Introduction
Prostate cancer (PC) is a prevalent male cancer in
European and American countries recognized as the second leading
cause of cancer-associated mortality globally (1,2). A
previous study has detected novel molecular targets for PC therapy,
however, the key initiating events regulating the pathogenesis and
progression of PC remain elusive (3). The diagnosis of PC is mainly based on
monitoring serum prostate-specific antigen (PSA) levels (4). Although the introduction of PSA
testing facilitates early PC diagnosis, the clinical accuracy of
PSA remains suboptimal to its lack of specificity and high false
positive rate (5). Identifying
novel biomarkers could improve the diagnosis accuracy in the early
stages of PC, which may be beneficial for early intervention and
for the prevention of progression to advanced stages of the disease
(6).
Advances in whole-genome sequencing has facilitated
the identification of non-coding RNA (ncRNA) in cancer, which has
demonstrated that abnormal expression of ncRNA is associated with
to PC progression (7). MicroRNA
(miRNA/miR) is a type of endogenous ncRNA ~21 nucleotides in
length, which can bind to mRNA targets and regulate their
expression at the post-transcriptional level (8). Increasing evidence suggests that
several miRNA molecules can act as tumor suppressors or oncogenes
in carcinogenesis (9,10). Profiling the miRNA expression in
different cancer types has identified abnormally expressed miRNA
molecules which may serve as biomarkers for diagnosis and prognosis
(11). Abnormal miRNA expression
has also been observed in PC, and complex miRNA regulatory networks
and modules have been proposed to regulate PC progression (12,13).
Aberrant miRNA expression frequently dysregulates the target genes
implicated in carcinogenesis of PC (14,15).
Among them, miR-1182 was identified as a tumor suppressor and
implicated in the tumorigenesis of different tumor types, such as
bladder (16), ovarian (17), gastric (18) and PC (19). However, its regulatory mechanism of
action in PC progression remains elusive.
The Akt serine/threonine kinase kinase family
comprises three highly homologous members: AKT1 (PKBα), AKT2 (PKBβ)
and AKT3 (PKBγ) (20). AKT3 is
known to play role in carcinogenesis (21,22).
AKT3 is abnormally expressed in PC, and its upregulation may
promote PC cell survival and proliferation (23,24).
Small interfering RNA (siRNA)-mediated AKT3 silencing inhibits PC
cell growth (25). Therefore, AKT3
signaling may play a vital role in PC tumorigenesis.
Located on chromosome 4q32, long intergenic
non-protein coding RNA 1207 (LINC01207) consists of two introns and
three exons (26–28). Recent studies have suggested that
LINC01207 is highly expressed in lung adenocarcinoma and promotes
cell proliferation and migration (26,29).
However, whether LINC01207 is involved in PC progression remains
unclear.
The present study aimed to detect the role of
LINC01207 in PC progression. The results demonstrated that
LINC01207 was significantly upregulated in PC compared with normal
prostate tissue samples. Mechanistic experiments demonstrated that
LINC01207 played its oncogenic role by sustaining AKT3 levels in PC
cells and sponging miR-1182. LINC01207 knockdown reduced AKT3
levels and induced apoptosis in PC cells. These findings identified
a novel mechanism underlying the oncogenic role of LINC01207 by
targeting the miR-1182/AKT3 axis in PC.
Materials and methods
Cell lines and cell culture
The human PC cell lines, PC-3, LNCaP, Du-145 and
C4-2B, and the human normal prostate cell line, RWPE1, were
purchased from American Type Culture Collection. These cell lines
were cultured in DMEM (Gibco; Thermo Fisher Scientific, Inc.)
containing 10% FBS (Gibco; Thermo Fisher Scientific, Inc.) and 1%
penicillin-streptomycin, and incubated in a humidified cell culture
incubator at 37°C with 5% CO2.
Cell transfection
The day prior to transfection, cells were
trypsinized, counted and seeded into 6-well plates at a density of
2×105 cells/well. When the cells reached 80% confluence,
small interfering RNA (siRNA), miR-8112 mimic or inhibitor, AKT3
expression vector or negative control (NC) were transfected into
cells using Lipofectamine® 2000 (Invitrogen; Thermo
Fisher Scientific, Inc.), according to the manufacturer's
instructions. In a 6-well plate, cells at 80% confluence were
transfected with 50 nM miRs/siRNAs (miR-1182 mimic, inhibitor or
NCs, siRNA and si-NC) or 6 µg plasmids (AKT3 expression vector,
sh-LINC01207 expression vector or empty vector control). After 16-h
incubation at 37°C with 5% CO2, the transfection
reagents were removed and replaced with fresh medium. Transfected
cells were incubated for 48 h post-transfection in the cell culture
incubator at 37°C with 5% CO2 before being used in
subsequent experiments.
Plasmids AKT3 overexpression or shRNA
transfection
The cDNA sequence of AKT3 was cloned into the pcDNA
3.1 plasmid (Suzhou GenePharma Co., Ltd.). The empty pcDNA 3.1
plasmid was used as NC. For sh-LINC01207 transfection, pLKO.1-Puro
vector containing the sh-LINC01207 sequence or sh-NC were
constructed by Guangzhou RiboBio Co., Ltd.
siRNA, miR-1182 mimic, miR-1182
inhibitor transfection
siRNA (si-NC, si-LINC01207#1 and si-LINC01207#2),
miR-1182 mimic, miR-1182 inhibitor and mimic-NC were obtained from
Guangzhou RiboBio Co., Ltd. The following sequences were used: i)
si-LINC01207#1 sense, 5′-CCAGCTAAGACATTAGTAA-3′ and antisense,
5′-TTACTAATGTCTTAGCTGG-3′; ii) si-LINC01207#2 sense,
5′-GCAGGAAGGAATCCACAAT-3′ and antisense, 5′-ATTGTGGATTCCTTCCTGC-3′
(Guangzhou RiboBio Co., Ltd.); iii) si-NC sense,
5′-UGCUGACUCCAAAGCUCUGdTdT-3′ and antisense,
5′-CAGAGCUUUGGAGUCAGCAdTdT-3′ (Invitrogen; Thermo Fisher
Scientific, Inc.); iv) miR-1182 mimic,
5′-GAGGGUCUUGGGAGGGAUGUGAC-3′; v) mimic-NC,
5′-CGGUACGAUCGCGGCGGGAUAUC-3′; vi) miR-1182 inhibitor sense,
5′-GAGGGUCUUGGGAGGGAUGUGAC-3′ and antisense,
5′-GUCACAUCCCUCCCAAGACCCUC-3′; inhibitor-NC sense,
5′-GGUUCGUACGUACACUGUUCA-3′ and antisense,
5′-UGAACAGUGUACGUACGAACC-3′ (Sigma-Aldrich; Merck KGaA).
Reverse transcription-quantitative PCR
(RT-qPCR)
TRIzol® reagent (Invitrogen; Thermo
Fisher Scientific, Inc.) was used to isolate total RNA according to
the manufacturer's instructions. For cellular fractionation,
Subcellular Fractionation Kit for Cultured Cells (Thermo Fisher
Scientific, Inc.; cat. no. 78840) was used according to the
manufacturer's instructions. Total RNA (1 µg) was used to perform
reverse transcription using Primer Script RT Master Mix kit (Takara
Bio, Inc.; cat. no. RR036A) under following conditions: 30 min at
37°C, 5 min at 85°C and holding at 4°C. qPCR analysis was performed
using SYBR Select Master Mix (Applied Biosystems, Inc.; cat. no.
4472908) on the ABI 7900 system (Applied Biosystems, Inc.). The
thermocycling conditions for the qPCR consisted of 30 sec at 95°C,
followed by 40 cycles of 5 sec at 95°C and 60 sec at 60°C. The
relative gene expression levels were normalized against GAPDH gene
using the 2−ΔΔCq method (30). The following PCR primers were used
in the study: i) GAPDH forward, 5′-CCACATCGCTCAGACACCAT-3′ and
reverse, 5′-ACCAGGCGCCCAATACG-3′; ii) LINC01207 forward,
5′-CAGACACAGGCCATTCAGTC-3′ and reverse,
5′-CTTCTTCACCAGAAGCATTCC-3′; iii) miR-1182 forward,
5′-GGGGAGGGTCTTGGGAGGGA-3′ and reverse, 5′-GTGCAGGGTCCGAGGT-3′; iv)
AKT3 forward, 5′-AATGGACAGAAGCTATCCAGGC-3′ and reverse,
5′-TGATGGGTTGTAGAGGCATCC-3′; U6 forward, 5′-CTCGCTTCGGGCAGCACA-3′
and reverse, 5′-AACGCCTTCCACGAATTTGCGT-3′.
Dual luciferase reporter assay
The bioinformatics website lncBase V.2 (https://rnacentral.org/expert-database/lncbase) was
used to identify LINC01207 targets. The TargetScan database 7.2
(http://www.targetscan.org/vert_72/)
was used to predict the target of miR-1182. The binding
relationships between the identified targets were confirmed using
dual luciferase reporter assays. The sequences of miR-1182 binding
sites at the 3′ untranslated regions (UTRs) of LINC01207 and AKT3
were amplified and individually inserted into the pmirGlO
luciferase reporter vector (Promega Corporation).
Wild-type (WT) and mutated (MUT) fragments
containing the binding site were synthesized and cloned into a
pMIR-REPORT plasmid (Guangzhou RiboBio Co., Ltd.). The plasmids
were then co-transfected into PC-3 cells with miR-1182 mimic or
mimic NC together with Renilla luciferase (hRlucneo) control
plasmid (Promega Corporation). Before transfection, cells were
seeded into a 96-well plate at a density of 2×103
cells/well. The target plasmid (0.5 µg) and miR-1182 mimic (0.5 µg)
were mixed in 50 µl serum-free Opti-MEM (Gibco; Thermo Fisher
Scientific, Inc.). In a separate preparation, 1 µl Lipofectamine
2000 was mixed in 50 µl serum-free Opti-MEM medium. The two
mixtures were combined together and incubated for 20 min, before
adding to the cells. A total of 48 h post-transfection, the
relative luciferase activities were measured using Dual-Luciferase
Reporter Assay kit (Promega Corporation; cat. no. E1910) on a
luminescence microplate reader (Infinite 200 PRO; Tecan). The
relative firefly luciferase activity in the reporter plasmid was
normalized to that of Renilla luciferase in the hRlucneo
control plasmid.
Cell Counting Kit-8 (CCK-8)
proliferation assay
A CCK-8 assay was performed to detect changes in
cell proliferation. PC-3 cells or PC-3-siLINC01207 cells were
seeded into a 96-well plate at a density of 1×104
cells/well, and incubated at 37°C in 5% CO2 for 48 h.
The CCK-8 (10 µl) reagent was added to each well at 0, 24, 48 and
72 h and incubated at 37°C for 1 h. The GX71 microplate reader
(Olympus Corporation) was used to detect the absorbance value
(optical density value) in each well at 450 nm.
Colony formation assay
PC-3 cells or PC-3-si-LINC01207 cells were seeded
into a 6-well plate at a density of 500 cells/well after cell
counting. The culture medium was changed every 2 days. After 14
days of culture, the cells were fixed in 100% methanol for 30 min
at room temperature and stained for 10 min using Giemsa Stain kit
(Abcam; cat. no. ab150670) according the manufacturer's
instructions. After 14 days, an image of the cell colony was
captured with a digital camera using an AM6000 microscope (Leica
Microsystems, Inc.). Colonies consisting of >50 cells were
counted manually.
Apoptosis detection using flow
cytometry
The Annexin-V-FITC apoptosis assay kit (BD
Biosciences) was utilized to analyze cell apoptosis. PC3 cells
(2×105) transfected with si-LINC01207#1, si-LINC01207#2
or si-NC were harvested at 48 h after transfection. Cells were
collected after trypsinization and resuspended in Annexin V binding
buffer. The solution containing 1X Annexin-V-FITC /propidium iodide
(PI) staining reagent was used to stain cells for 15 min at 37°C.
The FACSCalibur flow cytometer (BD Biosciences) was utilized to
detect cell apoptosis. The data were analyzed using FlowJo software
7.6 (FlowJo LLC).
Western blot analysis
Total protein lysates were harvested in cold NP40
lysis buffer (1 mM EDTA, 150 mM NaCl, 1% NP-40, 20 mM Tris-HCl,
phosphatase/protease inhibitor cocktail, pH 7.4). A Bradford assay
was used to measure protein concentration. Total protein (30 µg)
was separated by 4–12% gradient SDS-PAGE and transferred onto a
polyvinylidene fluoride membranes. The membrane was blocked with 5%
skimmed milk for 2 h at room temperature, then incubated overnight
with primary antibodies (dilution, 1:1,000 in PBS) at 4°C. The
membranes were then washed with TBS with 5% Tween-20 three times,
then incubated with horseradish peroxidase (HRP)-conjugated goat
anti-mouse IgG secondary antibody at room temperature for 90 min.
The antibodies used in the study were anti-AKT3 (dilution, 1:1,000;
cat. no. ab152157), anti-GAPDH (6C5 Loading Control; dilution,
1:1,000; cat. no. ab8245) and rabbit anti-mouse IgG H&L
(dilution, 1:2,500; cat. no. ab6728) (all from Abcam). Protein
bands were developed using Pierce™ ECL Plus-Chemiluminescent HRP
Substrates kit (Thermo Fisher Scientific, Inc.) followed by
visualization using the iBright™ FL1500 Imaging System (Thermo
Fisher Scientific, Inc.). Relative protein expression was
normalized to that of GAPDH using ImageJ software V 1.8.0 (National
Institutes of Health).
Lentivirus-mediated shRNA stable
expression cell line generation
The recombinant lentivirus vectors expressing shRNA
targeting LINC01207 and the 0.45-µm nitrocellulose (NC) membrane
filter were prepared by Shanghai GeneChem Co., Ltd. A total of 12
µg third generation packaging vectors pVSV-G and pGag-Pol (Addgene,
Inc.), and 10 µg shRNA expression vector pLenti-puro-shLINC0120
(Addgene, Inc.) were transfected into 293T cells (American Type
Culture Collection) in a 10-cm cell culture dish with Lipofectamine
2000 according to the manufacturer's instruction. The same amount
of empty vector pLenti-puro (Addgene, Inc.) was used to produce
negative control lentivirus (sh-NC group). Transfected cells were
incubated for 48 h at 37°C. The cell culture supernatant containing
the lentivirus was collected 48 h after transfection and was
filtered with a 0.45-µm nitrocellulose membrane filter. The
lentiviral supernatant was stored at −80°C until further use. To
generate stable cell lines, PC3 cells were infected with the
recombinant lentivirus at a MOI of 5 in the presence of 10 µg
polybrene (Sigma-Aldrich; Merck KGaA; cat. no. tr-1003-g). Infected
cells were selected with 1.0 µg/ml puromycin for two weeks to
eliminate the uninfected cells before further experiment. qPCR was
performed to confirm the efficient knockdown of LINC01207.
In vivo xenograft model
BALB/c immunodeficient nude mice (4–6 weeks old;
18–25 g) were provided by Experimental Animal Center, College of
Veterinary Medicine, Yangzhou University (Yangzhou, China). Mice
was raised under the pathogen-free condition. The Institutional
Committee for Animal Care and Use approved the present study
protocols. All the animal procedures were conducted strictly
following the approved protocols. PC-3-sh-NC and PC-3-sh-LINC01207
cells (2×106) were re-suspended in Matrigel™, and
injected subcutaneously into the flanks of BALB/c mice. Tumor
volume and mouse body weight were recorded on 7, 14, 21, 28, 35, 42
day. On day 42, all mice were euthanized and tumors were collected,
weighed and imaged. A total of 6 mice were included in sh-NC and
sh-LINC01207 group. Carbon dioxide was used for euthanasia.
Briefly, a euthanizing chamber was connected to a carbon dioxide
cylinder and the flow rate was adjusted to displace 20% of the cage
volume per min. Mice were placed into the euthanizing chamber for
10 min until no movement was observed. Death was assured by
subsequent by cervical dislocation. The tumors of terminally dead
mice were resected for weight measurement. All animal experiments
followed the Guide for the Care and Use of Laboratory Animals (US
and National Institutes of Health) (31). All animal procedures were approved
by the Animal Care and Use Ethical Committee of Central Hospital of
Enshi Tujia and Miao Autonomous Prefecture, Wuhan, China (approval
no. 20176209).
The Cancer Genome Atlas (TCGA)
datasets
TCGA data consisting of 492 PC tumor tissues and 152
normal prostate tissues was downloaded from GEPIA database
(http://gepia.cancer-pku.cn/). Gene
expression profiles from TCGA were utilized to compare LINC01207
gene expression between PC tumor and normal prostate tissue.
Patient tissue collection
A total of 50 PC tumor samples and 50 matched
non-carcinoma tissue samples (~2 cm away from the primary tumor
site) were obtained from 50 male patients by surgery from January
2018 to December 2019 in Enshi Central Hospital (Wuhan, China). All
patients were initially diagnosed by biopsy examination by
experienced pathologists. The age of the 50 enrolled patients
ranged between 35 and 61 years, with a median age of 48 years.
Patients meeting the following criteria were enrolled: i) Patients
confirmed by tumor pathology and genetics; ii) patients without
other histories of urologic neoplasms or tumor; and iii) patients
who received no chemotherapy or radiotherapy before the operation.
Patients meeting the following criteria were excluded: i) Patients
with severe impairment of vital organs, such as heart, liver and
lung; ii) patients with autoimmune history; iii) patients with
autoimmune history; and iv) patients with chronic infectious
disease or acute contagious diseases. The tissues were harvested
and kept in liquid nitrogen before further experiment. All patients
involved in the study were informed of the details and completed
written informed consent forms. The patients did not experience
radiotherapy and chemotherapy before surgery. This study was
approved by the Medical Ethics Committee of Central Hospital of
Enshi Tujia and Miao Autonomous Prefecture, Wuhan, China (approval
no. 20176209).
Statistical analysis
GraphPad Prism 5 (GraphPad Software, Inc.) was used
for statistical analysis. All experiments were repeated three
times, and the results are presented as the mean ± SEM. Differences
between PC and adjacent normal tissue samples were analyzed using
paired two-tailed Student's t-tests. In all other cases where two
groups were compared, two unpaired groups were compared using
unpaired t-tests. Comparisons of continuous variables among
multiple groups were analyzed using one-way ANOVA followed by
Tukey's post hoc test for pairwise comparisons. Comparisons of data
with effect of two nominal predictor variables were performed using
two-way ANOVA. Kaplan-Meier and log-rank tests were used to compare
the cumulative survival rates. Pearson's correlation analysis was
used to analyze correlation between gene expression levels.
P<0.05 was considered to indicate a statistically significant
difference.
Results
Expression of LINC01207 is upregulated
and correlated with poor prognosis in PC patients
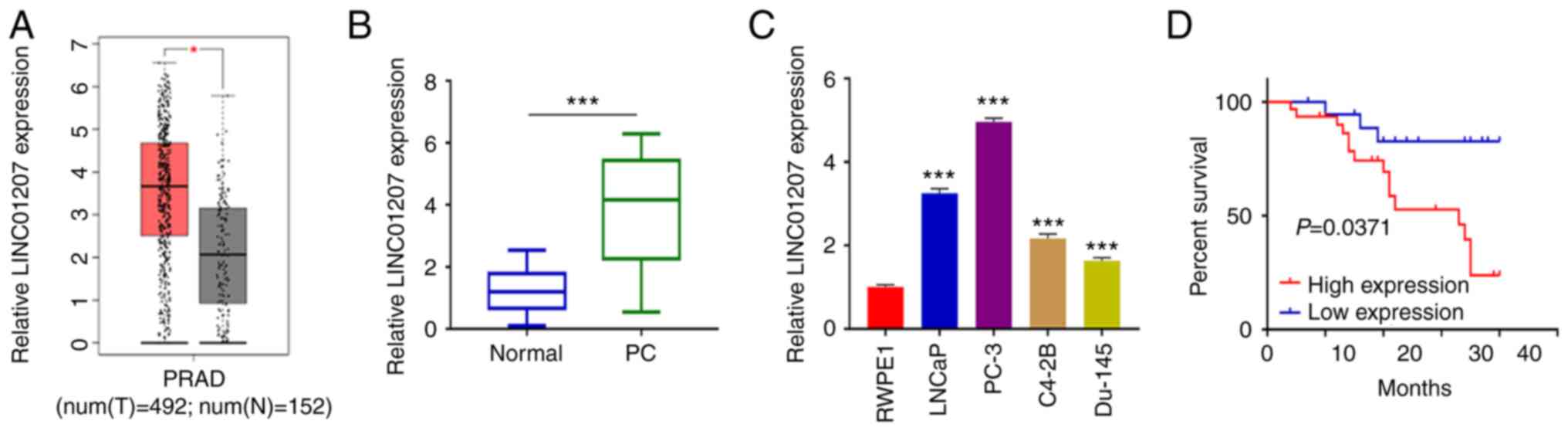
To determine LINC01207 expression level in PC
tumors, TCGA data consisting of 492 PC patients and 152 normal
prostate tissues were downloaded. Expression analysis demonstrated
that LINC01207 expression was significantly increased in PC tumor
samples compared with non-carcinoma tissue samples (Fig. 1A). To confirm this result, 50 PC
tumor samples and 50 matched normal samples were collected, and
LINC01207 expression using RT-qPCR. LINC01207 showed a
significantly higher expression level in PC samples relative to the
matched normal samples (Fig. 1B).
LINC01207 expression levels were also examined in human PC cell
lines (LNCaP, PC-3, C4-2B and Du-145) and a normal prostate cell
line (RWPE1). RT-qPCR analysis showed that LINC01207 was
upregulated in all PC cell lines compared with the RWPE1 cell line.
Among the PC cell lines, LINC01207 showed highest expression in
PC-3 cells (Fig. 1C). Furthermore,
the median expression value of LINC01207 in PC tissues was used as
the cut-off to divide the 50 patients with PC into high- (n=25) and
low-expression (n=25) groups. Kaplan-Meier survival curves were
used to evaluate overall survival rate of two groups of patients.
Our analysis showed that high levels of LINC01207 was associated
with prognosis (Fig. 1D).
Collectively, these data showed that LINC01207 expression was
significantly increased in PC tumor tissue, suggesting an oncogenic
role of LINC01207 in PC.
Knockdown of LINC01207 inhibits the
aggressive phenotype of PC cell
LINC01207 expression showed highest upregulation in
PC-3 cell line as indicated in Fig.
1C; therefore, the PC-3 cell line was chosen for further study.
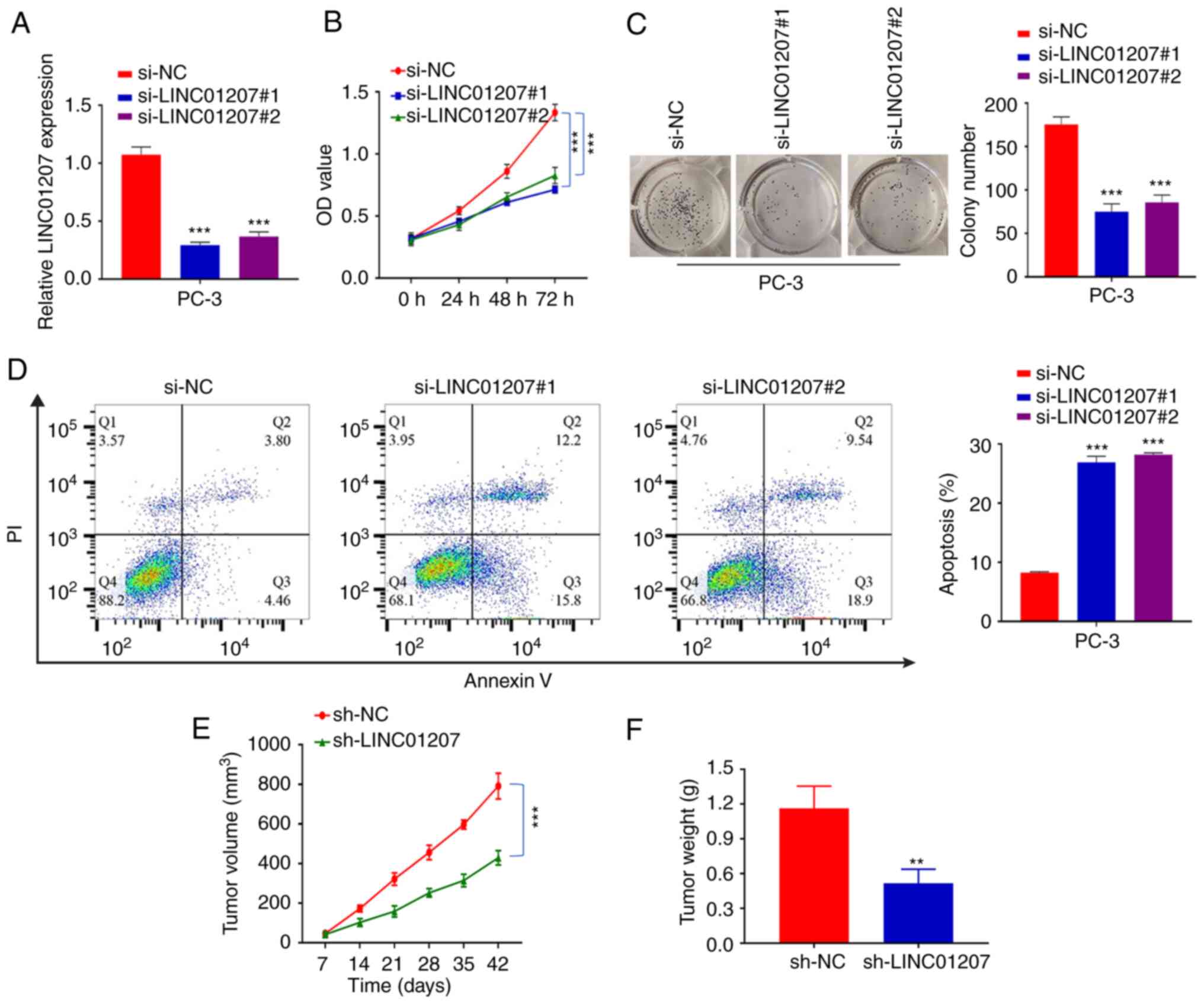
LINC01207 was knocked down in PC-3 cells using siRNA to assess its
biological function. RT-qPCR analysis demonstrated that
si-LINC01207#1 and si-LINC01207#2 could efficiently decrease
LINC01207 expression in PC-3 cells (Fig. 2A). CCK-8 assays were then used to
determine the role of LINC01207 in PC-3 proliferation. Both
LINC01207#1 and si-LINC01207#2 significantly suppressed the growth
of PC-3 cells (Fig. 2B).
Consistently, LINC01207#1 and si-LINC01207#2 impaired the colony
formation ability in PC-3 cells (Fig.
2C). Furthermore, Annexin V/PI staining showed that silencing
LINC01207 significantly increases the percentage of apoptotic
events in PC-3 cells. To further evaluate the role LINC01207 in
tumorigenesis, stable cell lines expressing shRNA were generated
(PC-3-shNC and PC-3-shLINC01207 cells), which were inoculated
subcutaneously in nude mice and the tumor volume was continuously
monitored. The silencing efficiency of shLINC01207 was validated by
RT-qPCR (Fig. S1A). Silencing of
LINC01207 significantly suppressed the tumor growth rate (Fig. 2E). Meanwhile, tumor weight was
significantly smaller in LINC01207 silenced group (Fig. 2F).
Taken together, these results indicate that
LINC01207 is indispensable for proliferation, colony formation and
cell survival of PC-3 cells in vitro. It is also required to
support the tumorigenesis in the xenograft mouse model.
LINC01207 sponges miR-1182
To explore the molecular mechanisms of LINC01207 in
regulating the aggressive behavior of PC cells, its target miRNA
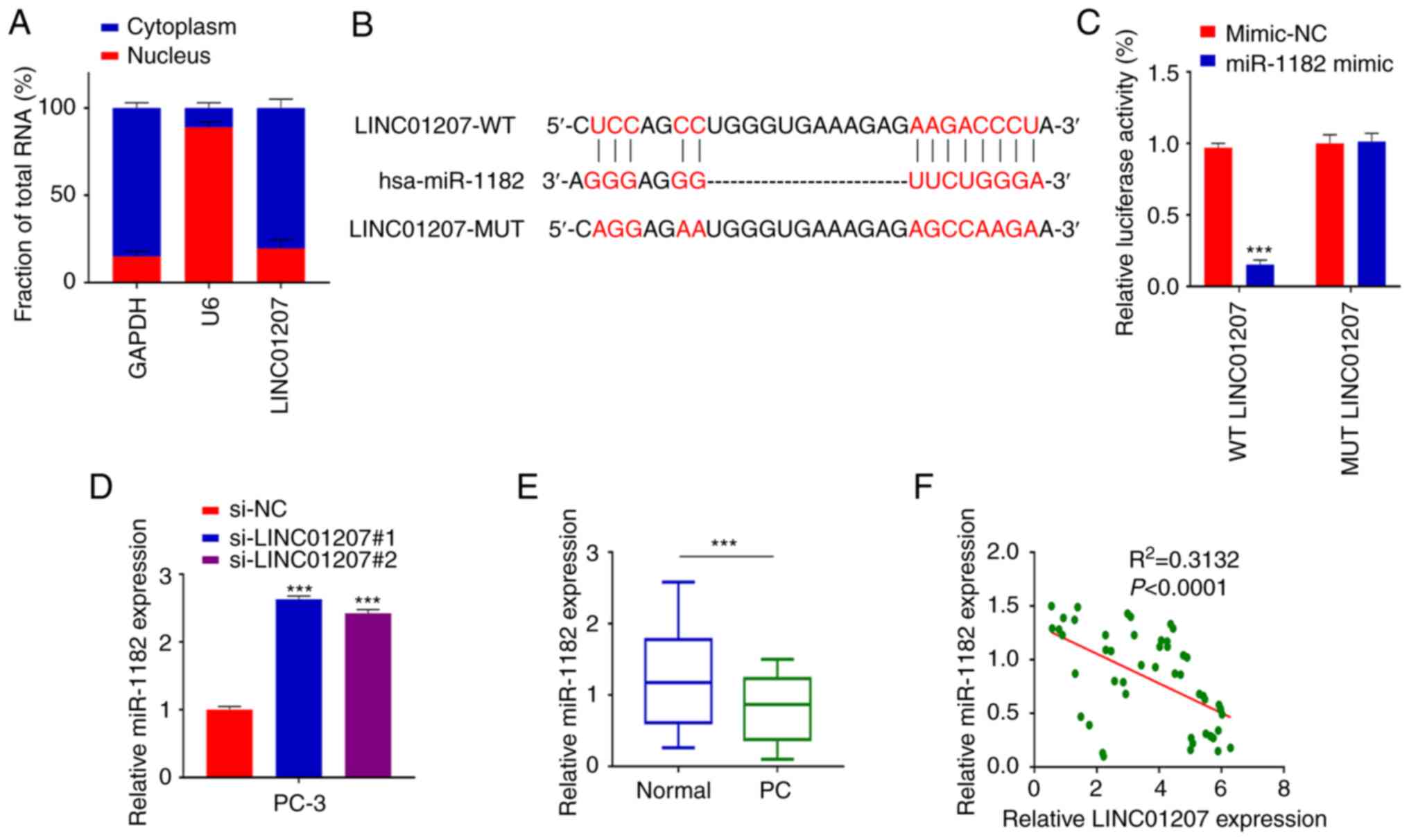
was examined. First, nuclear and cytoplasmic fractions were
separated in PC-3 cells to detect the cellular sub-localization of
LINC01207. As revealed by RT-qPCR analysis, LINC01207 was mostly
enriched in the cytoplasm (Fig.
3A). Next, the miRNA targets of LINC01207 were predicted using
the lncBase online tool, which identified miR-1182 as a potential
target binding to LINC01207 (Fig.
3B). To confirm their functional interaction, a dual luciferase
reporter assay was performed in the presence of miR-1182 mimic or
mimic-NC. Transfection of miR-1182 mimic significantly increased
intracellular miR-1182 level in PC-3 cells (Fig. S1B). miR-1182 mimic significantly
suppressed the luciferase reporter activity when compared with
mimic-NC, which was abrogated in the luciferase reporter with
mutated binding site (Fig. 3C).
Moreover, silencing LINC01207 could downregulate miR-1182
expression in PC-3 cells (Fig.
3D). In PC tumor samples and matched normal samples, miR-1182
expression showed significantly lower level in PC tumor tissues
(Fig. 3E). Furthermore, Pearson's
correlation analysis revealed a negative correlation between the
expression levels of LINC01207 and miR-1182 (Fig. 3F). Taken together, these data
indicate that LINC01207 interacts with and negatively regulates
miR-1182.
AKT3 is downstream target gene of
miR-1182
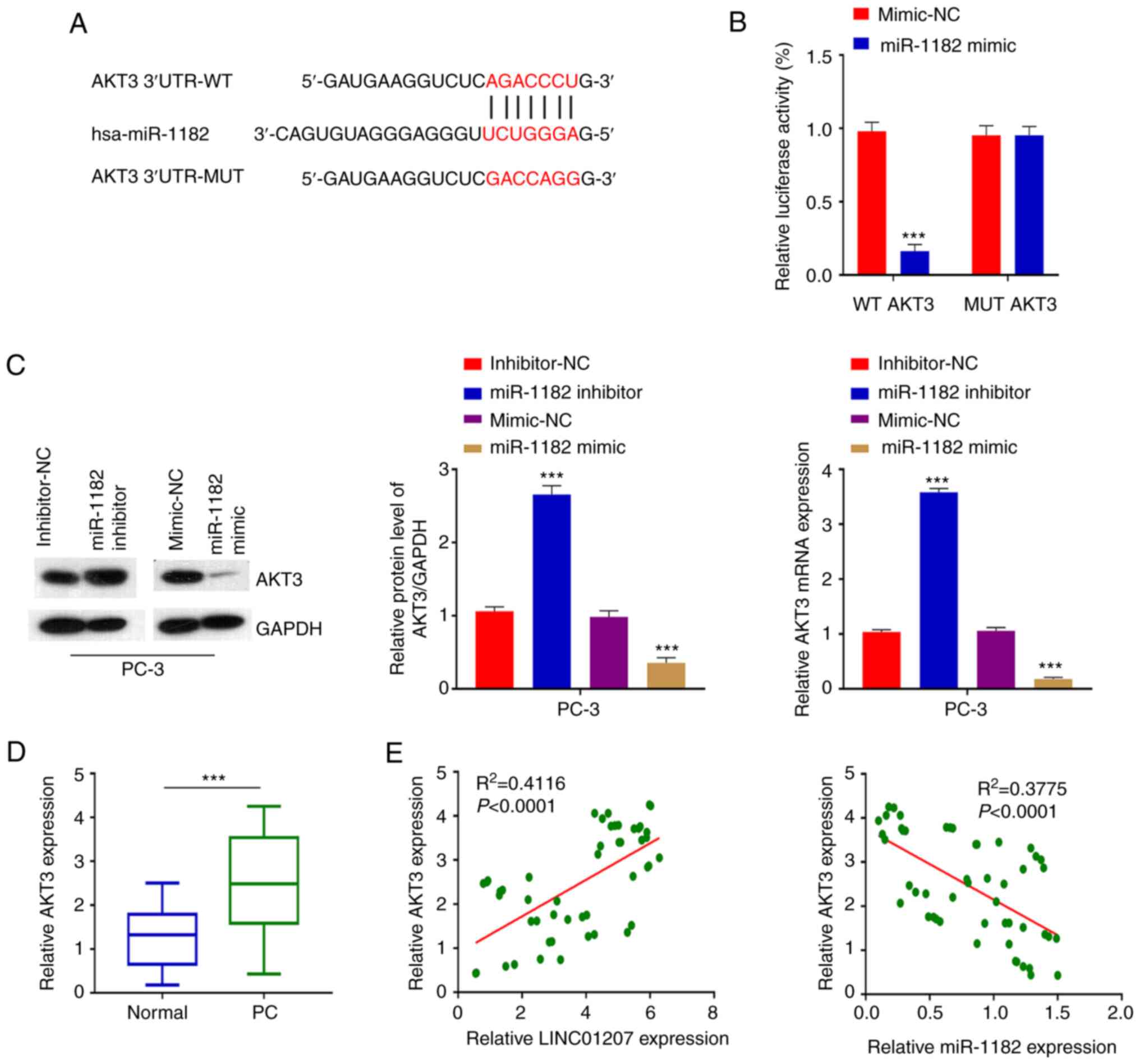
The gene target of miR-1182 was then identified.
TargetScan was used to predict the miR-1182 target genes. As a
result, AKT3 was identified as a potential miR-1182 target gene
(Fig. 4A). To confirm their
functional interaction, a dual luciferase reporter assay was
performed in the presence of miR-1182 mimic or mimic-NC. miR-1182
mimic significantly inhibited luciferase reporter of wild-type AKT3
3′UTR but not the reporter containing mutated binding site
(Fig. 4B). miR-1182 mimic also
decreased AKT3 expression at both mRNA and protein level (Fig. 4C). Consistently, knockdown of
miR-1182 by inhibitor significantly lowered the level of miR-1182
(Fig. S1C), and increased AKT3
expression at both mRNA and protein level (Fig. 4C). AKT3 was also upregulated in PC
tumor tissue compared with matched normal tissue samples (Fig. 4D). Of note, the expression levels
of AKT3 and miR-1182 were negatively correlated. By contrast, the
expression levels of AKT3 and LINC01207 were positively correlated
in 50 pairs of PC tumor tissues and matched normal tissues.
Altogether, these data suggest that ATK3 is a target of miR-1182,
and the expression levels of these two molecules are negatively
correlated in PC.
LINC01207 sponges miR-1182 to sustain
AKT3 level in PC cells
The above data showed that LINC01207, miR-1182 and
AKT3 expression are inter-correlated with each other in PC tumors.
Transfection with si-NC, si-LINC01207#1, miR-1182 inhibitor,
si-LINC01207#1+miR-1182 inhibitor and si-LINC01207#1 +AKT3 was used
in PC3 cells to validate a functional role of LINC01207/
miR-1182/AKT3 axis. qPCR analysis showed that transfection of AKT3
expression vector in PC-3 cells increased intracellular AKT3 mRNA
level by about 2.5 times (Fig.
S1D). As revealed by WB analysis, LINC01207 silencing decreased
AKT3 protein expression; conversely, AKT3 level was elevated in the
presence of miR-1182 inhibitor. The presence of miR-1182 inhibitor
or AKT3 expression vector partially rescued AKT3 level
downregulated by si-LINC01207#1 (Fig.
5A). LINC01207 and miR-1182 expression levels were also
examined. LINC01207 silencing increased miR-1182 mRNA expression,
while LINC01207 expression was dramatically increased when cell was
treated with miR-1182 inhibitor (Fig.
5B). Moreover, both LINC01207 and miR-1182 levels were slightly
increased when cells were co-transfected si-LINC01207#1 with
miR-1182 inhibitor or AKT3 (Fig.
5B). CCK8 proliferation assay showed that the presence of
miR-1182 inhibitor or AKT3 expression vector partially rescued the
cellular proliferation which was suppressed by si-LINC01207#1
(Fig. 5C). Consistently, the
presence of miR-1182 inhibitor or AKT3 expression vector also
rescued the colony formation ability when suppressed LINC01207 was
silenced (Fig. 5D). Finally,
apoptosis assay detected by FACS revealed that the presence of
miR-1182 inhibitor or AKT3 expression vector significantly recued
the apoptotic events induced by LINC01207 silencing (Fig. 5E). Collectively, these data imply
that LINC01207 plays an oncogenic role in PC cells by negatively
regulating miR-1182 to maintain a high level of AKT3.
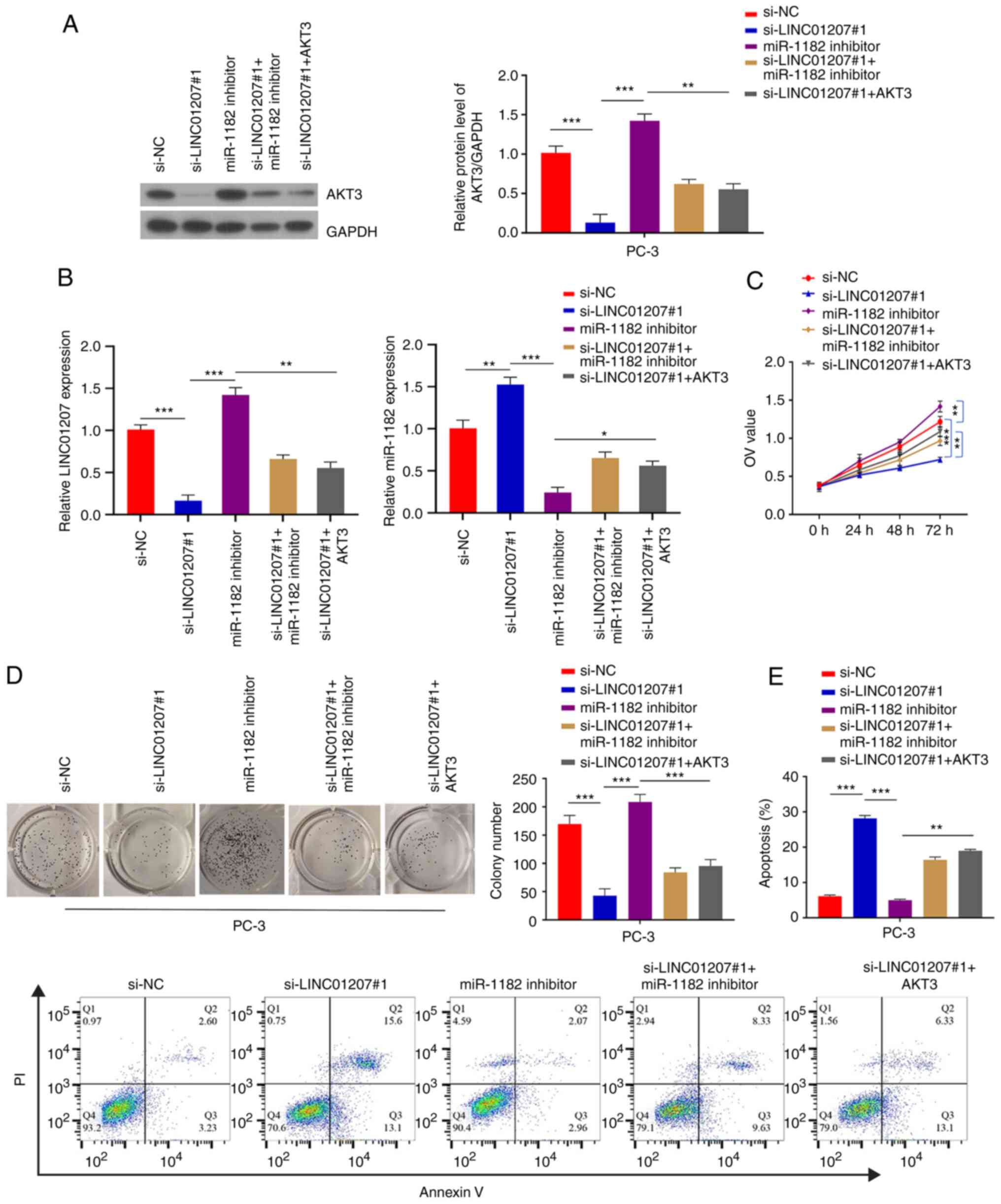 | Figure 5.LINC01207 sponges miR-1182 to
maintain AKT3 in prostate cancer cells. PC-3 cells were transfected
with si-NC, si-LINC01207#1, miR-1182 inhibitor, si-LINC01207#1 +
miR-1182 inhibitor or si-LINC01207#1 + AKT3. (A) AKT3 protein
expression levels following transfection. (B) LINC01207 and
miR-1182 expression following transfection. (C) Proliferation
following transfection. (D) Colony-forming following transfection.
(E) Apoptosis rate in PC-3 cells following transfection.
*P<0.05, **P<0.01, ***P<0.001. LINC01207, long intergenic
non-protein coding RNA 1207; miR, microRNA; NC, negative control;
si, small interfering RNA; OD, optical density; PI, propidium
iodide. |
Discussion
PC is the most prevalent malignancy in male patient
globally and the 4th leading mortality in cancer (32). Accumulating evidence has shown that
lncRNA is involved in tumorigenesis and cancer progression
(33,34). Abnormal miRNA expression is another
factor contributing to tumorigenesis by dysregulating target genes
related to PC progression (14,15).
Meanwhile, several studies have revealed that lncRNA promotes PC
growth mainly by regulating genes and pathways supporting cell
proliferation (1,3–5).
Several miRNA molecules and other proteins such as kallikreins have
been proposed as PC biomarkers (6). In addition, LINC01207 was previously
reported to be highly expressed in prostate cancer, as it could
downregulate miR-1972 and upregulate LIM and SH3 protein 1 to
promote PC progression (35). In
the present study, LINC01207 seems to act as an oncogenic gene
through modulating miR-1182/AKT3 axis. Collectively, previous
search and our results support the notion that LINC01207 plays a
functional role for the progression of PC.
Aberrant expression of LINC01207 has been reported
in numerous cancer types, such as pancreatic cancer and lung
adenocarcinoma (26,29). The present study adds novel
evidence of the abnormal LINC01207 expression in patients with PC,
which is associated with the poor prognosis. A previous study has
suggested that several upregulated lncRNA molecules seem to predict
a dismal prognostic outcome of PC (33). LINC01207 seems to be required to
support the proliferation and survival of PC cells both in
vitro and in vivo. Therefore, the present study and a
previous report (33) suggest that
a high level of LINC01207 expression could be used as a biomarker
to predict poor prognosis in PC patients.
The present study identified miR-1182 as a target of
LINC01207. miR-1182 dysregulation has been widely reported in a
variety of cancers, such as bladder cancer (16), ovarian cancer (17), gastric cancer (24) and PC (19). Therefore, miR-1182 may plays a
ubiquitous role in the regulating tumorigenesis of different types
of cancer. However, indifferent type of cancer, the exact molecular
mechanisms underlying its regulation can be different. In PC cells,
the present study demonstrated that the upregulation of LINC01207
negatively impacted on miR-1182 expression. Impaired expression of
miR-1182 was also observed in patients with PC. These results
suggest that miR-1182 may function as a tumor-suppressor gene,
which antagonizes the oncogenic effect of LINC01207.
Previous studies also showed high expression of AKT3
in PC tumors, which highlights AKTs as oncogenic factors in PC
progression (23,24). AKT signaling is an important
metabolic modulator coordinating cell survival and growth, which is
frequently dysregulated in tumor cells (23,24).
The present study showed miR-1182 targets and negatively regulates
AKT3 in PC cells. Since ATK3 expression positively correlates with
LINC01207 but negatively correlates with miR-8812, and AKT3 is
negatively regulated by miR-8812, it may be proposed that LINC01207
sustains AKT3 expression by sponging miR-8812 to support PC cell
proliferation and PC progression. Modulating the
LINC01207/miR-1182/AKT3 axis could be a novel strategy to control
PC progression, which needs to be further evaluated in mouse
models.
Overall, the findings of the present study indicate
that LINC01207 serves as an oncogenic factor that is upregulated PC
tumors. The aberrant upregulation of LINC01207 negatively impacts
on miR-1182 to sustain the expression of AKT3, which supports the
tumorigenesis of PC cells. LINC01207 knockdown suppresses
proliferation, induces apoptosis and impairs the tumorigenesis of
PC cells. Targeting the LINC01207/miR-1182/AKT3 axis could be
leveraged as a potential therapeutic strategy in the treatment of
PC.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
The datasets used and/or analysed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
DQ and CN designed and performed the experimental
work. BD and ZH performed literature research, research design and
manuscript editing. BT and SH performed the statistical analysis.
DQ and CN confirm the authenticity of all the raw data. All authors
read and approved the final manuscript.
Ethics approval and consent to
participate
All animal procedures were approved by the Animal
Care and Use Ethical Committee of Central Hospital of Enshi Tujia
and Miao Autonomous Prefecture (approval no. 20176209; Wuhan,
China). All patients enrolled in the study were informed of the
details and provided written informed consent. The use of the
clinical samples was approved by The Medical Ethics Committee of
Central Hospital of Enshi Tujia and Miao Autonomous Prefecture
(approval no. 20176209).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Edwards DR, Moroz K, Zhang H, Mulholland
D, Abdel-Mageed AB and Mondal D: PRL-3 increases the aggressive
phenotype of prostate cancer cells in vitro and its expression
correlates with high-grade prostate tumors in patients. Int J
Oncol. 52:402–412. 2018.PubMed/NCBI
|
|
2
|
Shukla ME, Yu C, Reddy CA, Stephans KL,
Klein EA, Abdel-Wahab M, Ciezki J and Tendulkar RD: Evaluation of
the current prostate cancer staging system based on cancer-specific
mortality in the surveillance, epidemiology, and end results
database. Clin Genitourin Cancer. 13:17–21. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Miyake H and Fujisawa M: Prognostic
prediction following radical prostatectomy for prostate cancer
using conventional as well as molecular biological approaches. Int
J Urol. 20:301–311. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Heidenreich A, Bellmunt J, Bolla M, Joniau
S, Mason M, Matveev V, Mottet N, Schmid HP, van der Kwast T, Wiegel
T, et al: EAU guidelines on prostate cancer. Part 1: Screening,
diagnosis, and treatment of clinically localised disease. Eur Urol.
59:61–71. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Dall'Era MA, Albertsen PC, Bangma C,
Carroll PR, Carter HB, Cooperberg MR, Freedland SJ, Klotz LH,
Parker C and Soloway MS: Active surveillance for prostate cancer: A
systematic review of the literature. Eur Urol. 62:976–983. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Egidi MG, Cochetti G, Guelfi G, Zampini D,
Diverio S, Poli G and Mearini E: Stability assessment of candidate
reference genes in urine sediment of prostate cancer patients for
miRNA applications. Dis Markers. 2015:9735972015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Cochetti G, Poli G, Guelfi G, Boni A,
Egidi MG and Mearini E: Different levels of serum microRNAs in
prostate cancer and benign prostatic hyperplasia: Evaluation of
potential diagnostic and prognostic role. Onco Targets Ther.
9:7545–7553. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Bartel DP: MicroRNAs: Genomics,
biogenesis, mechanism, and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Calin GA and Croce CM: MicroRNA signatures
in human cancers. Nat Rev Cancer. 6:857–866. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Gao Q and Zheng J: microRNA-323
upregulation promotes prostate cancer growth and docetaxel
resistance by repressing p73. Biomed Pharmacother. 97:528–534.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Kanwal R, Plaga AR, Liu X, Shukla GC and
Gupta S: MicroRNAs in prostate cancer: Functional role as
biomarkers. Cancer Lett. 407:9–20. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Cochetti G, Rossi de Vermandois JA, Maulà
V, Giulietti M, Cecati M, Del Zingaro M, Cagnani R, Suvieri C,
Paladini A and Mearini E: Role of miRNAs in prostate cancer: Do we
really know everything? Urol Oncol. 38:623–635. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Guelfi G, Cochetti G, Stefanetti V,
Zampini D, Diverio S, Boni A and Mearini E: Next generation
Sequencing of urine exfoliated cells: An approach of prostate
cancer microRNAs research. Sci Rep. 8:71112018. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Situ J, Zhang H, Jin Z, Li K, Mao Y and
Huang W: MicroRNA-939 directly targets HDGF to inhibit the
aggressiveness of prostate cancer via deactivation of the
WNT/β-catenin pathway. Onco Targets Ther. 13:4257–4270. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Yu C, Wang Y, Liu T, Sha K, Song Z, Zhao M
and Wang X: The microRNA miR-3174 suppresses the expression of
ADAM15 and inhibits the proliferation of patient-derived bladder
cancer cells. Onco Targets Ther. 13:4157–4168. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Zhou J, Dai W and Song J: miR-1182
inhibits growth and mediates the chemosensitivity of bladder cancer
by targeting hTERT. Biochem Biophys Res Commun. 470:445–452. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Hou XS, Han CQ and Zhang W: MiR-1182
inhibited metastasis and proliferation of ovarian cancer by
targeting hTERT. Eur Rev Med Pharmacol Sci. 22:1622–1628.
2018.PubMed/NCBI
|
|
18
|
Zhang D, Xiao YF, Zhang JW, Xie R, Hu CJ,
Tang B, Wang SM, Wu YY, Hao NB and Yang SM: miR-1182 attenuates
gastric cancer proliferation and metastasis by targeting the open
reading frame of hTERT. Cancer Lett. 360:151–159. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Huang C, Deng H, Wang Y, Jiang H, Xu R,
Zhu X, Huang Z and Zhao X: Circular RNA circABCC4 as the ceRNA of
miR-1182 facilitates prostate cancer progression by promoting FOXP4
expression. J Cell Mol Med. 23:6112–6119. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Dillon RL, White DE and Muller WJ: The
phosphatidyl inositol 3-kinase signaling network: Implications for
human breast cancer. Oncogene. 26:1338–1345. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Liby TA, Spyropoulos P, Buff Lindner H,
Eldridge J, Beeson C, Hsu T and Muise-Helmericks RC: Akt3 controls
vascular endothelial growth factor secretion and angiogenesis in
ovarian cancer cells. Int J Cancer. 130:532–543. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Nakatani K, Thompson DA, Barthel A, Sakaue
H, Liu W, Weigel RJ and Roth RA: Up-regulation of Akt3 in estrogen
receptor-deficient breast cancers and androgen-independent prostate
cancer lines. J Biol Chem. 274:21528–21532. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Lin HP, Lin CY, Huo C, Jan YJ, Tseng JC,
Jiang SS, Kuo YY, Chen SC, Wang CT, Chan TM, et al: AKT3 promotes
prostate cancer proliferation cells through regulation of Akt,
B-Raf, and TSC1/TSC2. Oncotarget. 6:27097–27112. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Zhang F and Wu Z: Significantly altered
expression of miR-511-3p and its target AKT3 has negative
prognostic value in human prostate cancer. Biochimie. 140:66–72.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Lin HP, Jiang SS and Chuu CP: Caffeic acid
phenethyl ester causes p21 induction, Akt signaling reduction, and
growth inhibition in PC-3 human prostate cancer cells. PLoS One.
7:e312862012. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Wang G, Chen H and Liu J: The long
noncoding RNA LINC01207 promotes proliferation of lung
adenocarcinoma. Am J Cancer Res. 5:3162–3173. 2015.PubMed/NCBI
|
|
27
|
Wang M, Lu X, Dong X, Hao F, Liu Z, Ni G
and Chen D: pERK1/2 silencing sensitizes pancreatic cancer BXPC-3
cell to gemcitabine-induced apoptosis via regulating Bax and Bcl-2
expression. World J Surg Oncol. 13:662015. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Wang Y, Li Z, Zheng S, Zhou Y, Zhao L, Ye
H, Zhao X, Gao W, Fu Z, Zhou Q, et al: Expression profile of long
non-coding RNAs in pancreatic cancer and their clinical
significance as biomarkers. Oncotarget. 6:35684–35698. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Liu C, Wang JO, Zhou WY, Chang XY, Zhang
MM, Zhang Y and Yang XH: Long non-coding RNA LINC01207 silencing
suppresses AGR2 expression to facilitate autophagy and apoptosis of
pancreatic cancer cells by sponging miR-143-5p. Mol Cell
Endocrinol. 493:1104242019. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
National Research Council (US) Committee
for the Update of the Guide for the Care, Use of Laboratory
Animals: Guide for the Care and Use of Laboratory Animals. (8th
edition). https://grants.nih.gov/grants/olaw/guide-for-the-care-and-use-of-laboratory-animals.pdfOctober
21–2021
|
|
32
|
Deng T, Yuan Y, Zhang C, Zhang C, Yao W,
Wang C, Liu R and Ba Y: Identification of circulating miR-25 as a
potential biomarker for pancreatic cancer diagnosis. Cell Physiol
Biochem. 39:1716–1722. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Cai B, Song XQ, Cai JP and Zhang S:
HOTAIR: A cancer-related long non-coding RNA. Neoplasma.
61:379–391. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Pang EJ, Yang R, Fu XB and Liu YF:
Overexpression of long non-coding RNA MALAT1 is correlated with
clinical progression and unfavorable prognosis in pancreatic
cancer. Tumor Biol. 36:2403–2407. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Wang S, Qiu J, Wang L, Wu Z, Zhang X, Li Q
and Jiang F: Long non-coding RNA LINC01207 promotes prostate cancer
progression by downregulating microRNA-1972 and upregulating LIM
and SH3 protein 1. IUBMB Life. 72:1960–1975. 2020. View Article : Google Scholar : PubMed/NCBI
|