|
1
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2018. CA Cancer J Clin. 68:7–30. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Torre L, Bray F, Siegel R, Ferlay J,
Lortet-Tieulent J and Jemal A: Global cancer statistics, 2012. CA
Cancer J Clin. 65:87–108. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Goldstraw P, Ball D, Jett J, Le Chevalier
T, Lim E, Nicholson AG and Shepherd FA: Non-small-cell lung cancer.
Lancet. 378:1727–1740. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Ramalingam S, Owonikoko T and Khuri F:
Lung cancer: New biological insights and recent therapeutic
advances. CA Cancer J Clin. 61:91–112. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Chaffer C and Weinberg R: A perspective on
cancer cell metastasis. Science. 331:1559–1564. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Ramalingam S and Belani C: Systemic
chemotherapy for advanced non-small cell lung cancer: Recent
advances and future directions. Oncologist. 13 (Suppl 1):S5–S13.
2008. View Article : Google Scholar
|
|
7
|
Detterbeck F, Boffa D and Tanoue L: The
new lung cancer staging system. Chest. 136:260–271. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Lothrop AP, Torres MP and Fuchs SM:
Deciphering post-translational modification codes. FEBS Lett.
587:1247–1257. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Husnjak K and Dikic I: Ubiquitin-binding
proteins: Decoders of ubiquitin-mediated cellular functions. Annu
Rev Biochem. 81:291–322. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Komander D, Clague M and Urbé S: Breaking
the chains: Structure and function of the deubiquitinases. Nat Rev
Mol Cell Biol. 10:550–563. 2009. View
Article : Google Scholar : PubMed/NCBI
|
|
11
|
Clague M, Barsukov I, Coulson J, Liu H,
Rigden D and Urbé S: Deubiquitylases from genes to organism.
Physiol Rev. 93:1289–1315. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Swatek KN and Komander D: Ubiquitin
modifications. Cell Res. 26:399–422. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Chen ZJ and Sun LJ: Nonproteolytic
functions of ubiquitin in cell signaling. Molecular cell.
33:275–286. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Sippl W, Collura V and Colland F:
Ubiquitin-specific proteases as cancer drug targets. Future Oncol.
7:619–632. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
McClurg UL and Robson CN: Deubiquitinating
enzymes as oncotargets. Oncotarget. 6:9657–9668. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Kee Y and Huang TT: Role of
Deubiquitinating enzymes in DNA repair. Mol Cell Biol. 36:524–544.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Clague MJ, Coulson JM and Urbé S: Cellular
functions of the DUBs. J Cell Sci. 125:277–286. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Yang JM: Emerging roles of
deubiquitinating enzymes in human cancer. Acta Pharmacol Sin.
28:1325–1330. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Schwickart M, Huang X, Lill JR, Liu J,
Ferrando R, French DM, Maecker H, O'Rourke K, Bazan F,
Eastham-Anderson J, et al: Deubiquitinase USP9X stabilizes MCL1 and
promotes tumour cell survival. Nature. 463:103–107. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Priolo C, Tang D, Brahamandan M, Benassi
B, Sicinska E, Ogino S, Farsetti A, Porrello A, Finn S, Zimmermann
J, et al: The isopeptidase USP2a protects human prostate cancer
from apoptosis. Cancer Res. 66:8625–8632. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Popov N, Wanzel M, Madiredjo M, Zhang D,
Beijersbergen R, Bernards R, Moll R, Elledge SJ and Eilers M: The
ubiquitin-specific protease USP28 is required for MYC stability.
Nat Cell Biol. 9:765–774. 2007. View
Article : Google Scholar : PubMed/NCBI
|
|
22
|
Coyne ES and Wing SS: The business of
deubiquitination-location, location, location. F1000Res. 5:F1000
Faculty Rev-163. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Mevissen TE, Hospenthal MK, Geurink PP,
Elliott PR, Akutsu M, Arnaudo N, Ekkebus R, Kulathu Y, Wauer T, El
Oualid F, et al: OTU deubiquitinases reveal mechanisms of linkage
specificity and enable ubiquitin chain restriction analysis. Cell.
154:169–184. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Carneiro AP, Reis CF, Morari EC, Maia YC,
Nascimento R, Bonatto JM, de Souza MA, Goulart LR and Ward LS: A
putative OTU domain-containing protein 1 deubiquitinating enzyme is
differentially expressed in thyroid cancer and identifies
less-aggressive tumours. Br J Cancer. 111:551–558. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Zhang Z, Fan Y, Xie F, Zhou H, Jin K, Shao
L, Shi W, Fang P, Yang B, van Dam H, et al: Breast cancer
metastasis suppressor OTUD1 deubiquitinates SMAD7. Nat Commun.
8:21162017. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Piao S, Pei HZ, Huang B and Baek SH:
Ovarian tumor domain-containing protein 1 deubiquitinates and
stabilizes p53. Cell Signal. 33:22–29. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Yuan L, Lv Y, Li H, Gao H, Song S, Zhang
Y, Xing G, Kong X, Wang L, Li Y, et al: Deubiquitylase OTUD3
regulates PTEN stability and suppresses tumorigenesis. Nat Cell
Biol. 17:1169–1181. 2015. View
Article : Google Scholar : PubMed/NCBI
|
|
28
|
Zhao Y, Majid MC, Soll JM, Brickner JR,
Dango S and Mosammaparast N: Noncanonical regulation of alkylation
damage resistance by the OTUD4 deubiquitinase. EMBO J.
34:1687–1703. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
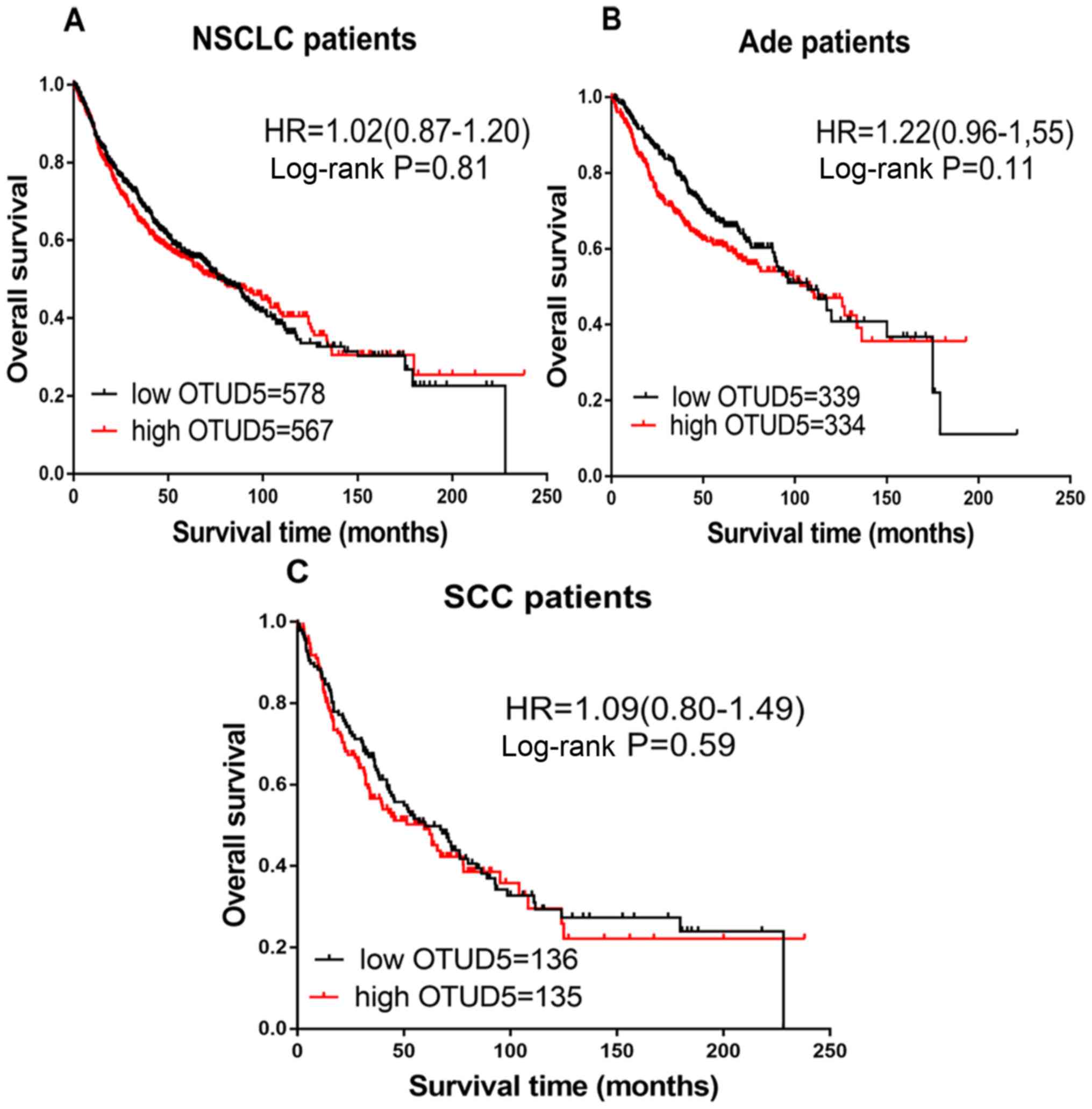
Luo J, Lu Z, Lu X, Chen L, Cao J, Zhang S,
Ling Y and Zhou X: OTUD5 regulates p53 stability by
deubiquitinating p53. PLoS One. 8:e776822013. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
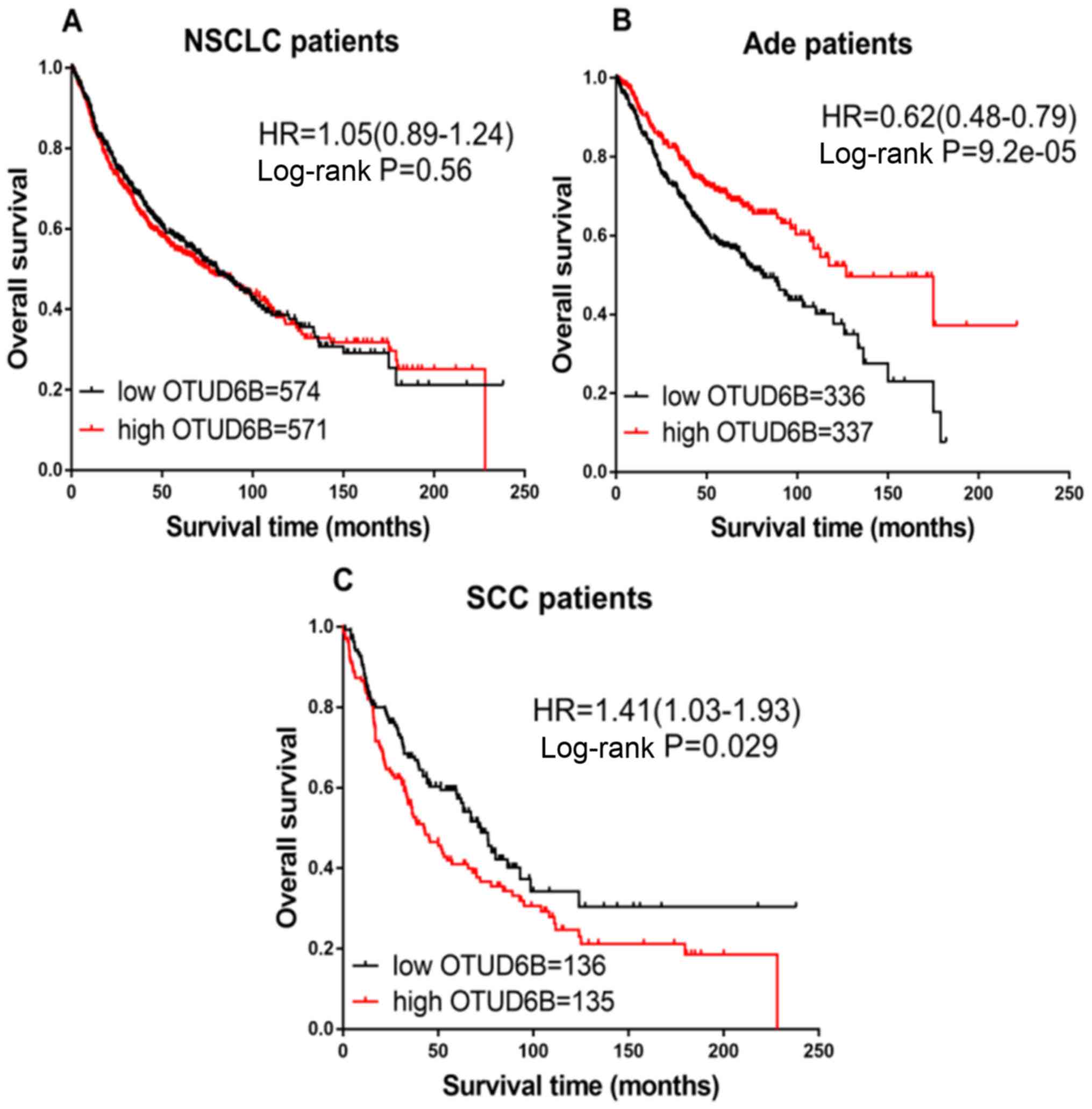
Sobol A, Askonas C, Alani S, Weber MJ,
Ananthanarayanan V, Osipo C and Bocchetta M: Deubiquitinase OTUD6B
isoforms are important regulators of growth and proliferation. Mol
Cancer Res. 15:117–127. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Rhodes D, Yu J, Shanker K, Deshpande N,
Varambally R, Ghosh D, Barrette T, Pandey A and Chinnaiyan AM:
ONCOMINE: A cancer microarray database and integrated data-mining
platform. Neoplasia. 6:1–6. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Rhodes D, Kalyana-Sundaram S, Mahavisno V,
Varambally R, Yu J, Briggs BB, Barrette TR, Anstet MJ, Kincead-Beal
C, Kulkarni P, et al: Oncomine 3.0: Genes, pathways, and networks
in a collection of 18,000 cancer gene expression profiles.
Neoplasia. 9:166–180. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Györffy B, Lanczky A, Eklund AC, Denkert
C, Budczies J, Li Q and Szallasi Z: An online survival analysis
tool to rapidly assess the effect of 22,277 genes on breast cancer
prognosis using microarray data of 1,809 patients. Breast Cancer
Res Treat. 123:725–731. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Győrffy B, Surowiak P, Budczies J and
Lánczky A: Online survival analysis software to assess the
prognostic value of biomarkers using transcriptomic data in
non-small-cell lung cancer. PLoS One. 8:e822412013. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Ivanova L, Zandberga E, Siliņa K, Kalniņa
Z, Ābols A, Endzeliņš E, Vendina I, Romanchikova N, Hegmane A,
Trapencieris P, et al: Prognostic relevance of carbonic anhydrase
IX expression is distinct in various subtypes of breast cancer and
its silencing suppresses self-renewal capacity of breast cancer
cells. Cancer Chemother Pharmacol. 75:235–246. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Wertz IE, Newton K, Seshasayee D¸, Kusam
S, Lam C, Zhang J, Popovych N, Helgason E, Schoeffler A, Jeet S, et
al: Phosphorylation and linear ubiquitin direct A20 inhibition of
inflammation. Nature. 528:370–375. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Xu Z, Zheng Y, Zhu Y, Kong X and Hu L:
Evidence for OTUD-6B participation in B lymphocytes cell cycle
after cytokine stimulation. PLoS One. 6:e145142011. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Ettinger DS, Wood DE, Aisner DL, Akerley
W, Bauman J, Chirieac LR, D'Amico TA, DeCamp MM, Dilling TJ,
Dobelbower M, et al: Non-small cell lung cancer, version 5.2017,
NCCN Clinical Practice Guidelines in Oncology. J Natl Compr Canc
Netw. 15:504–535. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Hou J, Aerts J, den Hamer B, van Ijcken W,
den Bakker M, Riegman P, van der Leest C, van der Spek P, Foekens
JA, Hoogsteden HC, et al: Gene expression-based classification of
non-small cell lung carcinomas and survival prediction. PLoS One.
5:e103122010. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Garber ME, Troyanskaya OG, Schluens K,
Petersen S, Thaesler Z, Pacyna-Gengelbach M, van de Rijn M, Rosen
GD, Perou CM, Whyte RI, et al: Diversity of gene expression in
adenocarcinoma of the lung. Proc Natl Acad Sci USA. 98:13784–13789.
2001. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Yamagata N, Shyr Y, Yanagisawa K, Edgerton
M, Dang TP, Gonzalez A, Nadaf S, Larsen P, Roberts JR, Nesbitt JC,
et al: A training-testing approach to the molecular classification
of resected non-small cell lung cancer. Clin Cancer Res.
9:4695–4704. 2003.PubMed/NCBI
|
|
42
|
Selamat SA, Chung BS, Girard L, Zhang W,
Zhang Y, Campan M, Siegmund KD, Koss MN, Hagen JA, Lam WL, et al:
Genome-scale analysis of DNA methylation in lung adenocarcinoma and
integration with mRNA expression. Genome Res. 22:1197–1211. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Makarova KS, Aravind L and Koonin EV: A
novel superfamily of predicted cysteine proteases from eukaryotes,
viruses and Chlamydia pneumoniae. Trends Biochem Sci. 25:50–52.
2000. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Cancer Genome Atlas Research Network, .
Comprehensive molecular characterization of urothelial bladder
carcinoma. Nature. 507:315–322. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Flierman D, van der Heden van Noort GJ,
Ekkebus R, Geurink PP, Mevissen TE, Hospenthal MK, Komander D and
Ovaa H: Non-hydrolyzable diubiquitin probes reveal linkage-specific
reactivity of deubiquitylating enzymes mediated by S2 pockets. Cell
Chem Biol. 23:472–482. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Rumpf S and Jentsch S: Functional division
of substrate processing cofactors of the ubiquitin-selective Cdc48
chaperone. Mol Cell. 21:261–269. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Li X, Wang S, Zhu R, Li H, Han Q and Zhao
RC: Lung tumor exosomes induce a pro-inflammatory phenotype in
mesenchymal stem cells via NFκB-TLR signaling pathway. J Hematol
Oncol. 9:422016. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Lu Z, Li Y, Wang J, Che Y, Sun S, Huang J,
Chen Z and He J: Long non-coding RNA NKILA inhibits migration and
invasion of non-small cell lung cancer via NF-κB/Snail pathway. J
Exp Clin Cancer Res. 36:542017. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Richardson JSM, Aminudin N and Abd Malek
SN: Chalepin: A compound from Ruta angustifolia L. pers
exhibits cell cycle arrest at S phase, suppresses nuclear
factor-kappa B (NF-κB) pathway, signal transducer and activation of
transcription 3 (STAT3) phosphorylation and extrinsic apoptotic
pathway in non-small cell lung cancer carcinoma (A549). Pharmacogn
Mag. 13 (Suppl 3):S489–S498. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Tang X, Sun L, Wang G, Chen B and Luo F:
RUNX1: A regulator of NF-kB signaling in pulmonary diseases. Curr
Protein Pept Sci. 19:172–178. 2018.PubMed/NCBI
|
|
51
|
Dhara A and Sinai AP: A cell
cycle-regulated toxoplasma deubiquitinase, TgOTUD3A, targets
polyubiquitins with specific lysine linkages. mSphere. 1:e00085–16.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Behnke MS, Wootton JC, Lehmann MM, Radke
JB, Lucas O, Nawas J, Sibley LD and White MW: Coordinated
progression through two subtranscriptomes underlies the tachyzoite
cycle of Toxoplasma gondii. PLoS One. 5:e123542010. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Huang OW, Ma X, Yin J, Flinders J, Maurer
T, Kayagaki N, Phung Q, Bosanac I, Arnott D, Dixit VM, et al:
Phosphorylation-dependent activity of the deubiquitinase DUBA. Nat
Struct Mol Biol. 19:171–175. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Kayagaki N, Phung Q, Chan S, Chaudhari R,
Quan C, O'Rourke KM, Eby M, Pietras E, Cheng G, Bazan JF, et al:
DUBA: A deubiquitinase that regulates type I interferon production.
Science. 318:1628–1632. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Santiago-Sim T, Burrage LC, Ebstein F,
Tokita MJ, Miller M, Bi W, Braxton AA, Rosenfeld JA, Shahrour M,
Lehmann A, et al: Biallelic variants in OTUD6B cause an
intellectual disability syndrome associated with seizures and
dysmorphic features. Am J Hum Genet. 100:676–688. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
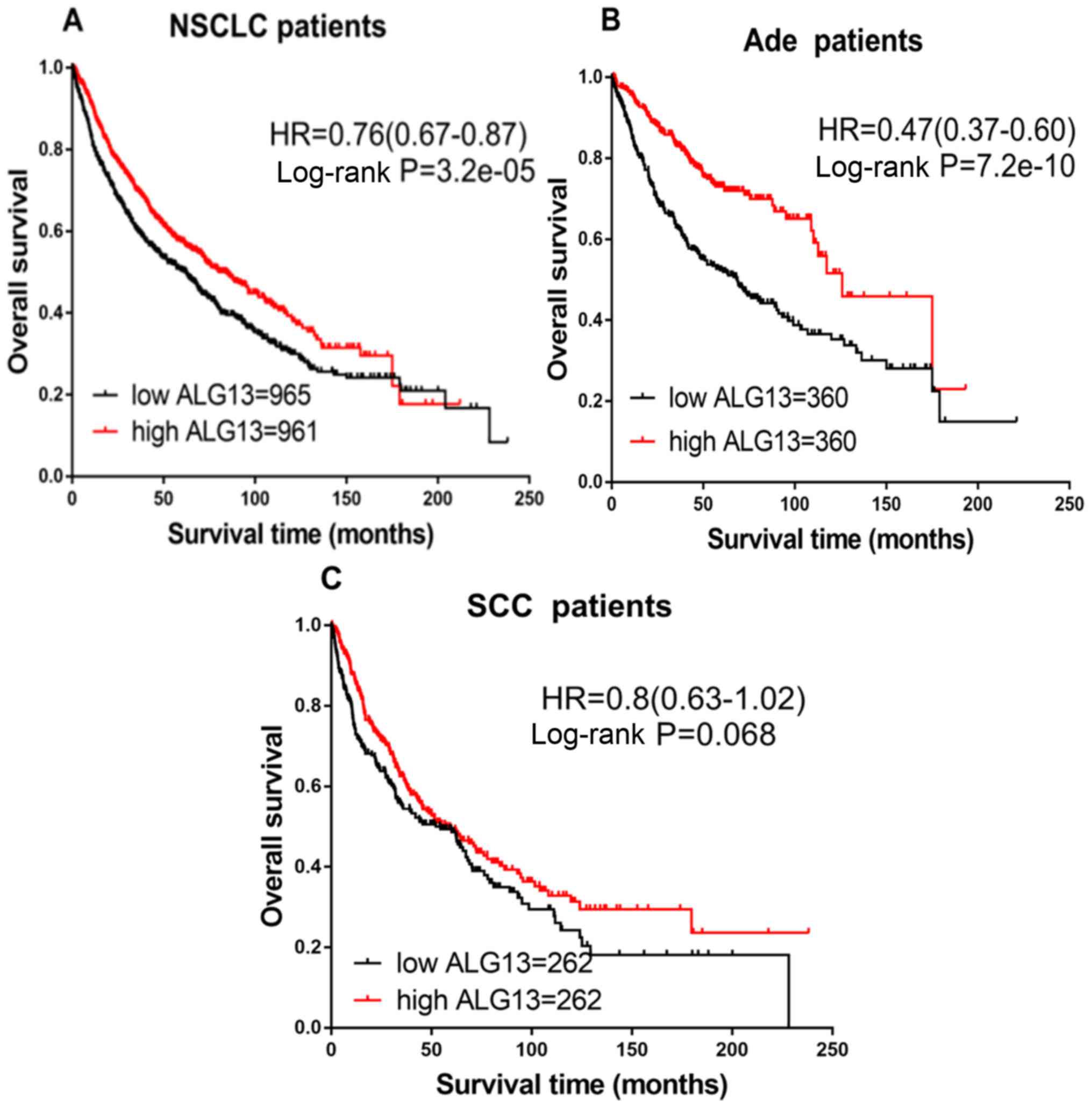
Wang X, Weldeghiorghis T, Zhang G,
Imperiali B and Prestegard JH: Solution structure of Alg13: The
sugar donor subunit of a yeast N-acetylglucosamine transferase.
Structure. 16:965–975. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
De Antonellis P, Carotenuto M,
Vandenbussche J, De Vita G, Ferrucci V, Medaglia C, Boffa I,
Galiero A, Di Somma S, Magliulo D, et al: Early targets of miR-34a
in neuroblastoma. Mol Cell Proteomics. 13:2114–2131. 2014.
View Article : Google Scholar : PubMed/NCBI
|