|
1
|
Yang JY, Cheng FW, Wong KC, Lee V, Leung
WK, Shing MM, Kumta SM and Li CK: Initial presentation and
management of osteosarcoma, and its impact on disease outcome. Hong
Kong Med J. 15:434–439. 2009.PubMed/NCBI
|
|
2
|
Ottaviani G and Jaffe N: The epidemiology
of osteosarcoma. Cancer Treat Res. 152:3–13. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Liang W, Gao B, Fu P, Xu S, Qian Y and Fu
Q: The miRNAs in the pathgenesis of osteosarcoma. Front Biosci
(Landmark Ed). 18:788–794. 2013. View
Article : Google Scholar : PubMed/NCBI
|
|
4
|
Hu H, Zhang Y, Cai XH, Huang JF and Cai L:
Changes in microRNA expression in the MG-63 osteosarcoma cell line
compared with osteoblasts. Oncol Lett. 4:1037–1042. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Ouyang L, Liu P, Yang S, Ye S, Xu W and
Liu X: A three-plasma miRNA signature serves as novel biomarkers
for osteosarcoma. Med Oncol. 30:3402013. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Sarver AL, Thayanithy V, Scott MC,
Cleton-Jansen AM, Hogendoorn PC, Modiano JF and Subramanian S:
MicroRNAs at the human 14q32 locus have prognostic significance in
osteosarcoma. Orphanet J Rare Dis. 8:72013. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Jianwei Z, Fan L, Xiancheng L, Enzhong B,
Shuai L and Can L: MicroRNA 181a improves proliferation and
invasion, suppresses apoptosis of osteosarcoma cell. Tumour Biol.
34:3331–3337. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Zhou G, Shi X, Zhang J, Wu S and Zhao J:
MicroRNAs in osteosarcoma: From biological players to clinical
contributors, a review. J Int Med Res. 41:1–12. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Ma W, Zhang X, Chai J, Chen P, Ren P and
Gong M: Circulating miR-148a is a significant diagnostic and
prognostic biomarker for patients with osteosarcoma. Tumour Biol.
35:12467–12472. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Pan W, Wang H, Jianwei R and Ye Z:
MicroRNA-27a promotes proliferation, migration and invasion by
targeting MAP2K4 in human osteosarcoma cells. Cell Physiol Biochem.
33:402–412. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Li E, Zhang J, Yuan T and Ma B: MiR-145
inhibits osteosarcoma cells proliferation and invasion by targeting
ROCK1. Tumour Biol. 35:7645–7650. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Xu JQ, Zhang WB, Wan R and Yang YQ:
MicroRNA-32 inhibits osteosarcoma cell proliferation and invasion
by targeting Sox9. Tumour Biol. 35:9847–9853. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Namløs HM, Meza-Zepeda LA, Baroy T,
Østensen IH, Kresse SH, Kuijjer ML, Serra M, Bürger H,
Cleton-Jansen AM and Myklebost O: Modulation of the osteosarcoma
expression phenotype by microRNAs. PLoS One. 7:e480862012.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Gautier L, Cope L, Bolstad BM and Irizarry
RA: Affy-analysis of affymetrix genechip data at the probe level.
Bioinformatics. 20:307–315. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Huang DW, Sherman BT, Tan Q, Collins JR,
Alvord WG, Roayaei J, Stephens R, Baseler MW, Lane HC and Lempicki
RA: The DAVID gene functional classification tool: A novel
biological module-centric algorithm to functionally analyze large
gene lists. Genome Biol. 8:R1832007. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Kawano M, Tanaka K, Itonaga I, Ikeda S,
Iwasaki T and Tsumura H: microRNA-93 promotes cell proliferation
via targeting of PTEN in Osteosarcoma cells. J Exp Clin Cancer Res.
5:762015. View Article : Google Scholar
|
|
18
|
Buddingh EP, Ruslan SEN, Reijnders CMA,
Szuhai K, Kuijjer ML, Roelofs H, Hogendoorn PCW, Maarten Egeler R,
Cleton-Jansen AM and Lankester AC: Mesenchymal stromal cells of
osteosarcoma patients do not show evidence of neoplastic changes
during long-term culture. Clin Sarcoma Res. 23:5–16. 2015.
|
|
19
|
Wang JY, Wu PK, Chen PC, Yen CC, Hung GY,
Chen CF, Hung SC, Tsai SF, Liu CL, Chen TH and Chen WM:
Manipulation therapy prior to diagnosis induced primary
osteosarcoma metastasis-from clinical to basic research. PLoS One.
7:e965712014. View Article : Google Scholar
|
|
20
|
Kresse SH, Rydbeck H, Skårn M, Namløs HM,
Barragan-Polania AH, Cleton-Jansen AM, Serra M, Liestøl K,
Hogendoorn PC, Hovig E, et al: Integrative analysis reveals
relationships of genetic and epigenetic alterations in
osteosarcoma. PLoS One. 7:e482622012. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
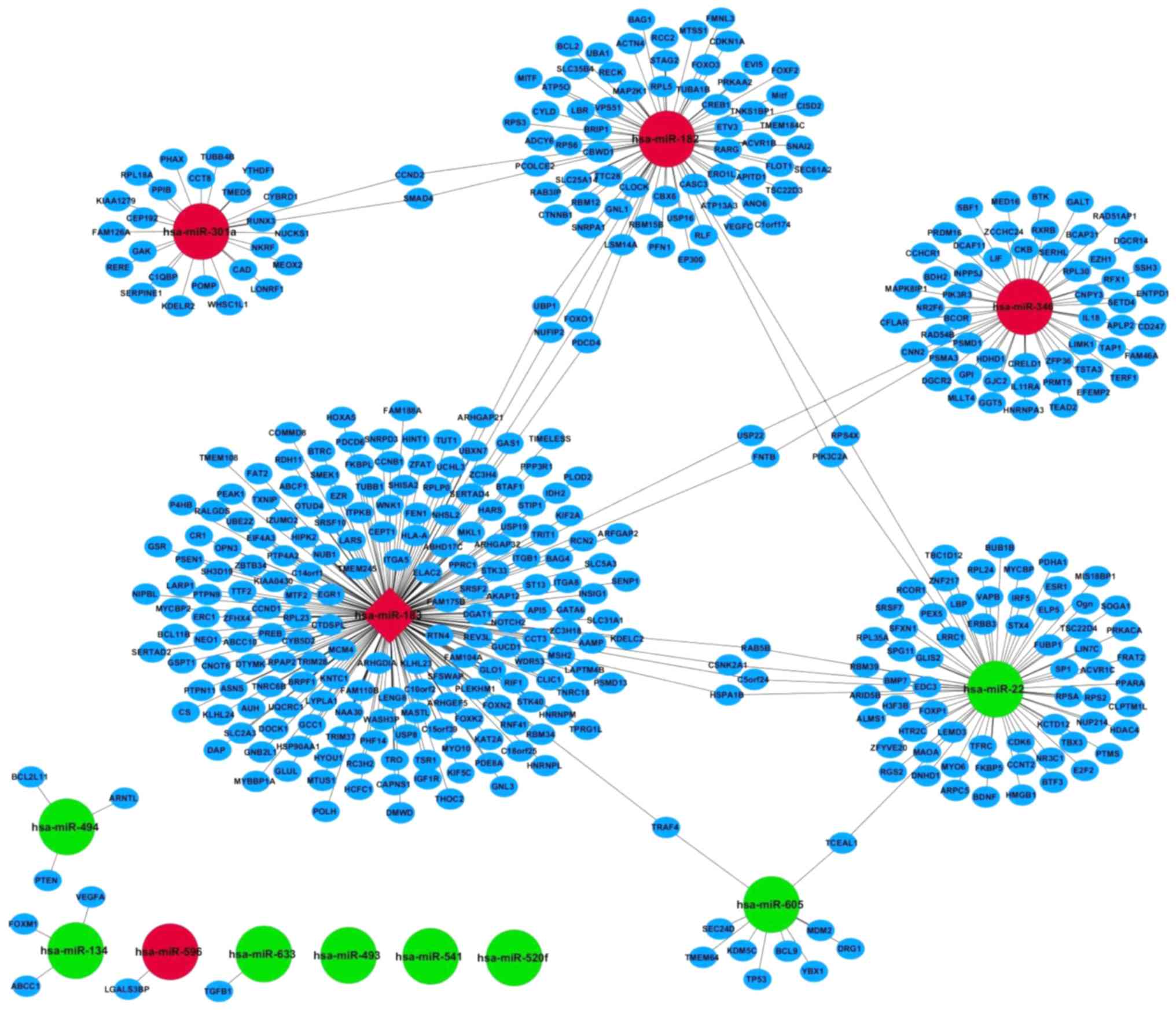
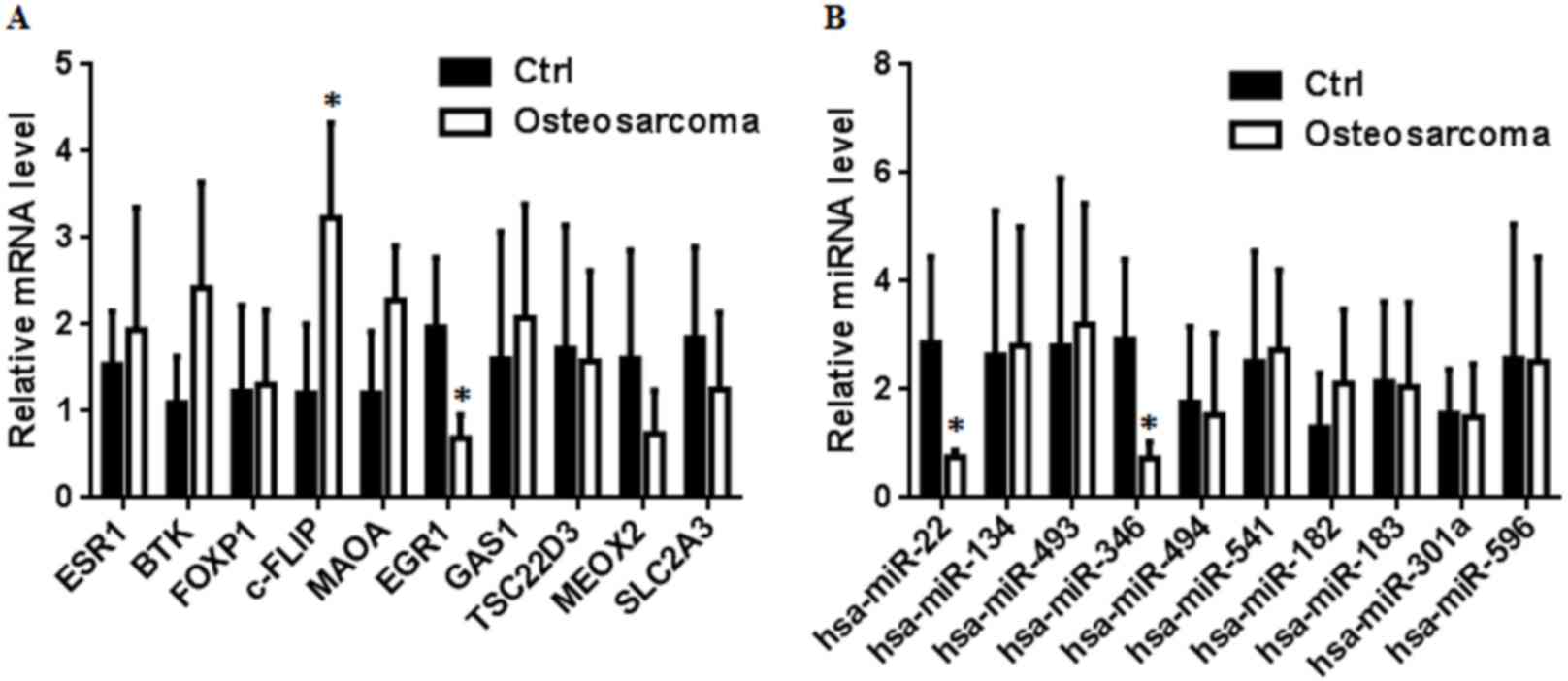
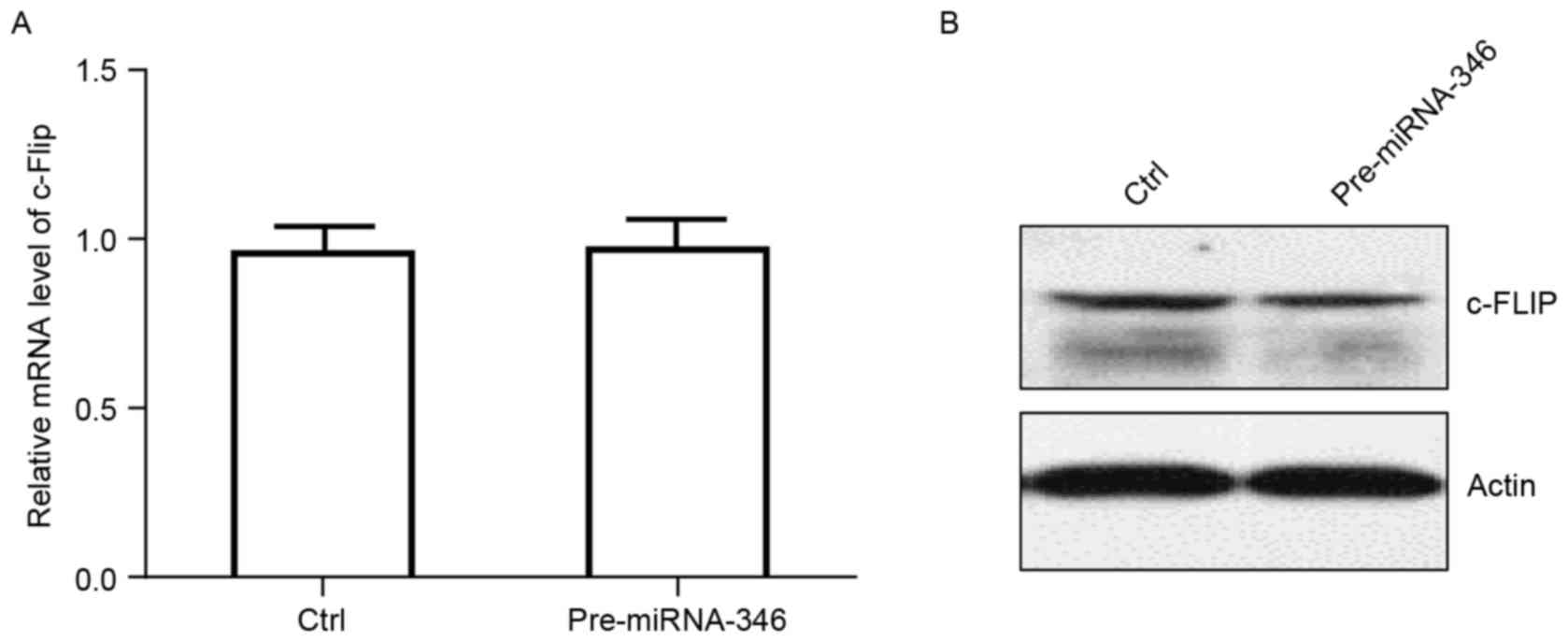
Wang H, Tang M, Ou L, Hou M, Feng T, Huang
YE, Jin Y, Zhang H and Zuo G: Biological analysis of cancer
specific microRNAs on function modeling in osteosarcoma. Sci Rep.
7:53822017. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Grimaldi A, Zarone MR, Irace C, Zappavigna
S, Lombardi A, Kawasaki H, Caraglia M and Misso G: Non-coding RNAs
as a new dawn in tumor diagnosis. Semin Cell Dev Biol. 78:37–50.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Liu X, Liu X, Wu Y, Wu Q, Wang Q, Yang Z
and Li L: MicroRNAs in biofluids are novel tools for bladder cancer
screening. Oncotarget. 8:32370–32379. 2017.PubMed/NCBI
|
|
24
|
Hutanu D, Popescu R, Stefanescu H, Pirtea
L, Candea A, Sarau C, Boruga O, Mehdi L, Ciuca I and Tanasescu S:
The molecular genetic expression as a novel biomarker in the
evaluation and monitoring of patients with osteosarcoma-subtype
bone cancer disease. Biochem Genet. 55:291–299. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Smolle MA, Leithner A, Posch F, Szkandera
J, Liegl-Atzwanger B and Pichler M: MicroRNAs in different
histologies of soft tissue sarcoma: A comprehensive review. Int J
Mol Sci. 18(pii): E19602017. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Sampson VB, Yoo S, Kumar A, Vetter NS and
Kolb EA: MicroRNAs and potential targets in osteosarcoma: Review.
Front Pediatr. 3:692015. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Wang G, Shen N, Cheng L, Lin J and Li K:
Downregulation of miR-22 acts as an unfavorable prognostic
biomarker in osteosarcoma. Tumour Biol. 36:7891–7895. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Lee JH, Park SJ, Jeong SY, Kim MJ, Jun S,
Lee HS, Chang IY, Lim SC, Yoon SP, Yong J and You HJ: MicroRNA-22
suppresses DNA repair and promotes genomic instability through
targeting of MDC1. Cancer Res. 75:1298–1310. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Hu J, Lv G, Zhou S, Zhou Y, Nie B, Duan H,
Zhang Y and Yuan X: The downregulation of MiR-182 is associated
with the growth and invasion of osteosarcoma cells through the
regulation of TIAM1 expression. PLoS One. 10:e01211752015.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Liu Y, Li Y, Liu J, Wu Y and Zhu Q:
MicroRNA-132 inhibits cell growth and metastasis in osteosarcoma
cell lines possibly by targeting Sox4. Int J Oncol. 47:1672–1684.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Zhu J, Feng Y, Ke Z, Yang Z, Zhou J, Huang
X and Wang L: Down-regulation of miR-183 promotes migration and
invasion of osteosarcoma by targeting Ezrin. Am J Pathol.
180:2440–2451. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Ma F, Zhang J, Zhong L, Wang L, Liu Y,
Wang Y, Peng L and Guo B: Upregulated microRNA-301a in breast
cancer promotes tumor metastasis by targeting PTEN and activating
Wnt/β-catenin signaling. Gene. 535:191–197. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Wang M, Li C, Yu B, Su L, Li J, Ju J, Yu
Y, Gu Q, Zhu Z and Liu B: Overexpressed miR-301a promotes cell
proliferation and invasion by targeting RUNX3 in gastric cancer. J
Gastroenterol. 48:1023–1033. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Lee YH, Na HS, Jeong SY, Jeong SH, Park HR
and Chung J: Comparison of inflammatory microRNA expression in
healthy and periodontitis tissues. Biocell. 35:43–49.
2011.PubMed/NCBI
|
|
35
|
Zhang Y, Duan G and Feng S: MicroRNA-301a
modulates doxorubicin resistance in osteosarcoma cells by targeting
AMP-activated protein kinase alpha 1. Biochem Biophys Res Commun.
459:367–373. 2015. View Article : Google Scholar : PubMed/NCBI
|