Introduction
Osteosarcoma (OS) is a malignant bone tumor
characterized by tumor cells directly forming bone or bone tissues.
It often occurs in adolescents or children under the age of 20 and
the prognosis is poor (1–3). The current treatment strategy focuses on
primary tumors, but it is limited in the treatment of metastatic OS
(4). Thus, treating metastatic OS is
now a great challenge for oncologists. It is necessary for us to
understand the pathogenesis and biological characteristics of
metastatic OS and to find new targets and biomarkers for the
treatment of OS patients is urgent (5,6).
It is known that, miRNAs can extensively disrupt
various functions of different cancers by targeting mRNA (7), and the target mRNA is also proven to
regulate cell proliferation, invasion and apoptosis (8). New evidence suggests that miRNAs are
involved in human carcinogenesis, including promoting or inhibiting
tumor development (9,10). For example, miR-365, miR-124a, miR-625
and miR-141 acted as tumor suppressors in OS (11–14), while
miR-504, miR-19 and miR-93 promoted OS cell progression (15–17).
The miR-29 family contains three mature members
(miR-29a, miR-29b and miR-29c). An increasing number of studies
have shown the decreased expression of miR-29 family in various
types of cancer and that it has the ability to suppress tumors,
including breast, bladder and pancreatic cancer (18–20). Shin
et al found that miR-29b effect on glioblastoma was
suppressive (21). Ma et al
stated that, miR-29 suppressed schwannoma cell proliferation and
motility by regulating CDK6 (22). In
colon cancer, miR-29 showed inhibitory effect on cell invasion and
migration by regulating MMP2 (23).
Additionally, miR-29 suppressed lung cancer development by
targeting DNMT3A and DNMT3B (24).
However, role of miR-29 in OS and its mechanism have rarely been
reported.
It is well known that phosphatase and tensin homolog
(PTEN) acts as a tumor suppressor and PTEN expression was proved
abnormal in many cancers (25). For
instance, PTEN functioned as a tumor inhibitor in regulating
pancreatic cancer progression regulated by miR-32 (26). miR-21 modulated gastric cancer
development via targeting PTEN (27)
and PTEN was also a target of miR-20b in modulating prostate cancer
cell viability and migratory ability (28). However, the effect of PTEN on OS cell
migration and proliferation and whether PTEN is regulated by miR-29
in OS have yet to be reported.
In our study, miR-29 and PTEN in OS and its
potential mechanism in modulating OS cell migration and
proliferation was investigated. Our results may provide important
insight into the prognosis and treatment of OS.
Materials and methods
OS specimens
We collected conventional OS and adjacent
non-cancerous tissues from 60 patients who underwent resection
surgery in the First Affiliated Hospital of Zhengzhou University
(Zhengzhou, China) between February 2010 and August 2016. None of
OS patients had received radiotherapy or chemotherapy before
surgery and the tissues were diagnosed as OS by pathologists. All
tissue specimens were placed into liquid nitrogen immediately and
stored at −80°C in a refrigerator for use in subsequent
experiments. The Ethics Committee of Zhengzhou University approved
this study and the patients signed informed consent prior to
surgery.
Cell lines and cell culture
The purchased OS cell lines (MG-63, U2OS, 143B,
Saos-2) were cultured in RPMI-1640 medium at 37°C and the medium
contained 20% FBS and penicillin (100 U/ml) and streptomycin (100
µg/ml). hFOB1.19 (normal human osteoblast) cells were cultured in
DMEM supplemented with 10% FBS and G418 (0.03 mg/ml) at 34°C with
5% CO2. OS cell lines (MG-63, U2OS, 143B, Saos-2) and
hFOB1.19 (normal human osteoblast) cells were obtained from ATCC
(Manassas, VA, USA).
Cell transfection
miR-29 mimic or inhibitor provided by Suzhou
GenePharma Co., Ltd. (Suzhou, China) was transfected into MG-63
cells to facilitate or inhibit miR-29 expression and control mimic
were used as control (con). We added MG-63 cells into 24-well
plates containing medium and we performed transfection using
Lipofectamine 3000 reagent (Invitrogen; Thermo Fisher Scientific,
Inc., Waltham, MA, USA) for 48 h. PTEN vector and con vector were
synthesized by GenePharma Co., Ltd. and they were used to enforce
PTEN expression and acted as a control separately.
RNA extraction and reverse
transcription-quantitative polymerase chain reaction (RT-qPCR)
Total RNA and microRNA were extracted from OS
tissues and cells using TRIzol reagent (Invitrogen; Thermo Fisher
Scientific, Inc.) according to the manufacturer's protocol.
Complementary DNA was synthesized using PrimeScript RT reagent kit
and RT-qPCR was performed using SYBR Premix Ex Taq (both from
Takara Biotechnology Co., Ltd., Dalian, China) with the Stratagene
Mx3000P real-time PCR system (Agilent Technologies, Inc., Santa
Clara, CA, USA). The primer sequences used were: miR-29-F:
TGCCAGGAGCTGGTGATTTCCT, miR-29-R: ACGGGCGTACAGAGGATCCCC. PTEN-F:
GTGCAG ATAATGACAAG, PTEN-R: GATTTGACGGCTCCTCT. proliferating cell
nuclear antigen (PCNA)-F: GGTGTTG GAGGCACTCAAGG, PCNA-R:
CAGGGTGAGCTGCACC AAAG. U6-F: CTCGCTTCGGCAGCAC, U6-R: ACGCTTC
ACGAATTTGC. β-actin-F: GA TCATTGCTCCTCCTGAGC; β-actin-R:
ACTCCTGCTTGCTGATCCAC. U6 and β-actin were used as internal
controls. Relative expression of miR-29, PTEN and PCNA was
calculated by the 2−ΔΔCq method (29).
Western blot analysis
Lysis buffer containing protease inhibitor and
phenylmethanesulfonyl fluoride (PMSF) was added into OS tissues or
cells and then homogenized on ice (the written informed consent
from patients were obtained). After centrifugation at 12,000 × g,
at 4°C for 30 min, the supernatant was measured by BCA kit. Then,
50 µg protein specimens at the same concentration were added onto
sodium dodecyl sulfate-polyacrylamide gel electrophoresis
(SDS-PAGE). Proteins were then transferred to polyvinylidene
fluoride (PVDF) membranes and blocked with skim milk (5–10%) at
room temperature for 2 h subsequently. Then primary antibodies,
monoclonal rabbit anti-PTEN (1:1,000, cat. no. 9188; Cell Signaling
Technology, Inc., Danvers, MA, USA), monoclonal mouse anti-actin
(1:1,000, cat. no. sc-58673; Santa Cruz Biotechnology, Inc.,
Dallas, TX, USA) were added to incubate the membranes at 4°C
overnight and secondary antibodies, monoclonal goat anti-rabbit or
anti-mouse IgG-HRP (1:2,000, cat. no. sc-2004; Santa Cruz
Biotechnology, Inc.) were later added to incubate the membranes at
room temperature (25°C) for 2 h. Finally, the enhanced
chemiluminescence kit (ECL; Merck Millipore, Billerica, MA, USA)
was used to detect the signals. The relative expression of target
proteins was evaluated using target protein/β-actin.
Transwell assay
Transwell assay was used to measure cell migration.
Transwell inserts with 8 µm pore size polycarbonic membrane
(Corning Costar Corp., Cambridge, MA, USA) were used to divide
Transwell chamber into upper and lower chambers. MG-63 cells
(5×105/well) with different transfection were placed
into the top chambers and DMEM containing 10% FBS into lower
chambers and then cultured for 48 h at 37°C under CO2
atmosphere. The cells from the top chambers migrated into the lower
chambers, and the cells on the top chambers were removed with
cotton. The cells were fixed on lower chambers using 4%
formaldehyde for 30 min at room temperature (25°C), cells were
stained with 0.1% crystal violet for 15 min, and images of the
cells were captured using an inverted microscope (Nikon 80i,
Feasterville, PA, USA). Cells were then counted using Living Image
Version 4.5 Software (PerkinElmer, Inc., Waltham, MA, USA) image
software.
MTT assay
Fresh culture (100 µl) medium was added into 96-well
plates to replace the culture medium when the MG-63 cell population
reached optimal densities. Then, MTT reagent (20 µl) was added and
cultured at 37°C for 4 h. Subsequently, the MTT medium was sucked
out and 100 µl of DMSO was added for additional 10 min, the plates
were read at a wavelength of 570 nm using the multilabel plate
reader (PerkinElmer, Inc.) at 1, 2, 3, 4 days to measure the
absorbance of each well.
Luciferase reporter assay
TargetScan Human 7.1 was firstly carried out to
identify the possible targets of miR-29. We used Lipofectamine 2000
(Thermo Fisher Scientific, Inc.) to co-transfect miR-29 mimic or
control mimic with 3′-UTR of wild or mutated PTEN in MG-63 cells.
The dual luciferase reporter system (GeneCopoeia, Inc., Rockville,
MD, USA) was then used to measure the luciferase activity of MG-63
cells treated with different transfection.
Statistical analysis
All experiments were repeated three times
independently. SPSS v.19.0 software (IBM Corp., Armonk, NY, USA)
was used to perform statistical analyses and GraphPad Prism 5.02
software (GraphPad Software, Inc., La Jolla, CA, USA) to complete
graph presentations. Results are presented as the mean ± standard
deviation, and the data were evaluated using Student's t-test or
Tukey's post hoc test after ANOVA in SPSS. P<0.05 was considered
to indicate a statistically significant difference.
Results
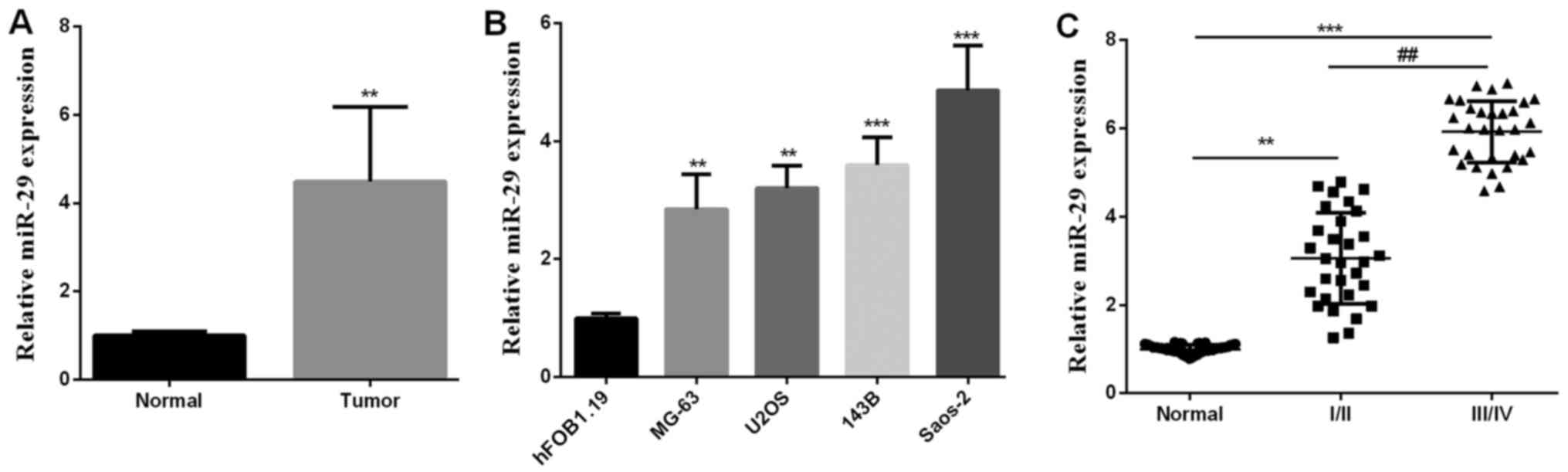
miR-29 is overexpressed in OS
Our primary aim was to explore miR-29 role in OS.
First, we detected miR-29 level in OS tissues and cells using
RT-qPCR. As shown in Fig. 1A, miR-29
average level in OS tissues was significantly increased compared
with normal tissues. miR-29 expression in OS cells (MG-63, U2OS,
143B, Saos-2) was also elevated compared to normal cells
(hFOB1.19), as expected (Fig. 1B).
miR-29 expression was also detected in various stages of OS tumors
and the results showed that the more advanced the tumor stage, the
higher the expression of miR-29 (Fig.
1C), indicating that miR-29 may participate in OS malignant
progression.
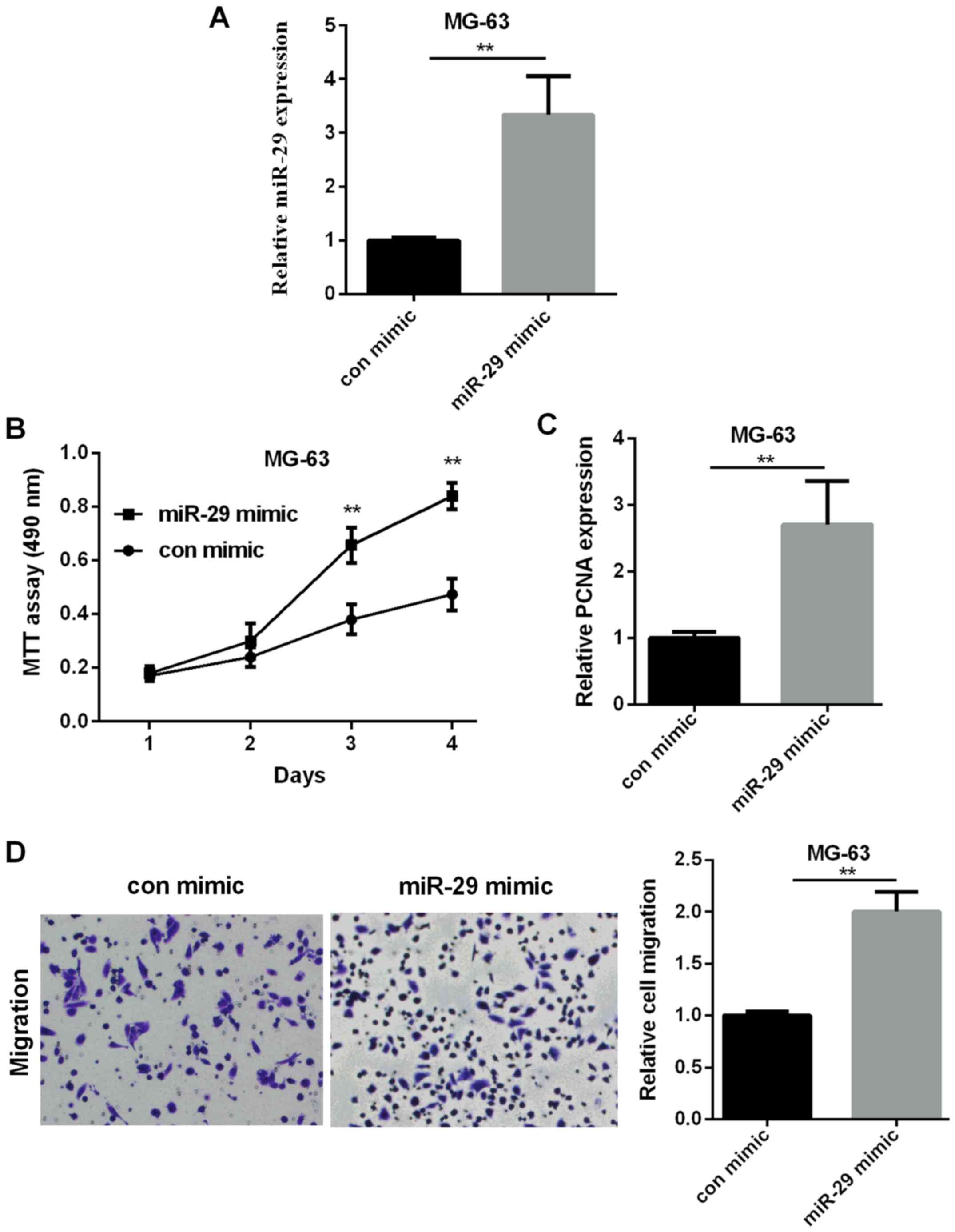
miR-29 mimic promotes OS cell
proliferation and migration
Secondly, we measured miR-29 effect on OS
progression. We overexpressed miR-29 in MG-63 cells using miR-29
mimic, and mRNA expression was increased significantly (Fig. 2A). Cell proliferation was measured by
MTT assay and RT-qPCR. MTT results showed that cell viability in
miR-29 mimic group was enhanced in comparison with control mimic
group (Fig. 2B). PCNA is a marker for
cell proliferation (30,31). RT-qPCR analysis found that PCNA
expression was increased in miR-29 mimic group (Fig. 2C). Cell migration measured by
Transwell demonstrated that miR-29 mimic promoted OS cell migratory
ability (Fig. 2D). The results above
demonstrated a promotion effect of miR-29 on OS progression.
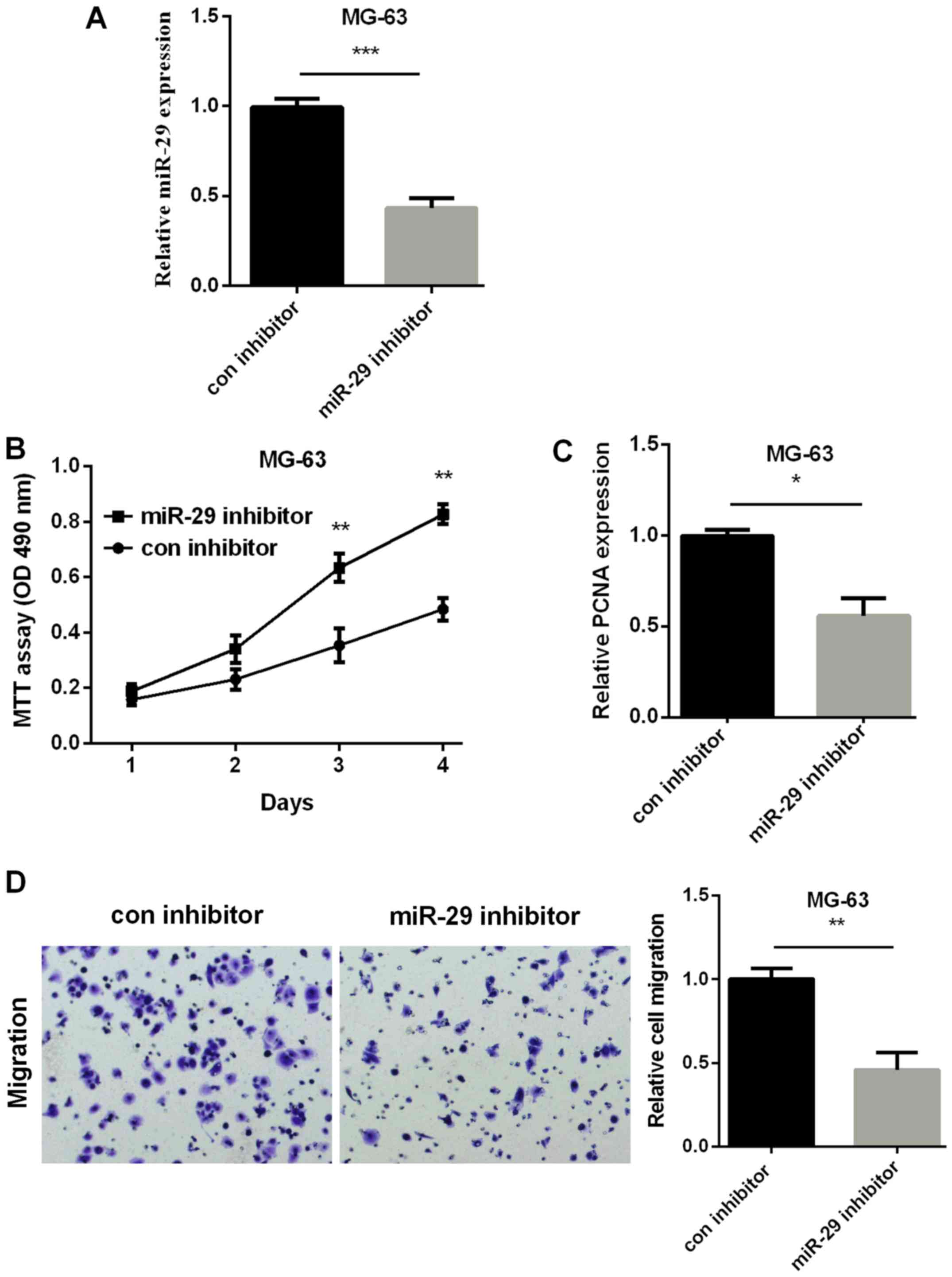
Silencing miR-29 curbed OS cell
proliferation and migration
We then silenced miR-29 using miR-29 inhibitor to
further confirm its effect on OS. The transfection efficiency is
shown in Fig. 3A, miR-29 inhibitor
group decreased miR-29 expression. As we expected, MTT results
showed the decreased cell viability in miR-29 inhibitor group in
comparison with the con-inhibitor group (Fig. 3B). RT-qPCR analysis revealed the
increased PCNA expression in MG-63 cells after silencing miR-29
(Fig. 3C). Cell migration measured by
Transwell showed enhanced cell migratory ability in miR-29
inhibitor group than the con-inhibitor group (Fig. 3D). The results above indicated that
silencing miR-29 inhibited the development of OS.
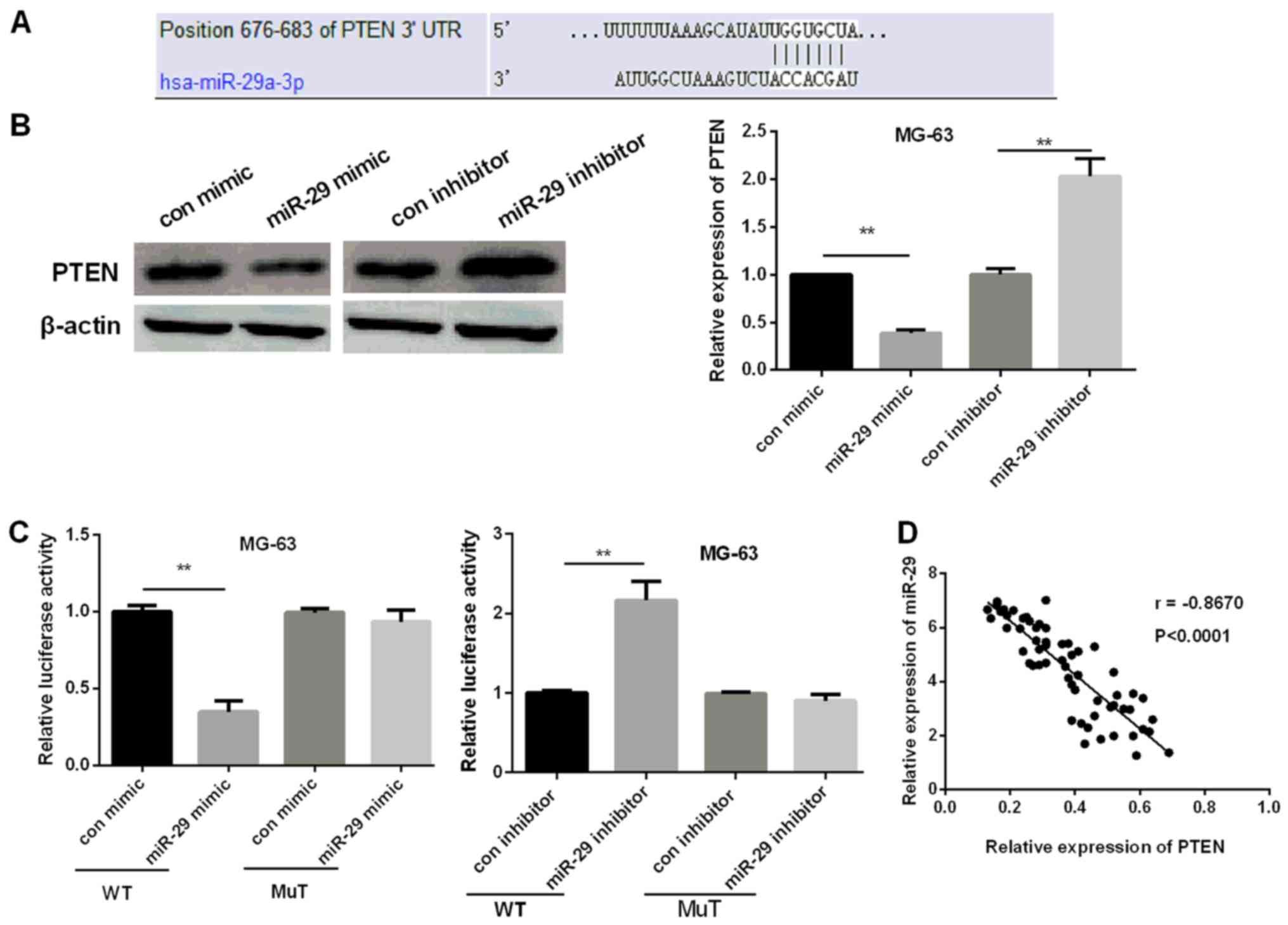
miR-29 targets PTEN in OS
The next step was to investigate miR-29 molecular
mechanism in OS. TargetScan Human 7.1 was carried out to identify
the possible targets of miR-29 and results are shown in Fig. 4A. As further confirmation, we assessed
PTEN protein and mRNA level after MG-63 cell restoration or
inhibiting miR-29 via western blot analysis and RT-qPCR. PTEN
expression was reduced when miR-29 was restored while increased
when inhibiting miR-29 (Fig. 4B).
Then, luciferase reporter assay was carried out to determine
whether PTEN was a direct target of miR-29 in OS. The luciferase
activity of miR-29 mimic group was decreased but increased in
miR-29 inhibitor group in wild-type. However, there was no
significant effect in mut type (Fig.
4C). The results verified PTEN as a direct target of miR-29. In
addition, miR-29 and PTEN expression were negatively correlated
(Fig. 4D).
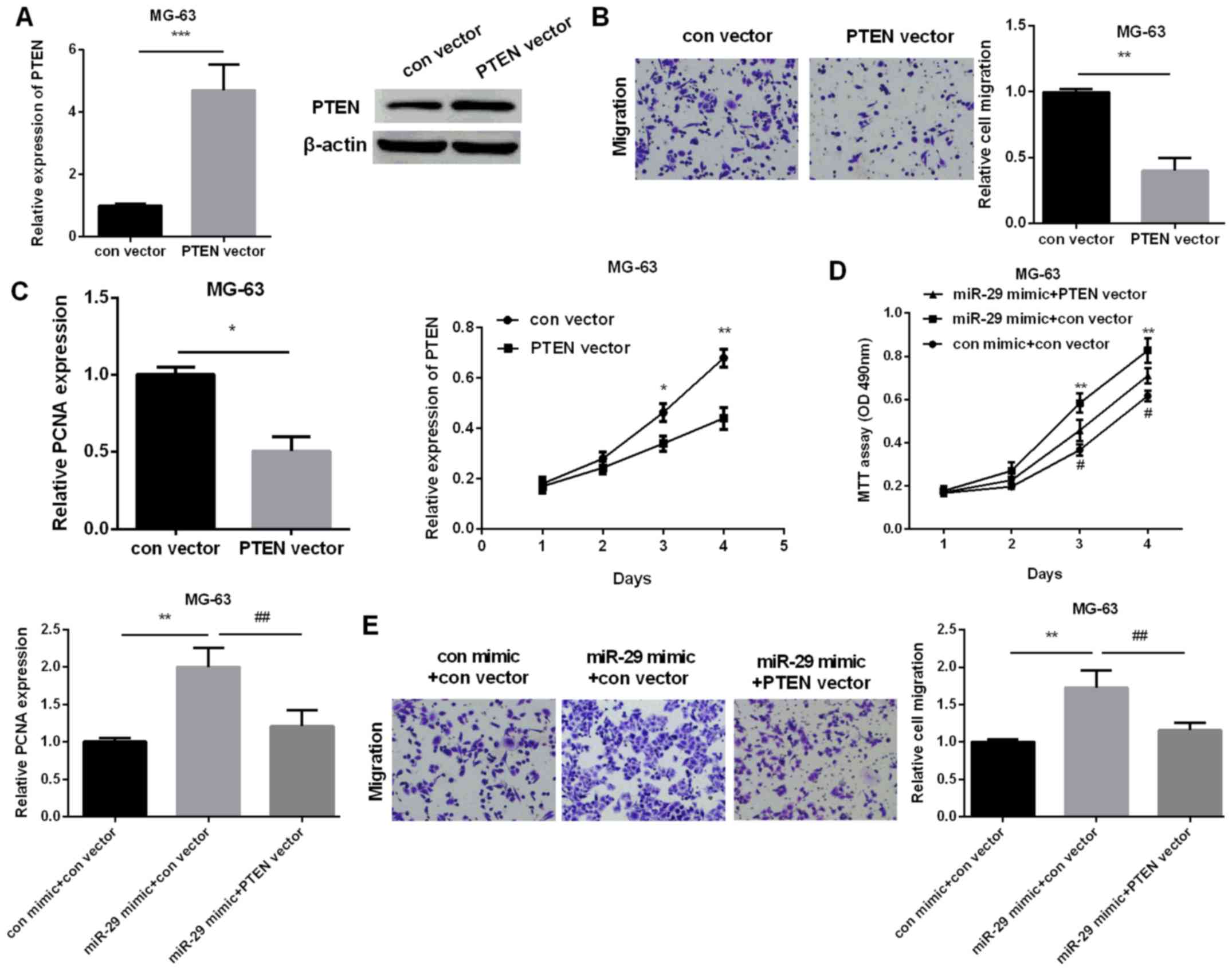
PTEN attenuates miR-29 promotion
effect on OS progression
Then, we investigated the role of PTEN in OS and
whether PTEN affected the miR-29 role in regulating OS progression.
PTEN was overexpressed by PTEN vector due to its lower expression
in OS. The results showed that PTEN expression was higher after
cells were transfected with PTEN vector (Fig. 5A). PTEN overexpressed could inhibit OS
cell migration and proliferation (Fig. 5B
and C). Subsequently, miR-29 mimic and PTEN vector were
co-transfected into MG-63 cells to test the role of PTEN in OS
regulated by miR-29. The cell viability and PCNA expression were
both increased obviously by miR-29 mimic in comparison with control
(Fig. 5D). By contrast,
overexpression of both miR-29 and PTEN, cell proliferation was
reduced compared with overexpression of miR-29 alone, indicating
that PTEN could attenuate miR-29 promotion effect on cell
proliferation. Additionally, the migration results suggested that
miR-29 promotion increased migration in OS cells compared with
control, whereas promoting both miR-29 and PTEN reduced cell
migration in comparison with promoting miR-127 alone, suggesting
that PTEN could reverse miR-29 promotion effect on OS cell
migration. In summary, miR-29 could facilitate OS cell migration
and proliferation by targeting PTEN.
Discussion
An increasing number of studies have reported the
important roles of miRNAs in tumorigenesis and tumor development
(32). Furthermore, the ability to
control cell growth, metastasis and survival may serve essential
roles in preventing and treating various human malignancies,
including OS (33). miR-29 was
reported to have significant inhibitory effect on various cancers.
For example, it inhibited hepatocellular carcinoma cell migration
and proliferation by regulating IGF1R (34). miR-29 played a tumor suppressive role
in modulating gliomas through targeting CDC42 (35). However, Gao et al reported that
miR-29 played a promoting role in OS (36) and Hong et al showed the same
discovery that miR-29 expression was higher in OS patients
(37). Those results were in line
with our findings that miR-29 expression was upregulated in OS
tissues and cells, but these studies did not show the function of
miR-29 in OS tumorigenesis and development. In the present study,
we first demonstrated that restoration of miR-29 facilitated cell
migration and proliferation, while inhibiting miR-29 showed
inhibitory effect.
PTEN was the first tumor suppressor gene found with
a bispecific phosphatase activity and closely associated with
carcinoma (38). PTEN suppressed the
migration and genetic deletion of the PTEN tumor suppressor gene,
promoting cell motility and overexpression or reconstitution of
PTEN and inhibiting cell motility in a variety of cell types
(39,40). Hundreds of genes were found to be the
target of miRNA, which harbors the target sequence in their 3′-UTR
complementary to the seed region of the miRNA. PTEN was also proven
to be a target of miRNAs in modulating cancer progression. PTEN was
the target of miR-28 and suppressed gastric cancer cell
proliferation and invasion (41).
Additionally, it suppressed renal cancer cell proliferation as a
target of miR-30a (42). Zhao et
al reported that PTEN regulated OS cell growth and apoptosis
and acted as a target of miR-19 (43). We also found that the PTEN mRNA
expression was downregulated in MG63 cells compared with the
hFOB1.19 cells (data not shown). However, the underlying mechanism
between miR-29 and PTEN was still unclear in OS. In our study, we
found that miR-29 could negatively regulate the expression of PTEN
in MG63 cells. Further investigation was conducted to explore the
molecular mechanism how PTEN promoted OS cell proliferation and
migration. In addition, PTEN overexpressed was proved to inhibit OS
cell proliferation and migration, PTEN could reverse miR-29
promotion effect on OS progression as well. Therefore, to the best
of our knowledge, the present study provided an effective treatment
target in OS.
In conclusion, miR-29 facilitated OS migration and
proliferation and this is the first report that miR-29 targeted
PTEN and regulated OS development. Furthermore, PTEN could reverse
miR-29 promotion effect on OS providing a new idea for OS
therapy.
Acknowledgements
Not applicable.
Funding
This study received the specific grant from the
Co-construction project of Henan Provincial Medical Science and
Technology Key Project (grant no. 201401007).
Availability of data and materials
The datasets generated or analyzed during the
present study are available from the corresponding author on
reasonable request.
Authors' contributions
QL was a major contributor in writing the manuscript
and contributed to the conception of the study. PG contributed
significantly to data analysis and manuscript preparation. LS
performed the data analyses and wrote the manuscript. QW and PW
helped perform the data analysis with constructive discussions. All
authors read and approved the final study.
Ethics approval and consent to
participate
The study was approved by the Ethics Committee of
The First Affiliated Hospital of Zhengzhou University (Zhengzhou,
China). Signed informed consents were obtained from all
participants.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Endo-Munoz L, Evdokiou A and Saunders NA:
The role of osteoclasts and tumour-associated macrophages in
osteosarcoma metastasis. Biochim Biophys Acta. 1826:434–442.
2012.PubMed/NCBI
|
|
2
|
Harting MT and Blakely ML: Management of
osteosarcoma pulmonary metastases. Semin Pediatr Surg. 15:25–29.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Jia S and Li B: Osteosarcoma of the jaws:
Case report on synchronous multicentric osteosarcomas. J Clin Diagn
Res. 8:ZD01–ZD03. 2014.PubMed/NCBI
|
|
4
|
He JP, Hao Y, Wang XL, Yang XJ, Shao JF,
Guo FJ and Feng JX: Review of the molecular pathogenesis of
osteosarcoma. Asian Pac J Cancer Prev. 15:5967–5976. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Egas-Bejar D, Anderson PM, Agarwal R,
Corrales-Medina F, Devarajan E, Huh WW, Brown RE and Subbiah V:
Theranostic profiling for actionable aberrations in advanced high
risk osteosarcoma with aggressive biology reveals high molecular
diversity: The human fingerprint hypothesis. Oncoscience.
1:167–179. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Poletajew S, Fus L and Wasiutyński A:
Current concepts on pathogenesis and biology of metastatic
osteosarcoma tumors. Ortop Traumatol Rehabil. 13:537–545. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Xu C, Ping Y and Li X, Zhao H, Wang L, Fan
H, Xiao Y and Li X: Prioritizing candidate disease miRNAs by
integrating phenotype associations of multiple diseases with
matched miRNA and mRNA expression profiles. Mol Biosyst.
10:2800–2809. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Gaál Z and Oláh E: MicroRNA-s and their
role in malignant hematologic diseases. Orv Hetil. 153:2051–2059.
2012.(In Hungarian). View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Lin S, Pan L, Guo S, Wu J, Jin L, Wang JC
and Wang S: Prognostic role of microRNA-181a/b in hematological
malignancies: A meta-analysis. PLoS One. 8:e595322013. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Kalinowski FC, Brown RA, Ganda C, Giles
KM, Epis MR, Horsham J and Leedman PJ: microRNA-7: A tumor
suppressor miRNA with therapeutic potential. Int J Biochem Cell
Biol. 54:312–317. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Xu Y, Chu H, Zhou Y, Wang J, Dong C and
Yin R: miR-365 functions as a tumor suppressor by directly
targeting CYR61 in osteosarcoma. Biomed Pharmacother. 98:531–537.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Yu L, Wang S, Lin X, Lu Y and Gu P:
MicroRNA-124a inhibits cell proliferation and migration in liver
cancer by regulating interleukin-11. Mol Med Rep. 17:3972–3978.
2018.PubMed/NCBI
|
|
13
|
Luo Z, Wu G, Zhang D, Liu J and Ran R:
microRNA 625 targets Yes associated protein 1 to suppress cell
proliferation and invasion of osteosarcoma. Mol Med Rep.
17:2005–2011. 2018.PubMed/NCBI
|
|
14
|
Wang N, Li P, Liu W, Wang N, Lu Z, Feng J,
Zeng X, Yang J, Wang Y and Zhao W: miR-141-3p suppresses
proliferation and promotes apoptosis by targeting GLI2 in
osteosarcoma cells. Oncol Rep. 39:747–754. 2018.PubMed/NCBI
|
|
15
|
He Y and Yu B: MicroRNA-93 promotes cell
proliferation by directly targeting P21 in osteosarcoma cells. Exp
Ther Med. 13:2003–2011. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Sun Z, Liu Q, Hong H, Zhang H and Zhang T:
miR-19 promotes osteosarcoma progression by targeting SOCS6.
Biochem Biophys Res Commun. 495:1363–1369. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Cai Q, Zeng S, Dai X, Wu J and Ma W:
miR-504 promotes tumour growth and metastasis in human osteosarcoma
by targeting TP53INP1. Oncol Rep. 38:2993–3000. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Wu Z, Huang X, Huang X, Zou Q and Guo Y:
The inhibitory role of Mir-29 in growth of breast cancer cells. J
Exp Clin Cancer Res. 32:982013. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Lv M, Zhong Z, Huang M, Tian Q, Jiang R
and Chen J: lncRNA H19 regulates epithelial-mesenchymal transition
and metastasis of bladder cancer by miR-29b-3p as competing
endogenous RNA. Biochim Biophys Acta. 1864:1887–1899. 2017.
View Article : Google Scholar
|
|
20
|
Kwon JJ, Willy JA, Quirin KA, Wek RC, Korc
M, Yin XM and Kota J: Novel role of miR-29a in pancreatic cancer
autophagy and its therapeutic potential. Oncotarget. 7:71635–71650.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Shin J, Shim HG, Hwang T, Kim H, Kang SH,
Dho YS, Park SH, Kim SJ and Park CK: Restoration of miR-29b exerts
anti-cancer effects on glioblastoma. Cancer Cell Int. 17:1042017.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Ma J, Li T, Yuan H, Han X, Shui S, Guo D
and Yan L: MicroRNA-29a inhibits proliferation and motility of
schwannoma cells by targeting CDK6. J Cell Biochem. 119:2617–2626.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Ding Q, Chang CJ, Xie X, Xia W, Yang JY,
Wang SC, Wang Y, Xia J, Chen L, Cai C, et al: APOBEC3G promotes
liver metastasis in an orthotopic mouse model of colorectal cancer
and predicts human hepatic metastasis. J Clin Invest.
121:4526–4536. 2011. View
Article : Google Scholar : PubMed/NCBI
|
|
24
|
Fabbri M, Garzon R, Cimmino A, Liu Z,
Zanesi N, Callegari E, Liu S, Alder H, Costinean S,
Fernandez-Cymering C, et al: MicroRNA-29 family reverts aberrant
methylation in lung cancer by targeting DNA methyltransferases 3A
and 3B. Proc Natl Acad Sci USA. 104:15805–15810. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Boosani CS and Agrawal DK: PTEN
modulators: A patent review. Expert Opin Ther Pat. 23:569–580.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Gao ZQ, Wang JF, Chen DH, Ma XS, Wu Y,
Tang Z and Dang XW: Long non-coding RNA GAS5 suppresses pancreatic
cancer metastasis through modulating miR-32-5p/PTEN axis. Cell
Biosci. 7:662017. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Wang P, Guan Q, Zhou D, Yu Z, Song Y and
Qiu W: miR-21 inhibitors modulate biological functions of gastric
cancer cells via PTEN/PI3K/mTOR pathway. DNA Cell Biol. 37:38–45.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Guo J, Xiao Z, Yu X and Cao R: miR-20b
promotes cellular proliferation and migration by directly
regulating phosphatase and tensin homolog in prostate cancer. Oncol
Lett. 14:6895–6900. 2017.PubMed/NCBI
|
|
29
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Hall PA, Levison DA, Woods AL, Yu CC,
Kellock DB, Watkins JA, Barnes DM, Gillett CE, Camplejohn R, Dover
R, et al: Proliferating cell nuclear antigen (PCNA)
immunolocalization in paraffin sections: An index of cell
proliferation with evidence of deregulated expression in some
neoplasms. J Pathol. 162:285–294. 1990. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Moldovan GL, Pfander B and Jentsch S:
PCNA, the maestro of the replication fork. Cell. 129:665–679. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Li Y, Zeng C, Tu M, Jiang W, Dai Z, Hu Y,
Deng Z and Xiao W: MicroRNA-200b acts as a tumor suppressor in
osteosarcoma via targeting ZEB1. Onco Targets Ther. 9:3101–3111.
2016.PubMed/NCBI
|
|
33
|
Yin Z, Ding H, He E, Chen J and Li M:
Up-regulation of microRNA-491-5p suppresses cell proliferation and
promotes apoptosis by targeting FOXP4 in human osteosarcoma. Cell
Prolif. 50:e123082017. View Article : Google Scholar
|
|
34
|
Wang X, Liu S, Cao L, Zhang T, Yue D, Wang
L, Ping Y, He Q, Zhang C, Wang M, et al: miR-29a-3p suppresses cell
proliferation and migration by downregulating IGF1R in
hepatocellular carcinoma. Oncotarget. 8:86592–86603.
2017.PubMed/NCBI
|
|
35
|
Shi C, Ren L, Sun C, Yu L, Bian X, Zhou X,
Wen Y, Hua D, Zhao S, Luo W, et al: miR-29a/b/c function as
invasion suppressors for gliomas by targeting CDC42 and predict the
prognosis of patients. Br J Cancer. 117:1036–1047. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Gao S, Cheng C, Chen H, Li M, Liu K and
Wang G: IGF1 3′UTR functions as a ceRNA in promoting angiogenesis
by sponging miR-29 family in osteosarcoma. J Mol Histol.
47:135–143. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Hong Q, Fang J, Pang Y and Zheng J:
Prognostic value of the microRNA-29 family in patients with primary
osteosarcomas. Med Oncol. 31:372014. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Li J, Yen C, Liaw D, Podsypanina K, Bose
S, Wang SI, Puc J, Miliaresis C, Rodgers L, McCombie R, et al:
PTEN, a putative protein tyrosine phosphatase gene mutated in human
brain, breast, and prostate cancer. Science. 275:1943–1947. 1997.
View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Yang YK, Xi WY, Xi RX, Li JY, Li Q and Gao
YE: MicroRNA-494 promotes cervical cancer proliferation through the
regulation of PTEN. Oncol Rep. 33:2393–2401. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Hodakoski C, Fine B, Hopkins B and Parsons
R: Analysis of intracellular PTEN signaling and secretion. Methods.
77-78:164–171. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Li L, Zhu X, Shou T, Yang L, Cheng X, Wang
J, Deng L and Zheng Y: MicroRNA-28 promotes cell proliferation and
invasion in gastric cancer via the PTEN/PI3K/AKT signalling
pathway. Mol Med Rep. 17:4003–4010. 2018.PubMed/NCBI
|
|
42
|
Li J, Li C, Li H, Zhang T, Hao X, Chang J
and Xu Y: MicroRNA 30a 5p suppresses tumor cell proliferation of
human renal cancer via the MTDH/PTEN/AKT pathway. Int J Mol Med.
41:1021–1029. 2018.PubMed/NCBI
|
|
43
|
Zhao D, Chen Y, Chen S, Zheng C, Hu J and
Luo S: MiR-19a regulates the cell growth and apoptosis of
osteosarcoma stem cells by targeting PTEN. Tumour Biol.
39:10104283177053412017. View Article : Google Scholar : PubMed/NCBI
|