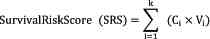Introduction
With estimated 2.3 million new cases and 670,000
deaths globally, breast cancer accounted for ~25% of new female
cancer diagnoses and 15.5% of cancer-related mortality in 2022
(1). Breast cancer is classified
histopathologically and molecularly into luminal A/B,
HER-2-enriched and triple negative breast cancer (TNBC). TNBC
accounts for 10–20% of global breast cancer cases (2). TNBC is characterised by a poor
prognosis that is associated with its aggressive behaviour,
intrinsic chemoresistance and tendency to early relapse/metastatic
spread (3). Further molecular
sub-stratification has separated TNBC into basal-like,
immunomodulatory, mesenchymal, mesenchymal stem-like and luminal
androgen receptor (4). TNBC
exhibits higher immunogenicity compared with other subtypes,
characterized by increased tumour-infiltrating lymphocytes (TILs),
programmed death-ligand 1 (PD-L1) expression and tumour mutational
burden (TMB) (5). Key
immune-related genes (IRGs) influencing prognosis and response to
immunotherapy include programmed cell death protein 1 (PD-1),
PD-L1, cytotoxic T-lymphocyte-associated protein 4 (CTLA4), and
genes involved in antigen presentation (human leukocyte antigen),
cytokine signalling [interferon-γ (IFNG) and CXCL chemokines] and
T-cell function (granzyme B and perforin-1), highlighting potential
therapeutic targets (6).
Treatment strategies for breast cancer have made
certain notable advancements in the past two decades. Fibroblast
growth factor receptor 4 promotes TNBC progression by activating
the AKT/ryanodine receptor type 2 axis, reprogramming fatty acid
metabolism (7). Separately,
circXPO6 stabilizes c-Myc to enhance glycolysis (glucose
transporter type 1/hexokinase 2/monocarboxylate transporter 4),
driving tumour growth (8). Both
pathways are associated with poor prognosis and offer therapeutic
targets. However, for patients with metastatic TNBC,
immune-checkpoint blockade, particularly targeting PD-1/PD-L1,
demonstrated promising results (9).
TILs are key effectors and regulatory factors of the
tumour microenvironment and they have been closely associated with
clinical outcomes in patients with breast cancer, particularly
those achieving complete pathological response after neoadjuvant
chemotherapy (10,11). In TNBC, increased TIL density is
associated with chemosensitivity and lower likelihood of recurrence
(12,13). While CD8+ cytotoxic T
cells comprise one component, one of the strongest relationships
with survival in breast cancer cohorts is the frequency of
CD8+ TILs (14). TIL
levels can also predict sensitivity to immune-checkpoint inhibitors
(ICIs): PD-1 expression is frequently high in the presence of rich
lymphocyte infiltration, which enhances therapeutic response
benefit (15–17).
TMB is defined as the number of somatic sequence
variants per megabase of tumour DNA. A higher TMB should create
neo-antigens that sharpen immune recognition, increase sensitivity
to immune-checkpoint blockade and provide improved response rates
and survival (18–20). As with TILs, TMB is an accepted
biological marker in predicting the effectiveness of ICIs (21,22).
Assessments of transcriptomic data on a large scale also suggest
that groups of co-expressed genes predict the number and function
of TILs and are prognostic markers (23–25).
Recently, ICIs have exhibited promising activity in TNBC treatment
(26,27). Furthermore, previous studies
(28,29) have demonstrated that high TMB is
notably associated with enhanced T cell infiltration, improved
response to ICIs and prolonged survival in TNBC. Considering the
prognostic role of TMB and TILs in cancer outcome and immunotherapy
response, the present study chose TMB and TILs as the entry point
(28,30).
Despite progress in therapies and prognostic
markers, the majority of patients with TNBC still have a poor
outcome. Therefore, the present study aimed to identify
differentially expressed IRGs (DEIRGs), associating with survival,
by using a combination of TMB and the density of CD8+ T
lymphocytes; ii) develop a personalised, TIL-based prognostic
model; iii) create a list of IRGs to inform clinicians of possible
outcome predictions; and iv) generate a clinically relevant
estimate of potential benefit from immunotherapy in TNBC.
Material and methods
Data acquisition and processing
RNA-sequencing data and clinical details for TNBC
were downloaded from The Cancer Genome Atlas (TCGA; http://portal.gdc.cancer.gov/) database. After
quality control (selecting ‘RNA-sequence’, ‘prognosis’ and ‘TNBC’
in TCGA database), 130 cases (age range, 26–90 years; all female)
were evaluated. Tumours were stratified according to the median
values for both TMB and CD8+ T-cell infiltration into
TMBhigh/CD8high and
TMBlow/CD8low. The baseline characteristics
are shown in Table I. IRGs from the
Immunology Database and Analysis Portal (ImmPort) database
(https://immport.niaid.nih.gov/home)
were used for the present analysis (31). For external validation, the
expression profiles from 107 TNBC samples in the Gene Expression
Omnibus (GEO) database (GSE58812) were analysed
(ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE58812 (32).
 | Table I.Clinical features of patients with
TNBC in TCGA. |
Table I.
Clinical features of patients with
TNBC in TCGA.
| Clinical
features | Patients (n=130), n
(%) |
|---|
| Median age (range),
years | 53 (26–90) |
| T stage |
|
| T1 | 33 (25.38) |
| T2 | 82 (63.08) |
| T3 | 11 (8.46) |
| T4 | 4 (3.08) |
| N stage |
|
| 0 | 87 (66.92) |
| 1 | 28 (21.54) |
| 2 | 10 (7.69) |
| 3 | 5 (3.85) |
| G3 | 39 (30.00) |
| AJCC stage |
|
| 1 | 22 (16.92) |
| 2 | 87 (66.92) |
| 3 | 21 (16.16) |
Identification of differentially
expressed genes (DEGs), DEIRGs and survival-associated DEIRGs
The ‘edgeR’ package (http://bioconductor.org/packages/edgeR/) in R software
(R Development Core Team, version 4.2.2) was applied to screen out
DEGs. Statistical significance was defined based on criteria such
as log2 (fold-change) >0.5 and a false-discovery
rate-adjusted P<0.05 (33).
Volcano plots and heatmaps were produced using the ‘gplots’
(cran.r-project.org/web/packages/gplots/) and ‘heatmap’
(stat.ethz.ch/R-manual/R-devel/library/stats/html/heatmap.html)
functions. By mapping the significant genes in the ImmPort
catalogue, DEIRGs were derived. Univariate Cox proportional-hazards
modelling was used to identify DEIRGs which were associated with
overall survival (OS) and cross-validated in the GSE58812
dataset.
Building the immune-related gene
prognostic index (IRGPI)
Survival-associated IRGs were refined using the
least absolute shrinkage and selection operator-penalized Cox
regression and transformed into a composite risk score. Patients
from the GSE58812 dataset were split into high- and low-risk groups
according to the median IRGPI and Kaplan-Meier curves were used to
compare survival between strata. The risk-score formula was
calculated as follows (34):
In the formula, ‘k’ represents the number of
prognostic indicators involved in the model, ‘Ci’
represents the coefficient of the prognostic indicators in
multivariate Cox regression analysis and ‘Vi’ represents
the specific value of the prognostic indicator.
Relationship between
survival-associated IRGs and immune cell infiltration
Using the Tumour IMmune Estimation Resource (TIMER)
database (cistrome.shinyapps.io/timer/) (35), the infiltration landscape of immune
cells in tumour-resident immune cells was investigated. The TIMER
repository consists of 10,897 TCGA samples across 15 tumour types
and quantifies the densities of eight immune subsets [B cells,
CD4+ and CD8+ T cells, natural-killer (NK)
cells, regulatory T cells (Tregs), macrophages, dendritic cells
(DCs) and monocytes]. For each patient with TNBC, the infiltration
values were derived and correlated with the IRGPI.
Relationship between the immune
prognostic model and clinical characteristics and outcome
The present study evaluated whether the IRGPI
offered prognostic value over standard variables by performing both
univariate and multivariate Cox regression for age, American Joint
Committee on Cancer (AJCC) stage and tumour (T) and lymph node (N)
status (36).
Tumour dysfunction and exclusion
(TIDE) score analysis
Potential for immune-escape was assessed using TIDE
analysis (tide.dfci.harvard.edu/login), which integrates T-cell
dysfunction and exclusion markers to predict the response to
checkpoint blockade.
Immunohistochemistry (IHC)
TNBC tissues collected from 10 female patients at
Suzhou Municipal Hospital from June 2016 to May 2018 (Suzhou,
China; aged 30–78 years; median age of 53 years; initial diagnosis
of TNBC in postoperative or puncture specimens; not received any
treatment and provided written informed consent) were fixed for 24
h in 4% paraformaldehyde/PBS and at 4°C. The tissues were sliced
into 3–5 µm-thick sections. The slides were treated with 0.3%
H2O2/PBS at room temperature for 10 min to
eliminate endogenous peroxidase activity. The slides were treated
with 5% goat serum (cat. no. 16210064; Thermo Fisher Scientific,
Inc.) for 30 min at room temperature. The slides were further
stained for CD8, C-X-C motif chemokine ligand 13 (CXCL13), TAP
binding protein like (TAPBPL), latent TGF-β binding protein 2
(LTBP2) and placental growth factor (PGF). TNBC tissues were
incubated overnight at 4°C with primary antibodies against CD8
(1:1,000; cat. no. GB12068; Wuhan Servicebio Technology Co., Ltd.),
CXCL13 (1:1,000; cat. no. GB11919; Wuhan Servicebio Technology Co.,
Ltd.), TAPBPL (1:200; cat. no. 201030-T08; Sino Biological, Inc.),
LTBP2 (1:200; cat. no. OACD05363; Beijing Aovia Biotechnology Co.,
Ltd.) and PGF (1:500; cat. no. 10642-1-AP; Proteintech Group,
Inc.), which was followed by incubation with HRP-conjugated
secondary antibodies (goat anti-rabbit IgG, 1:1,000; cat. no.
G1301; Wuhan Servicebio Technology Co., Ltd.) at room temperature
for 30 min. Tissue sections were stained with 3,3′-diaminobenzidine
at room temperature for 10 min, counterstained with haematoxylin at
room temperature for 15 min, dehydrated using a series of ethanol
and then mounted. Two pathologists, blinded to clinical
information, scored staining intensity using the histochemistry
Score) semi-quantitative system by light microscope (Olympus CX33
Corporation) (37).
Statistical analysis
Survival curves were generated using the ‘survival’
R package and differences were assessed using the log-rank test.
Time-dependent areas under the curves were generated using the
‘survivalROC’ package (v1.0.3,
cran.r-project.org/web/packages/survivalROC/index.html). A
two-sided P<0.05 was considered to indicate a statistically
significant difference. The present study conducted association
analyses of the risk score, CXCL13, TAPBPL, PGF and LTBP2 with
immune cell infiltration separately, employing the unpaired t-test,
Mann-Whitney U test and Pearson's correlation analysis. Each
independent experiment was repeated three times, and normality test
was conducted using the Kolmogorov-Smirnov test. Data are presented
as the mean.
Results
Patients in the
TMBlow/CD8low group exhibited worse
outcomes
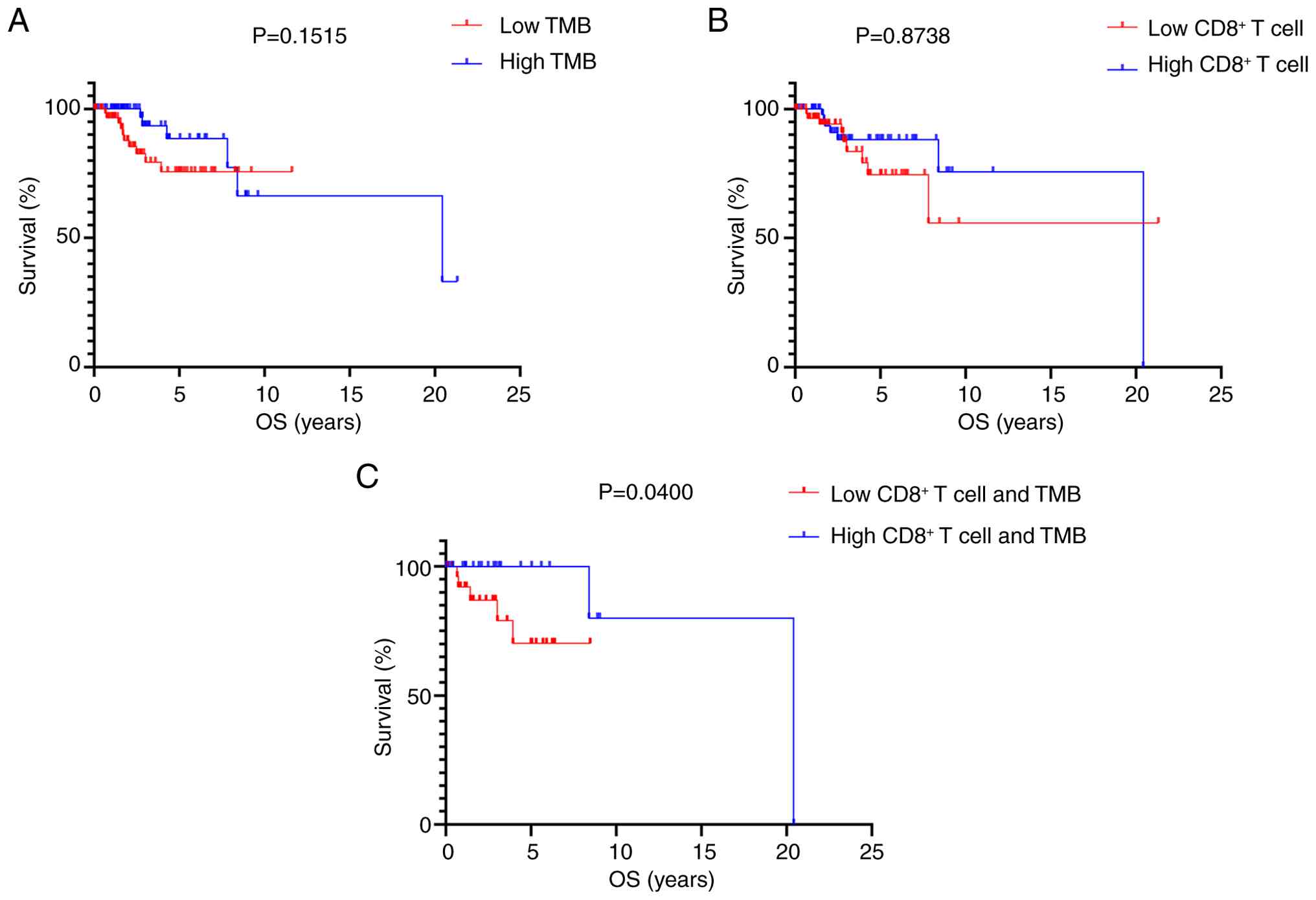
By using median values as cut-offs, there was no
significant survival difference between the TMBhigh and
TMBlow (Fig. 1A), and
between the CD8high and CD8low cohorts
(Fig. 1B). Patients classified as
TMBlow/CD8low were associated with
significantly worse OS compared with the
TMBhigh/CD8high cohort (Fig. 1C).
Identification of DEIRGs
A total of 225 immune genes with altered expression
were identified using ‘edgeR’, with 88 IRGs identified as being
upregulated and 137 IRGs identified as being downregulated
(Fig. S1).
Screening for prognosis-related
genes
Out of a total of 63 IRGs, univariate Cox analysis
was performed on 14 IRGs that were significantly associated with
survival in TCGA dataset (Table
II).
 | Table II.Univariate Cox regression analysis of
TCGA and GEO databases. |
Table II.
Univariate Cox regression analysis of
TCGA and GEO databases.
|
| Overall
survival |
|---|
|
|
|
|---|
|
| TCGA | GEO |
|---|
|
|
|
|
|---|
| Gene | OR (95% CI) | P-value | OR (95% CI) | P-value |
|---|
| CXCL13 | 0.352
(0.122–0.979) | 0.047a | 0.253
(0.108–0.593) | 0.002b |
| CXCL10 | 0.328
(0.114–0.949) | 0.040a | 0.434
(0.201–0.934) | 0.033a |
| TAPBPL | 0.191
(0.054–0.678) | 0.010a | 0.426
(0.194–0.937) | 0.034a |
| IDO1 | 0.334
(0.116–0.965) | 0.043a | 0.356
(0.162–0.783) | 0.010a |
| IFNG | 0.265
(0.085–0.827) | 0.022a | 0.390
(0.177–0.857) | 0.019a |
| APOBEC3C | 0.304
(0.098–0.944) | 0.039a | 0.509
(0.240–1.078) | 0.078 |
| CCL5 | 0.339
(0.118–0.978) | 0.045a | 0.519
(0.245–1.100) | 0.087 |
| CCL13 | 0.150
(0.034–0.978) | 0.013a | 0.537
(0.254–1.139) | 0.105 |
| FABP7 | 0.284
(0.090–0.896) | 0.032a | 1.485
(0.709–3.110) | 0.294 |
| FOS | 3.514
(1.107–11.151) | 0.033a | 1.339
(0.644–2.784) | 0.434 |
| LTBP2 | 3.328
(1.041–10.644) | 0.043a | 0.821
(0.395–1.706) | 0.597 |
| PDCD1 | 0.283
(0.091–0.879) | 0.029a | 0.353
(0.707–1.470) | 0.353 |
| PGF | 3.361
(1.080–10.460) | 0.036a | 1.285
(0.618–2.672) | 0.502 |
| PSME1 | 0.192
(0.054–0.673) | 0.010a | 0.629
(0.300–1.318) | 0.219 |
Low risk-score group demonstrates
improved OS in both prognostic and validation model
In TCGA prognostic model, the low risk-score group
had improved OS compared with the high risk-score group (Fig. S2A; OS limited to 20 years). In the
validation model, the high-risk group had worse OS compared with
the low-risk score group (Fig.
S2B). The formula of risk score was as follows: (Expression
level of CXCL13 ×-0.009833) + (expression level of TAPBPL ×
−0.016158) + (expression level of LTBP2 × 0.016080) + (expression
level of PGF × 0.096733). The present study separated patients with
TNBC into two groups (high- and low-risk score groups) with median
risk score in TCGA model (Fig. S3A, C
and E). The prognostic model was verified by a validation
prognostic model based on the GEO database (Fig. S3B, D and F). Patients with a lower
T stage (T1-2) had a significantly lower risk score compared with
patients with stage T3-4 (Fig.
S4).
Univariate and multivariate Cox
regression analysis of prognostic model
Using univariate Cox regression analysis, advanced T
stage [hazard ratio (HR)=4.384; 95% CI, 1.388–13.850], N Stage
(HR=3.543; 95% CI, 1.282–9.794), AJCC stage (HR=7.210; 95% CI,
2.586–20.104) and risk score (HR=6.302; 95% CI, 1.786–22.235) were
associated with an unfavourable prognosis (Table III). Multivariate analysis
determined that the risk score was an independent prognostic factor
for overall survival (HR=4.944; 95 % CI, 1.323–18.479; Table III).
 | Table III.Univariate and multivariate Cox
regression analysis of TCGA prognostic model. |
Table III.
Univariate and multivariate Cox
regression analysis of TCGA prognostic model.
|
| Overall
survival |
|---|
|
|
|
|---|
|
| Univariate | Multivariate |
|---|
|
|
|
|
|---|
| Clinical
feature | OR (95% CI) | P-value | OR (95% CI) | P-value |
|---|
| Age | 0.517
(0.179–1.494) | 0.223 | - | - |
| T stage | 4.384
(1.388–13.850) | 0.012a | 0.917
(0.218–3.851) | 0.562 |
| N stage | 3.543
(1.282–9.794) | 0.015a | 2.135
(0.612–7.450) | 0.234 |
| AJCC stage | 7.210
(2.586–20.104) |
<0.01a | 3.015
(0.690–13.176) | 0.142 |
| Risk score | 6.302
(1.786–22.235) | 0.004a | 4.944
(1.323–18.479) | 0.018a |
Validation of the accuracy of the
prognostic model
To substantiate the model, a nomogram was
established from the multivariate Cox model that could compare
predictive performance across age, AJCC, T and N stages and risk
score. The receiver-operating characteristic curves shown in
Fig. S5A demonstrated improved
discrimination for the composite model. The nomogram, which
included the five variables outlined earlier, generated a point
value for each variable and higher total scores indicated lower 3-,
5- and 10-year survival probabilities (Fig. S5B).
Analysis of risk score and immune cell
infiltration
Individuals classified into the low-risk cohort had
a significantly higher immune score (IS; Fig. 2A). However, TMB values were not
statistically different between high- and low-risk groups (Fig. 2B). Low-risk patients also had
significantly higher CD8+ T lymphocyte infiltration
(Fig. 2C), M1 macrophages (Fig. 2F), resting DCs (Fig. 2M) and activated CD4+ T
lymphocytes (Fig. 2L), while both
M0 (Fig. 2E) and M2 (Fig. 2G) macrophage densities were
significantly lower (both P<0.001). Tregs (Fig. 2D), resting NK cells (Fig. 2H), risk score (Fig. 2I), naïve B lymphocytes (Fig. 2J), resting CD4+ T cells
(Fig. 2K), activated DCs (Fig. 2N) and monocytes (Fig. 2O) infiltration did not differ
between risk groups.
 | Figure 2.Risk score and immune cell
infiltration association analysis using Mann-Whitney U test.
Association of high- and low-risk groups with (A) immune score, (B)
TMB, (C) CD8+ T cells, (D) Treg cells, (E) M0
macrophages, (F) M1 macrophages, (G) M2 macrophages, (H) NK cells
(resting), (I) NK cells (activated), (J) B cells, (K)
CD4+ T cells (resting), (L) CD4+ T cells
(activated), (M) DC cells (resting), (N) DCs (activated) and (O)
monocytes. TMB, tumour mutation burden; Treg, regulatory T cell;
NK, natural killer; DC, dendritic cell. |
Higher transcription levels of CXCL13 were
accompanied by significantly higher IS (Fig. 3A) but were not associated with TMB
(Fig. 3B). Samples with high CXCL13
levels had significantly greater recruitments of CD8+ T
cells (Fig. 3C), activated
CD4+ T cells (Fig. 3L)
and M1 macrophages (Fig. 3F) and
significantly lower levels of M0 (Fig.
3E) and M2 macrophages compared with those with low CXCL13
levels (Fig. 3G). Treg cells
(Fig. 3D), resting NK cells
(Fig. 3H), risk score (Fig. 3I), naïve B cells (Fig. 3J), resting CD4+ T cells
(Fig. 3K), resting DCs (Fig. 3M), activated DCs (Fig. 3N) and monocytes (Fig. 3O) concentrations did not appear to
be impacted by the level of CXCL13.
 | Figure 3.CXCL13 and immune cell infiltration
association analysis using Mann-Whitney U test. Association of
CXCL13 expression with (A) immune score, (B) TMB, (C)
CD8+ T cells, (D) Treg cells, (E) M0 macrophages, (F) M1
macrophages, (G) M2 macrophages, (H) NK cells (resting), (I) NK
cells (activated), (J) B cells, (K) CD4+ T cells
(resting), (L) CD4+ T cells (activated), (M) DC cells
(resting), (N) DCs (activated) and (O) monocytes. CXCL13, C-X-C
motif chemokine ligand 13; TMB, tumour mutation burden; Treg,
regulatory T cells; NK, natural killer; DC, dendritic cell. |
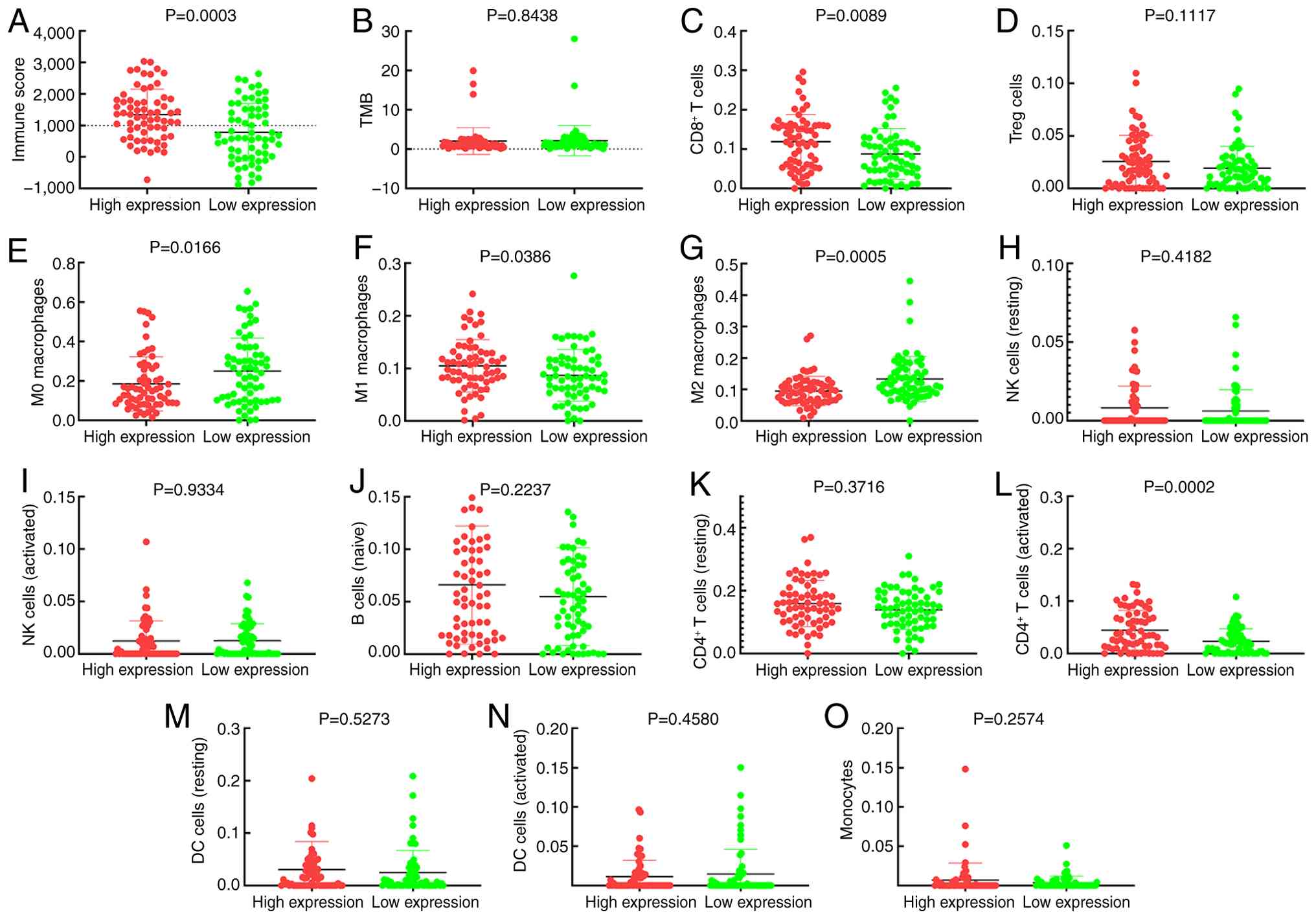
Similarly, enhanced levels of TAPBPL were
significantly associated with higher IS (Fig. 4A) and not associated with TMB
(Fig. 4B). Increased levels of
TAPBPL transcripts were significantly associated with greater
infiltration of CD8+ T lymphocytes (Fig. 4C) and with lower proportions of M0
(Fig. 4E) and M2 (Fig. 4G) macrophages. Treg cells (Fig. 4D), M1 macrophages (Fig. 4F), resting NK cells (Fig. 4H), risk score (Fig. 4I), naïve B cells (Fig. 4J), resting CD4+ T cells
(Fig. 4K), activated
CD4+ T cells (Fig. 4L),
resting DCs (Fig. 4M), activated
DCs (Fig. 4N) and monocytes
(Fig. 4O) concentrations were not
affected by whether TAPBPL was expressed.
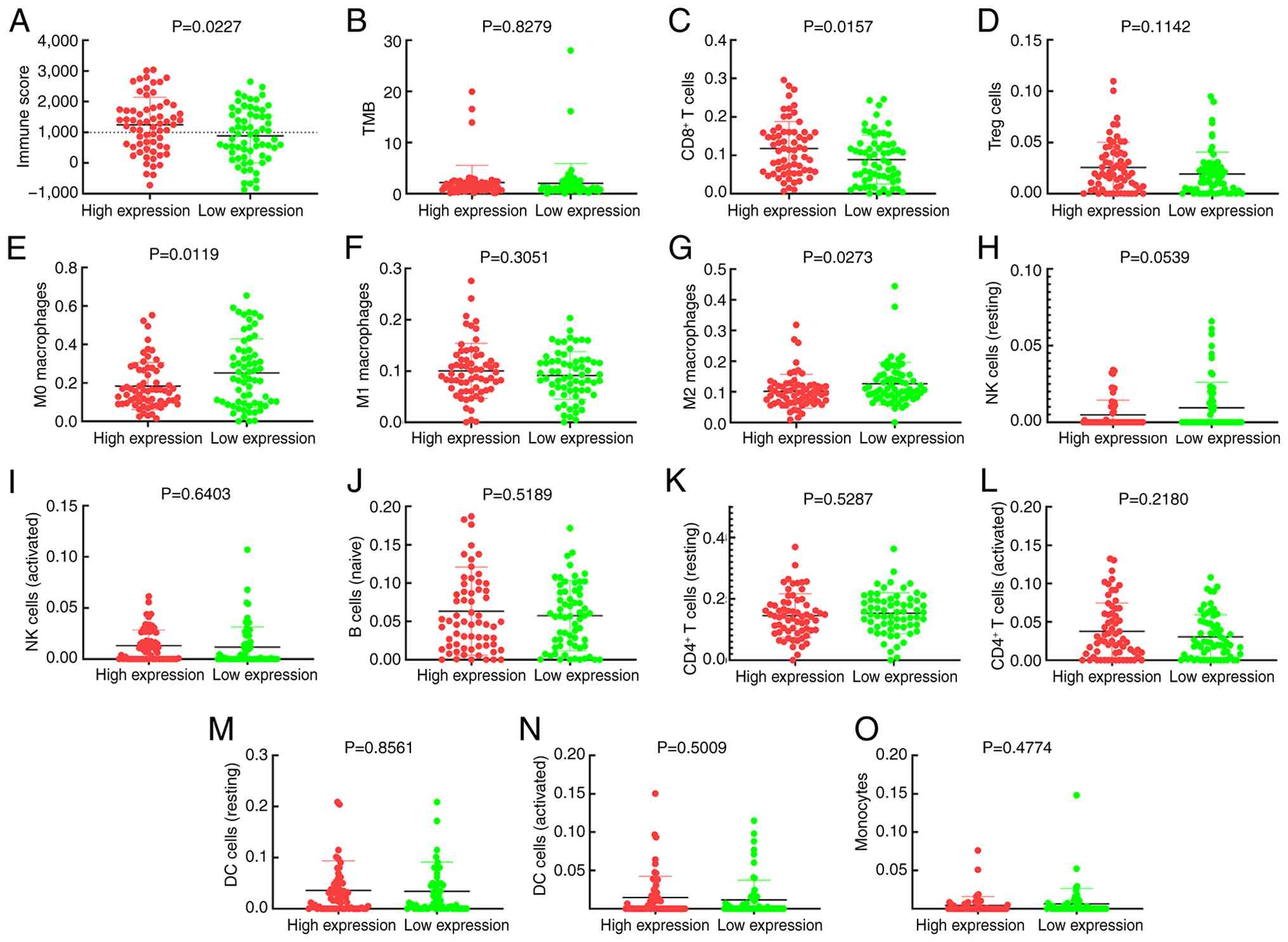 | Figure 4.TAPBPL and immune cell infiltration
association analysis by Mann-Whitney U test. Association of TAPBPL
expression with (A) immune score, (B) TMB, (C) CD8+ T
cells, (D) Treg cells, (E) M0 macrophages, (F) M1 macrophages, (G)
M2 macrophages, (H) NK cells (resting), (I) NK cells (activated),
(J) B cells, (K) CD4+ T cells (resting), (L)
CD4+ T cells (activated), (M) DC cells (resting), (N)
DCs (activated) and (O) monocytes. TAPBPL, transporter associated
with antigen processing binding protein like; TMB, tumour mutation
burden; Treg, regulatory T cells; NK, natural killer; DC, dendritic
cell. |
Increased PGF expression, by contrast, was
significantly associated with greater infiltration of M0 (Fig. 5E) and lower infiltration of M1
macrophages (Fig. 5F). PGF levels
were not associated with IS (Fig.
5A) or TMB (Fig. 5B). PGF
expression was also not associated with CD8+ T cells
(Fig. 5C), Treg cells (Fig. 5D), M2 macrophages (Fig. 5G), resting NK cells (Fig. 5H), risk score (Fig. 5I), naïve B cells (Fig. 5J), resting CD4+ T cells
(Fig. 5K), activated
CD4+ T cells (Fig. 5L),
resting DCs (Fig. 5M), activated
DCs (Fig. 5N) and monocytes
(Fig. 5O) as well.
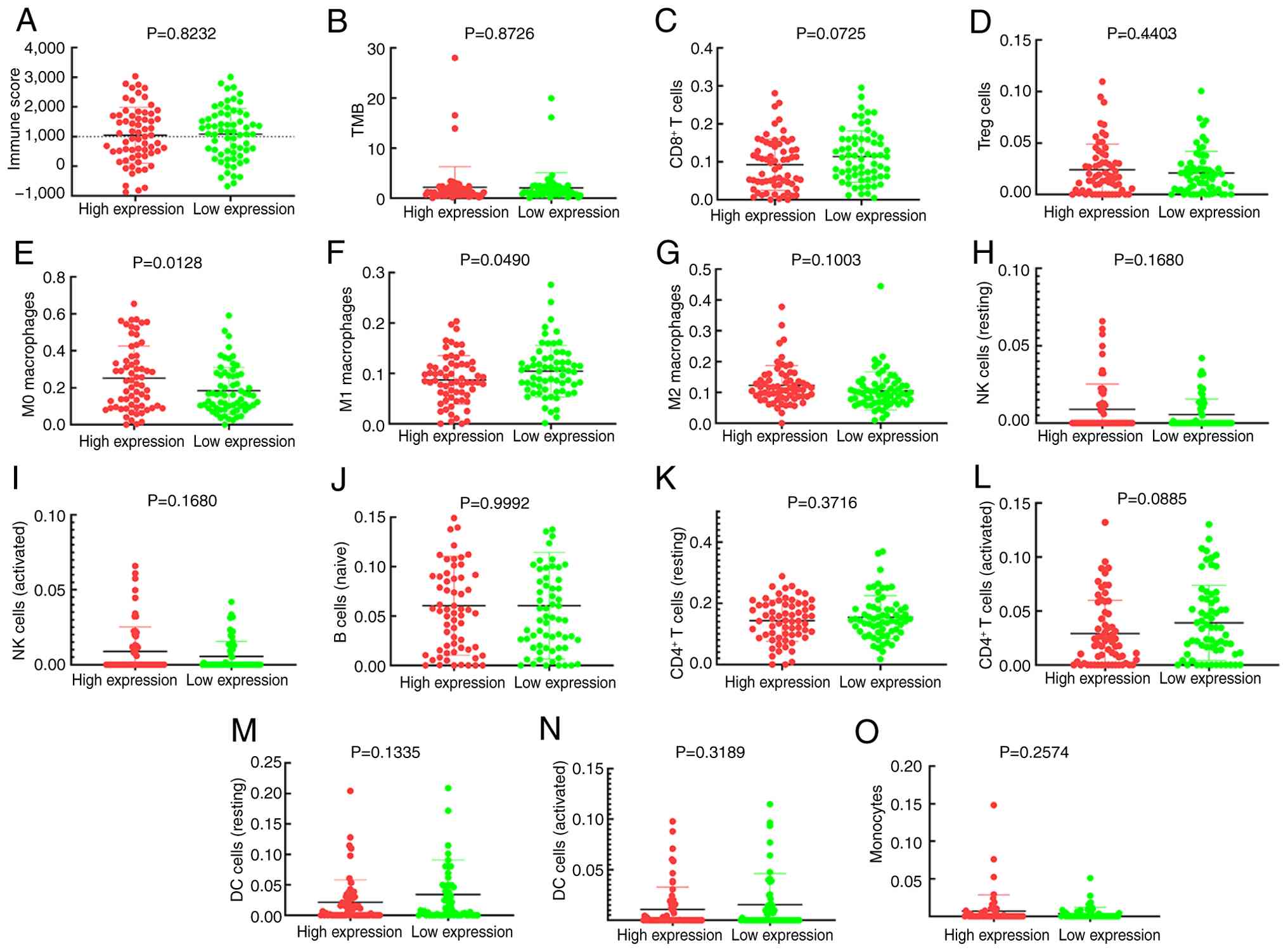 | Figure 5.PGF and immune cell infiltration
association analysis by Mann-Whitney U test. Association of PGF
expression with (A) immune score, (B) TMB, (C) CD8+ T
cells, (D) Treg cells, (E) M0 macrophages, (F) M1 macrophages, (G)
M2 macrophages, (H) NK cells (resting), (I) NK cells (activated),
(J) B cells, (K) CD4+ T cells (resting), (L)
CD4+ T cells (activated), (M) DC cells (resting), (N)
DCs (activated) and (O) monocytes. PGF, placental growth factor;
TMB, tumour mutation burden; Treg, regulatory T cells; NK, natural
killer; DC, dendritic cell. |
Increased LTBP2 expression was significantly
associated with greater infiltration of resting CD4+ T
lymphocytes (Fig. 6K) and
independent of IS (Fig. 6A) and TMB
(Fig. 6B). LTBP2 levels did not
influence infiltration of CD8+ T lymphocytes (Fig. 6C), Treg cells (Fig. 6D), M0 macrophages (Fig. 6E), M1 macrophages (Fig. 6F), M2 macrophages (Fig. 6G), resting NK cells (Fig. 6H), risk score (Fig. 6I), naïve B cells (Fig. 6J), activated CD4+ T cells
(Fig. 6L), resting DCs (Fig. 6M), activated DCs (Fig. 6N) or monocytes (Fig. 6O).
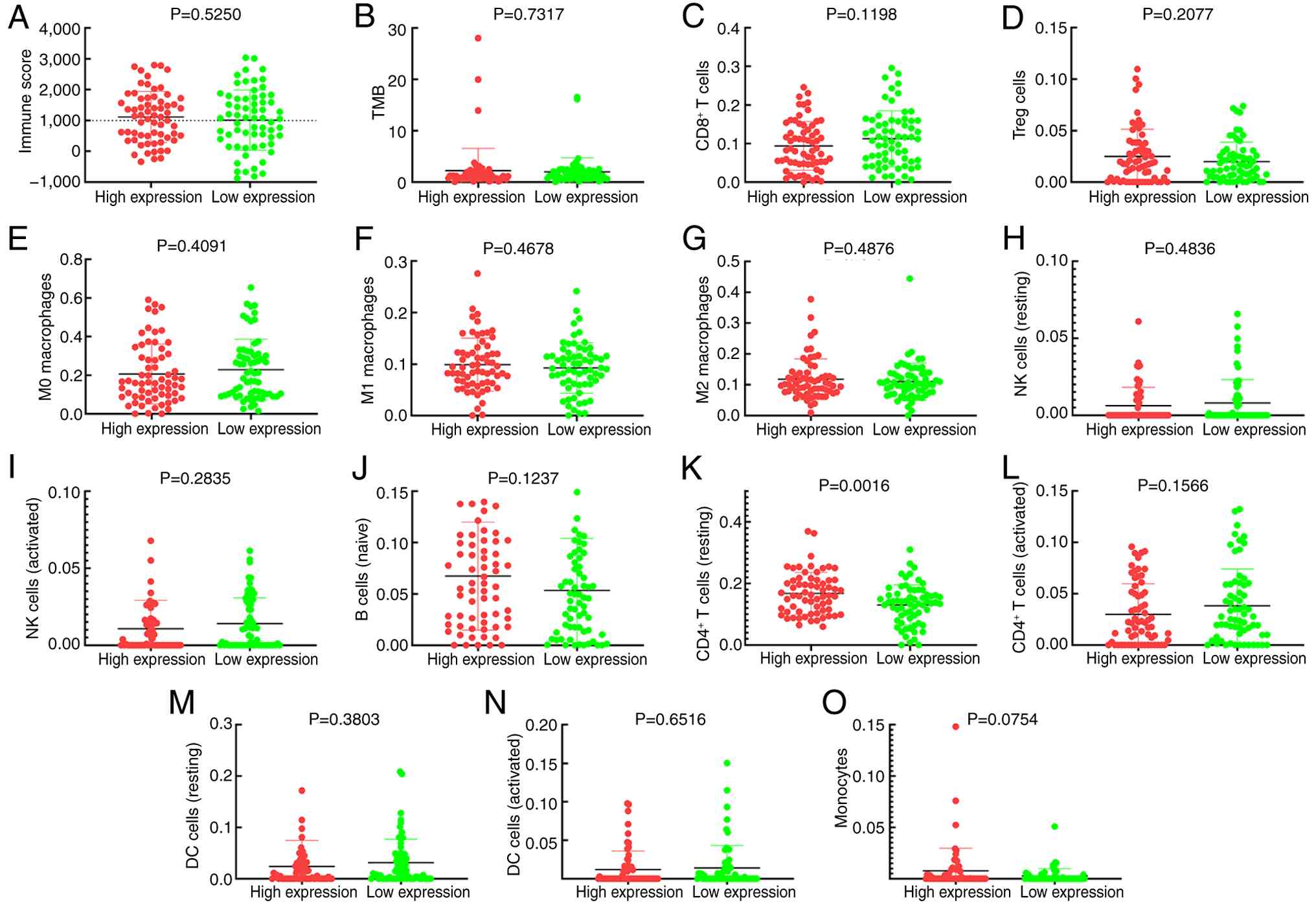 | Figure 6.LTBP2 and immune cell infiltration
association analysis by Mann-Whitney U test. Association of LTBP2
expression with (A) immune score, (B) TMB, (C) CD8+ T
cells, (D) Treg cells, (E) M0 macrophages, (F) M1 macrophages, (G)
M2 macrophages, (H) NK cells (resting), (I) NK cells (activated),
(J) B cells, (K) CD4+ T cells (resting), (L)
CD4+ T cells (activated), (M) DC cells (resting), (N)
DCs (activated) and (O) monocytes. LTBP2, latent TGF-β binding
protein 2; TMB, tumour mutation burden; Treg, regulatory T cell;
NK, natural killer; DC, dendritic cell. |
Immune checkpoint-related genes
correlation analysis
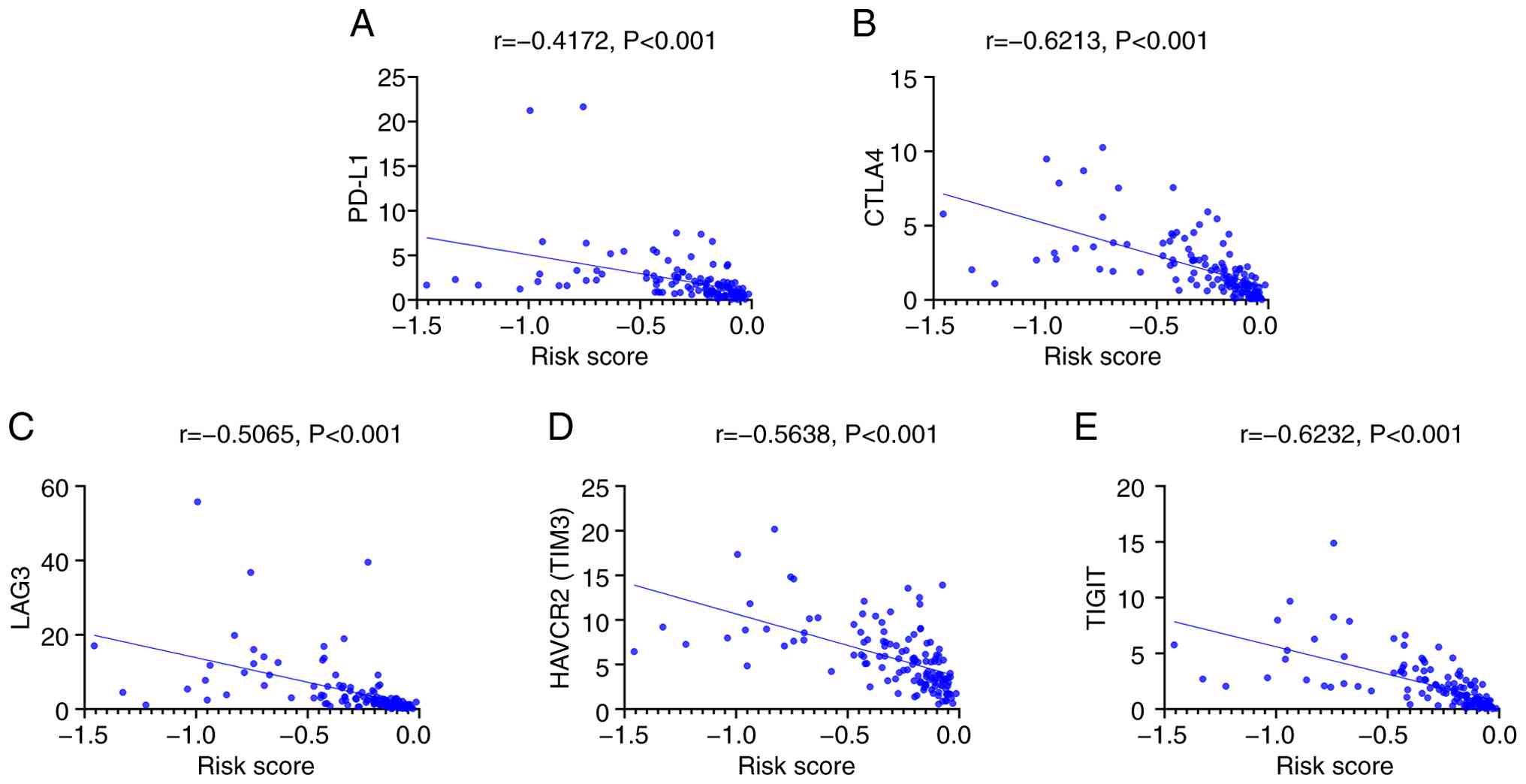
The composite risk score indicated a positive
correlation with immune-checkpoint genes PD-L1, T-cell
immunoglobulin and cytotoxic T-lymphocyte antigen 4 (CTLA-4) and
TIGIT, but negatively with lymphocyte activation gene 3 (LAG3)
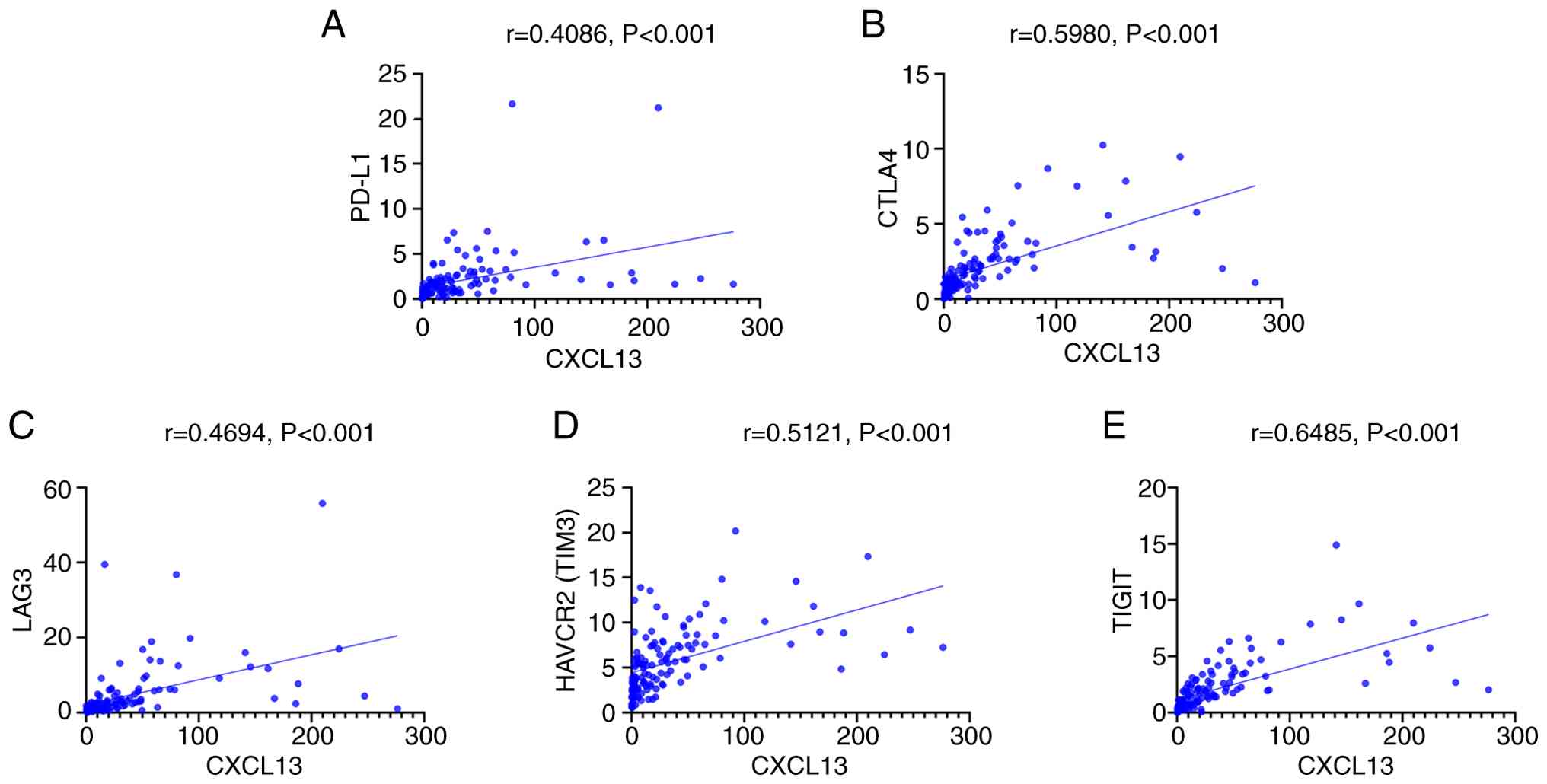
(Fig.7). CXCL13 levels were also
positively correlated with each of these immune-checkpoint genes
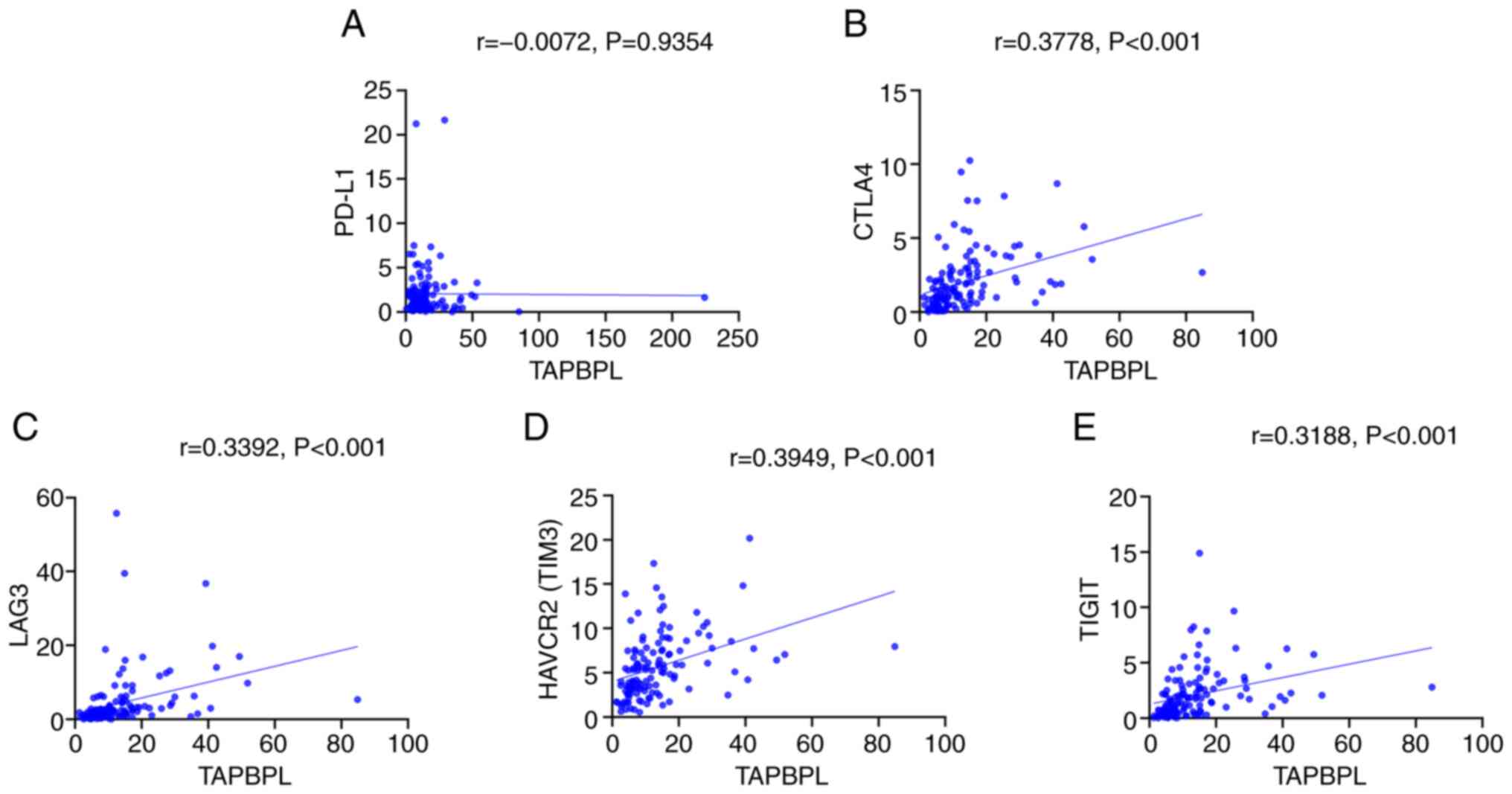
(Fig. 8). TAPBPL levels correlated
positively with mucin domain-containing protein 3 (TIM3), CTLA-4
and TIGIT, negatively with LAG3, (Fig.
9). PGF levels were not significantly correlated with PD-L1,
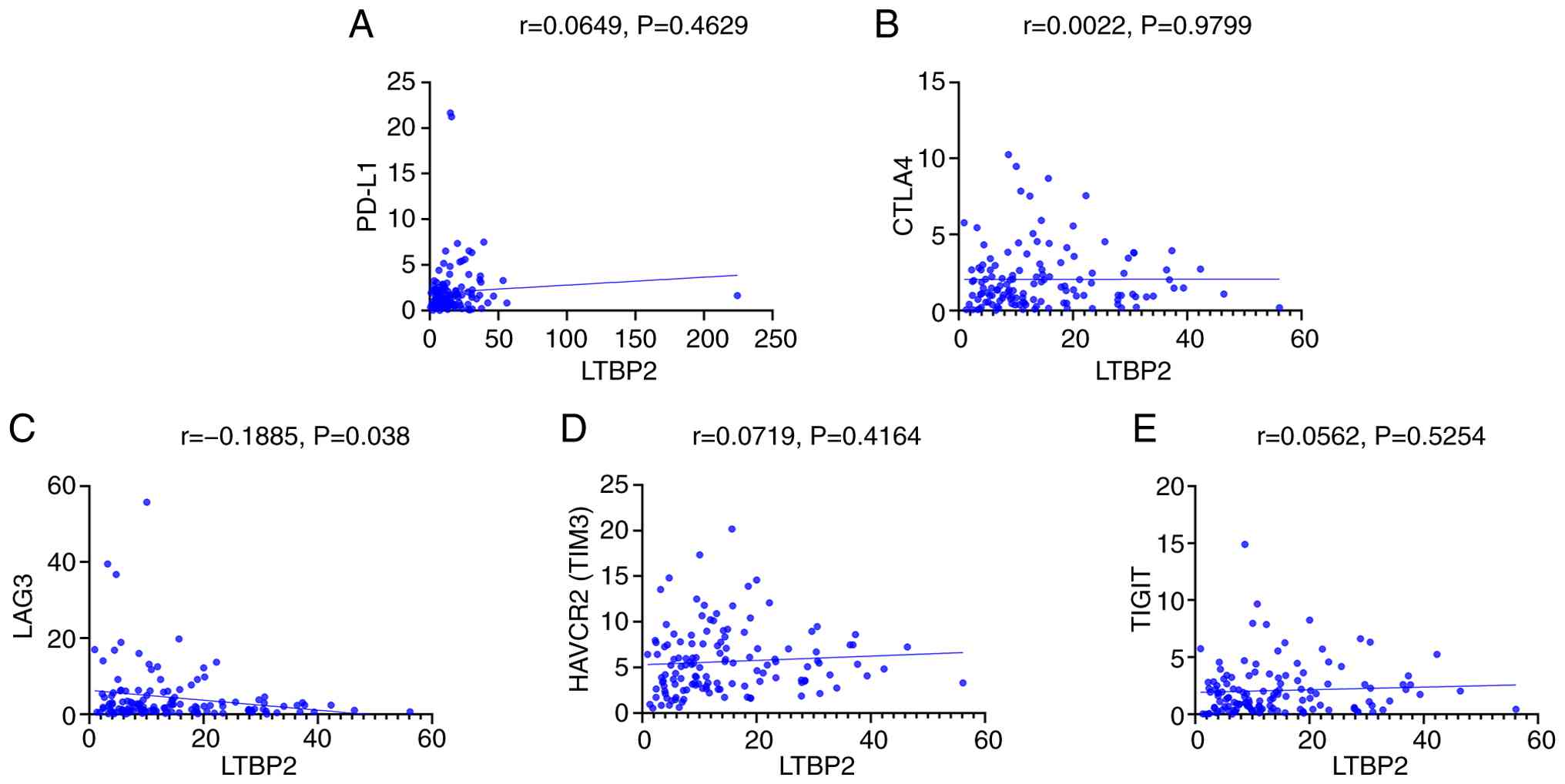
TIM3, LAG3, CTLA-4 or TIGIT transcript levels either (Fig. 10). Lastly, LTBP2 levels correlated
negatively with LAG3 levels, while no correlation was observed
between PD-L1, TIM3, TIGIT, CTLA-4 and LTBP2 (Fig. 11).
TIDE score analysis of
survival-associated IRGs
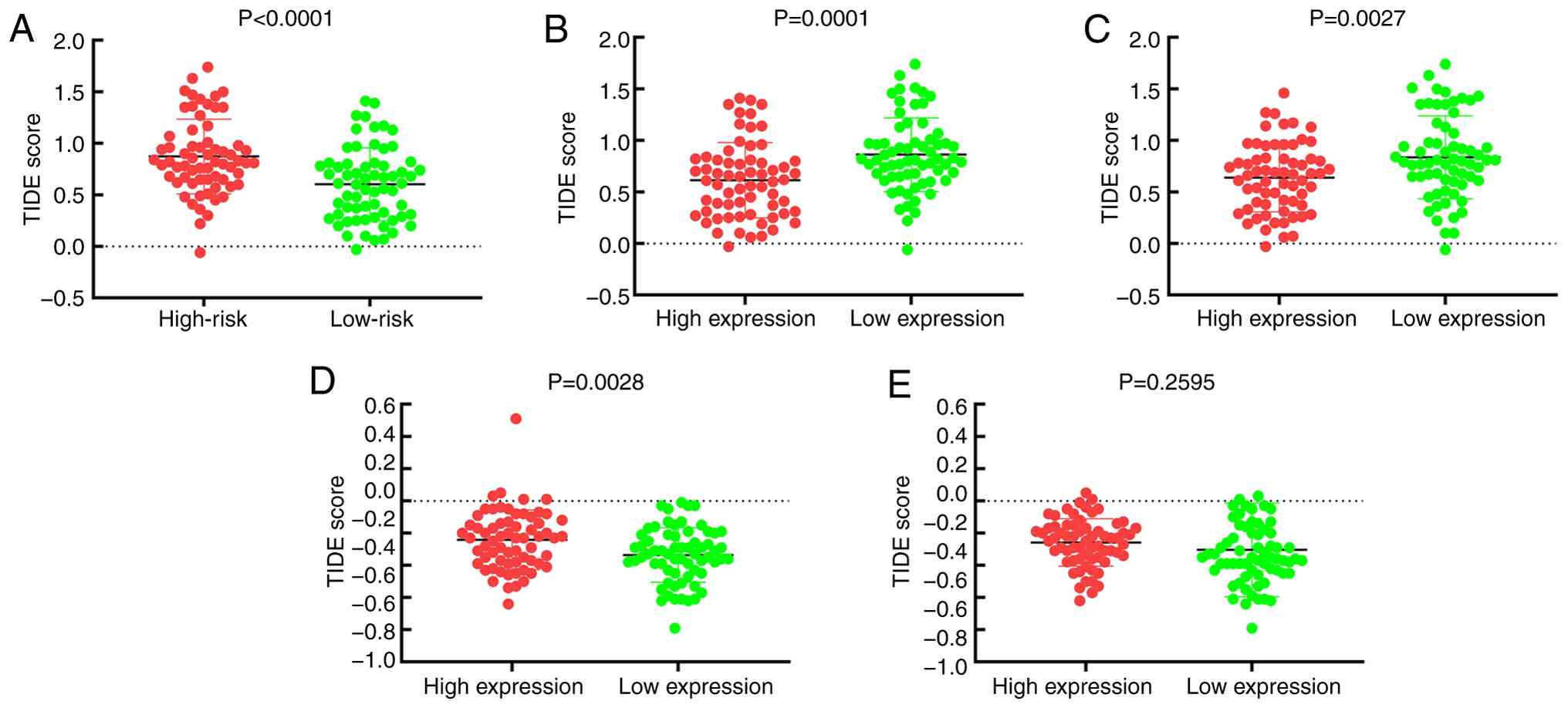
Individuals classified as low-risk had a markedly
lower TIDE score compared with the high-risk group (Fig. 12A). Higher levels of CXCL13
expression were associated with lower TIDE scores (Fig. 12B). Similarly, individuals with
higher levels of TAPBPL expression had lower TIDE scores compared
with low levels of TAPBPL expression (Fig. 12C). By contrast, higher levels of
PGF expression were associated with higher TIDE scores (Fig. 12D). There was no significant
difference in the TIDE score between risk categories with LTBP2
expression (Fig. 12E).
Validation of survival-associated IRGs
expression in TNBC tissues
To assess protein expression levels of CXCL13,
TAPBPL, LTBP2 and PGF, IHC analysis was performed on TNBC samples.
In support of the immune cell infiltrating data, TNBC tissues with
high CD8 expression levels demonstrated strong CXCL13 and TAPBPL
expression, while LTBP2 and PGF demonstrated weak IHC signals
(Fig. S6).
Discussion
The host immune system demonstrates a
well-established double-edged sword effect on neoplasia. Immune
elements can potentially drive malignancy yet provide strong
antitumour and cytotoxic functions (38). Therefore, ICIs have become standard
treatment for certain relapsed or metastatic diseases, such as
TNBC. Clinical trials such as KEYNOTE-522 (NCT03036488) and
IMpassion-130 (NCT02425891) led to the regulatory approval of
pembrolizumab and atezolizumab for recurrent or metastatic TNBC
(39–41); however, while there were notable
clinical benefits in these studies, they were mostly confined to
tumours with a high combined positive score (CPS). Since CPS-high
lesions are a small subset of all breast cancer types, it is key to
find novel predictive biomarkers to improve the precision and
population-wide benefit of treatment in TNBC. Notable evidence has
demonstrated that a high TMB is notably correlated with enhanced
T-cell infiltration, higher neoantigen burden, improved response to
ICIs and prolonged survival in TNBC (42,43).
Considering the value of TMB and TILs in evaluating the efficacy of
ICIs (44,45), the present study therefore focused
on their prognostic value in TNBC.
In the present study, 130 patients with TNBC were
assigned to TMBhigh/CD8+
T-cellhigh or TMBlow/CD8+
T-celllow groups. Kaplan-Meier analysis indicated that
the TMBhigh/CD8+ T-cellhigh cohort
had a significantly improved survival outcome compared with the
TMBlow/CD8+ T-celllow group.
Higher TMB and CD8+ T cell levels may create
neo-antigens that enhance immune recognition, increase sensitivity
to immune-checkpoint blockade and provide improved response rates
and survival. Therefore, individuals with high TMB and
CD8+ expression have improved prognosis (46,47).
Further investigation revealed 14 genes were differentially
expressed, immune-related and had prognostic value. To corroborate
these observations, external validation was performed using the GEO
dataset GSE58812 to reduce the applicable genes to obtain a
prognostic model based on five IRGs.
Subsequently, differential transcripts between
defined risk strata were used to perform classification across
these two groups to create a prognostic signature. Tumour immune
microenvironment (TIME) characterisation revealed that low-risk
tumours had a denser intratumoral CD8+ T-cell
infiltration with a higher global IS but richer M2-polarised
macrophages in the high-risk tumours. Previous studies established
that CD8+ T effector cells are the predominant
lymphocyte infiltrate in breast tumours (14,48).
Furthermore, heightened T-cell infiltration is associated with
objective response rates among patients with TNBC who receive ICI
therapy (49) and the intratumoral
vs. stromal compartment of CD8+ T-lymphocytes is
associated with enhanced relapse-free survival (50). By contrast, IL-6, IL-17 and TGF-β
cytokines have demonstrated the ability to activate
tumour-associated macrophages and Tregs to inhibit CD8+
T cell proliferation, migration and cytolytic function by releasing
immunosuppressive mediators (51–53).
Macrophages themselves exhibit two main activation states: i)
Classically activated (M1), providing antimicrobial defence; and
ii) alternatively activated (M2), promoting metastases, an
immunosuppressive environment and angiogenesis (54–57).
Therefore, the accumulation of M1 macrophages and CD8+ T
cells from the low-risk cluster could explain their improved
clinical course.
The present study identified four immune-related
prognostic genes including CXCL13, TAPBPL, LTBP2 and PGF. CXCL13,
also known as B-lymphocyte attracting chemokine 1, attracts CXC
receptor 5+ B cells and promotes germinal-centre
activity (58). Several studies
associate elevated CXCL13 levels with improved disease-free
survival and higher pathologic complete response rates after
neoadjuvant chemotherapy treatment in patients with TNBC (59,60).
Lv et al (61) demonstrated
that there was a greater chemotherapy response in patients with
high expression of CXCL13 than those with low expression of CXCL13
in TNBCs. TAPBPL is expressed by resting and activated T cells, B
cells, DCs and monocytes. Experimental investigations demonstrated
that TAPBPL inhibits T cell activation in vitro and its
inhibition enhances antitumour immunity in culture and mice
(62). CXCL10 was recognised as a
strong T cell chemoattractant and orchestrates migration,
differentiation and activation of immune cells (63,64).
Yi et al (65) demonstrated
CXCL10 is a key regulatory immune molecule and prognostic factor
for TNBC; Katsuta et al (66) identified high CXCL10 expression
levels facilitated antitumour immunity and improved survival.
Indoleamine 2,3-dioxygenase 1 (IDO1) mRNA is upregulated in several
cancer types, inhibiting T-cell infiltration and limiting
pharmacological approaches targeting IDO1 (67,68).
In TNBC, high IDO1 transcription is associated with recurrence
<5 years after chemotherapy administration (69). IFNG remains one of the original
effector cytokines for antitumour immunity and one of the most
continuously associated predictors for ICI candidate-responding
tumours (70,71). Increased transcription of IFNG was
reported to be strongly associated with improved disease-free
survival in TNBC (HR=0.38) (72).
The present risk-score analysis indicated that high
CXCL13 expression in TNBC samples is associated with significantly
higher IS scores, increased overall CD8+ T cells and
increased M1 macrophages and significantly lower M2 macrophages,
indicating a more pro-immunogenic antitumour immune environment.
CXCL13 demonstrated strong positive correlations with multiple
immune-checkpoint genes (PD-L1, TIM3, LAG3, CTLA-4 and TIGIT),
indicating that CXCL13 reflects TIME state and predicts ICI
response. Increased TAPBPL expression was also positively
correlated with increased IS score and CD8+ T-cell
density and inversely correlated with increased M0 and M2
macrophages. Furthermore, increased TAPBPL expression demonstrated
positive correlations with CTLA4, TIM3 and LAG3. PGF is a member of
the vascular endothelial growth factor family that promotes
neovascularisation and inflammation (72); here, increased PGF expression was
significantly associated with greater infiltration of M0 and lower
infiltration of M1 macrophages. LTBP2, which has been implicated in
various malignancies (73),
indicated significantly inverse correlations with LAG3. Due to the
contrasting pro- and anti-differentiation functions of M1 and M2
macrophages in cancer biology, these IRGs might represent
actionable biomarkers that could be used for treatment decisions
and prognosis in TNBC.
Finally, to validate the accuracy of this
prognostic model, we constructed ROC curves and a nomogram to
compare the predictive efficacy across different ages, AJCC, T
stages, N stages, and risk scores. In the ROC analysis, both the
risk score and N stage yielded AUC values greater than 0.8,
demonstrating excellent diagnostic performance. Meanwhile, the
nomogram clearly indicates that a lower total score corresponds to
higher 3, 5, and 10-year survival rates, which aligns with clinical
expectations. In summary, compared with conventional methods, the
present prognostic model exhibits superior predictive accuracy.
To further explore immune evasion mechanisms, the
present study also calculated TIDE scores, with larger scores
representing an increased propensity to immune escape and
resistance to ICI therapy. Low-risk TNBC tumours and tumours with
high CXCL13, TAPBPL or PGF expression demonstrated statistically
lower TIDE scores, indicating these tumours might be more
susceptible to checkpoint blockade.
To the best of our knowledge, the present study is
the first to combine TMB status with CD8+ T-cell
infiltration in an overall immunogenomic context to produce an
overall risk model of IRGs for TNBC. In addition, the present study
demonstrated that the risk signature is closely associated multiple
immune-checkpoint mediators, advancing the current understanding of
the TNBC immune landscape and providing a potential biomarker for
the prospective prediction of immunotherapy benefit in this
aggressive subtype of breast cancer.
There are still certain limitations in the present
study. Firstly, the prognostic index in the present study was based
on gene expression data provided by TCGA, and the high price and
long testing time might limit the application of the prognostic
index in clinical practice. Secondly, despite the approval of ICIs
for the first-line treatment of recurrent and metastatic TNBC, the
number of patients receiving ICI treatment is currently limited.
Thus, more cases undergoing ICI treatment are warranted in future
investigations. Furthermore, due to the relatively low incidence
rate of TNBC, the present study did not have sufficient samples to
quantitatively analyse the association between the expression
levels of PD-L1 and immune-related genes in the tumour immune
microenvironment. Future studies may include a larger sample size
to verify the present study findings.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
The present study was supported by Science and Technology
Project Foundation of Suzhou (grant no. SKJY2021126) and Clinical
Oncology Research Foundation of Beijing CSCO (grant no.
Y-Young2021-0087).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
WW, YW and YoM designed the present study. YW and
YoM provided administrative support. WW, LL and JZ provided study
materials and recruited patients. ZH, YinL, YH and ZZ collected the
data. ZH, YuM, YifL and HW conducted the data analysis and
interpretation. WW and YuM confirm the authenticity of all the
data. All authors wrote the main manuscript. All authors read and
approved the final manuscript.
Ethics approval and consent to
participate
The research protocol was approved by the Ethics
Committee of Suzhou Municipal Hospital (approval no.
K-2022-019-K01; Suzhou, China) and conforms to the provisions of
the Declaration of Helsinki. Written informed consent was provided
by all patients prior to the surgical operation.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Lei H, Fu J, Gu W, Qiao H, Guo H, Chen Z,
Wang SM and Chen T: Breast cancer: Molecular pathogenesis, targeted
therapy, screening, and prevention. MedComm (2020). 7:e705602026.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Cancer Genome Atlas Network, .
Comprehensive molecular portraits of human breast tumours. Nature.
490:61–70. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Dent R, Trudeau M, Pritchard KI, Hanna WM,
Kahn HK, Sawka CA, Lickley LA, Rawlinson E, Sun P and Narod SA:
Triple-negative breast cancer: Clinical features and patterns of
recurrence. Clin Cancer Res. 13:4429–4434. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Lehmann BD, Bauer JA, Chen X, Sanders ME,
Chakravarthy AB, Shyr Y and Pietenpol JA: Identification of human
triple-negative breast cancer subtypes and preclinical models for
selection of targeted therapies. J Clin Invest. 121:2750–2767.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Palma M: Advancing breast cancer
treatment: The role of immunotherapy and cancer vaccines in
overcoming therapeutic challenges. Vaccines (Basel). 13:3442025.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Lyons TG, Dickler MN and Comen EE:
Checkpoint inhibitors in the treatment of breast cancer. Curr Oncol
Rep. 20:512018. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Ye J, Wu S, Quan Q, Ye F, Zhang J, Song C,
Fan Y, Cao H, Tang H and Zhao J: Fibroblast growth factor receptor
4 promotes triple-negative breast cancer progression via regulating
fatty acid metabolism through the AKT/RYR2 signaling. Cancer Med.
13:e704392024. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Xu A, Li X, Cai Q, Yang C, Yang M, Gao H,
Cheng M, Chen X, Ji F, Tang H and Wang K: CircXPO6 promotes breast
cancer progression through competitively inhibiting the
ubiquitination degradation of c-Myc. Mol Cell Biochem.
480:1731–1745. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Emens LA: Breast cancer immunotherapy:
Facts and hopes. Clin Cancer Res. 24:511–520. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Loi S: Tumor-infiltrating lymphocytes,
breast cancer subtypes and therapeutic efficacy. Oncoimmunology.
2:e247202013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Adams S, Gray RJ, Demaria S, Goldstein L,
Perez EA, Shulman LN, Martino S, Wang M, Jones VE, Saphner TJ, et
al: Prognostic value of tumor-infiltrating lymphocytes in
triple-negative breast cancers from two phase III randomized
adjuvant breast cancer trials: ECOG 2197 and ECOG 1199. J Clin
Oncol. 32:2959–2966. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Dieci MV, Criscitiello C, Goubar A, Viale
G, Conte P, Guarneri V, Ficarra G, Mathieu MC, Delaloge S,
Curigliano G and Andre F: Prognostic value of tumor-infiltrating
lymphocytes on residual disease after primary chemotherapy for
triple-negative breast cancer: A retrospective multicenter study.
Ann Oncol. 25:611–618. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Adams S, Goldstein LJ, Sparano JA, Demaria
S and Badve SS: Tumor infiltrating lymphocytes (TILs) improve
prognosis in patients with triple negative breast cancer (TNBC).
Oncoimmunology. 4:e9859302015. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Mahmoud SMA, Paish EC, Powe DG, Macmillan
RD, Grainge MJ, Lee AH, Ellis IO and Green AR: Tumor-infiltrating
CD8+ lymphocytes predict clinical outcome in breast cancer. J Clin
Oncol. 29:1949–1955. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Taube JM, Klein A, Brahmer JR, Xu H, Pan
X, Kim JH, Chen L, Pardoll DM, Topalian SL and Anders RA:
Association of PD-1, PD-1 ligands, and other features of the tumor
immune microenvironment with response to anti-PD-1 therapy. Clin
Cancer Res. 20:5064–5074. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Tumeh PC, Harview CL, Yearley JH, Shintaku
IP, Taylor EJ, Robert L, Chmielowski B, Spasic M, Henry G, Ciobanu
V, et al: PD-1 blockade induces responses by inhibiting adaptive
immune resistance. Nature. 515:568–571. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Liu L, Bai X, Wang J, Tang XR, Wu DH, Du
SS, Du XJ, Zhang YW, Zhu HB, Fang Y, et al: Combination of TMB and
CNA stratifies prognostic and predictive responses to immunotherapy
across metastatic cancer. Clin Cancer Res. 25:7413–7423. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Brown SD, Warren RL, Gibb EA, Martin SD,
Spinelli JJ, Nelson BH and Holt RA: Neo-antigens predicted by tumor
genome meta-analysis correlate with increased patient survival.
Genome Res. 24:743–750. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Kumari N, Dwarakanath BS, Das A and Bhatt
AN: Role of interleukin-6 in cancer progression and therapeutic
resistance. Tumour Biol. 37:11553–11572. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Warburg O: On the origin of cancer cells.
Science. 123:309–314. 1956. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Chalmers ZR, Connelly CF, Fabrizio D, Gay
L, Ali SM, Ennis R, Schrock A, Campbell B, Shlien A, Chmielecki J,
et al: Analysis of 100,000 human cancer genomes reveals the
landscape of tumor mutational burden. Genome Med. 9:342017.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Samstein RM, Lee CH, Shoushtari AN,
Hellmann MD, Shen R, Janjigian YY, Barron DA, Zehir A, Jordan EJ,
Omuro A, et al: Tumor mutational load predicts survival after
immunotherapy across multiple cancer types. Nat Genet. 51:202–206.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Rody A, Holtrich U, Pusztai L, Liedtke C,
Gaetje R, Ruckhaeberle E, Solbach C, Hanker L, Ahr A, Metzler D, et
al: T-cell metagene predicts a favorable prognosis in estrogen
receptor-negative and HER2-positive breast cancers. Breast Cancer
Res. 11:R152009. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Iglesia MD, Parker JS, Hoadley KA, Serody
JS, Perou CM and Vincent BG: Genomic analysis of immune cell
infiltrates across 11 tumor types. J Natl Cancer Inst.
108:djw1442016. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Bedognetti D, Hendrickx W, Marincola FM
and Miller LD: Prognostic and predictive immune gene signatures in
breast cancer. Curr Opin Oncol. 27:433–444. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Liu Y, Hu Y, Xue J, Li J, Yi J, Bu J,
Zhang Z, Qiu P and Gu X: Advances in immunotherapy for
triple-negative breast cancer. Mol Cancer. 22:1452023. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Villacampa G, Navarro V, Matikas A,
Ribeiro JM, Schettini F, Tolosa P, Martínez-Sáez O, Sánchez-Bayona
R, Ferrero-Cafiero JM, Salvador F, et al: Neoadjuvant immune
checkpoint inhibitors plus chemotherapy in early breast cancer: A
systematic review and meta-analysis. JAMA Oncol. 10:1331–1341.
2024. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Das R, Deb S and Suresh PK: TMB as a
predictive biomarker for ICI response in TNBC: Current evidence and
future directions for augmented anti-tumor responses. Clin Exp Med.
26:252025. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Wang X and Chen H: Prognosis prediction
through an integrated analysis of single-cell and bulk
RNA-sequencing data in triple-negative breast cancer. Front Genet.
13:9281752022. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Thomas A, Routh ED, Pullikuth A, Jin G, Su
J, Chou JW, Hoadley KA, Print C, Knowlton N, Black MA, et al: Tumor
mutational burden is a determinant of immune-mediated survival in
breast cancer. Oncoimmunology. 7:e14908542018. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Bhattacharya S, Andorf S, Gomes L, Dunn P,
Schaefer H, Pontius J, Berger P, Desborough V, Smith T, Campbell J,
et al: ImmPort: Disseminating data to the public for the future of
immunology. Immunol Res. 58:234–239. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Jézéquel P, Loussouarn D,
Guérin-Charbonnel C, Campion L, Vanier A, Gouraud W, Lasla H,
Guette C, Valo I, Verrièle V and Campone M: Gene-expression
molecular subtyping of triple-negative breast cancer tumours:
Importance of immune response. Breast Cancer Res. 17:432015.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Robinson MD, McCarthy DJ and Smyth GK:
edgeR: A bioconductor package for differential expression analysis
of digital gene expression data. Bioinformatics. 26:139–140. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Liu J, Lichtenberg T, Hoadley KA, Poisson
LM, Lazar AJ, Cherniack AD, Kovatich AJ, Benz CC, Levine DA, Lee
AV, et al: An integrated TCGA pan-cancer clinical data resource to
drive high-quality survival outcome analytics. Cell.
173:400–416.e11. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Li T, Fan J, Wang B, Traugh N, Chen Q, Liu
JS, Li B and Liu XS: TIMER: A web server for comprehensive analysis
of tumor-infiltrating immune cells. Cancer Res. 77:e108–e110. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Giuliano AE, Connolly JL, Edge SB,
Mittendorf EA, Rugo HS, Solin LJ, Weaver DL, Winchester DJ and
Hortobagyi GN: Breast cancer-major changes in the American joint
committee on cancer eighth edition cancer staging manual. CA Cancer
J Clin. 67:290–303. 2017.PubMed/NCBI
|
|
37
|
McCarty KS Jr, Szabo E, Flowers JL, Cox
EB, Leight GS, Miller L, Konrath J, Soper JT, Budwit DA, Creasman
WT, et al: Use of a monoclonal anti-estrogen receptor antibody in
the immunohistochemical evaluation of human tumors. Cancer Res. 46
(8 Suppl):4244s–4248s. 1986.PubMed/NCBI
|
|
38
|
Vikas P, Borcherding N and Zhang W: The
clinical promise of immunotherapy in triple-negative breast cancer.
Cancer Manag Res. 10:6823–6833. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Halama A, Takada K, Takamori S, Haratake
N, Akamine T, Kinoshita F, Ono Y, Wakasu S, Tanaka K, Oku Y, et al:
Clinical significance of preoperative inflammatory markers in
non-small cell lung cancer patients: A multicenter retrospective
study. PLoS One. 15:e02415802020. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Dent R, Cortés J, Pusztai L, McArthur H,
Kümmel S, Bergh J, Denkert C, Park YH, Hui R, Harbeck N, et al:
Neoadjuvant pembrolizumab plus chemotherapy/adjuvant pembrolizumab
for early-stage triple-negative breast cancer: Quality-of-life
results from the randomized KEYNOTE-522 study. J Natl Cancer Inst.
116:1654–1663. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Schmid P, Rugo HS, Adams S, Schneeweiss A,
Barrios CH, Iwata H, Diéras V, Henschel V, Molinero L, Chui SY, et
al: Atezolizumab plus nab-paclitaxel as first-line treatment for
unresectable, locally advanced or metastatic triple-negative breast
cancer (IMpassion130): Updated efficacy results from a randomised,
double-blind, placebo-controlled, phase 3 trial. Lancet Oncol.
21:44–59. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Sukumar J, Gast K, Quiroga D, Lustberg M
and Williams N: Triple-negative breast cancer: promising prognostic
biomarkers currently in development. Expert Rev Anticancer Ther.
21:135–148. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Dvir K, Giordano S and Leone JP:
Immunotherapy in breast cancer. Int J Mol Sci. 25:75172024.
View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Ravelli A, Roviello G, Cretella D,
Cavazzoni A, Biondi A, Cappelletti MR, Zanotti L, Ferrero G, Ungari
M, Zanconati F, et al: Tumor-infiltrating lymphocytes and breast
cancer: Beyond the prognostic and predictive utility. Tumour Biol.
39:10104283176950232017. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Dushyanthen S, Beavis PA, Savas P, Teo ZL,
Zhou C, Mansour M, Darcy PK and Loi S: Relevance of
tumor-infiltrating lymphocytes in breast cancer. BMC Med.
13:2022015. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Rizvi NA, Hellmann MD, Snyder A, Kvistborg
P, Makarov V, Havel JJ, Lee W, Yuan J, Wong P, Ho TS, et al: Cancer
immunology. Mutational landscape determines sensitivity to PD-1
blockade in non-small cell lung cancer. Science. 348:124–128. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Naing A, Infante JR, Papadopoulos KP, Chan
IH, Shen C, Ratti NP, Rojo B, Autio KA, Wong DJ, Patel MR, et al:
PEGylated IL-10 (Pegilodecakin) induces systemic immune activation,
CD8+ T cell invigoration and polyclonal T cell expansion
in cancer patients. Cancer Cell. 34:775–791.e3. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Bense RD, Sotiriou C, Piccart-Gebhart MJ,
Haanen JBAG, van Vugt MATM, de Vries EGE, Schröder CP and Fehrmann
RSN: Relevance of tumor-infiltrating immune cell composition and
functionality for disease outcome in breast cancer. J Natl Cancer
Inst. 109:djw1922016. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Zou Y, Zou X, Zheng S, Tang H, Zhang L,
Liu P and Xie X: Efficacy and predictive factors of immune
checkpoint inhibitors in metastatic breast cancer: A systematic
review and meta-analysis. Ther Adv Med Oncol.
12:17588359209409282020. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Egelston CA, Avalos C, Tu TY, Rosario A,
Wang R, Solomon S, Srinivasan G, Nelson MS, Huang Y, Lim MH, et al:
Resident memory CD8+ T cells within cancer islands mediate survival
in breast cancer patients. JCI Insight. 4:e1300002019. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Gatti-Mays ME, Balko JM, Gameiro SR, Bear
HD, Prabhakaran S, Fukui J, Disis ML, Nanda R, Gulley JL, Kalinsky
K, et al: If we build it they will come: Targeting the immune
response to breast cancer. NPJ Breast Cancer. 5:372019. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Doedens AL, Stockmann C, Rubinstein MP,
Liao D, Zhang N, DeNardo DG, Coussens LM, Karin M, Goldrath AW and
Johnson RS: Macrophage expression of hypoxia-inducible factor-1
alpha suppresses T-cell function and promotes tumor progression.
Cancer Res. 70:7465–7475. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Ruffell B, Chang-Strachan D, Chan V,
Rosenbusch A, Ho CM, Pryer N, Daniel D, Hwang ES, Rugo HS and
Coussens LM: Macrophage IL-10 blocks CD8+ T cell-dependent
responses to chemotherapy by suppressing IL-12 expression in
intratumoral dendritic cells. Cancer Cell. 26:623–637. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Montes VN, Turner MS, Subramanian S, Ding
Y, Hayden-Ledbetter M, Slater S, Goodspeed L, Wang S, Omer M, Den
Hartigh LJ, et al: T cell activation inhibitors reduce CD8+ T cell
and pro-inflammatory macrophage accumulation in adipose tissue of
obese mice. PLoS One. 8:e677092013. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Xu Y, Romero R, Miller D, Kadam L, Mial
TN, Plazyo O, Garcia-Flores V, Hassan SS, Xu Z, Tarca AL, et al: An
M1-like macrophage polarization in decidual tissue during
spontaneous preterm labor that is attenuated by rosiglitazone
treatment. J Immunol. 196:2476–2491. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Mao Y, Keller ET, Garfield DH, Shen K and
Wang J: Stromal cells in tumor microenvironment and breast cancer.
Cancer Metastasis Rev. 32:303–315. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Qian BZ and Pollard JW: Macrophage
diversity enhances tumor progression and metastasis. Cell.
141:39–51. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Daou HN: Exercise as an anti-inflammatory
therapy for cancer cachexia: A focus on interleukin-6 regulation.
Am J Physiol Regul Integr Comp Physiol. 318:R296–R310. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Denkert C, Loibl S, Noske A, Roller M,
Müller BM, Komor M, Budczies J, Darb-Esfahani S, Kronenwett R,
Hanusch C, et al: Tumor-associated lymphocytes as an independent
predictor of response to neoadjuvant chemotherapy in breast cancer.
J Clin Oncol. 28:105–113. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Lee HJ, Lee JJ, Song IH, Park IA, Kang J,
Yu JH, Ahn JH and Gong G: Prognostic and predictive value of
NanoString-based immune-related gene signatures in a neoadjuvant
setting of triple-negative breast cancer: Relationship to
tumor-infiltrating lymphocytes. Breast Cancer Res Treat.
151:619–627. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Lv Y, Lv D, Lv X, Xing P, Zhang J and
Zhang Y: Immune cell infiltration-based characterization of
triple-negative breast cancer predicts prognosis and chemotherapy
response markers. Front Genet. 12:6164692021. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Lin Y, Cui C, Su M, Silbart LK, Liu H,
Zhao J, He L, Huang Y, Xu D, Wei X, et al: Identification of TAPBPL
as a novel negative regulator of T-cell function. EMBO Mol Med.
13:e134042021. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Yu Y, Li J, Zhu X, Tang X, Bao Y, Sun X,
Huang Y, Tian F, Liu X and Yang L: Humanized CD7 nanobody-based
immunotoxins exhibit promising anti-T-cell acute lymphoblastic
leukemia potential. Int J Nanomedicine. 12:1969–1983. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Tokunaga R, Zhang W, Naseem M, Puccini A,
Berger MD, Soni S, McSkane M, Baba H and Lenz HJ: CXCL9, CXCL10,
CXCL11/CXCR3 axis for immune activation-a target for novel cancer
therapy. Cancer Treat Rev. 63:40–47. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Yi J, Shuang Z, Zhong W, Wu H, Feng J,
Zouxu X, Huang X, Li S and Wang X: Identification of immune-related
risk characteristics and prognostic value of immunophenotyping in
TNBC. Front Genet. 12:7304422021. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Katsuta E, Yan L, Opyrchal M, Kalinski P
and Takabe K: Cytotoxic T-lymphocyte infiltration and chemokine
predict long-term patient survival independently of tumor
mutational burden in triple-negative breast cancer. Ther Adv Med
Oncol. 13:175883592110066802021. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Li F, Zhang R, Li S and Liu J: IDO1: An
important immunotherapy target in cancer treatment. Int
Immunopharmacol. 47:70–77. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Cheong JE and Sun L: Targeting the
IDO1/TDO2-KYN-AhR pathway for cancer immunotherapy-challenges and
opportunities. Trends Pharmacol Sci. 39:307–325. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Pérez-Pena J, Tibor Fekete J, Páez R,
Baliu-Piqué M, García-Saenz JÁ, García-Barberán V, Manzano A,
Pérez-Segura P, Esparis-Ogando A, Pandiella A, et al: A
Transcriptomic immunologic signature predicts favorable outcome in
neoadjuvant chemotherapy treated triple negative breast tumors.
Front Immunol. 10:28022019. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Castro F, Cardoso AP, Gonçalves RM, Serre
K and Oliveira MJ: Interferon-gamma at the crossroads of tumor
immune surveillance or evasion. Front Immunol. 9:8472018.
View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Burke JD and Young HA: IFN-γ: A cytokine
at the right time, is in the right place. Semin Immunol.
43:1012802019. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Yeong J, Lim JCT, Lee B, Li H, Ong CCH,
Thike AA, Yeap WH, Yang Y, Lim AYH, Tay TKY, et al: Prognostic
value of CD8 + PD-1+ immune infiltrates and PDCD1 gene expression
in triple negative breast cancer. J Immunother Cancer. 7:342019.
View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Chen J, Gao G, Wang H, Ye X, Zhou J and
Lin J: Expression and clinical significance of latent-transforming
growth factor beta-binding protein 2 in primary hepatocellular
carcinoma. Medicine (Baltimore). 98:e172162019. View Article : Google Scholar : PubMed/NCBI
|