|
1
|
DeSantis C, Siegel R, Bandi P and Jemal A:
Breast cancer statistics, 2011. CA Cancer J Clin. 61:409–418. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Fan L, Strasser-Weippl K, Li JJ, St LJ,
Finkelstein DM, Yu KD, Chen WQ, Shao ZM and Goss PE: Breast cancer
in China. Lancet Oncol. 15:e279–e289. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2015. CA Cancer J Clin. 65:5–29. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Onitilo AA, Engel JM, Greenlee RT and
Mukesh BN: Breast cancer subtypes based on ER/PR and Her2
expression: Comparison of clinicopathologic features and survival.
Clin Med Res. 7:4–13. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Du SJ, Tan X and Zhang J: SMYD proteins:
Key regulators in skeletal and cardiac muscle development and
function. Anat Rec (Hoboken). 297:1650–1662. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Doughan M, Spellmon N, Li C and Yang Z:
SMYD proteins in immunity: Dawning of a new era. AIMS Biophys.
3:450–455. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Leinhart K and Brown M: SET/MYND lysine
methyltransferases regulate gene transcription and protein
activity. Genes (Basel). 2:210–218. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Spellmon N, Holcomb J, Trescott L,
Sirinupong N and Yang Z: Structure and function of SET and MYND
domain-containing proteins. Int J Mol Sci. 16:1406–1428. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Sakamoto LH, Andrade RV, Felipe MS,
Motoyama AB and Pittella SF: SMYD2 is highly expressed in pediatric
acute lymphoblastic leukemia and constitutes a bad prognostic
factor. Leuk Res. 38:496–502. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Hu L, Zhu YT, Qi C and Zhu YJ:
Identification of Smyd4 as a potential tumor suppressor gene
involved in breast cancer development. Cancer Res. 69:4067–4072.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Sealfon SC and Chu TT: RNA and DNA
microarrays. Methods Mol Biol. 671:3–34. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Rhodes DR, Yu J, Shanker K, Deshpande N,
Varambally R, Ghosh D, Barrette T, Pandey A and Chinnaiyan AM:
ONCOMINE: A cancer microarray database and integrated data-mining
platform. Neoplasia. 6:1–6. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Jézéquel P, Campone M, Gouraud W,
Guérin-Charbonnel C, Leux C, Ricolleau G and Campion L:
bc-GenExMiner: An easy-to-use online platform for gene prognostic
analyses in breast cancer. Breast Cancer Res Treat. 131:765–775.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Jézéquel P, Frénel JS, Campion L,
Guérin-Charbonnel C, Gouraud W, Ricolleau G and Campone M:
bc-GenExMiner 3.0: New mining module computes breast cancer gene
expression correlation analyses. Database (Oxford).
2013:bas0602013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Györffy B, Lanczky A, Eklund AC, Denkert
C, Budczies J, Li Q and Szallasi Z: An online survival analysis
tool to rapidly assess the effect of 22,277 genes on breast cancer
prognosis using microarray data of 1,809 patients. Breast Cancer
Res Treat. 123:725–731. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Li Q, Birkbak NJ, Gyorffy B, Szallasi Z
and Eklund AC: Jetset: Selecting the optimal microarray probe set
to represent a gene. BMC Bioinformatics. 12:4742011. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Cancer Genome Atlas Network, .
Comprehensive molecular portraits of human breast tumours. Nature.
490:61–70. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Gao J, Aksoy BA, Dogrusoz U, Dresdner G,
Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, et al:
Integrative analysis of complex cancer genomics and clinical
profiles using the cBioPortal. Sci Signal. 6:2013. View Article : Google Scholar
|
|
19
|
Cerami E, Gao J, Dogrusoz U, Gross BE,
Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, et
al: The cBio cancer genomics portal: An open platform for exploring
multidimensional cancer genomics data. Cancer Discov. 2:401–404.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
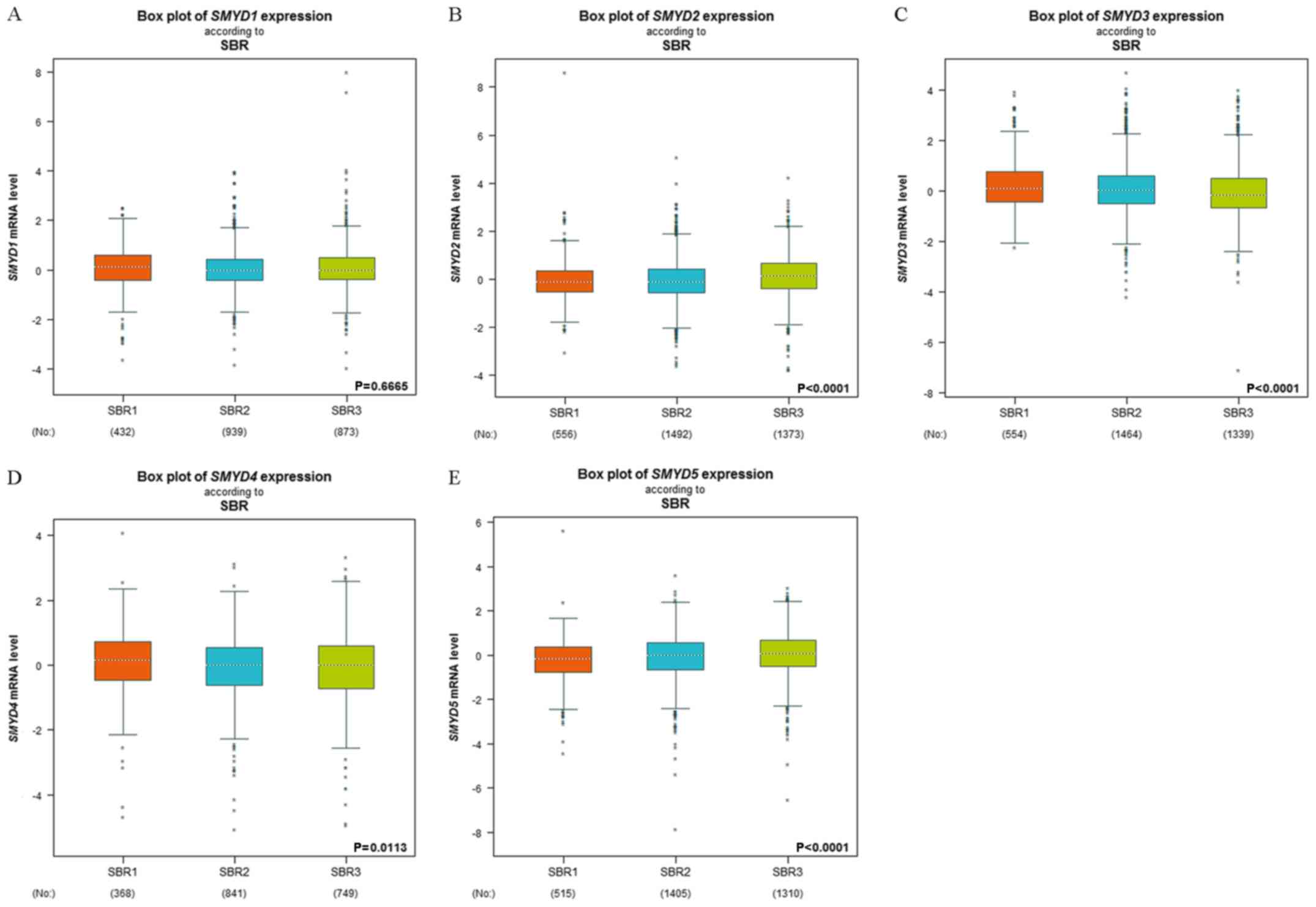
Martinazzi M, Zampatti C, Crivelli F,
Zampieri A and Martinazzi S: Scarff-bloom-richardson
histoprognostic grading correlates with the immunohistochemical
expressions of genomic alterations in infiltrating ductal
carcinomas (nos) of the breast. Oncol Rep. 1:1087–1091.
1994.PubMed/NCBI
|
|
21
|
Nicolai P, Redaelli de Zinis LO, Tomenzoli
D, Barezzani MG, Bertoni F, Bignardi M and Antonelli AR: Prognostic
determinants in supraglottic carcinoma: Univariate and Cox
regression analysis. Head Neck. 19:323–34. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Hsu CL and Lee WC: Detecting
differentially expressed genes in heterogeneous diseases using half
student's t-test. Int J Epidemiol. 39:1597–1604. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Curtis C, Shah SP, Chin SF, Turashvili G,
Rueda OM, Dunning MJ, Speed D, Lynch AG, Samarajiwa S, Yuan Y, et
al: The genomic and transcriptomic architecture of 2,000 breast
tumours reveals novel subgroups. Nature. 486:346–352. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Gottlieb PD, Pierce SA, Sims RJ, Yamagishi
H, Weihe EK, Harriss JV, Maika SD, Kuziel WA, King HL, Olson EN, et
al: Bop encodes a muscle-restricted protein containing MYND and SET
domains and is essential for cardiac differentiation and
morphogenesis. Nat Genet. 31:25–32. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Everett AD: Identification, cloning, and
developmental expression of hepatoma-derived growth factor in the
developing rat heart. Dev Dyn. 222:450–458. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Al-Shar'i NA and Alnabulsi SM: Explaining
the autoinhibition of the SMYD enzyme family: A theoretical study.
J Mol Graph Model. 68:147–157. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Yang J and Everett AD: Hepatoma-derived
growth factor represses SET and MYND domain containing 1 gene
expression through interaction with C-terminal binding protein. J
Mol Biol. 386:938–950. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Hu TH, Huang CC, Liu LF, Lin PR, Liu SY,
Chang HW, Changchien CS, Lee CM, Chuang JH and Tai MH: Expression
of hepatoma-derived growth factor in hepatocellular carcinoma.
Cancer. 98:1444–1456. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Chen SC, Kung ML, Hu TH, Chen HY, Wu JC,
Kuo HM, Tsai HE, Lin YW, Wen ZH, Liu JK, et al: Hepatoma-derived
growth factor regulates breast cancer cell invasion by modulating
epithelial-mesenchymal transition. J Pathol. 228:158–169. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Li LX, Zhou JX, Calvet JP, Godwin AK,
Jensen RA and Li X: Lysine methyltransferase SMYD2 promotes triple
negative breast cancer progression. Cell Death Dis. 9:3262018.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Huang J, Perez-Burgos L, Placek BJ,
Sengupta R, Richter M, Dorsey JA, Kubicek S, Opravil S, Jenuwein T
and Berger SL: Repression of p53 activity by Smyd2-mediated
methylation. Nature. 444:629–632. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Nakakido M, Deng Z, Suzuki T, Dohmae N,
Nakamura Y and Hamamoto R: Dysregulation of AKT pathway by
SMYD2-mediated lysine methylation on PTEN. Neoplasia. 17:367–373.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Wang T, Wu H, Liu S, Lei Z, Qin Z, Wen L,
Liu K, Wang X, Guo Y, Liu Q, et al: SMYD3 controls a Wnt-responsive
epigenetic switch for ASCL2 activation and cancer stem cell
maintenance. Cancer Lett. 430:11–24. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Mazur PK, Reynoird N, Khatri P, Jansen PW,
Wilkinson AW, Liu S, Barbash O, Van Aller GS, Huddleston M, Dhanak
D, et al: SMYD3 links lysine methylation of MAP3K2 to Ras-driven
cancer. Nature. 510:283–287. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Kim JM, Kim K, Schmidt T, Punj V, Tucker
H, Rice JC, Ulmer TS and An W: Cooperation between SMYD3 and PC4
drives a distinct transcriptional program in cancer cells. Nucleic
Acids Res. 43:8868–8883. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Tsai CH, Chen YJ, Yu CJ, Tzeng SR, Wu IC,
Kuo WH, Lin MC, Chan NL, Wu KJ and Teng SC: SMYD3-Mediated H2A.Z.1
Methylation promotes cell cycle and cancer proliferation. Cancer
Res. 76:6043–6053. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Hamamoto R, Silva FP, Tsuge M, Nishidate
T, Katagiri T, Nakamura Y and Furukawa Y: Enhanced SMYD3 expression
is essential for the growth of breast cancer cells. Cancer Sci.
97:113–118. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Frank B, Hemminki K, Wappenschmidt B,
Klaes R, Meindl A, Schmutzler RK, Bugert P, Untch M, Bartram CR and
Burwinkel B: Variable number of tandem repeats polymorphism in the
SMYD3 promoter region and the risk of familial breast cancer. Int J
Cancer. 118:2917–2918. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Kidder BL, He R, Wangsa D, Padilla-Nash
HM, Bernardo MM, Sheng S, Ried T and Zhao K: SMYD5 Controls
heterochromatin and chromosome integrity during embryonic stem cell
differentiation. Cancer Res. 77:6729–6745. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Kidder BL, Hu G, Cui K and Zhao K: SMYD5
regulates H4K20me3-marked heterochromatin to safeguard ES cell
self-renewal and prevent spurious differentiation. Epigenetics
Chromatin. 10:82017. View Article : Google Scholar : PubMed/NCBI
|