|
1
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 6:394–424. 2018. View Article : Google Scholar
|
|
2
|
Ng L, Poon RT and Pang R: Biomarkers for
predicting future metastasis of human gastrointestinal tumors. Cell
Mol Life Sci. 70:3631–3656. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Fidler MM, Gupta S, Soerjomataram I,
Ferlay J, Steliarova-Foucher E and Bray F: Cancer incidence and
mortality among young adults aged 20–39 years worldwide in 2012: A
population-based study. Lancet Oncol. 18:1579–1589. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Kim JL, Cho KH, Park EC and Cho WH: A
single measure of cancer burden combining incidence with mortality
rates for worldwide application. Asian Pac J Cancer Prev.
15:433–439. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Qin L, Chen C, Chen L, Xue R, Ou-Yang M,
Zhou C, Zhao S, He Z, Xia Y, He J, et al: Worldwide malaria
incidence and cancer mortality are inversely associated. Infect
Agent Cancer. 12:142017. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Dhamija S and Diederichs S: From junk to
master regulators of invasion: lncRNA functions in migration, EMT
and metastasis. Int J Cancer. 139:269–280. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Bhan A and Mandal SS: LncRNA HOTAIR: A
master regulator of chromatin dynamics and cancer. Biochim Biophys
Acta. 1856:151–164. 2015.PubMed/NCBI
|
|
8
|
Yang G, Lu X and Yuan L: LncRNA: A link
between RNA and cancer. Biochim Biophys Acta. 1839:1097–1109. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Lin C and Yang L: Long noncoding RNA in
cancer: Wiring signaling circuitry. Trends Cell Biol. 28:287–301.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Li CH and Chen Y: Targeting long
non-coding RNAs in cancers: Progress and prospects. Int J Biochem
Cell Biol. 45:1895–1910. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Zhou M, Zhao H, Wang Z, Cheng L, Yang L,
Shi H, Yang H and Sun J: Identification and validation of potential
prognostic lncRNA biomarkers for predicting survival in patients
with multiple myeloma. J Exp Clin Cancer Res. 34:1022015.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Giulietti M, Righetti A, Principato G and
Piva F: lncRNA Co-expression network analysis reveals novel
biomarkers for pancreatic cancer. Carcinogenesis. 39:1016–1025.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Ning L, Li Z, Wei D, Chen H and Yang C:
lncRNA, NEAT1 is a prognosis biomarker and regulates cancer
progression via epithelial-mesenchymal transition in clear cell
renal cell carcinoma. Cancer Biomark. 19:75–83. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
White NM, Cabanski CR, Silva-Fisher JM,
Dang HX, Govindan R and Maher CA: Transcriptome sequencing reveals
altered long intergenic non-coding RNAs in lung cancer. Genome
Biol. 15:4292014. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Liu AN, Qu HJ, Yu CY and Sun P: Knockdown
of LINC01614 inhibits lung adenocarcinoma cell progression by
up-regulating miR-217 and down-regulating FOXP1. J Cell Mol Med.
22:4034–4044. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Vishnubalaji R, Shaath H, Elkord E and
Alajez NM: Long non-coding RNA (lncRNA) transcriptional landscape
in breast cancer identifies LINC01614 as non-favorable prognostic
biomarker regulated by TGF β and focal adhesion kinase (FAK)
signaling. Cell Death Discov. 5:1092019. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Sun Y and Ling C: Analysis of the long
non-coding RNA LINC01614 in non-small cell lung cancer. Medicine
(Baltimore). 98:e164372019. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Wang Y, Song B, Zhu L and Zhang X: Long
non-coding RNA, LINC01614 as a potential biomarker for prognostic
prediction in breast cancer. PeerJ. 7:e79762019. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Guo Y, Sheng Q, Li J, Ye F, Samuels DC and
Shyr Y: Large scale comparison of gene expression levels by
microarrays and RNAseq using TCGA data. PLoS One. 8:e714622013.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Budczies J, Klauschen F, Sinn BV, Győrffy
B, Schmitt WD, Darb-Esfahani S and Denkert C: Cutoff Finder: A
comprehensive and straightforward Web application enabling rapid
biomarker cutoff optimization. PLoS One. 7:e518622012. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
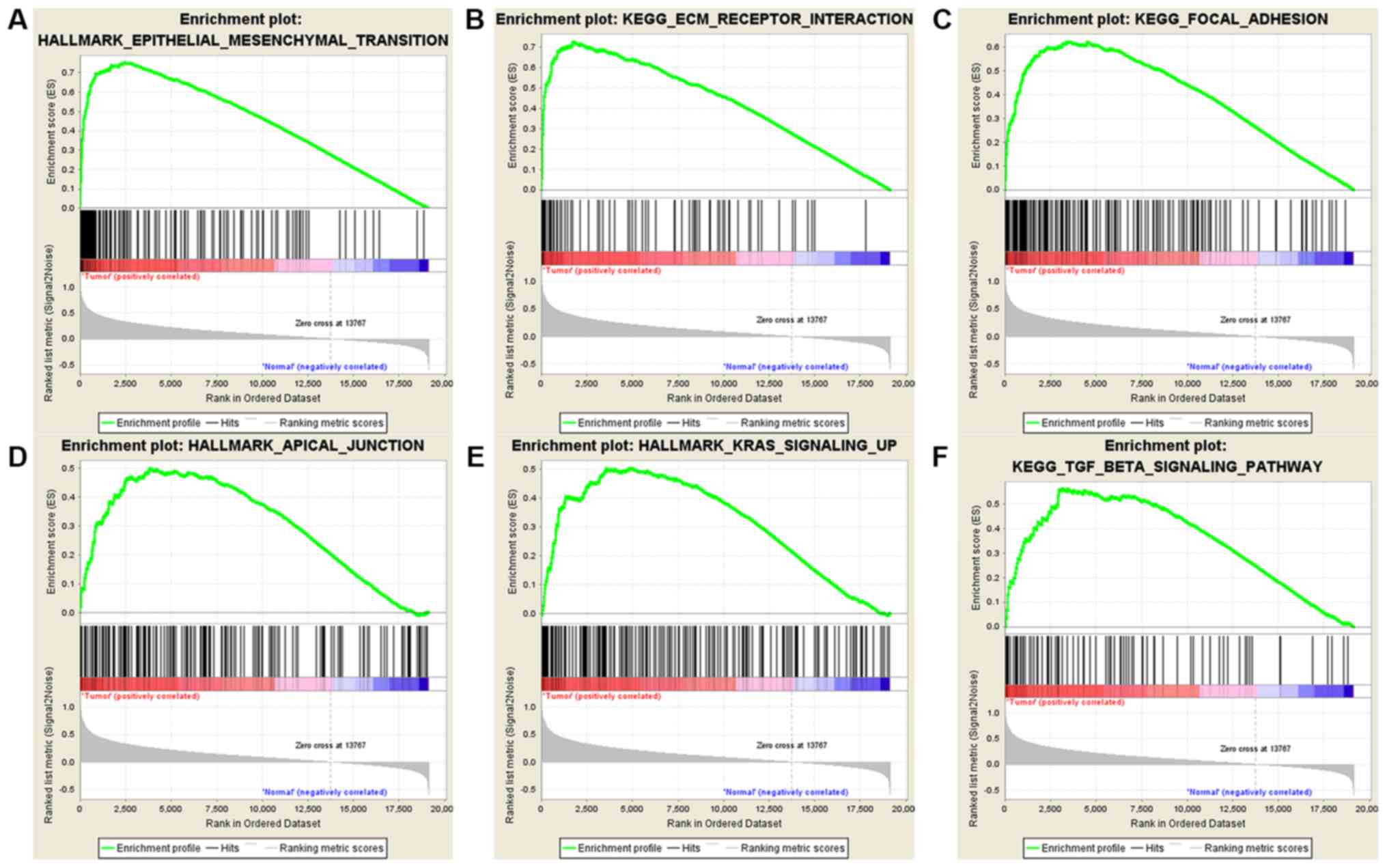
Subramanian A, Tamayo P, Mootha VK,
Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub
TR, Lander ES and Mesirov JP: Gene set enrichment analysis: A
knowledge-based approach for interpreting genome-wide expression
profiles. Proc Natl Acad Sci USA. 102:15545–15550. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Liberzon A, Birger C, Thorvaldsdottir H,
Ghandi M, Mesirov JP and Tamayo P: The Molecular Signatures
Database (MSigDB) hallmark gene set collection. Cell Syst.
1:417–425. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
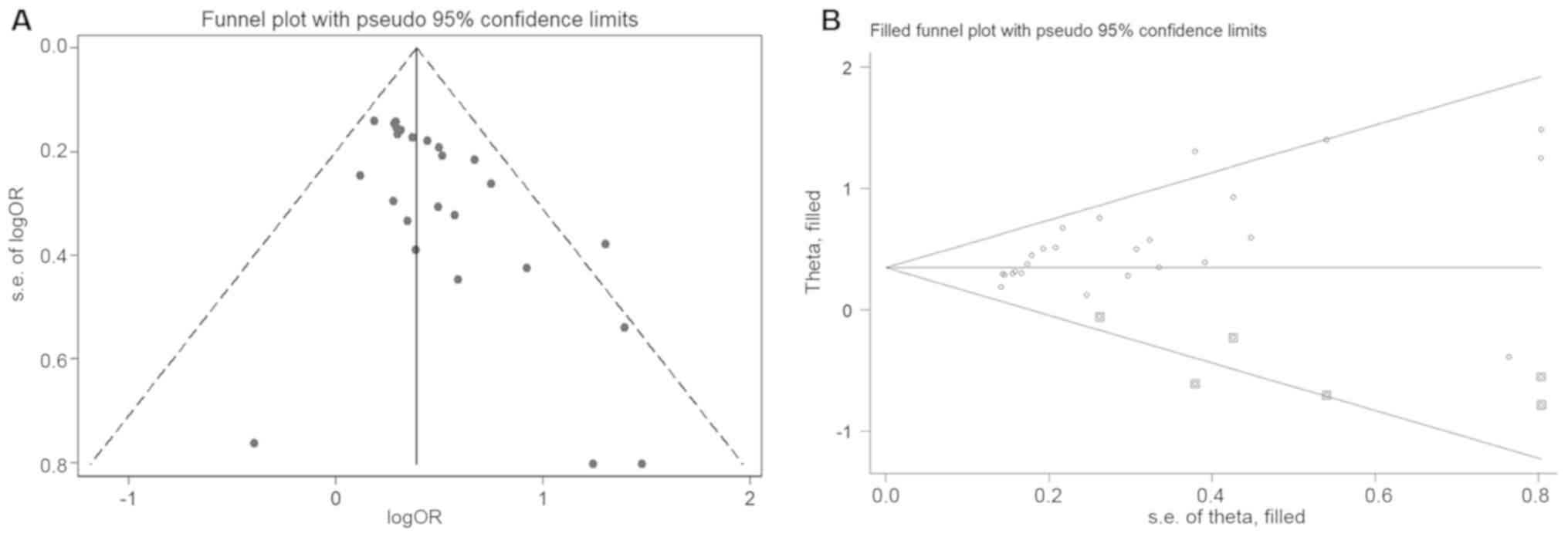
Duval S and Tweedie R: Trim and fill: A
simple funnel-plot-based method of testing and adjusting for
publication bias in meta-analysis. Biometrics. 56:455–463. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Iwakiri J, Hamada M and Asai K:
Bioinformatics tools for lncRNA research. Biochim Biophys Acta.
1859:23–30. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Bassett AR, Akhtar A, Barlow DP, Bird AP,
Brockdorff N, Duboule D, Ephrussi A, Ferguson-Smith AC, Gingeras
TR, Haerty W, et al: Considerations when investigating lncRNA
function in vivo. ELife. 3:e030582014. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Joung J, Engreitz JM, Konermann S,
Abudayyeh OO, Verdine VK, Aguet F, Gootenberg JS, Sanjana NE,
Wright JB, Fulco CP, et al: Genome-scale activation screen
identifies a lncRNA locus regulating a gene neighbourhood. Nature.
548:343–346. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Shibayama Y, Fanucchi S, Magagula L and
Mhlanga MM: lncRNA and gene looping: What's the connection?
Transcription. 5:e286582014. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Galuschka C, Proynova R, Roth B, Augustin
HG and Muller-Decker K: Models in Translational Oncology: A public
resource database for preclinical cancer research. Cancer Res.
77:2557–2563. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Kim H, Watkinson J, Varadan V and
Anastassiou D: Multi-cancer computational analysis reveals
invasion-associated variant of desmoplastic reaction involving
INHBA, THBS2 and COL11A1. BMC Med Genomics. 3:512010. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Shi JW, Liu W, Zhang TT, Wang SC, Lin XL,
Li J, Jia JS, Sheng HF, Yao ZF, Zhao WT, et al: The enforced
expression of c-Myc in pig fibroblasts triggers
mesenchymal-epithelial transition (MET) via F-actin reorganization
and RhoA/Rock pathway inactivation. Cell Cycle. 12:1119–1127. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Zhang B, Zhang C, Yang X, Chen Y, Zhang H,
Liu J and Wu Q: Cytoplasmic collagen XIαI as a prognostic biomarker
in esophageal squamous cell carcinoma. Cancer Biol Ther.
19:364–372. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Zhu H, Chen H, Wang J, Zhou L and Liu S:
Collagen stiffness promoted non-muscle-invasive bladder cancer
progression to muscle-invasive bladder cancer. Onco Targets Ther.
12:3441–3457. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Liu J, Huang C, Peng C, Xu F, Li Y, Yutaka
Y, Xiong B and Yang X: Stromal fibroblast activation protein alpha
promotes gastric cancer progression via epithelial-mesenchymal
transition through Wnt/β-catenin pathway. BMC Cancer. 18:10992018.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Yang X, Lin Y, Shi Y, Li B, Liu W, Yin W,
Dang Y, Chu Y, Fan J and He R: FAP promotes immunosuppression by
Cancer-Associated fibroblasts in the tumor Microenvironment via
STAT3-CCL2 signaling. Cancer Res. 76:4124–4135. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Chen L, Tian X, Gong W, Sun B, Li G, Liu
D, Guo P, He Y, Chen Z, Xia Y, et al: Periostin mediates
epithelial-mesenchymal transition through the MAPK/ERK pathway in
hepatoblastoma. Cancer Biol Med. 16:89–100. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Choi KU, Yun JS, Lee IH, Heo SC, Shin SH,
Jeon ES, Choi YJ, Suh DS, Yoon MS and Kim JH: Lysophosphatidic
acid-induced expression of periostin in stromal cells: Prognoistic
relevance of periostin expression in epithelial ovarian cancer. Int
J Cancer. 128:332–342. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Shi Q, Bao S, Song L, Wu Q, Bigner DD,
Hjelmeland AB and Rich JN: Targeting SPARC expression decreases
glioma cellular survival and invasion associated with reduced
activities of FAK and ILK kinases. Oncogene. 26:4084–4094. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Zhang F, Zhang Y, Da J, Jia Z, Wu H and Gu
K: Downregulation of SPARC expression decreases cell migration and
invasion involving epithelial-mesenchymal transition through the
p-FAK/p-ERK pathway in esophageal squamous cell carcinoma. J
Cancer. 11:414–420. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Acloque H, Adams MS, Fishwick K,
Bronner-Fraser M and Nieto MA: Epithelial-mesenchymal transitions:
the importance of changing cell state in development and disease. J
Clin Invest. 119:1438–1449. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Thiery JP, Acloque H, Huang RY and Nieto
MA: Epithelial-mesenchymal transitions in development and disease.
Cell. 139:871–890. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Nieto MA: The ins and outs of the
epithelial to mesenchymal transition in health and disease. Annu
Rev Cell Dev Biol. 27:347–376. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Furuya S, Endo K, Takahashi A, Miyazawa K
and Saitoh M: Snail suppresses cellular senescence and promotes
fibroblast-led cancer cell invasion. FEBS Open Bio. 7:1586–1597.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Nieto MA: Context-specific roles of EMT
programmes in cancer cell dissemination. Nat Cell Biol. 19:416–418.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Lee SJ, Seol HJ, Lee HW, Kang WY, Kang BG,
Jin J, Jo MY, Jin Y, Lee JI, Joo KM and Nam DH: Gene silencing of
c-Met leads to brain metastasis inhibitory effects. Clin Exp
Metastasis. 30:845–854. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Chaffer CL and Weinberg RA: A perspective
on cancer cell metastasis. Science. 331:1559–1564. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Jie XX, Zhang XY and Xu CJ:
Epithelial-to-mesenchymal transition, circulating tumor cells and
cancer metastasis: Mechanisms and clinical applications.
Oncotarget. 8:81558–81571. 2017. View Article : Google Scholar : PubMed/NCBI
|