Introduction
Renal cell carcinoma is the second leading cause of
mortality among urinary tumors, accounting for 2% of adult
malignancies, of which the most common subtype is clear cell renal
cell carcinoma (ccRCC) (1). ccRCC
affects approximately 150,000 individuals each year and causes
approximately 78,000 mortalities globally with increasing incidence
and mortality. More than one-third of patients may have metastasis
when diagnosed, and 50% of patients may suffer recurrence even
after nephrectomy (2). In recent
years, specific prognostic markers for ccRCC, including Ki-67,
MCM2, SAT1, L1CAM and BIRC5, have emerged, but large-scale clinical
application is impossible. Despite the continual progress in
medical technology, the clinical characteristics of ccRCC remain
difficult to predict (3).
Therefore, novel diagnostic and prognostic markers of ccRCC could
be valuable in high-risk individuals and those with existing
disease.
Protein tyrosine phosphorylation plays a important
role in signal transduction. This process contains numerous
complicated processes. A recent study revealed the corelations
between abnormal protein tyrosine phosphorylation and
tumorigenesis, invasion and metastasis (4). Therefore, studies on tyrosine kinases
could be significant for acquiring insights into the underlying
mechanisms of oncogenesis, distant metastasis and recurrence.
FER, a 94-kDa protein, belongs to the subfamily of
non-receptor protein tyrosine kinases. Similar to other tyrosine
kinases, FER has a central Src-homology 2 (SH2) domain involved in
binding to phosphotyrosine-containing peptide sequences and a
highly conserved C-terminal kinase domain. It is distinguished from
other tyrosine kinases by an NH2-terminal FER/ClP4
homology and adjacent coiled-coil domains (5,6). FER
exists ubiquitously in human cells and participates in the
signaling processes of cell proliferation, differentiation,
apoptosis, movement and adhesion (5). However, a number of studies have
identified higher levels of FER in cancer cells compared with
adjacent normal cells (7,8); the underlying mechanisms of which have
remained unclear to date. Moreover, the correlation between FER
expression and prognosis status remains ambiguous. In the present
study, we examine the expression and clinical significance of FER
in ccRCC and explore the association between FER expression level
and prognosis status.
Materials and methods
Patients and tissue samples
The study was approved by the institutional review
board and ethical committee of Peking University Shenzhen Hospital,
Shenzhen, China. All patients participated after providing written
informed consent. For real-time qPCR, 48 paired samples of ccRCCs
and normal adjacent tissues (ADTs) were collected from patients who
underwent radical nephrectomy at the Department of Urology, Peking
University Shenzhen Hospital between November 2009 and December
2011. The ADTs were located 2.0 cm away from visible ccRCC lesions.
The fresh tissue samples were immediately immersed in RNAlater
(Qiagen, Hilden, Germany) following surgical resection and stored
at 4°C overnight and then frozen in liquid nitrogen and stored at
−80°C until analysis.
For immunohistochemical analysis of FER protein, a
total of 206 paraffin-embedded samples of pathologically verified
ccRCC and 87 adjacent normal renal tissue samples were collected.
All patients received radical nephrectomy at the Department of
Urology, Sun Yat-Sen University Cancer Center, Guangzhou, China
between June 2000 and September 2010. The histological and clinical
diagnoses of the tumors in all patients were performed by the
Pathology Department of Sun Yat-Sen University Cancer Center. The
characteristics of the 206 patients are listed in Table I. The survival information of the
206 patients was collected over the telephone. The clinical
characteristics of the patients were obtained from patient medical
records. Tumor stage was reclassified based on the 2011 Union for
International Cancer Control (UICC) TNM classification of malignant
tumors, and nuclear grading was performed according to Fuhrman’s
system (9).
 | Table I.Association between FER and
clinicopathological characteristics in ccRCC patients. |
Table I.
Association between FER and
clinicopathological characteristics in ccRCC patients.
| | FER
| | |
|---|
| Parameters | n | High | Low | χ2 | P-value |
|---|
| Total | 206 | 136 | 70 | 0.168 | 0.682 |
| Gender | | | | | |
| Male | 145 | 97 | 48 | | |
| Female | 61 | 39 | 22 | | |
| Age (years) | | | | | |
| ≥50 | 137 | 94 | 43 | 1.227 | 0.268 |
| <50 | 69 | 42 | 27 | | |
| Size (cm) | | | | 8.161 | 0.004 |
| ≥7 | 84 | 65 | 19 | | |
| <7 | 122 | 71 | 51 | | |
| T stage | | | | 8.542 | 0.014 |
| T1 | 139 | 87 | 52 | | |
| T2 | 34 | 20 | 14 | | |
| T3/4 | 33 | 29 | 4 | | |
| N stage | | | | 6.131 | 0.013 |
| N0 | 177 | 111 | 66 | | |
| N+ | 29 | 25 | 4 | | |
| Metastasis | | | | 6.680 | 0.010 |
| No (M0) | 180 | 113 | 67 | | |
| Yes (M1) | 26 | 23 | 3 | | |
| Recurrence | | | | 8.959 | 0.003 |
| No | 172 | 106 | 66 | | |
| Yes | 34 | 30 | 4 | | |
| Fuhrman | | | | 12.374 | 0.006 |
| 1 | 37 | 18 | 19 | | |
| 2 | 124 | 81 | 43 | | |
| 3 | 25 | 20 | 5 | | |
| 4 | 20 | 18 | 2 | | |
Real-time qPCR
Total RNA was extracted from 48 paired ccRCC samples
and normal tissue by TRIzol (Invitrogen Life Technologies,
Carlsbad, CA, USA) according the manufacturer’s instructions. Then
Omniscript RT kit (Qiagen) was used to synthesize the first-strand
cDNA. The total reaction volume was 20 μl including 1
μg RNA, and the reaction mixture was incubated at 42°C for
60 min, heated at 95°C for 10 min and then cooled on ice. The RNA
and cDNA were evaluated using an Agilent 2100 Bioanalyzer (Agilent
Technologies, Santa Clara, CA, USA). The corresponding primer
sequences were as follows: FER sense primer,
5′-TTCGAGGGCACTGGGTTTTC-3′; reverse primer,
5′-TTCCCTTGCCCAGTAATTCTCC-3′. GAPDH sense primer,
5′-GGAGTCCACTGGCGTCTTCACC-3′; reverse primer,
5′-GAGGAGTGGGTGTCGCTGTTG-3′. Real-time PCR was conducted using SYBR
Green dye in a 7000 Sequence Detection System (Applied Biosystems,
Carlsbad, CA, USA). The 20-μl real-time PCR mixture
contained 1 μl of cDNA (synthesized as described above), 10
μl SYBR Green Master mix (Invitrogen) and 1 μl of
each upstream and downstream primer. The amplification conditions
were 95°C (2 min) for 1 cycle and 95°C (5 sec), 57°C (30 sec) and
68°C (30 sec) for 40 cycles. Relative expression levels of FER were
normalized to the geometric mean of GAPDH (internal control gene).
The data were analysed via the comparative threshold cycle
(2−ΔCT, −
ΔCT=CTFER−CTGAPDH) method
(10).
Immunohistochemical analysis
Immunohistochemistry was performed to examine FER
expression in the 206 ccRCC samples and 87 paired samples of ADTs.
All procedures were performed using standard techniques. Briefly,
paraffin-embedded specimens were cut into 5-μm sections and
heated at 65°C for 30 min. The sections were deparaffnized in
xylene and rehydrated in a descending ethanol series. Endogenous
peroxidase activity was blocked with 3% hydrogen peroxide for 20
min. The sections were boiled in 10 mmol/l citrate buffer (pH 6.0)
to unmask the epitopes.Non-specific protein binding was performed
by incubations with 10% bovine serum albumin for 30 min. For the
detection of FER, the sections were incubated with the polyclonal
rabbit anti-human FER antibody (Abcam, Cambridge, MA, USA) diluted
at 1:400, and then incubated overnight at 4°C. The negative control
was performed by replacing the primary antibody with antibody
diluent. They were then rinsed with PBS and incubated using a
anti-Rabbit Immunohistochemistry kit (Maixin Bio., Fujian, China)
at 37°C for 20 min. After rinsing with PBS, the tissue sections
were stained for 5 min with 3,3′-diaminobenzidine
tetrahydrochloride (DAB), counterstained with hematoxylin,
dehydrated and then mounted in Crystal Mount (Maixin Technologies,
Fuzhou, China).
Staining evaluation
The stained sections were reviewed by two
independent observers who had no prior knowledge of the
clinicopathological data of the patients. The scoring was based
mainly on color intensity and extensity. The proportion of cells
expressing FER varied from 0 to 100%, and the intensity of staining
varied from weak to strong. The proportion of FER expression tumour
cells was scored at low magnification on a scale of 0–5 (0, no
cells positive; 1, 0–5% of cells positive; 2, 6–25%; 3, 26–50%; 4,
51–75%; 5, 76–100%). The intensity score was determined at high
magnification on a scale of 0–3 (0, negative staining; 1, weakly
positive staining; 2, moderately positive staining; 3, strongly
positive staining), and the final score was calculated by the
multiplication of the two parameters, with scores of 0, 1, 2, 3, 4,
5, 6, 8, 9, 10, 12 and 15. We set the optimal cut-off values for
FER level by measuring heterogeneity in overall survival rates via
the log-rank test method, and designated low expression as total
score <5, high expression as total score ≥5. Thus, the stained
sections were divided into two different groups.
Statistical analysis
Paired-sample t-tests were used in the RT-PCR and
immunohistochemistry assays to analyze the significance of the
differences in mRNA level and protein expression between ccRCCs and
the adjacent normal tissues. The correlation between FER expression
and clinical and pathological characteristics was assessed using
the χ2 test. Survival curves were plotted according to
the Kaplan-Meier method and compared by the log-rank test.
Multivariate analysis according to Cox’s proportional hazards
regression model adjusted for clinicopathological factors (age,
gender, tumor size, Fuhrman grade, TNM stage and FER expression)
was performed to assess which clinical variables were independently
correlated with overall survival. Statistical analysis was
performed using the SPSS 17.0 package. P<0.05 was considered to
indicate a statistically significant difference.
Results
RT-PCR analysis of FER mRNA in 48 ccRCC
tumor samples
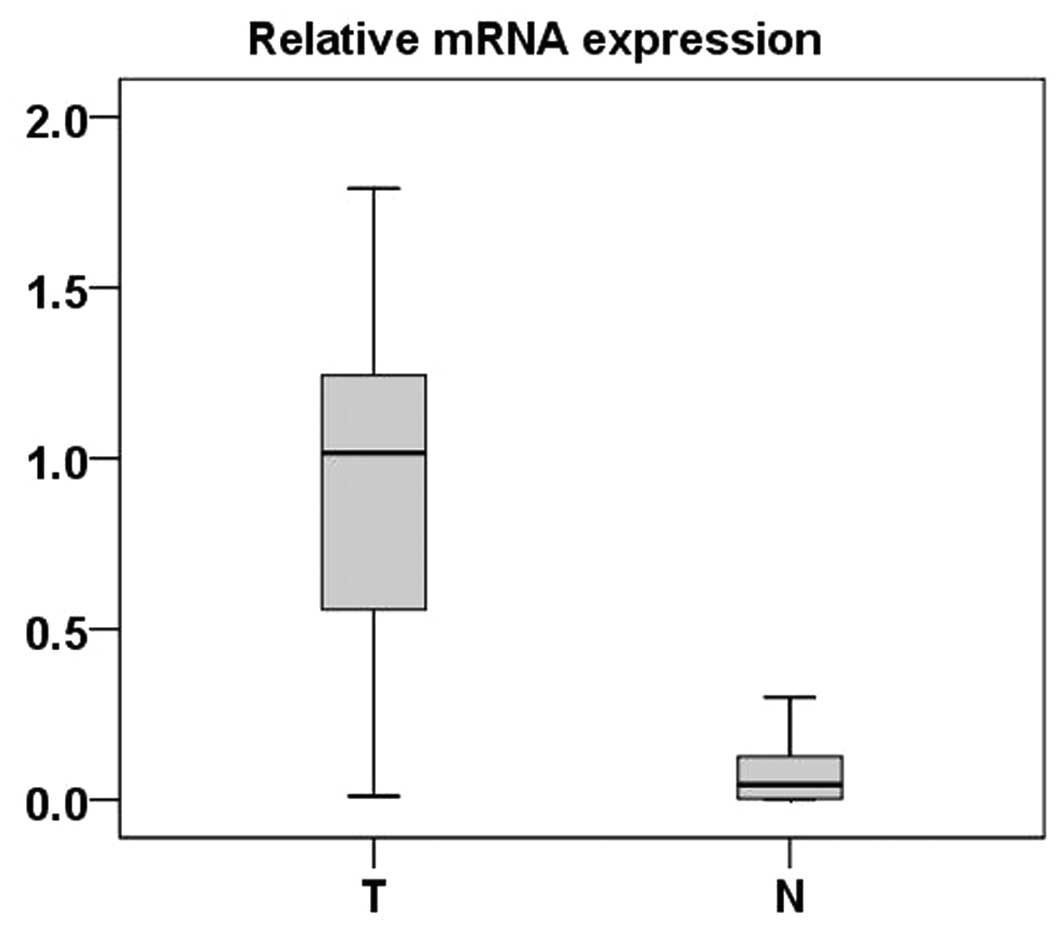
Real-time PCR was performed to measure the
expression of FER mRNA in 48 ccRCC tumor tissues and ADTs. Compared
with normal tissues, 46 ccRCC tumor tissues exhibited significantly
high expression of FER at the mRNA level (P<0.001, paired-sample
t-test) (Fig. 1).
Immunohistochemical analysis of FER
expression in 87 ccRCC samples and the paired ADTs
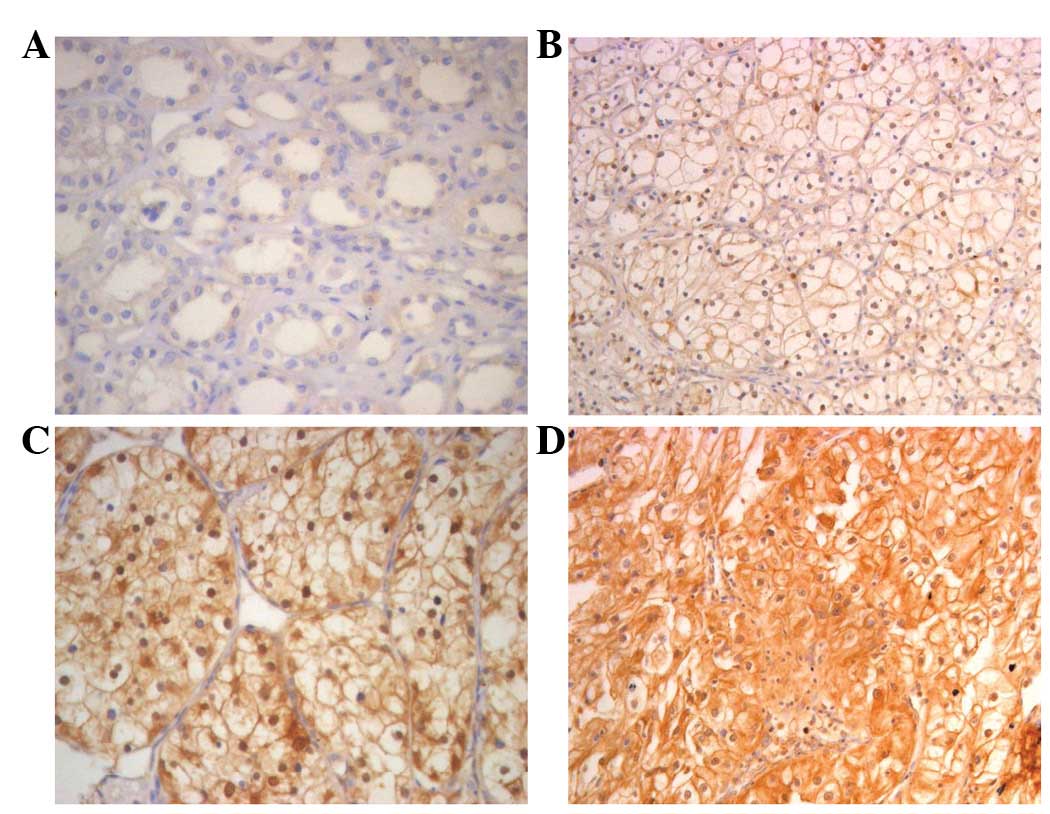
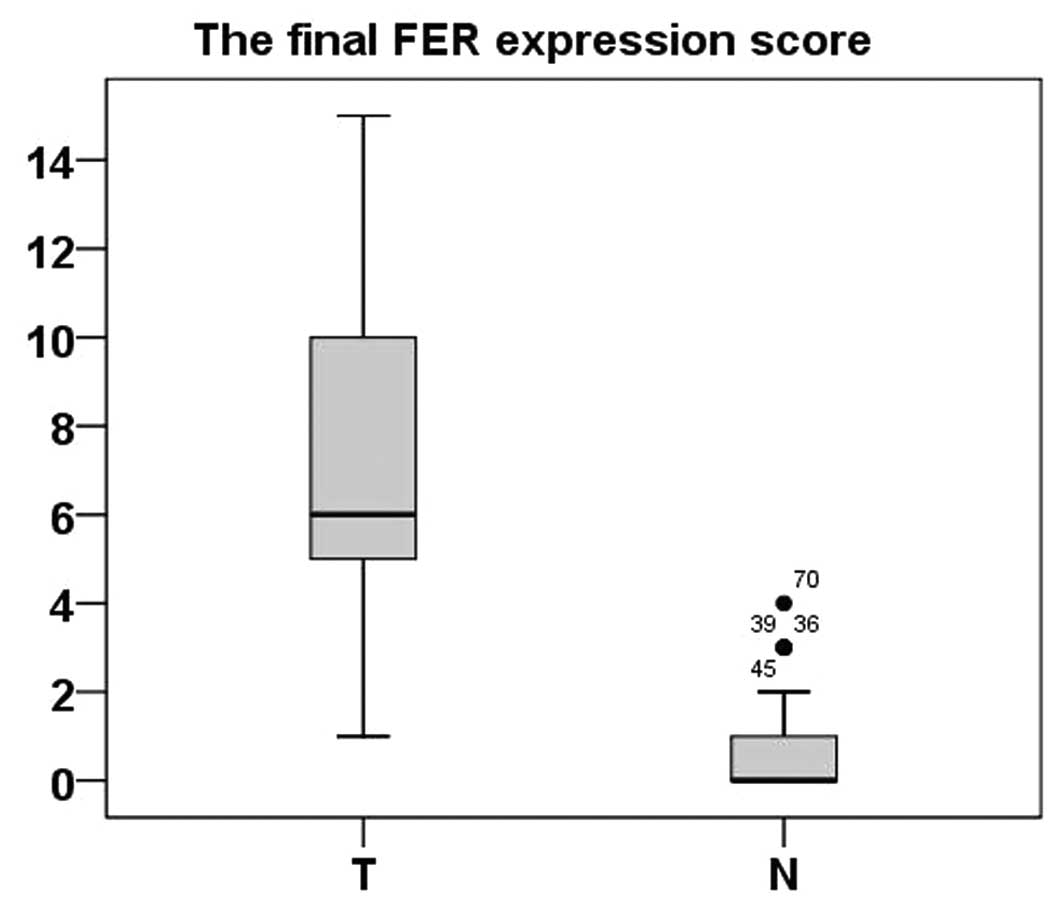
Immunohistochemistry was applied to assess the
expression and subcellular localization of FER protein in 87
paraffin-embedded ccRCC tissues and 87 paired ADTs. As shown in
Fig. 2, FER staining was present
mainly in the nuclei and cytoplasm (Fig. 2). In the normal renal tissues, FER
expression was negative (52/87; score, 0) or at a low level (35/87;
score, ≤5). FER expression in the 83 tumor tissue samples was
higher than the ADTs (P<0.001, paired-sample t-test) (Fig. 3).
Immunohistochemical analysis of the
correlation between FER protein expression and clinical features in
206 ccRCC tumor samples
Immunohistochemical analysis was performed in 206
paraffin-embedded ccRCC tumor samples to further assess the
correlation between FER expression and various clinicopathological
parameters. As shown in Table I,
low expression of FER (score, ≤4) was demonstrated in 70 of the 206
tumor samples, while high expression (score, ≥5) was demonstrated
in a further 136 samples. Increased expression of FER in tumor
samples was correlated with tumor size (χ2=8.161;
P=0.004), T stage (χ2=8.542; P=0.014), N classification
(χ2=6.131; P=0.013), metastasis (χ2=6.680;
P=0.010), recurrence (χ2=8.959; P=0.003) and Fuhrman
grade (χ2=12.374; P=0.006), while associations with age
(χ2=1.227; P=0.268) and gender (χ2=0.168;
P=0.682) were not identified. High expression of FER was revealed
in 45.8, 58.8 and 87.9% of T1, T2, and T3/4 stage ccRCCs,
respectively (P=0.014; χ2 test). High expression of FER
was observed in 58.2 and 77.4% of ccRCCs with size <7 cm and ≥7
cm, respectively (P=0.004; χ2 test). High expression of
FER was observed in 62.7 and 86.2% of N0 and N1/2 stage ccRCCs,
respectively (P=0.013, χ2 test). High expression of FER
was observed in 88.5 and 62.8% of ccRCCs with or without metastasis
respectively (P=0.010, χ2 test). High expression of FER
protein was observed in 88.2 and 61.6% of ccRCCs with or without
recurrence, respectively (P=0.003; χ2 test).
Survival analysis
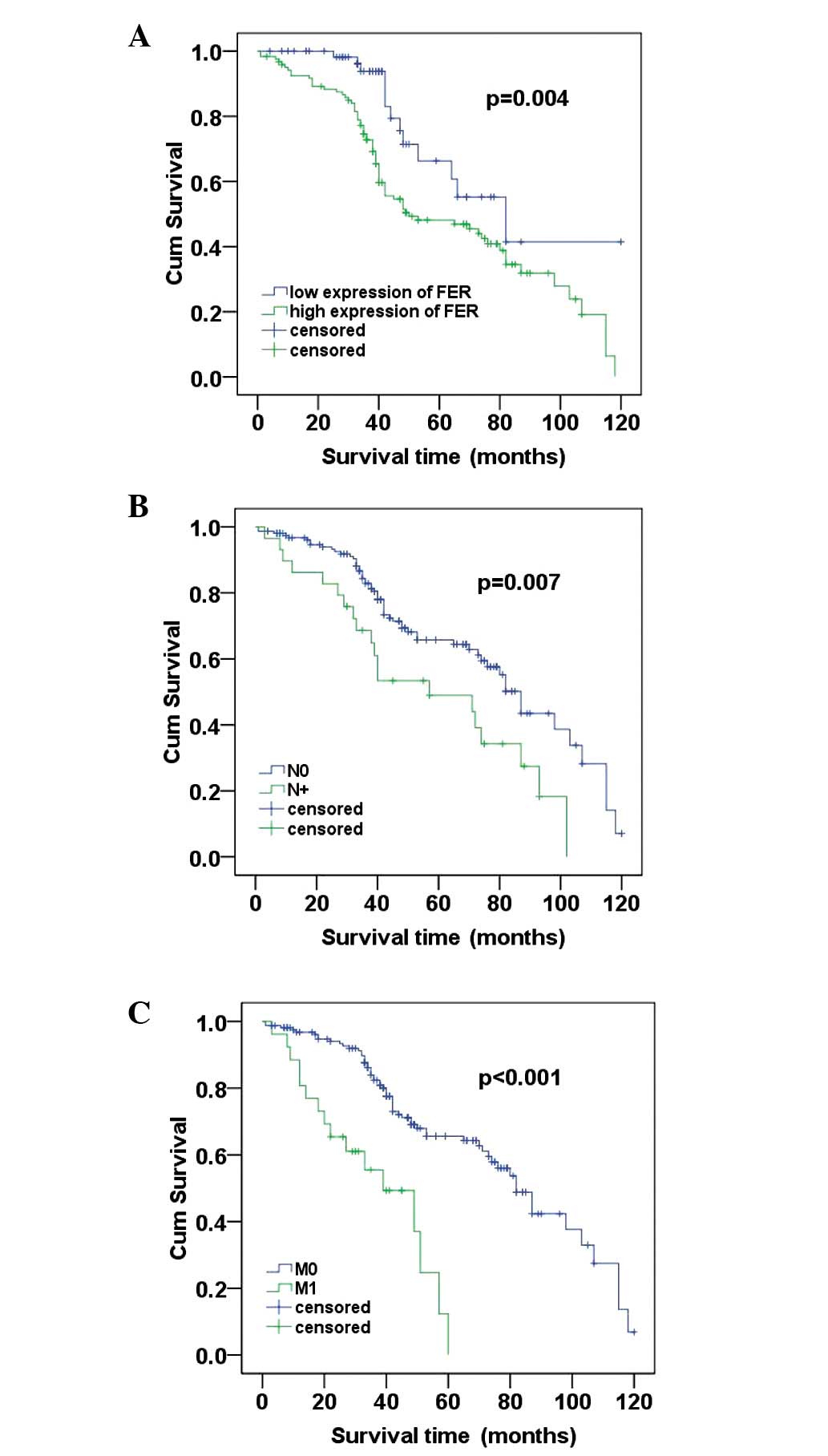
To further investigate the prognostic value of FER
expression in ccRCC, Kaplan-Meier analysis and the log-rank test
were applyed to assess the correlation between FER expression level
in ccRCC and prognosis status. We identified that the level of FER
expression was correlated with the overall survival of ccRCC
patients. Individuals with a higher level of FER expression had
poorer survival rates compared with those with a lower level. The
mean survival time in the group of highly expressed FER patients
was 64.840 months and the median survival time was 50 months, but
the mean and median survival time in the low expression group time
were 90.331 and 89 months, respectively. The results of the
log-rank test demonstrated that the survival rates were
significantly different between these two groups (log-rank,
P=0.004) (Fig. 4A). Furthermore,
patients with no regional lymph node involvement (N0) had a better
prognosis than those with regional lymph node involvement (N+;
log-rank, P=0.007; (Fig. 4B).
Similarly, patients with no metastasis (M0) had a high cumulative
survival rates compared with patients with metastasis (M1;
log-rank, P<0.001) (Fig.
4C).
In addition, the multivariate Cox regression
analysis indicated that FER expression (P=0.028), N stage (P=0.009)
and distant metastasis (P<0.001) were independent prognosis
factors for overall survival of ccRCC patients (Table II).
 | Table II.Multivariate Cox regression analysis
for the overall survival rates of ccRCC patients. |
Table II.
Multivariate Cox regression analysis
for the overall survival rates of ccRCC patients.
| Risk factors | Relative risk | 95% Confidence
interval | P-value |
|---|
| T stage | 1.291 | 0.545–1.143 | 0.266 |
| N stage | 1.993 | 1.185–3.417 | 0.009 |
| M stage | 4.257 | 2.264–9.863 | <0.001 |
| Age | 0.621 | 0.505–1.002 | 0.137 |
| Size | 0.840 | 0.603–1.578 | 0.740 |
| Gender | 0.714 | 0.481–1.173 | 0.306 |
| Fuhrman Grade | 9.108 | 1.398–7.601 | 0.122 |
| FER expression | 0.560 | 0.437–0.964 | 0.028 |
Discussion
ccRCC is the most common subtype of renal tumor,
accounting for 70–80% of all RCCs. Incidence of ccRCC has increased
markedly in the last two decades, and the annual mortality has
become significantly higher than other tumors in the genitourinary
tract (11).
The clinical outcome of ccRCC remains poor despite
advances in clinical technologies (12). The TNM staging system and other
clinicopathological parameters have limited value in predicting the
prognosis of individuals with ccRCC, and approximately 50% of those
patients with less advanced disease are thought to experience
metastasis even after nephrectomy (13). Therefore, it is extremely
significant to identify specific molecular biomarkers of ccRCC for
early diagnosis and evaluation of prognosis.
FER is a member of the non-receptor protein tyrosine
kinases subfamily. FER is expressed ubiquitously in a variety of
tissues and cells and is overexpressed at a markedly high level in
malignant tumors (5,7), including prostate cancer and
hepatocarcinoma (7,8). Overexpression of Drosophila FER
may induce rodent fibroblasts canceration (14). However, since its discovery in 1988
(15), studies on the functional
regulation mechanism of FER in malignancy have been relatively
limited, and the focus has been mainly set on cell adhesion. The
part FER plays in the regulation of integrin-mediated focal
adhesion and cadherin-mediated intercellular adhesion has been
confirmed. FER may promote CTNNB1 dephosphorylation and
E-cadherin-mediated adhesion stability under normal circumstances.
However, high expression of FER may directly induce CTNNB1
phosphorylation, resulting in a disintegration of cadherin-mediated
adhesion (16,17). FER may integrate with the integrin
and cadherin complex via phosphorylation of cortactin, a crucial
molecule in tumor cell metastasis (18,19).
Notably, FER dissociated from the cadherin complex could be
recruited to the integrin complex, leading to p130CAS
dephosphorylation and blocking of integrin-mediated adhesion
(17) .
In addition, overexpressed FER could phosphorylate
EGF receptor and activate the EGF-mediated NF-κB signaling pathway,
which is crucial for cancer cell survival and proliferation
(20). Downregulation of FER via
antisense cDNA impairs prostate cancer cells growth and colony
formation in vitro(21). RNA
interference against FER may also arrest the mitotic cycle in G0–G1
phasein vitro(22).
Thus a high level of FER may disintegrate
integrin/cadherin-mediated intercellular adhesion and promote
metastasis. In accordance with this, our results demonstrate a high
distant metastasis and recurrence rate in ccRCC patients with a
high FER level, which suggests a greater metastasis and invasion
ability in tumor cells with overexpressed FER.
To the best of our knowledge, this is the first
study to indicate the clinical significance of FER in ccRCC.
Real-time PCR in 48 ccRCC tumour tissues and paired ADT samples
revealed a significant increase in FER mRNA in ccRCC samples.
Further immunohistochemical analysis in 87 paired samples of ccRCCs
and ADTs confirmed overexpression of FER protein in tumor tissue.
These results indicate that FER may play important roles in the
initiation and progression of malignancies.
To further investigate the prognostic value of FER,
immunohistochemical analysis was performed to evaluate the
correlation between FER expression and various clinicopathological
parameters. In the present study, we demonstrated that an increased
level of FER expression was significantly correlated with tumor
size, Fuhrman grade, stage, N classification, metastasis and
recurrence. According to the results of Kaplan-Meier analysis, the
FER protein expression level in ccRCC was significantly correlated
with overall survival. Patients with a high FER expression level
had a shorter survival time than those with a low FER level. The
log-rank test revealed that the group with a lower expression of
FER had a more favorable prognosis than the higher expression
group. The TNM stage of ccRCC was closely correlated with prognosis
(23). Consistent with this, in
this study, FER expression, N classification and distant metastasis
were independent prognosis factors for overall survival of ccRCC
patients by multivariate Cox regression analysis. Therefore, this
study reveals that there are significant correlations between FER
expression level and clinicopathological parameters and may be a
potential prognostic marker and therapeutic target for ccRCC.
Fuhrman’s nuclear grading system is considered to be
a reliable prognostic indicator for ccRCC (9). However, the multivariate Cox
regression analysis in our study did not reveal any correlations
between Fuhrman grade and prognosis, this may due to our limited
sample size or observation error made by the pathologists.
This is the first study aimed at evaluating the
possibility of using FER as a clinically potential indicator for
disease progression, as well as a prognostic marker for patient
survival in ccRCC. However, it should be acknowledged that this
study was a single hospital-based, retrospective study, and
therefore, multicenter or community-based prospective studies are
required.
In conclusion, our findings indicate that the
expression levels of FER in ccRCC tissues are significantly higher
than those in ADTs. Moreover, high levels of FER were associated
with poor survival in patients with advanced ccRCC. Multivariate
survival analysis showed that FER is a potential prognostic marker
for ccRCC. FER, therefore, may provide a valuable marker in the
prognostic evaluation of ccRCC.
Acknowledgements
This study was supported by the
National Natural Science Foundation of China (No. 81101922), the
Biobank of Complex Diseases in Shenzhen (CXC201005260001A), the
Technology Program of Shenzhen (201003099) and the Technology
Program of Guangdong Province (2011B061300046). We thank the
Department of Urology, Sun Yat-Sen University Cancer Center for
providing paraffin-embedded samples and patient prognosis
information.
References
|
1.
|
Jemal A, Bray F, Center MM, Ferlay J, Ward
E and Forman D: Global cancer statistics. CA Cancer J Clin.
61:69–90. 2011. View Article : Google Scholar
|
|
2.
|
Zbar B, Klausner R and Linehan WM:
Studying cancer families to identify kidney cancer genes. Annu Rev
Med. 54:217–233. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
3.
|
Wei X, Zhou L, Hu L and Huang Y:
Tanshinone IIA arrests cell cycle and induces apoptosis in 786-O
human renal cell carcinoma cells. Oncol Lett. 3:1144–1148.
2012.PubMed/NCBI
|
|
4.
|
Chong PK, Lee H, Kong JW, Loh MC, Wong CH
and Lim YP: Phosphoproteomics, oncogenic signaling and cancer
research. Proteomics. 8:4370–4382. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
5.
|
Greer P: Closing in on the biological
functions of Fps/Fes and Fer. Nat Rev Mol Cell Biol. 3:278–289.
2002. View
Article : Google Scholar : PubMed/NCBI
|
|
6.
|
Tsujita K, Suetsugu S, Sasaki N, Furutani
M, Oikawa T and Takenawa T: Coordination between the actin
cytoskeleton and membrane deformation by a novel membrane
tubulation domain of PCH proteins is involved in endocytosis. J
Cell Biol. 172:269–279. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
7.
|
Pasder P, Salem Y, Yaffe E, Shpungin S and
Nir U: FER as a novel target for cancer therapy. Drugs Fut.
32:61–70. 2007. View Article : Google Scholar
|
|
8.
|
Li H, Ren Z, Kang X, Zhang L, Li X, Wang
Y, Xue T, Shen Y and Liu Y: Identification of
tyrosine-phosphorylated proteins associated with metastasis and
functional analysis of FER in human hepatocellular carcinoma cells.
BMC Cancer. 9:3662009. View Article : Google Scholar
|
|
9.
|
Fuhrman SA, Lasky LC and Limas C:
Prognostic significance of morphologic parameters in renal cell
carcinoma. Am J Surg Pathol. 6:655–663. 1982. View Article : Google Scholar : PubMed/NCBI
|
|
10.
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2-[Delta][Delta] CT method. Methods. 25:402–408. 2001.
|
|
11.
|
Schrader A, Sevinc S, Olbert P, Hegele A,
Varga Z and Hofmann R: Gender-specific characteristics and survival
of renal cell carcinoma. Urologe A. 47:1184–1186. 2008.PubMed/NCBI
|
|
12.
|
Lane BR, Babineau D, Kattan MW, Novick AC,
Gill IS, Zhou M, Weight CJ and Campbell SC: A preoperative
prognostic nomogram for solid enhancing renal tumors 7 cm or less
amenable to partial nephrectomy. J Urology. 178:429–434. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
13.
|
Tamaskar I, Choueiri TK, Sercia L, Rini B,
Bukowski R and Zhou M: Differential expression of caveolin-1 in
renal neoplasms. Cancer. 110:776–782. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
14.
|
Paulson R, Jackson J, Immergluck K and
Bishop JM: The DFer gene of Drosophila melanogaster encodes
two membrane-associated proteins that can both transform vertebrate
cells. Oncogene. 14:641–652. 1997.PubMed/NCBI
|
|
15.
|
Letwin K, Yee SP and Pawson T: Novel
protein-tyrosine kinase cDNAs related to fps/fes and eph cloned
using anti-phosphotyrosine antibody. Oncogene. 3:621–627.
1988.PubMed/NCBI
|
|
16.
|
Piedra J, Miravet S, Castaño J, Pálmer HG,
Heisterkamp N, García de Herreros A and Duñach M: p120
Catenin-associated Fer and Fyn tyrosine kinases regulate β-catenin
Tyr-142 phosphorylation and β-catenin-α-catenin Interaction. Mol
Cell Biol. 23:2287–2297. 2003.PubMed/NCBI
|
|
17.
|
Xu G, Craig AWB, Greer P, Miller M,
Anastasiadis PZ, Lilien J and Balsamo J: Continuous association of
cadherin with β-catenin requires the non-receptor tyrosine-kinase
Fer. J Cell Sci. 117:3207–3219. 2004.
|
|
18.
|
Daly RJ: Cortactin signalling and dynamic
actin networks. Biochem J. 382:13–25. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
19.
|
El Sayegh TY, Arora PD, Laschinger CA, Lee
W, Morrison C, Overall CM, Kapus A and McCulloch CA: Cortactin
associates with N-cadherin adhesions and mediates intercellular
adhesion strengthening in fibroblasts. J Cell Sci. 117:5117–5131.
2004.PubMed/NCBI
|
|
20.
|
Lu HY, Mao WM, Cheng QY, Chen B, Cai JF,
Wang XJ, Wang Z and Xie FJ: Mutation status of epidermal growth
factor receptor and clinical features of patients with combined
small cell lung cancer who received surgical treatment. Oncol Lett.
3:1288–1292. 2012.PubMed/NCBI
|
|
21.
|
Ben-Dor I, Bern O, Tennenbaum T and Nir U:
Cell cycle-dependent nuclear accumulation of the p94fer tyrosine
kinase is regulated by its NH2 terminus and is affected by kinase
domain integrity and ATP binding. Cell Growth Differ. 10:113–129.
1999.PubMed/NCBI
|
|
22.
|
Pasder O, Shpungin S, Salem Y, Makovsky A,
Vilchick S, Michaeli S, Malovani H and Nir U: Downregulation of Fer
induces PP1 activation and cell-cycle arrest in malignant cells.
Oncogene. 25:4194–4206. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
23.
|
Levi F, Ferlay J, Galeone C, Lucchini F,
Negri E, Boyle P and La Vecchia C: The changing pattern of kidney
cancer incidence and mortality in Europe. BJU Int. 101:949–958.
2008. View Article : Google Scholar : PubMed/NCBI
|