|
1
|
Jankowsky E, Gross CH, Shuman S and Pyle
AM: Active disruption of an RNA-protein interaction by a DExH/D RNA
helicase. Science. 291:121–125. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Fairman-Williams ME, Guenther UP and
Jankowsky E: SF1 and SF2 helicases: Family matters. Curr Opin
Struct Biol. 20:313–324. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
He Y, Andersen GR and Nielsen KH:
Structural basis for the function of DEAH helicases. EMBO Rep.
11:180–186. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Smith WA, Schurter BT, Wong-Staal F and
David M: Arginine methylation of RNA helicase a determines its
subcellular localization. J Biol Chem. 279:22795–22798. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Singleton MR, Dillingham MS and Wigley DB:
Structure and mechanism of helicases and nucleic acid translocases.
Annu Rev Biochem. 76:23–50. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Marchat LA, Arzola-Rodríguez SI,
Hernandez-de la Cruz O, Lopez-Rosas I and Lopez-Camarillo C:
DEAD/DExH-Box RNA Helicases in Selected Human Parasites. Korean J
Parasitol. 53:583–595. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Du Pont KE, Davidson RB, McCullagh M and
Geiss BJ: Motif V regulates energy transduction between the
flavivirus NS3 ATPase and RNA-binding cleft. J Biol Chem.
295:1551–1564. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Robert F and Pelletier J: Perturbations of
RNA helicases in cancer. Wiley Interdiscip Rev RNA. 4:333–349.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Abdelhaleem M: RNA helicases: Regulators
of differentiation. Clin Biochem. 38:499–503. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Godbout R, Li L, Liu RZ and Roy K: Role of
DEAD box 1 in retinoblastoma and neuroblastoma. Future Oncol.
3:575–587. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Taunk NK, Goyal S, Wu H, Moran MS, Chen S
and Haffty BG: DEAD box 1 (DDX1) expression predicts for local
control and overall survival in early stage, node-negative breast
cancer. Cancer. 118:888–898. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Eberle J, Krasagakis K and Orfanos CE:
Translation initiation factor eIF-4A1 mRNA is consistently
overexpressed in human melanoma cells in vitro. Int J Cancer.
71:396–401. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Shuda M, Kondoh N, Tanaka K, Ryo A,
Wakatsuki T, Hada A, Goseki N, Igari T, Hatsuse K, Aihara T, et al:
Enhanced expression of translation factor mRNAs in hepatocellular
carcinoma. Anticancer Res. 20:2489–2494. 2000.PubMed/NCBI
|
|
14
|
Heerma van Voss MR, Schrijver WAME, Ter
Hoeve ND, Hoefnagel LD, Manson QF, van der Wall E, Raman V and van
Diest PJ; Dutch Distant Breast Cancer Metastases Consortium, : The
prognostic effect of DDX3 upregulation in distant breast cancer
metastases. Clin Exp Metastasis. 34:85–92. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Wang Z, Shen GH, Xie JM, Li B and Gao QG:
Rottlerin upregulates DDX3 expression in hepatocellular carcinoma.
Biochem Biophys Res Commun. 495:1503–1509. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Xia Q, Kong XT, Zhang GA, Hou XJ, Qiang H
and Zhong RQ: Proteomics-based identification of DEAD-box protein
48 as a novel autoantigen, a prospective serum marker for
pancreatic cancer. Biochem Biophys Res Commun. 330:526–532. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Lin J, Chen Q, Yang J, Qian J, Deng ZQ,
Qian W, Chen XX, Ma JC, Xiong DS, Ma YJ, et al: DDX43 promoter is
frequently hypomethylated and may predict a favorable outcome in
acute myeloid leukemia. Leuk Res. 38:601–607. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Ma N, Xu HE, Luo Z, Zhou J, Zhou Y and Liu
M: Expression and significance of DDX43 in lung adenocarcinoma. Pak
J Pharm Sci. 30 (Suppl 4):1491–1496. 2017.PubMed/NCBI
|
|
19
|
Dai L, Pan G, Liu X, Huang J, Jiang Z, Zhu
X, Gan X, Xu Q and Tan N: High expression of ALDOA and DDX5 are
associated with poor prognosis in human colorectal cancer. Cancer
Manag Res. 10:1799–1806. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Taniguchi K, Sugito N, Kumazaki M,
Shinohara H, Yamada N, Matsuhashi N, Futamura M, Ito Y, Otsuki Y,
Yoshida K, et al: Positive feedback of DDX6/c-Myc/PTB1 regulated by
miR-124 contributes to maintenance of the Warburg effect in colon
cancer cells. Biochim Biophys Acta. 1852:1971–1980. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
He Y, Zhang D, Yang Y, Wang X, Zhao X,
Zhang P, Zhu H, Xu N and Liang S: A double-edged function of DDX3,
as an oncogene or tumor suppressor, in cancer progression (Review).
Oncol Rep. 39:883–892. 2018.PubMed/NCBI
|
|
22
|
Kim KH, Kang YJ, Jo JO, Ock MS, Moon SH,
Suh DS, Yoon MS, Park ES, Jeong N, Eo WK, et al: DDX4 (DEAD box
polypeptide 4) colocalizes with cancer stem cell marker CD133 in
ovarian cancers. Biochem Biophys Res Commun. 447:315–322. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Schudrowitz N, Takagi S, Wessel GM and
Yajima M: Germline factor DDX4 functions in blood-derived cancer
cell phenotypes. Cancer Sci. 108:1612–1619. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Janknecht R: Multi-talented DEAD-box
proteins and potential tumor promoters: p68 RNA helicase (DDX5) and
its paralog, p72 RNA helicase (DDX17). Am J Transl Res. 2:223–234.
2010.PubMed/NCBI
|
|
25
|
Taniguchi K, Iwatsuki A, Sugito N,
Shinohara H, Kuranaga Y, Oshikawa Y, Tajirika T, Futamura M,
Yoshida K, Uchiyama K, et al: Oncogene RNA helicase DDX6 promotes
the process of c-Myc expression in gastric cancer cells. Mol
Carcinog. 57:579–589. 2018. View
Article : Google Scholar : PubMed/NCBI
|
|
26
|
Bhattacharya C, Wang X and Becker D: The
DEAD/DEAH box helicase, DDX11, is essential for the survival of
advanced melanomas. Mol Cancer. 11:822012. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Li J, Liu L, Liu X, Xu P, Hu Q and Yu Y:
The role of upregulated DDX11 as a potential prognostic and
diagnostic biomarker in lung adenocarcinoma. J Cancer.
10:4208–4216. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Vychytilova-Faltejskova P, Svobodova
Kovarikova A, Grolich T, Prochazka V, Slaba K, Machackova T,
Halamkova J, Svoboda M, Kala Z, Kiss I, et al: MicroRNA biogenesis
pathway genes are deregulated in colorectal cancer. Int J Mol Sci.
20:202019. View Article : Google Scholar
|
|
29
|
Zhang T, Ma Z, Liu L, Sun J, Tang H, Zhang
B, Zou Y and Li H: DDX39 promotes hepatocellular carcinoma growth
and metastasis through activating Wnt/β-catenin pathway. Cell Death
Dis. 9:6752018. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Sugiura T, Nagano Y and Noguchi Y: DDX39,
upregulated in lung squamous cell cancer, displays RNA helicase
activities and promotes cancer cell growth. Cancer Biol Ther.
6:957–964. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Lee S, Baek M, Yang H, Bang YJ, Kim WH, Ha
JH, Kim DK and Jeoung DI: Identification of genes differentially
expressed between gastric cancers and normal gastric mucosa with
cDNA microarrays. Cancer Lett. 184:197–206. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Hellman K, Alaiya AA, Becker S, Lomnytska
M, Schedvins K, Steinberg W, Hellström AC, Andersson S, Hellman U
and Auer G: Differential tissue-specific protein markers of vaginal
carcinoma. Br J Cancer. 100:1303–1314. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Cho B, Lee H, Jeong S, Bang YJ, Lee HJ,
Hwang KS, Kim HY, Lee YS, Kang GH and Jeoung DI: Promoter
hypomethylation of a novel cancer/testis antigen gene CAGE is
correlated with its aberrant expression and is seen in premalignant
stage of gastric carcinoma. Biochem Biophys Res Commun. 307:52–63.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Park SY, Kim WJ, Byun JH, Lee JJ, Jeoung
D, Park ST and Kim Y: Role of DDX53 in taxol-resistance of cervix
cancer cells in vitro. Biochem Biophys Res Commun. 506:641–647.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Cao S, Sun R, Wang W, Meng X, Zhang Y,
Zhang N and Yang S: RNA helicase DHX9 may be a therapeutic target
in lung cancer and inhibited by enoxacin. Am J Transl Res.
9:674–682. 2017.PubMed/NCBI
|
|
36
|
Ito S and Koso H: RNA helicase DHX15 acts
as a tumor suppressor in glioma. Cancer Science. Wiley; Hoboken,
NJ, USA: 109. pp. 782018
|
|
37
|
Abdelhaleem M: The novel helicase
homologue DDX32 is down-regulated in acute lymphoblastic leukemia.
Leuk Res. 26:945–954. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Alli Z, Ho M and Abdelhaleem M: Expression
of DHX32 in lymphoid tissues. Exp Mol Pathol. 79:219–223. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Landry JR, Mager DL and Wilhelm BT:
Complex controls: The role of alternative promoters in mammalian
genomes. Trends Genet. 19:640–648. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Tai CL, Pan WC, Liaw SH, Yang UC, Hwang LH
and Chen DS: Structure-based mutational analysis of the hepatitis C
virus NS3 helicase. J Virol. 75:8289–8297. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Caruthers JM and McKay DB: Helicase
structure and mechanism. Curr Opin Struct Biol. 12:123–133. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Schneider S, Hotz HR and Schwer B:
Characterization of dominant-negative mutants of the DEAH-box
splicing factors Prp22 and Prp16. J Biol Chem. 277:15452–15458.
2002. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Alli Z, Ackerley C, Chen Y, Al-Saud B and
Abdelhaleem M: Nuclear and mitochondrial localization of the
putative RNA helicase DHX32. Exp Mol Pathol. 81:245–248. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Di Liegro CM, Schiera G and Di Liegro I:
Regulation of mRNA transport, localization and translation in the
nervous system of mammals (Review). Int J Mol Med. 33:747–762.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Alli Z, Ho M, Ackerley C and Abdelhaleem
M: Characterization of murine Dhx32. Exp Mol Pathol. 83:115–118.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Iborra FJ, Kimura H and Cook PR: The
functional organization of mitochondrial genomes in human cells.
BMC Biol. 2:92004. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Chen Y, Alli Z, Ackerley C, Al-Saud B and
Abdelhaleem M: Altered distribution of heat shock protein 60
(Hsp60) with dysregulated expression of DHX32. Exp Mol Pathol.
82:256–261. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Abdelhaleem M, Sun TH and Ho M: DHX32
expression suggests a role in lymphocyte differentiation.
Anticancer Res. 25:2645–2648. 2005.PubMed/NCBI
|
|
49
|
McNeer NA, Philip J, Geiger H, Ries RE,
Lavallée VP, Walsh M, Shah M, Arora K, Emde AK, Robine N, et al:
Genetic mechanisms of primary chemotherapy resistance in pediatric
acute myeloid leukemia. Leukemia. 33:1934–1943. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Alli Z, Nam EH, Beimnet K and Abdelhaleem
M: The activation-induced expression of DHX32 in Jurkat T cells is
specific and involves calcium and nuclear factor of activated T
cells. Cell Immunol. 237:141–146. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Alli Z, Chen Y, Abdul Wajid S, Al-Saud B
and Abdelhaleem M: A role for DHX32 in regulating T-cell apoptosis.
Anticancer Res. 27:373–377. 2007.PubMed/NCBI
|
|
52
|
Huang C, Liang X, Huang R and Zhang Z:
Up-regulation and clinical relevance of novel helicase homologue
DHX32 in colorectal cancer. Journal of experimental & clinical
cancer research. CR (East Lansing Mich). 28:112009.
|
|
53
|
Lin H, Liu W, Fang Z, Liang X, Li J, Bai
Y, Lin L, You H, Pei Y, Wang F, et al: Overexpression of DHX32
contributes to the growth and metastasis of colorectal cancer. Sci
Rep. 5:92472015. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Lin H, Fang Z, Su Y, Li P, Wang J, Liao H,
Hu Q, Ye C, Fang Y, Luo Q, et al: DHX32 promotes angiogenesis in
colorectal cancer through augmenting β-catenin signaling to induce
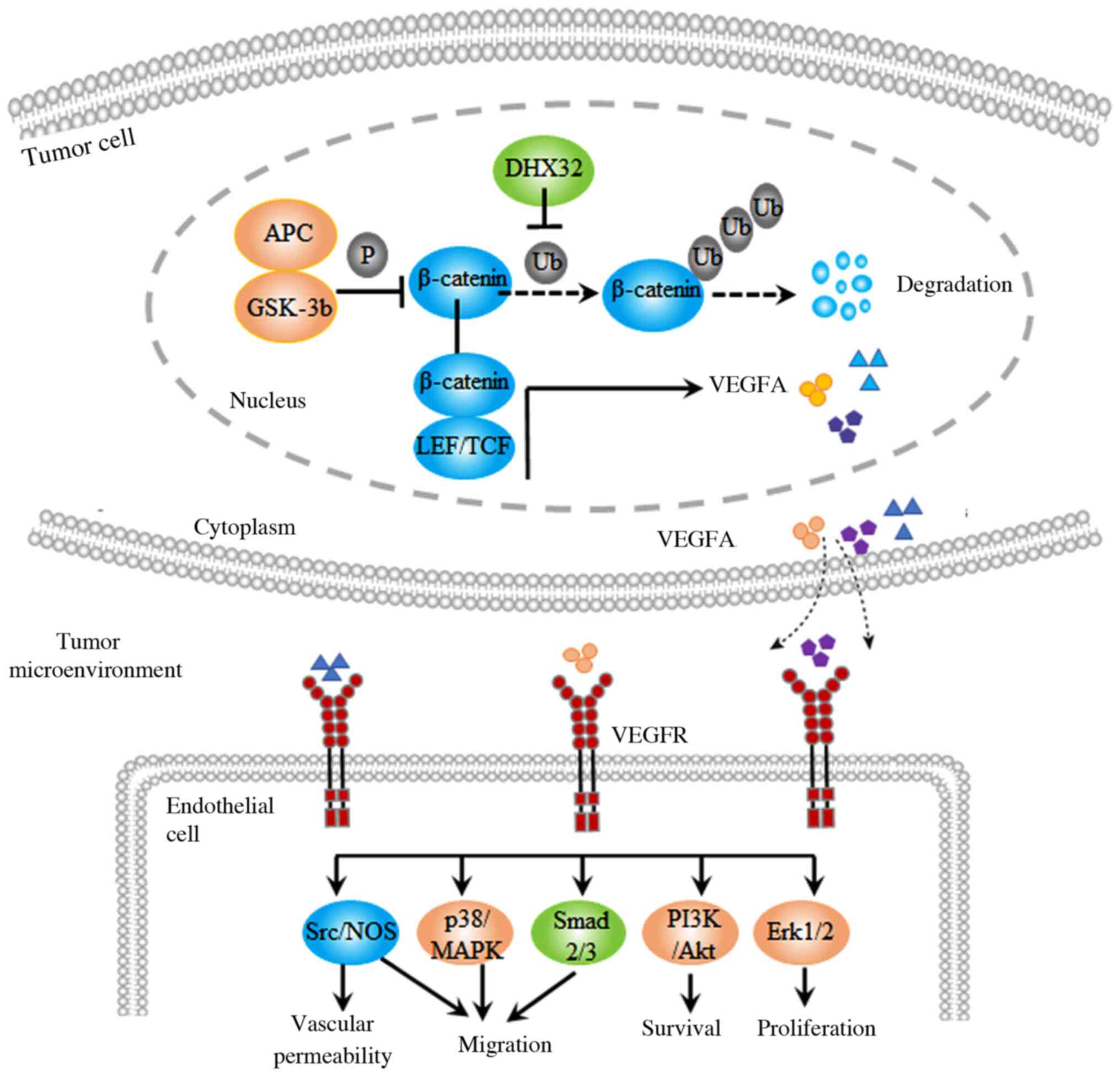
expression of VEGFA. EBioMedicine. 18:62–72. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Wang M, Zhang G, Wang Y, Ma R, Zhang L, Lv
H, Fang F and Kang X: DHX32 expression is an indicator of poor
breast cancer prognosis. Oncol Lett. 13:942–948. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Pajares MJ, Ezponda T, Catena R, Calvo A,
Pio R and Montuenga LM: Alternative splicing: An emerging topic in
molecular and clinical oncology. Lancet Oncol. 8:349–357. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Zhan R, Leng X, Liu X, Wang X, Gong J, Yan
L, Wang L, Wang Y, Wang X and Qian LJ: Heat shock protein 70 is
secreted from endothelial cells by a non-classical pathway
involving exosomes. Biochem Biophys Res Commun. 387:229–233. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Lancaster GI and Febbraio MA:
Exosome-dependent trafficking of HSP70: A novel secretory pathway
for cellular stress proteins. J Biol Chem. 280:23349–23355. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Bol GM, Khan R, Heerma van Voss MR,
Tantravedi S, Korz D, Kato Y and Raman V: PLGA nanoparticle
formulation of RK-33: An RNA helicase inhibitor against DDX3.
Cancer Chemother Pharmacol. 76:821–827. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Heerma van Voss MR, Vesuna F, Bol GM,
Afzal J, Tantravedi S, Bergman Y, Kammers K, Lehar M, Malek R,
Ballew M, et al: Targeting mitochondrial translation by inhibiting
DDX3: A novel radiosensitization strategy for cancer treatment.
Oncogene. 37:63–74. 2018. View Article : Google Scholar : PubMed/NCBI
|