphadenectomy.
Seventy-four patients were diagnosed with LNM. Thirty-nine patients
had an expansive growth pattern, whereas 62 had an infiltrative
classification. With respect to TNM tumor staging, gastric cancer
LNs with a higher expression of Runx3 were 20.8% (5 out of
24) at stages I and II, and 28.6% (22 out of 77) at stages III and
IV, respectively. Univariate analysis showed that the level of
Runx3 mRNA expression was inversely correlated with advanced
clinical stage (P<0.001), distant metastasis (P<0.05), LNM
(P<0.001), infiltrative depth (P<0.05) and histological grade
(P<0.001). However, no significant associations were observed
with patient age or gender (P>0.05), or with the growth pattern
of the tumor (P>0.05) (Table
I).
Survival analysis
The patients had a median follow-up duration of 35.6
months (range 3–84) following radical gastrectomy. Seventy-nine
patients succumbed due to recurrence or metastasis of the tumor and
22 patients survived. According to the median of the gray scale
bands (0.362; range 0.088–1.016) for Runx3 mRNA expression
in LNs, the 101 cases were separated into two groups: the
lower-expressing group (≤0.362) and the over-expressing group
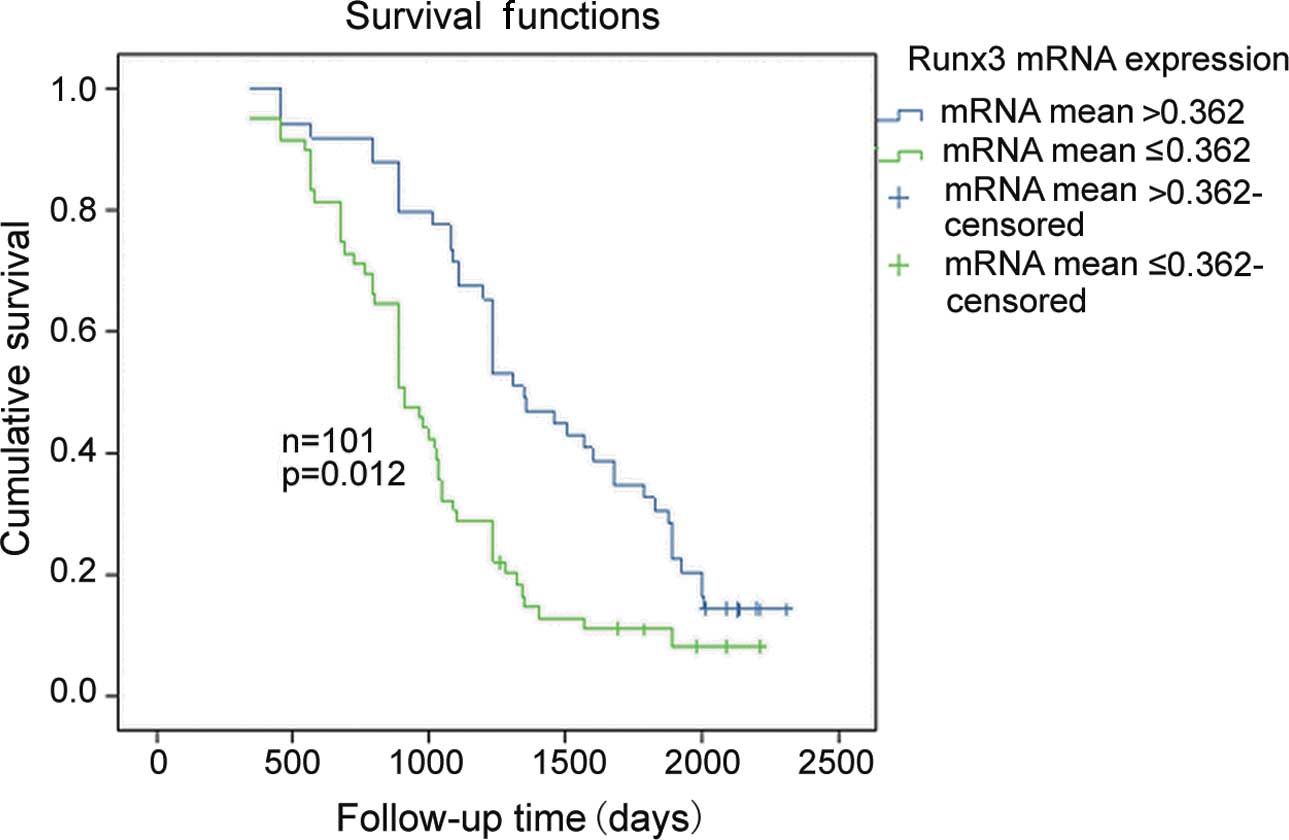
(>0.362). Kaplan-Meier analysis confirmed that patients with a
higher Runx3 expression (>0.362) had significantly better
survival outcomes compared to the lower group (≤0.362). The median
OS for the high and low Runx3 mRNA expression groups were
1,234 and 1,021 days, with a 5-year survival rate of 16 and 28%,
respectively (log-rank test, P=0.012; Fig. 3).
Discussion
In the present study, Runx3 expression was
first examined at the RNA and protein levels in the LN tissues of
gastric cancer using RT-PCR and Western blotting. The results
indicate that the expression of Runx3 mRNA was more frequent
in the LNs without metastasis than in the metastatic regions.
Western blotting demonstrated similar findings. Loss of
Runx3 expression was associated with a lower 5-year survival
rate and poorer prognosis of patients.
Runx3 is a transcription factor that
regulates lineage- specific gene expression in developmental
processes and is involved in the formation of a variety of cancers.
Runx3 elicits its tumor suppressor role by controlling the
expression of numerous genes involved in the growth, apoptosis and
differentiation of gastric epithelial cells (15–19),
as well as genes involved in angiogenesis and cell junction
formation (20,21). Mounting evidence suggests that
Runx3 is a tumor suppressor. Runx3 is expressed in
glandular stomach epithelial cells, and loss of Runx3
expression is causally related to the genesis and progression of
gastric cancer and correlates with differentiation, metastasis and
poor prognosis of gastric cancer (22).
In primary gastric cancer, much is known about the
expression of Runx3, but less is known about the
relationship between Runx3 expression in LNs and the
prognosis of patients. In this study, various clinicopathological
factors were analyzed for Runx3 expression in LNs. A low
proportion of Runx3 mRNA expression in LNM was found to be
significantly associated with poor clinicopathological factors,
such as deep infiltration, distant organ metastasis, poor
differentiation, LNM and later clinicopathological stages. However,
no significant associations were observed with age, gender and the
growth pattern of the tumor (Table
I). Runx3 expression was also associated with a lower
5-year survival rate and poorer prognosis of patients (Fig. 3). These results support our previous
studies (23), which revealed that
the low expression of Runx3 in primary gastric cancer was
associated with a significantly shorter survival.
LNM is one of the most significant factors
predicting recurrence in patients who have undergone gastrectomy
for stomach carcinoma (24,25). In China, a gastrectomy along with an
extended LN dissection is an established procedure and widely
accepted as the standard for the surgical treatment of gastric
cancer. However, certain patients develop local or distant tumor
recurrence even if a curative resection of the primary tumor is
performed. In our studies, the rate of LNM in gastric cancer was
73.7%, which is higher than that previously reported in Japan and
similar to reports from China (26,27).
This finding may be explained due to the bias of histological
criteria employed in our study and those used in Japan. We also
found that the presence of metastasis is significantly correlated
with the postoperative prognosis of patients with gastric cancer
(data not shown).
Concerning LNM, Park et al (28) reported MGMT expression to be
significantly associated with LNM in patients with gastric cancer.
Gene analysis along with a protein examination is recommended to
increase the positive detection of LNM and prognosis, leading to a
more accurate diagnosis and therapy of the tumor in gastric cancer.
Furthermore, such an analysis may positively contribute to the
selection of optimal chemotherapeutic regimens based on the gene.
Compared to the data generated in this study, Runx3 gene
expression in the LNs was significantly associated with poor
clinicopathological factors and poorer prognosis of patients.
Consequently, a gene expression analysis using LN tissue specimens
may aid in the detection of the survival rate and prognosis of
stomach carcinoma. However, the precise link between Runx3
expression in LNs and LNM remains unclear and further biological
studies are required to explain this effect (29).
In conclusion, a loss or substantial decrease of
Runx3 expression was observed in the group presenting with
LN with metastasis as compared to the group without metastasis, and
a low expression of Runx3 was significantly associated with
unfavorable clinicopathological variables and a shorter survival in
gastric carcinoma patients. LNM is also associated with a poorer
prognosis of gastric cancer. It may contribute to the detection of
gene expression and gastric cancer metastasis as a potential
molecular marker and a potential target in the therapy for gastric
cancer.
Acknowledgements
This study was supported by grants from the National
Natural Science Foundation of China (no. 30171039) and the Natural
Science Foundation of Jiangsu Province (no. BK2008115).
References
|
1
|
Kamangar F, Dores GM and Anderson WF:
Patterns of cancer incidence, mortality, and prevalence across five
continents: defining priorities to reduce cancer disparities in
different geographic regions of the world. J Clin Oncol.
24:2137–2150. 2006. View Article : Google Scholar
|
|
2
|
Rohatgi PR, Yao JC, Hess K, et al: Outcome
of gastric cancer patients after successful gastrectomy: influence
of the type of recurrence and histology on survival. Cancer.
107:2576–2580. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Nashimoto A, Nakajima T, Furukawa H, et
al: Randomized trial of adjuvant chemotherapy with mitomycin,
Fluorouracil, and Cytosine arabinoside followed by oral
Fluorouracil in serosa-negative gastric cancer: Japan Clinical
Oncology Group 9206-1. J Clin Oncol. 21:2282–2287. 2003. View Article : Google Scholar
|
|
4
|
Marrelli D, De Stefano A, de Manzoni G,
Morgagni P, Di Leo A and Roviello F: Prediction of recurrence after
radical surgery for gastric cancer: a scoring system obtained from
a prospective multicenter study. Ann Surg. 241:247–255. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Deng JY, Sun D, Liu XY, Pan Y and Liang H:
STAT-3 correlates with lymph node metastasis and cell survival in
gastric cancer. World J Gastroenterol. 16:5380–5387. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Tsang YH, Lamb A, Romero-Gallo J, et al:
Helicobacter pylori CagA targets gastric tumor suppressor
RUNX3 for proteasome-mediated degradation. Oncogene. 29:5643–5650.
2010. View Article : Google Scholar
|
|
7
|
Ito Y: Runx genes in development and
cancer: regulation of viral gene expression and the discovery of
Runx family genes. Adv Cancer Res. 99:33–76. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Ito K, Lim AC, Salto-Tellez M, et al:
Runx3 attenuates β-catenin/T cell factors in intestinal
tumourigenesis. Cancer Cell. 14:226–237. 2008.
|
|
9
|
Soong R, Shah N, Peh BK, et al: The
expression of Runx3 in colorectal cancer is associated with disease
stage and patient outcome. Br J Cancer. 100:676–679. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Mabuchi M, Kataoka H, Miura Y, et al:
Tumor suppressor, AT motif binding factor 1 (ATBF1), translocates
to the nucleus with runt domain transcription factor 3 (RUNX3) in
response to TGF-beta signal transduction. Biochem Biophys Res
Commun. 398:321–325. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Wei D, Gong W, Oh SC, et al: Loss of RUNX3
expression significantly affects the clinical outcome of gastric
cancer patients and its restoration causes drastic suppression of
tumor growth and metastasis. Cancer Res. 65:4809–4816. 2005.
View Article : Google Scholar
|
|
12
|
Li WQ, Pan KF, Zhang Y, et al:
Relationship between Runx3 expression and precancerous gastric
lesions in high-risk population. Beijing Da Xue Xue Bao.
41:348–352. 2009.PubMed/NCBI
|
|
13
|
Li QL, Ito K, Sakakura C, et al: Causal
relationship between the loss of Runx3 expression and gastric
cancer. Cell. 109:113–124. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Li WQ, Pan KF, Zhang Y, et al: RUNX3
methylation and expression associated with advanced precancerous
gastric lesions in a Chinese population. Carcinogenesis.
32:406–410. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Yano T, Ito K, Fukamachi H, et al: The
RUNX3 tumor suppressor upregulates Bim in gastric epithelial cells
undergoing transforming growth factor beta-induced apoptosis. Mol
Cell Biol. 26:4474–4488. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Ito K, Liu Q, Salto-Tellez M, et al:
RUNX3, a novel tumor suppressor, is frequently inactivated in
gastric cancer by protein mislocalization. Cancer Res.
65:7743–7750. 2005.PubMed/NCBI
|
|
17
|
Nakanishi Y, Shiraha H, Nishina SI, et al:
Loss of runt-related transcription factor 3 expression leads
hepatocellular carcinoma cells to escape apoptosis. BMC Cancer.
11:32011. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Subramaniam MM, Chan JY, Yeoh KG, Quek T,
Ito K and Salto-Tellez M: Molecular pathology of RUNX3 in human
carcinogenesis. Biochim Biophys Acta. 1796:315–331. 2009.PubMed/NCBI
|
|
19
|
Ogasawara N, Tsukamoto T, Mizoshita T, et
al: RUNX3 expression correlates with chief cell differentiation in
human gastric cancers. Histol Histopathol. 24:31–40.
2009.PubMed/NCBI
|
|
20
|
Peng Z, Wei D, Wang L, et al: RUNX3
inhibits the expression of vascular endothelial growth factor and
reduces the angiogenesis, growth, and metastasis of human gastric
cancer. Clin Cancer Res. 12:6386–6394. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Chang TL, Ito K, Ko TK, et al: Claudin-1
has tumor suppressive activity and is a direct target of RUNX3 in
gastric epithelial cells. Gastroenterology. 138:255–265. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Hsu PI, Hsieh HL, Lee J, et al: Loss of
RUNX3 expression correlates with differentiation, nodal metastasis,
and poor prognosis of gastric cancer. Ann Surg Oncol. 16:1686–1694.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Xu HW, Ren F, Chen W and Wang YJ: The
influence of survival analysis on RUNX3 gene expression in the
primary tumor of patients suffering from stomach carcinoma. Cancer
Research on Prevention and Treatment. 2011.(E-pub ahead of
print).
|
|
24
|
Ikeda Y, Saku M, Kishihara F and Maehara
Y: Effective follow-up for recurrence or a second primary cancer in
patients with early gastric cancer. Br J Surg. 92:235–239. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Furukawa H, Imamura H and Kodera Y: The
role of surgery in the current treatment of gastric carcinoma.
Gastric Cancer. 5:13–16. 2002. View Article : Google Scholar
|
|
26
|
Zhang XP, Wang ZL, Tang L, Sun YS, Cao K
and Gao Y: Support vector machine model for diagnosis of lymph node
metastasis in gastric cancer with multidetector computed
tomography: a preliminary study. BMC Cancer. 11:102011. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Tajima Y, Murakami M, Yamazaki K, et al:
Risk factors for lymph node metastasis from gastric cancers with
submucosal invasion. Ann Surg Oncol. 17:1597–604. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Park TJ, Han SU, Cho YK, Paik WK, Kim YB
and Lim IK: Methylation of O6-methylguaine-DNA methyltransferase
gene is associated significantly with K-ras mutation, lymph node
invasion, tumor staging, and disease free survival in patients with
gastric carcinoma. Cancer. 92:2760–2768. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Chen Y, Wei X, Guo C, et al: Runx3
suppresses gastric cancer metastasis through inactivation of MM9 by
up-regulation of TIMP-1. Int J Cancer. Dec;2010.(E-pub ahead of
print).
|