|
1
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2018. CA Cancer J Clin. 68:7–30. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Colzani E, Liljegren A, Johansson AL,
Adolfsson J, Hellborg H, Hall PF and Czene K: Prognosis of patients
with breast cancer: Causes of death and effects of time since
diagnosis, age, and tumor characteristics. J Clin Oncol.
29:4014–4021. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Avgeris M, Tsilimantou A, Levis PK, Tokas
T, Sideris DC, Stravodimos K, Ardavanis A and Scorilas A: Loss of
GAS5 tumour suppressor lncRNA: An independent molecular cancer
biomarker for short-term relapse and progression in bladder cancer
patients. Brit J Cancer. 119:1477–1486. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Chakraborty S, Andrieux G, Hasan AMM,
Ahmed M, Hosen MI, Rahman T, Hossain MA and Boerries M: Harnessing
the tissue and plasma lncRNA-peptidome to discover peptide-based
cancer biomarkers. Sci Rep. 9:123222019. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Yarmishyn AA, Batagov AO, Tan JZ, Sundaram
GM, Sampath P, Kuznetsov VA and Kurochkin IV: HOXD-AS1 is a novel
lncRNA encoded in HOXD cluster and a marker of neuroblastoma
progression revealed via integrative analysis of noncoding
transcriptome. BMC Genomics. (15 Suppl 9):S72014. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Sakaue S, Hirata J, Maeda Y, Kawakami E,
Nii T, Kishikawa T, Ishigaki K, Terao C, Suzuki K, Akiyama M, et
al: Integration of genetics and miRNA-target gene network
identified disease biology implicated in tissue specificity.
Nucleic Acids Res. 46:11898–11909. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Fu X, Mao X, Wang Y, Ding X and Li Y:
Let-7c-5p inhibits cell proliferation and induces cell apoptosis by
targeting ERCC6 in breast cancer. Oncol Rep. 38:1851–1856. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Hausser J, Syed AP, Bilen B and Zavolan M:
Analysis of CDS-located miRNA target sites suggests that they can
effectively inhibit translation. Genome Res. 23:604–615. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Shuwen H, Qing Z, Yan Z and Xi Y:
Competitive endogenous RNA in colorectal cancer: A systematic
review. Gene. 645:157–162. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Neumann P, Jaé N, Knau A, Glaser SF,
Fouani Y, Rossbach O, Kruger M, John D, Bindereif A, Grote P, et
al: The lncRNA GATA6-AS epigenetically regulates endothelial gene
expression via interaction with LOXL2. Nat Commun. 9:2372018.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Cerami E, Gao J, Dogrusoz U, Gross BE,
Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, et
al: The cBio cancer genomics portal: An open platform for exploring
multidimensional cancer genomics data. Cancer Discov. 2:401–404.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Gao J, Aksoy BA, Dogrusoz U, Dresdner G,
Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, et al:
Integrative analysis of complex cancer genomics and clinical
profiles using the cBioPortal. Sci Signal. 6:pl12013. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Huang da W, Sherman BT and Lempicki RA:
Systematic and integrative analysis of large gene lists using DAVID
bioinformatics resources. Nat Protoc. 4:44–57. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Huang DW, Sherman BT, Tan Q, Kir J, Liu D,
Bryant D, Guo Y, Stephens R, Baseler MW, Lane HC, et al: DAVID
Bioinformatics Resources: Expanded annotation database and novel
algorithms to better extract biology from large gene lists. Nucleic
Acids Res. 35:W169–W175. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Chang TH, Huang HY, Hsu JB, Weng SL, Horng
JT and Huang HD: An enhanced computational platform for
investigating the roles of regulatory RNA and for identifying
functional RNA motifs. BMC Bioinformatics. 14 (Suppl):S42013.
View Article : Google Scholar
|
|
16
|
Agarwal V, Bell GW, Nam JW and Bartel DP:
Predicting effective microRNA target sites in mammalian mRNAs.
ELife. 4:2015. View Article : Google Scholar
|
|
17
|
Shannon P, Markiel A, Ozier O, Baliga NS,
Wang JT, Ramage D, Amin N, Schwikowski B and Ideker T: Cytoscape: A
software environment for integrated models of biomolecular
interaction networks. Genome Res. 13:2498–2504. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Lang B, Armaos A and Tartaglia GG: RNAct:
Protein-RNA interaction predictions for model organisms with
supporting experimental data. Nucleic Acids Res. 47:D601–D606.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Guo J, Li P, Liu X and Li Y: NOTCH
signaling pathway and non-coding RNAs in cancer. Pathol Res Pract.
215:1526202019. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Youness RA and Gad MZ: Long non-coding
RNAs: Functional regulatory players in breast cancer. Non-coding
RNA Res. 4:36–44. 2019. View Article : Google Scholar
|
|
21
|
Kazan H, Ray D, Chan ET, Hughes TR and
Morris Q: RNAcontext: A new method for learning the sequence and
structure binding preferences of RNA-binding proteins. PLoS Comput
Boil. 6:e10008322010. View Article : Google Scholar
|
|
22
|
Weir BA, Woo MS, Getz G, Perner S, Ding L,
Beroukhim R, Lin WM, Province MA, Kraja A, Johnson LA, et al:
Characterizing the cancer genome in lung adenocarcinoma. Nature.
450:893–898. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Sanchez Calle A, Kawamura Y, Yamamoto Y,
Takeshita F and Ochiya T: Emerging roles of long non-coding RNA in
cancer. Cancer Sci. 109:2093–2100. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Gu J, Wang Y, Wang X, Zhou D, Shao C, Zhou
M and He Z: Downregulation of lncRNA GAS5 confers tamoxifen
resistance by activating miR-222 in breast cancer. Cancer Lett.
434:1–10. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
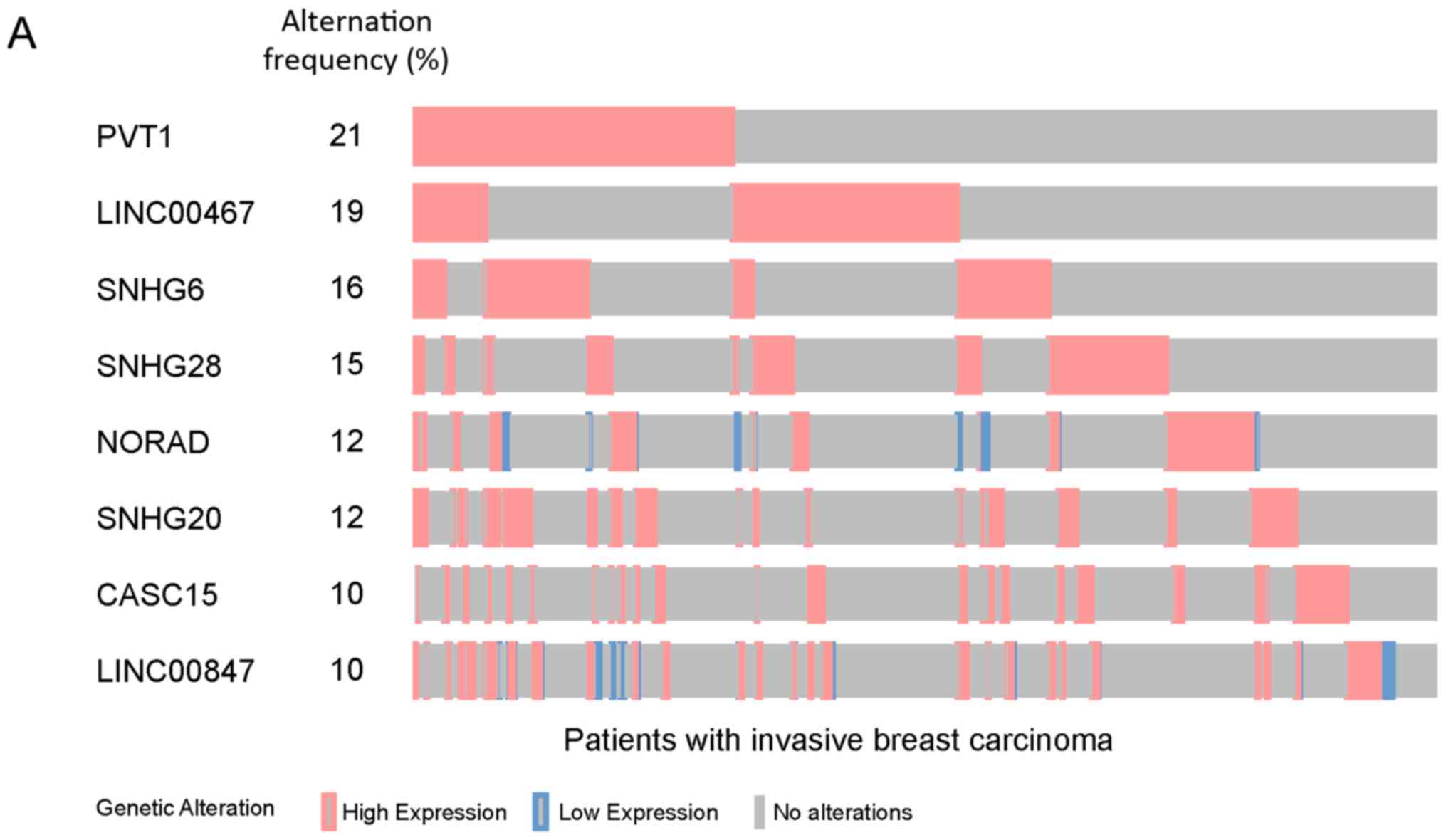
He Y, Li X, Meng Y, Fu S, Cui Y, Shi Y and
Du H: A prognostic 11 long noncoding RNA expression signature for
invasive breast carcinoma. J Cell Biochem. 120:16692–16702. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Sun J, Chen X, Wang Z, Guo M, Shi H, Wang
X, Cheng L and Zhou M: A potential prognostic long non-coding RNA
signature to predict metastasis-free survival of breast cancer
patients. Sci Rep. 5:165532015. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Liu H, Li J, Koirala P, Ding X, Chen B,
Wang Y, Wang Z, Wang C, Zhang X and Mo YY: Long non-coding RNAs as
prognostic markers in human breast cancer. Oncotarget.
7:20584–20596. 2016.PubMed/NCBI
|
|
28
|
Vishnubalaji R, Shaath H, Elkord E and
Alajez NM: Long non-coding RNA (lncRNA) transcriptional landscape
in breast cancer identifies LINC01614 as non-favorable prognostic
biomarker regulated by TGFβ and focal adhesion kinase (FAK)
signaling. Cell Death Discov. 5:1092019. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Zhou K, Ou Q, Wang G, Zhang W, Hao Y and
Li W: High long non-coding RNA NORAD expression predicts poor
prognosis and promotes breast cancer progression by regulating
TGF-β pathway. Cancer Cell Int. 19:632019. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Yu J, Yan Y, Hua C and Ming L:
Upregulation of lncRNA SNHG1 is associated with metastasis and poor
prognosis in cancers: A meta-analysis. Medicine. 98:e151962019.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Ma W, Zhang CQ, Dang CX, Cai HY, Li HL,
Miao GY, Wang JK and Zhang LJ: Upregulated long-non-coding RNA
DLEU2 exon 9 expression was an independent indicator of unfavorable
overall survival in patients with esophageal adenocarcinoma. Biomed
Pharmacother. 113:1086552019. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Chen Z, Ju H, Yu S, Zhao T, Jing X, Li P,
Jia J, Li N, Tan B and Li Y: Prader-Willi region non-protein coding
RNA 1 suppressed gastric cancer growth as a competing endogenous
RNA of miR-425-5p. Clin Sci (Lond). 132:1003–1019. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Su H, Sun T, Wang H, Shi G, Zhang H, Sun F
and Ye D: Decreased TCL6 expression is associated with poor
prognosis in patients with clear cell renal cell carcinoma.
Oncotarget. 8:5789–5799. 2017.PubMed/NCBI
|
|
34
|
Kong X, Duan Y, Sang Y, Li Y, Zhang H,
Liang Y, Liu Y, Zhang N and Yang Q: lncRNA-CDC6 promotes breast
cancer progression and function as ceRNA to target CDC6 by sponging
microRNA-215. J Cell Physiol. 234:9105–9117. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Xia F, Jin P, Ding Y, Li F and Shi C:
GON4L drives nasopharyngeal carcinoma growth and proliferation
through regulation of β-catenin/wnt singling pathway. Biomed Res.
28:4348–4353. 2017.
|
|
36
|
Lam S, Lodder K, Teunisse AF, Rabelink MJ,
Schutte M and Jochemsen AG: Role of Mdm4 in drug sensitivity of
breast cancer cells. Oncogene. 29:2415–2426. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Liu H, Ma Y, He HW, Wang JP, Jiang JD and
Shao RG: SLC9A3R1 stimulates autophagy via BECN1 stabilization in
breast cancer cells. Autophagy. 11:2323–2334. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Li Q and Lozano G: Molecular pathways:
Targeting Mdm2 and Mdm4 in cancer therapy. Clin Cancer Res.
19:34–41. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Busà R, Paronetto MP, Farini D,
Pierantozzi E, Botti F, Angelini DF, Attisani F, Vespasiani G and
Sette C: The RNA-binding protein Sam68 contributes to proliferation
and survival of human prostate cancer cells. Oncogene.
26:4372–4382. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Chen S, Wu DD, Sang XB, Wang LL, Zong ZH,
Sun KX, Liu BL and Zhao Y: The lncRNA HULC functions as an oncogene
by targeting ATG7 and ITGB1 in epithelial ovarian carcinoma. Cell
Death Dis. 8:e31182017. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Souchelnytskyi S: Proteomics of TGF-beta
signaling and its impact on breast cancer. Expert Rev Proteomics.
2:925–935. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Chano T, Ikebuchi K, Ochi Y, Tameno H,
Tomita Y, Jin Y, Inaji H, Ishitobi M, Teramoto K, Nishimura I, et
al: RB1CC1 activates RB1 pathway and inhibits proliferation and
cologenic survival in human cancer. PLoS One. 5:e114042010.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Young LC, Hartig N, Muñoz-Alegre M,
Oses-Prieto JA, Durdu S, Bender S, Vijayakumar V, Vietri Rudan M,
Gewinner C, Henderson S, et al: An MRAS, SHOC2, and SCRIB complex
coordinates ERK pathway activation with polarity and tumorigenic
growth. Mol Cell. 52:679–692. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Mathias C, Zambalde EP, Rask P, Gradia DF
and de Oliveira JC: Long non-coding RNAs differential expression in
breast cancer subtypes: What do we know? Clin Genet. 95:558–568.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Klinge CM: Non-coding RNAs in breast
cancer: Intracellular and intercellular communication. Non-Coding
RNA. 4(pii): E402018. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Zheng Y, Xu Q, Liu M, Hu H, Xie Y, Zuo Z
and Ren J: lnCAR: A comprehensive resource for lncRNAs from cancer
arrays. Cancer Res. 79:2076–2083. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Chen L, Sun F, Yang X, Jin Y, Shi M, Wang
L, Shi Y, Zhan C and Wang Q: Correlation between RNA-Seq and
microarrays results using TCGA data. Gene. 628:200–204. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Zheng S, Li M, Miao K and Xu H: SNHG1
contributes to proliferation and invasion by regulating miR-382 in
breast cancer. Cancer Manag Res. 11:5589–5598. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Zhang YL, Li XB, Hou YX, Fang NZ, You JC
and Zhou QH: The lncRNA XIST exhibits oncogenic properties via
regulation of miR-449a and Bcl-2 in human non-small cell lung
cancer. Acta Pharmacol Sin. 38:371–381. 2017. View Article : Google Scholar : PubMed/NCBI
|