Introduction
Lung cancer is the leading cause of cancer-related
mortality worldwide, with an estimated 2.5 million new cases and
1.8 million deaths annually (1).
Non-small cell lung cancer (NSCLC) is the most prevalent
histological subtype, accounting for ~85% of all lung cancer cases.
Among NSCLC subtypes, lung adenocarcinoma (LUAD) and lung squamous
cell carcinoma (LUSC/LSCC) are the most prevalent (2). Despite advancements in lung cancer
treatment strategies, the prognosis for patients with inoperable
NSCLC remains poor, with a 5-year survival rate ranging from 13 to
60% (3–5). Consequently, elucidation of the
molecular mechanisms underlying the development and progression
NSCLC is critical for improving the outcomes of patients with this
disease.
BUD23, also known as Williams-Beuren syndrome
critical region 22, is a member of the rRNA methyltransferase
family that functions as a ribosome maturation factor and was
initially identified in patients with Williams-Beuren syndrome
(6). Current evidence indicates
that that BUD23 acts as a methyltransferase and plays a crucial
role in RNA methylation (7,8). In addition, it has a confirmed
involvement in ribosome biogenesis (7,9).
Several studies have suggested that BUD23
contributes to tumorigenesis. For example, high BUD23 expression
has been associated with poor prognosis in patients with
glioblastoma (10). In addition,
BUD23 contributes to the survival of multiple myeloma cells, and
its knockdown significantly impairs cell proliferation and invasive
capacity, and its knockdown significantly impairs cell
proliferation and invasive capacity (11). In breast cancer, BUD23 promotes
metastasis by inhibiting Zac1/p53-dependent apoptosis (12). Conversely, BUD23 and tRNA
methyltransferase activator subunit 11-2 together inhibit
pancreatic cancer cell proliferation, invasion and tumorigenesis
through the transcriptional regulation of interferon-stimulated
gene 15 (13). In lung cancer
cells, BUD23 has been implicated in drug resistance, as the
knockdown of BUD23 reduces the sensitivity of H460 NSCLC cells to
the active camptothecin metabolite 7-ethyl-10-hydroxycamptothecin
and 5-fluorouracil (14). However,
despite its involvement in multiple cancer types, the precise role
of BUD23 in NSCLC remains to be elucidated and warrants further
investigation.
In the present study, transcriptomics and proteomics
profiling was performed to evaluate the expression of BUD23 in
NSCLC, and its association with patient outcomes was assessed.
Integrative immunogenomic analysis was conducted to evaluate the
association of BUD23 levels with tumor immune infiltration. Gene
Set Enrichment Analysis (GSEA), Kyoto Encyclopedia of Genes and
Genomes (KEGG) and single-cell pathway enrichment analyses were
also performed to identify the pathways associated with BUD23. In
addition, the effect of BUD23 knockdown on the proliferation and
migration of NSCLC cells was evaluated in vitro, and its
potential downstream genes were identified and verified. The
findings are intended to provide a basis for future research into
the role of BUD23 in the development and progression of NSCLC.
Materials and methods
Expression analysis of BUD23
Differences in the expression of BUD23 between tumor
and normal tissues were analyzed using standardized data from The
Cancer Genome Atlas (TCGA), including TCGA-LUAD and TCGA-LUSC
cohorts. The TCGA data downloaded from GEPIA2 (http://gepia2.cancer-pku.cn/#index) completed
standardization. Additionally, two proteomic datasets derived from
lung cancer samples were retrieved from the National Cancer
Institute's Clinical Proteomic Tumor Analysis Consortium (CPTAC:
http://pdc.cancer.gov/pdc), namely the
CPTAC LSCC Discovery Study and CPTAC LUAD Discovery Study (15,16).
Gene expression omnibus (GEO) dataset
processing
The datasets GSE30219 (17), GSE19188 (18), GSE40791 (19), GSE32665 (20), GSE40419 (21), GSE75037 (22), GSE7670 (23), GSE27262 (24), GSE63459 (25), GSE87340 (26), GSE31547 (27) and GSE31210 (28), including their matrix and platform
information, were obtained from the GEO database (http://www.ncbi.nlm.nih.gov/geo). We used the
gene identifiers provided by the probe platform, as well as the
expression values in the data matrix. All expression matrices have
undergone normalization by GEO submitters.
Survival analysis
The Kaplan-Meier plotter tool (http://kmplot.com/analysis/) was used to evaluate the
association of BUD23 expression with survival (29). BUD23 (Affymetrix ID 207628_s_at) was
also subjected to univariate and multivariate Cox regression
analyses within the lung cancer module. The patient population was
segmented using the ‘Auto select the best cutoff’ feature.
Kaplan-Meier survival curves were generated for overall survival
(OS), post-progression survival (PPS) and first progression
survival (FPS). P<0.05 was considered to indicate statistical
significance.
Immune infiltration analysis in NSCLC
from Sangerbox 2.0 (http://sangerbox.com/)
To assess the extent of immune infiltration
associated with UD23 expression in NSCLC, several online analysis
platforms utilizing TCGA-LUAD and -LUSC datasets were employed.
Specifically, correlations between BUD23 expression and immune cell
infiltration levels were analyzed using TIMER, EPIC, MCPcounter and
QUANTISEQ algorithms (30–33). The correlations of BUD23 expression
with stromal, immune and ESTIMATE scores were further investigated
using the ESTIMATE algorithm (34).
Additionally, the Immunophenoscore (IPS), calculated using the IPS
algorithm, was used to evaluate the correlation between BUD23
expression and various components associated with tumor
immunogenicity in NSCLC (35).
Correlation analysis using UALCAN
UALCAN (https://ualcan.path.uab.edu/) was used to identify the
genes that significantly correlated with BUD23 in the ‘TCGA’ module
(36). After inputting ‘BUD23’ as
the Gene Symbol and selecting the ‘Correlation analysis’ module,
the analysis results were downloaded for both LUAD and LUSC. Genes
with extremely low expression (median transcripts per million
<0.5) were excluded from the list, and only those with
R2≥0.3 and P<0.05 were included in the final sets.
This yielded 134 significantly correlated genes in LUAD and 165 in
LUSC.
Correlation analysis using
Metascape
The KEGG enrichment analysis was performed utilizing
Metascape (https://metascape.org/gp/index.html#/main/step1). All
enrichment parameters were set as the default values (Min Overlap:
3; P-value Cutoff: 0.01; Min Enrichment: 1.5), and the KEGG Pathway
database was selected as the enrichment dataset for subsequent
analysis.
GSEA pathway enrichment analysis
TCGA-LUAD and -LUSC data were analyzed by GSEA. The
patients were divided into BUD23 high and low groups according to
the median expression level of BUD23 within each cohort. The
Hallmark gene sets were used for GSEA analysis. GSEA software
(v3.0) was obtained from the official website
(DOI:10.1073/pnas.0506580102; http://software.broadinstitute.org/gsea/index.jsp).
Samples were stratified into high and low BUD23 expression groups
by the 50% cutoff. The h.all.v7.4.symbols.gmt gene set was
downloaded from MSigDB (DOI:10.1093/bioinformatics/btr260;
http://www.gsea-msigdb.org/gsea/downloads.jsp).
Parameters were set as: Minimum gene set size=5, maximum=5,000 and
1,000 permutations. P<0.05 and FDR <0.25 were considered
statistically significant. P<0.05 was considered to be
statistically significant.
Single-cell analysis of BUD23 in
NSCLC
The CancerSEA online database (http://biocc.hrbmu.edu.cn/CancerSEA/home.jsp) was used
to examine the association of BUD23 with 14 different functional
states of NSCLC at single-cell resolution in the E-MTAB-6149
dataset (37). These functional
states encompass angiogenesis, apoptosis, invasion,
epithelial-mesenchymal transition, differentiation, proliferation,
DNA damage, metastasis, hypoxia, inflammation, cell cycle
progression, DNA repair, stemness and quiescence.
Cell culture and cell
transfection
The HBE [full name: HBE4-E6/E7 (Human Bronchial
Epithelial Cells; cat. no. CRL-2078)] cell line, A549 lung
adenocarcinoma cell line (cat. no. CCL-185), H1299 lung large cell
carcinoma cell line (cat. no. CRL-5803), H460 lung large cell
carcinoma cell line (cat. no. HTB-177) and Jurkat T cells (cat. no.
TIB-152; a childhood T acute lymphoblastic leukemia T-cell line),
were purchased from the American Type Culture Collection. Cells
were cultured and maintained in RPMI-1640 (cat. no. PM150110;
Procell Life Science & Technology Co., Ltd.) supplemented with
10% fetal bovine serum (FBS; cat. no. 164210-50; Procell Life
Science & Technology Co., Ltd.), 100 U/ml penicillin and 100
µg/ml streptomycin, and were incubated in a humidified chamber at
37°C with 95% air and 5% CO2.
BUD23-small interfering RNAs (siRNAs) and negative
control siRNA were obtained from Guangzhou RiboBio. The BUD23
siRNAs were as follows: Si1 sense, 5′GTGGTAGACTACCCTAACA3′ and
antisense: 5′CAUCACCUGAGUCGAGUCUU3′; Si2 sense,
5′CAGTGGCTCTGTAATGCTA3′ and antisense, 5′GUCACCACAUCAUCAGCAU3′. The
specific sequence of the negative control siRNA (cat. no.
siN0000002-1-5) was not disclosed by the supplier. The A549 and
H1299 cells were transfected at 50–60% confluency. The transfection
was performed using Lipo8000 transfection reagent (cat. no. C0533;
Beyotime Institute of Biotechnology). The nucleic acid mass used
was 2 µg per well. Transfection was carried out at 37°C for 6 h and
the subsequent experiments were performed 24 h after
transfection.
Transwell co-culture assay
Transwell co-culture was conducted with 0.4 µm pore
Transwell inserts (cat. no. 3413; Corning, Inc.); this pore size
enables soluble factor exchange without direct cell contact.
A549/H1299 cells underwent BUD23 knockdown for 24 h and were then
seeded into the lower chamber at 5×104 cells/well with DMEM (10%
FBS). Jurkat T cells (cultured in RPMI 1640 with 10% FBS) were
added to the upper inserts at 1×105 cells/insert, followed by 48 h
co-culture at 37°C with 5% CO2. Control groups consisted of
BUD23-knockdown A549/H1299 cells cultured alone under the same
conditions. After co-culture, lower-chamber A549/H1299 cells were
washed with cold PBS and harvested for downstream assays.
Wound-healing assays
Following the knockdown of BUD23, A549 and H1299
cells were seeded on 6-well plates and allowed to reach 80–90%
confluence. Then, the cell monolayers were scratched with a 200-µl
pipette tip, and culture was maintained in RPMI-1640 supplemented
with 1% FBS. The migration distance of the cells was measured at 0,
24 and 48 h. Wound images were captured using an inverted
phase-contrast microscope. The wound closure rate was quantified by
measuring the wound width at 0, 24 and 48 h with ImageJ software
(version 1.54r; National Institutes of Health), and the relative
migration rate was calculated accordingly.
Cell counting kit-8 (CCK-8) assay
Cell viability was quantified using a CCK-8 assay
(Dojindo Laboratories, Inc.). Following the siRNA-mediated
knockdown of BUD23, A549 cells and H1299 were seeded in 96-well
plates (3,000 or 2,000 cells/well, respectively; 5 replicates per
group) and cultured for 24 or 48 h (the time when cells were
transferred into 48-well plates and allowed to adhere was set as 0
h). Subsequently, 10 µl CCK-8 reagent was added to each well and
the plate was incubated for 1 h at 37°C in 5% CO2. Absorbance at
450 nm was measured using a microplate reader (BioTek Synergy H1;
Agilent Technologies, Inc.).
Apoptosis assay
Following BUD23 knockdown, A549 and H1299 cells were
seeded in 6-well plates at 70% confluence for 8 h, and then
processed using an Annexin V-FITC Apoptosis Detection Kit (Cat. No.
AD10, lot SH680; Dojindo Laboratories, Inc.). Flow cytometric
analysis was performed using a BD FACSCalibur flow cytometer (BD
Biosciences). Data were analyzed using FlowJo software (version
10.8.1; BD Life Sciences). The percentages of early and late
apoptotic cells were calculated for each sample. The cells in
quadrants Q2 (late apoptosis) and Q3 (early apoptosis) were
considered to be apoptotic and were subjected to analysis.
Cell cycle analysis
Cell cycle distribution in H1299 and A549 cells was
detected using the Cell Cycle and Apoptosis Analysis Kit (cat. no.
C1052, Beyotime Institute of Biotechnology) in strict accordance
with the manufacturer's instructions. Cells were harvested, fixed
with pre-chilled 70% ethanol overnight at 4°C, washed with cold PBS
and stained with PI/RNase A staining buffer at 37°C for 30 min in
the dark. Flow cytometry was conducted for detection and data were
analyzed using FlowJo software (version 10.8.1) to calculate the
percentage of cells in G0/G1, S and G2/M phases.
Reverse transcription-quantitative PCR
(RT-qPCR)
RNA was extracted from the cells using
TRIzol® (Invitrogen; Thermo Fisher Scientific, Inc.). RT
was performed using the PrimeScript™ RT Reagent Kit with
gDNA Eraser (Perfect Real Time; cat. no. RR047A; TaKaRa Bio Inc.)
containing reverse transcriptase, RT buffer, dNTP mix, oligo(dT)
primers, random primers and gDNA Eraser. The cDNA synthesis was
performed strictly according to the manufacturer's protocol and
cDNA was stored at −20°C for qPCR analysis. The qPCR analyses were
performed using SYBR Premix Ex Taq (Takara Bio, Inc.) according to
the manufacturer's instructions [the thermal cycling protocol was
as follows: 42°C for 2 min (gDNA removal), 37°C for 15 min (cDNA
synthesis) and 85°C for 5 sec (enzyme inactivation). cDNA was
stored at −20°C for qPCR analysis) and quantified using a CFX96
Real-Time PCR System (Bio-Rad Laboratories, Inc.). Relative fold
changes in expression were calculated using the 2−ΔΔCq
method (38). The primer pairs used
for qPCR are listed in Table
SI.
Statistical analysis
All statistical analyses were performed using SPSS
software (version 29.0; IBM Corp.). Student's t-tests were used to
analyze the differences between two groups, with an unpaired t-test
used for independent samples and paired t-test for paired samples.
One-way analysis of variance followed by Tukey's post-hoc test was
used for the comparison of three or more groups. Pearson's
correlation analysis was performed to assess correlations among
variables. Data are presented as mean ± standard deviation from
three independent replicates. P<0.05 was considered to indicate
a statistically significant result.
Results
BUD23 is highly expressed in NSCLC
based on TCGA, GTEx and CPTAC data
The mRNA expression profile of BUD23 in various
cancers was examined using data on human cancer tissue from TCGA
database. This analysis revealed that BUD23 mRNA was significantly
upregulated in LUAD and LUSC tissues relative to that in normal
lung tissues (Fig. 1A). Given the
limited number of normal tissue samples in TCGA, the TCGA data were
supplemented with samples from the GTEx database through the GEPIA2
platform. This allowed a more comprehensive comparison of BUD23
expression between normal lung and lung tumor tissues. The results
indicated that BUD23 expression was significantly upregulated in
LUAD and LUSC (fold change >1.5, P<0.01; Fig. 1B and C). In addition, BUD23 protein
expression levels in LUAD and LUSC tissues were compared with those
in normal tissues using CPTAC proteomics data. Consistent with the
mRNA findings, BUD23 protein levels were significantly elevated in
tumor tissues compared with normal tissues (Fig. 1D and E). Collectively, these
analyses demonstrate that BUD23 is highly expressed in NSCLC.
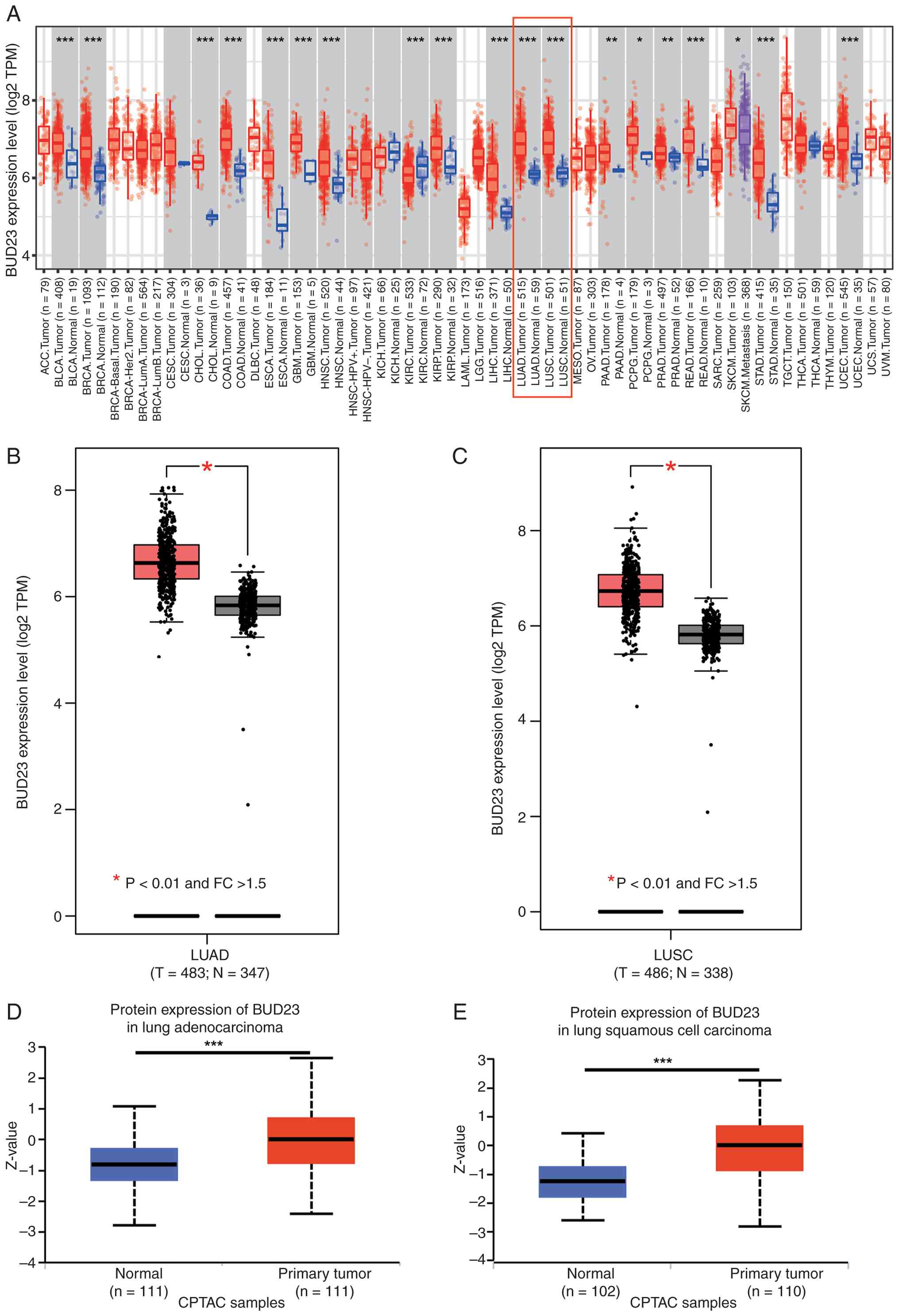 | Figure 1.BUD23 expression is upregulated in
NSCLC based on TCGA, GTEx and CPTAC data. (A) Pan-cancer analysis
of BUD23 expression in tumor and normal tissues based on TCGA data.
Expression analysis of BUD23 in (B) LUAD and (C) LUSC between
cancer and normal tissues based on integrated TCGA and GTEx
datasets. Protein expression analysis of BUD23 in (D) lung
adenocarcinoma or (E) lung squamous cell carcinoma compared with
that in normal tissues based on CPTAC data. *P<0.05, **P<0.01
and ***P<0.001 as indicated. NSCLC, non-small cell lung cancer;
TCGA, The Cancer Genome Atlas; GTEx, Genotype-Tissue Expression;
CPTAC, Clinical Proteomic Tumor Analysis Consortium; LUAD, lung
adenocarcinoma; LUSC, lung squamous cell carcinoma; TPM,
transcripts per million; FC, fold change; T, tumor; N, normal. |
BUD23 is highly expressed in NSCLC
based on GEO datasets
To examine the differential expression of BUD23 in
NSCLC, multiple publicly available gene expression datasets from
the GEO database were analyzed. BUD23 expression was confirmed to
be significantly upregulated in tumor tissues compared with normal
tissues in multiple NSCLC cohorts, namely those in the GSE19188,
GSE30219, GSE40791, GSE32665, GSE40419, GSE75037, GSE7670,
GSE27262, GSE63459 and GSE87340 datasets. This stable upregulation
was observed regardless of whether the samples were unmatched:
Tumor and adjacent normal tissues were from different patients
(Fig. 2A-C) or matched: Tumor and
adjacent normal tissues were from the same patient (Fig. 2D-J), indicating a consistent and
significant upregulation of BUD23 expression in NSCLC. These
analyses suggest that BUD23 may play a role in the development or
progression of NSCLC.
Elevated BUD23 levels are associated
with poor clinical outcomes in patients with NSCLC
The association between BUD23 expression and patient
prognosis was analyzed using the Kaplan-Meier plotter, which
encompasses 17 NSCLC datasets. The analysis verified that elevated
BUD23 expression was significantly associated with shorter OS [high
vs. low; hazard ratio (HR), 1.43; 95% CI, 1.25–1.64; P<0.05;
n=2,166; Fig. 3A], FPS (high vs.
low; HR, 1.61, 95% CI, 1.3–2.0; P<0.05; n=1,252; Fig. 3B) and PPS (high vs. low; HR, 1.58;
95%CI, 1.27–1.97; P<0.05; n=477; Fig. 3C). Furthermore, the association
between BUD23 expression and OS was explored in patients with NSCLC
stratified by N stage. The results revealed that BUD23 expression
was not associated with OS in patients with N0 stage NSCLC
(Fig. 3D). However, elevated BUD23
expression was significantly associated with poor OS in patients
with stages N1 (high vs. low; HR, 1.52; 95% CI, 1.05–2.19;
P<0.05; Fig. 3E) and N2 (high
vs. low; HR, 1.7; 95%CI, 1.05–2.74; P<0.05; Fig. 3F). Multivariate regression analysis
also indicated that BUD23 expression is associated with a poor
prognosis for OS (high vs. low; HR,1.61; 95% CI, 1.22–2.13;
P<0.05) and PPS (high vs. low; HR, 1.92; 95% CI, 1.36–2.72;
P<0.05) (Table I). The
association between BUD23 expression and clinical stage was also
investigated. The analysis of GSE40419, GSE31547 and GSE31210
datasets revealed a significant elevation in BUD23 expression as
the clinical stage advanced (Fig.
3G-I). In conclusion, elevated BUD23 levels are associated with
worse clinical outcomes in patients with NSCLC.
 | Figure 3.Elevated BUD23 levels are associated
with poor clinical outcomes in patients with NSCLC. Kaplan-Meier
survival curves comparing high and low BUD23 expression groups for
(A) OS (n=2,166), (B) FPS (n=1252) and (C) PPS (n=447), generated
using Kaplan-Meier Plotter. (D-F) Kaplan-Meier OS curves comparing
high and low BUD23 expression groups in NSCLC stratified by N
stage: (D) N0, (E) N1 and (F) N2. Association of BUD23 expression
with different clinical stages of NSCLC based on analysis of the
(G) GSE340419, (H) GSE31547 and (I) GSE31210 datasets. *P<0.05
and **P<0.01 as indicated. NSCLC, non-small cell lung cancer;
OS, overall survival; FPS, first progression survival; PPS,
post-progression survival; HR, hazard ratio (95% CI values). |
 | Table I.Multivariate analysis of BUD23
expression in patients with non-small cell lung cancer. |
Table I.
Multivariate analysis of BUD23
expression in patients with non-small cell lung cancer.
|
| Overall
survival | First progression
survival | Post-progression
survival |
|---|
|
|
|
|
|
|---|
| Factor | P-value | Hazard ratio | P-value | Hazard ratio | P-value | Hazard ratio |
|---|
| Histology | 0.5294 | 0.91
(0.67–1.22) | - | - | - | - |
| Grade | 0.9248 | 1.01
(0.82–1.24) | 0.4984 | 0.92
(0.73–1.17) | 0.8174 | 0.97
(0.75–1.26) |
| AJCC T stage | 0.0004 | 1.42
(1.17–1.72) | 0.0012 | 1.51
(1.18–1.93) | 0.0032 | 1.52
(1.15–2.01) |
| AJCC N stage | <0.0001 | 1.85
(1.57–2.19) | <0.0001 | 1.57
(1.27–1.93) | 0.0017 | 1.43
(1.14–1.78) |
| Sex (male vs.
female) | 0.1264 | 1.23
(0.94–1.60) | 0.5043 | 1.11
(0.81–1.53) | 0.8213 | 1.04
(0.74–1.47) |
| Smoking
history | 0.2346 | 0.75
(0.47–1.20) | 0.3323 | 0.79
(0.49–1.27) | 0.8330 | 1.06
(0.62–1.81) |
| BUD23 | 0.0007 | 1.61
(1.22–2.13) | 0.0989 | 1.32
(0.95–1.84) | 0.0002 | 1.92
(1.36–2.72) |
Immune characteristics of BUD23 in
NSCLC
Immune infiltration serves as an independent
prognostic indicator in tumors (39). Consequently, the correlations
between BUD23 expression and immune cell infiltration levels in
TCGA-LUAD and -LUSC cohorts were examined using the TIMER tool. The
results indicated that BUD23 expression is significantly negatively
correlated with B cell, CD8+ T cell and macrophage
counts in LUAD, while in LUSC, BUD23 expression is significantly
negatively correlated with B cell, CD8+ T cell,
neutrophil, macrophage and dendritic cell counts (Fig. 4A). In addition, immune
characterization analyses performed using EPIC (Fig. 4B), MCPcounter (Fig. 4C) and QUANTISEQ (Fig. 4D) further confirmed that BUD23
expression is negatively correlated with the infiltration of B
cells (all three methods; Fig.
4B-D), CD8+ T cell infiltration (MCPcounter and QUANTISEQ;
Fig. 4C and D) and macrophage
infiltration (QUANTISEQ; Fig. 4D).
The IPS is a metric used to evaluate tumor immunogenicity, with
higher IPS values indicating greater immunogenic potential.
Assessment of the correlation between BUD23 expression and IPS
revealed that BUD23 is negatively correlated with IPS in LUSC
(Fig. 4E), suggesting that LUSC
tumors with high BUD23 expression are less immunogenic.
Furthermore, the correlations of BUD23 expression with stromal and
immune cell levels were evaluated using the ESTIMATE algorithm.
High BUD23 expression was found to be negatively correlated with
stromal, immune and ESTIMATE scores in NSCLC (Fig. 4F-H), indicating that patients with
elevated BUD23 expression have higher tumor purity. Collectively,
these findings demonstrate that BUD23 is negatively associated with
immune cell infiltration.
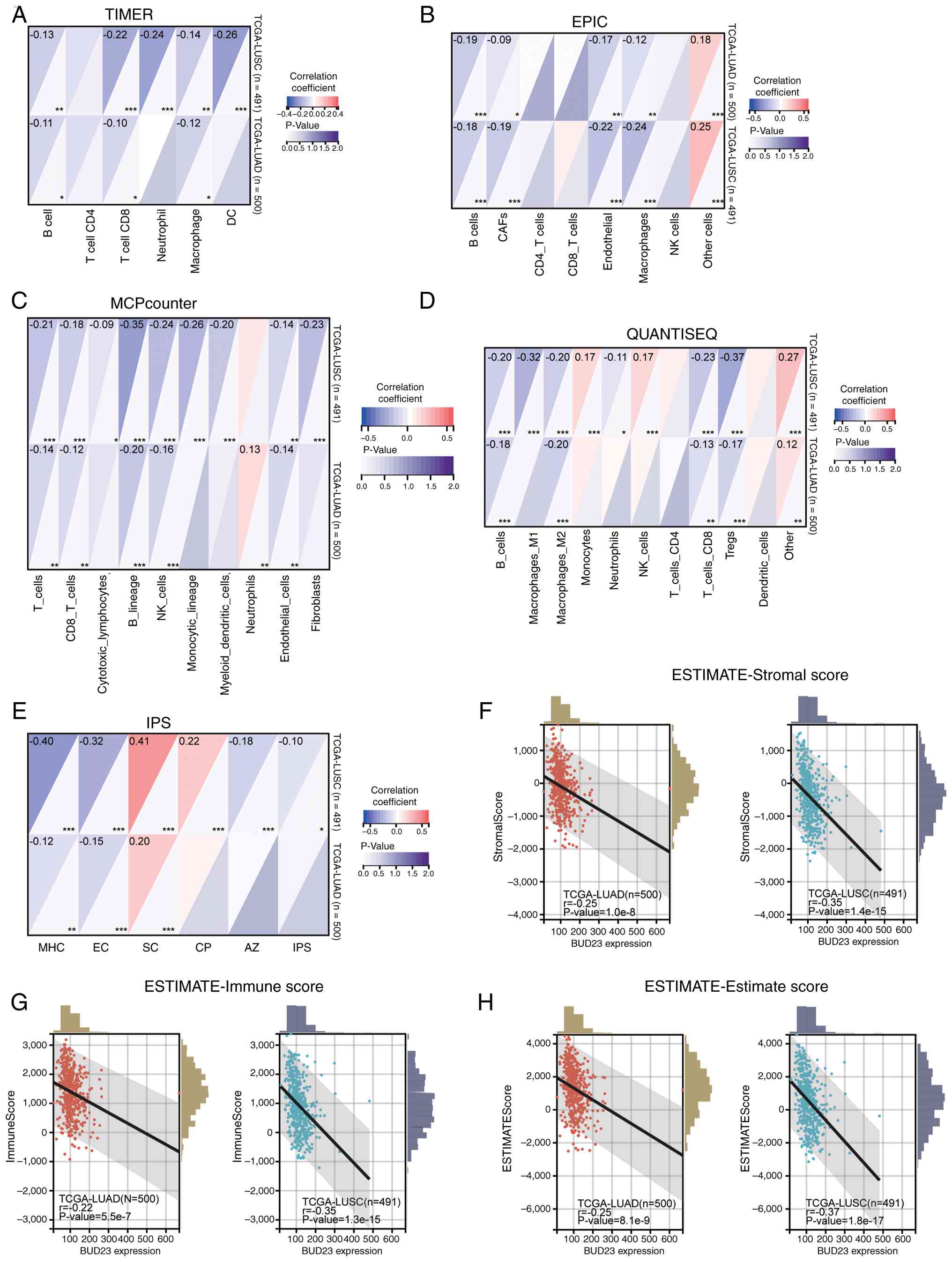 | Figure 4.Correlation between BUD23 and immune
characteristics in NSCLC analyzed using various algorithms based on
TCGA-LUAD and -LUSC data. Correlations of BUD23 expression with the
infiltration levels of (A) 6 immune cell types analyzed using
TIMER, (B) 8 immune cell types analyzed using EPIC, (C) 10 immune
cell types analyzed using MCPcounter and (D) 11 immune cell types
of analyzed using QUANTISEQ. Correlations of BUD23 expression with
(E) IPS score determined using the IPS algorithm and (F) stromal,
(G) immune and (H) ESTIMATE scores analyzed using ESTIMATE.
*P<0.05, **P<0.01 and ***P<0.001. NSCLC, non-small cell
lung cancer; TCGA, The Cancer Genome Atlas; LUAD, lung
adenocarcinoma; LUSC, lung squamous cell carcinoma; IPS,
immunophenoscore; DC, dendritic cell; CAFs, cancer-associated
fibroblasts; NK, natural killer; Tregs, regulatory T cells; MHC,
major histocompatibility complex; EC, effector cell; SC, suppressor
cell; CP, checkpoint molecule; AZ, antigen processing and
presentation machinery. |
BUD23 may be involved in DNA repair
and cell cycle pathways and associated with NSCLC malignancy
To elucidate the potential downstream effects and
molecular mechanisms of BUD23 in NSCLC, GSEA was performed to
compare patients with high and low BUD23 expression levels in
TCGA-LUAD and -LUSC cohorts. The analysis demonstrated that the
Hallmark ‘DNA repair’ gene set was significantly enriched in
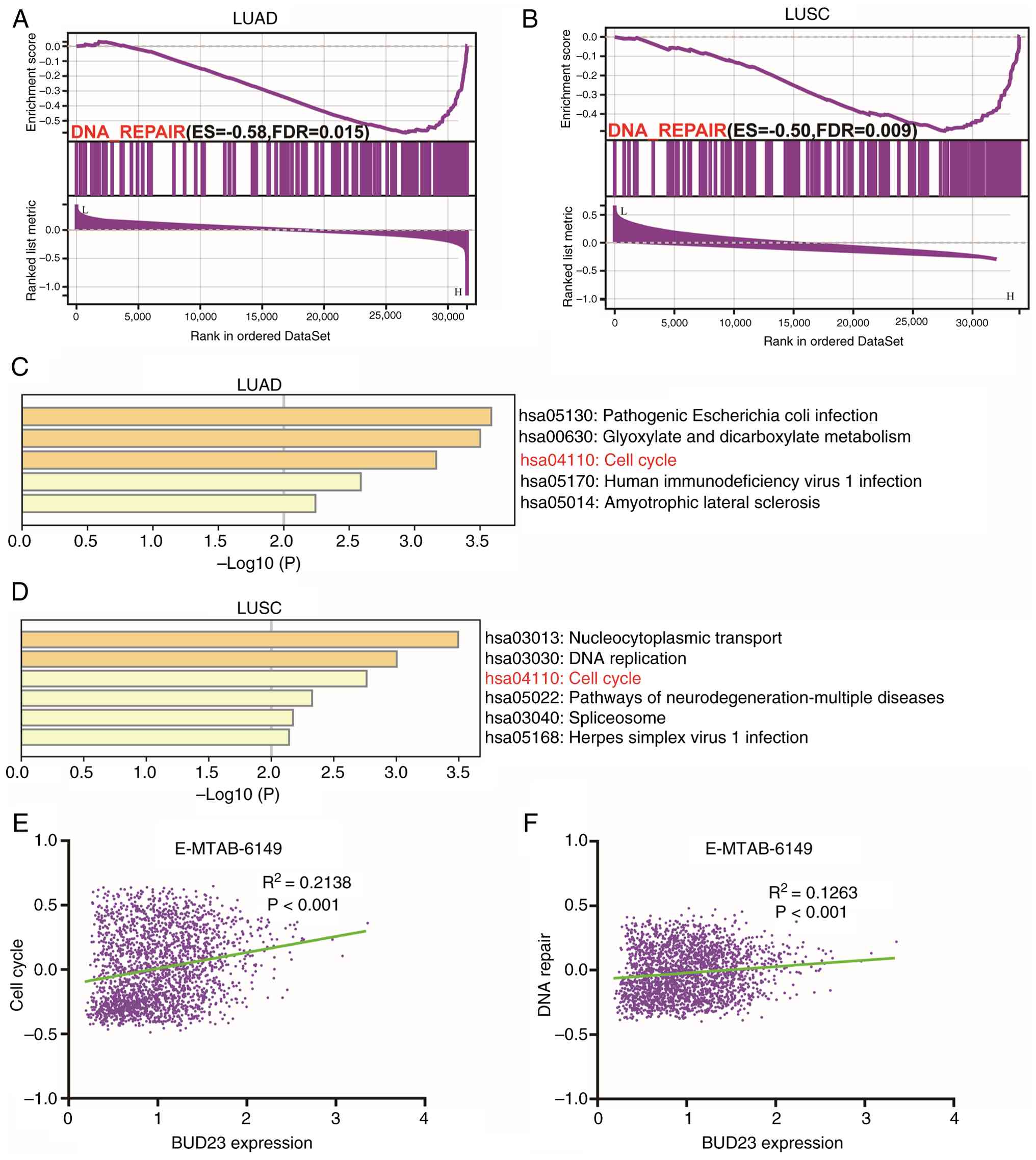
patients with elevated BUD23 expression in both cohorts (Fig. 5A and B). Correlation analyses were
performed in TCGA-LUAD and -LUSC cohorts to screen out the genes
that exhibit a positive correlation with BUD23 (R2≥0.3,
P<0.05), and KEGG enrichment analyses were conducted to identify
the pathways associated with BUD23-related genes. The results
indicate that BUD23-related genes are enriched in the ‘cell cycle’
signaling pathway in both LUAD and LUSC (Fig. 5C and D). Moreover, to elucidate the
correlation of BUD23 expression with cancer functional states in
NSCLC at single-cell resolution, an analysis was conducted via
CancerSEA based on the E-MTAB-6149 dataset. The expression of BUD23
was found to be positively correlated with cell cycle activity
scores (R2=0.2138, P<0.05) and DNA repair activity
scores (R2=0.1263, P<0.05; Fig. 5E and F, respectively). These
findings further suggest that BUD23 is involved in ‘DNA repair’ and
‘cell cycle’ pathways and is associated with NSCLC malignancy.
BUD23 knockdown suppresses the
proliferative and migratory potential of NSCLC cells
To further elucidate the oncogenic role of BUD23 in
NSCLC, its transcript levels in non-malignant human HBE cells and a
panel of three NSCLC cell lines were compared. RT-qPCR revealed a
significant upregulation of BUD23 expression in A549, H1299 and
H460 cells compared with that in HBE cells (P<0.001; Fig. 6A). The siRNA-mediated depletion of
BUD23 in A549 and H1299 cells significantly attenuated cell
viability at the 48-h timepoint (P<0.001; Fig. 6B-E). Wound healing assays
demonstrated that BUD23 knockdown also reduced the migratory
capacity of A549 and H1299 cells after 48 h (P<0.01; Fig. 6F and G). To investigate the
relationship between BUD23 and immune infiltration, Jurkat T cells
were co-cultured with A549 or H1299 cells and CCK-8 viability
assays were subsequently performed. The results indicate that
Jurkat T cells significantly increased the inhibitory effect of
BUD23 knockdown on A549 and H1299 cell viability (P<0.001;
Fig. 6H and I). By contrast,
Annexin V/PI flow cytometry detected no significant difference in
the apoptosis rates of A549 and H1299 following BUD23 knockdown
(Fig. 6J). Collectively, these
findings indicate that BUD23 silencing suppresses NSCLC cell
proliferation and migration without inducing apoptosis.
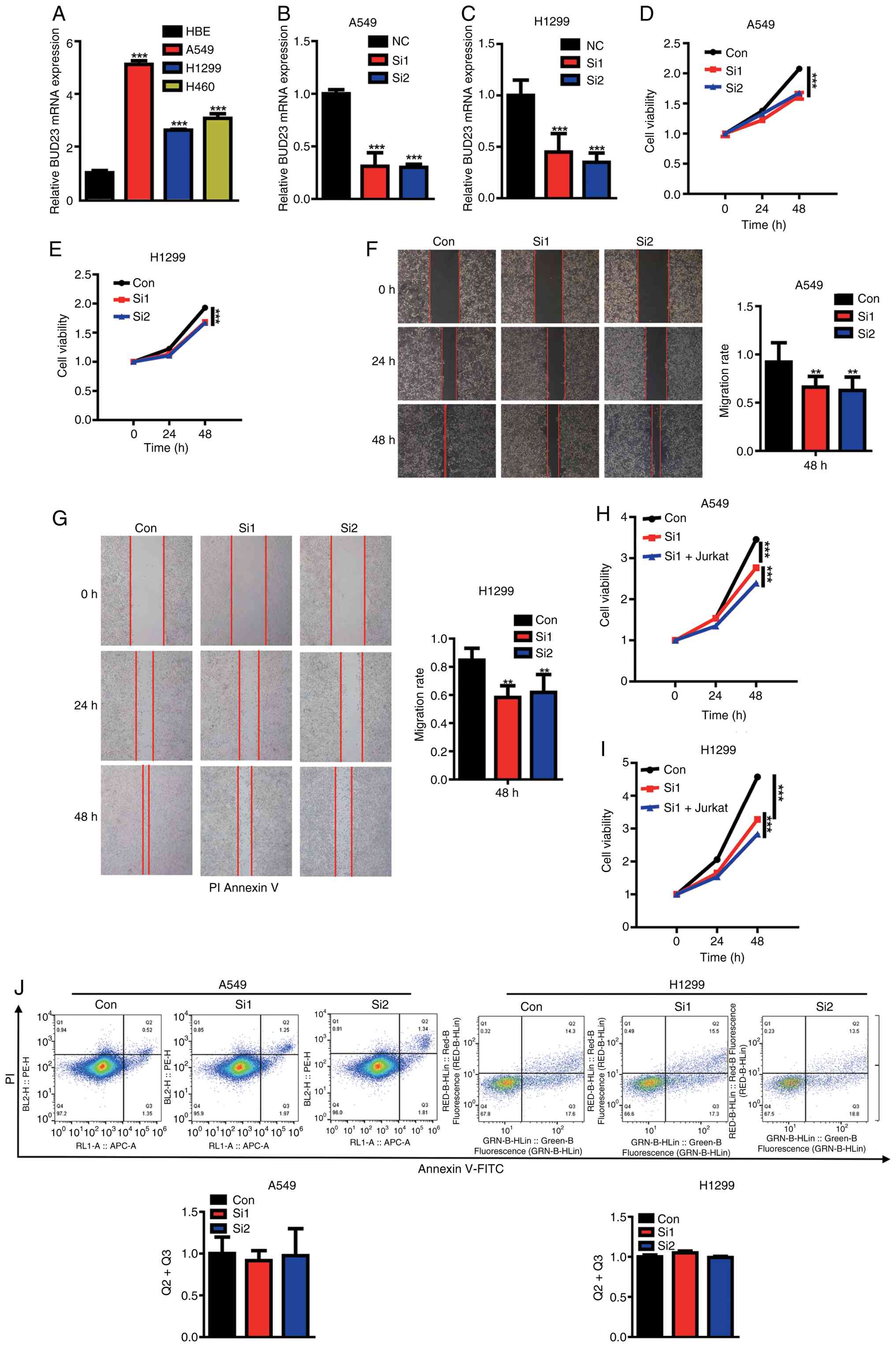 | Figure 6.BUD23 knockdown suppresses the
proliferative and migration of NSCLC cells. (A) RT-qPCR was used to
quantify BUD23 mRNA levels in HBE cells and a panel of NSCLC cell
lines. Validation of BUD23 knockdown efficiency in (B) A549 and (C)
H1299 cells by RT-qPCR. Assessment of cell viability in (D) A549
and (E) H1299 cells via CCK-8 assay and cell migration in (F) A549
and (G) H1299 cells by wound healing assay (magnification, ×10).
CCK-8 assays of (H) A549 and (I) H1299 cells 24 h after BUD23
knockdown with or without subsequent co-culture with Jurkat T cells
for 48 h in Transwell chambers. (J) Quantification of apoptosis in
A549 and H1299 cells via Annexin V/PI flow cytometry. **P<0.01
and ***P<0.001 vs. HBE, NC, Con or as indicated. NSCLC,
non-small cell lung cancer; RT-qPCR, reverse
transcription-quantitative PCR; HBE, human bronchial epithelial;
CCK-8, Cell Counting Kit-8; NC, negative control; Con, control;
Si1/2, small interfering RNA targeting BUD23. |
BUD23 knockdown significantly
downregulates POLR2J expression in NSCLC
To identify the critical downstream targets of
BUD23, a hierarchical intersection strategy was used. GSEA revealed
that BUD23 significantly modulates genes within the Hallmark DNA
repair gene set (Fig. 5A and B).
Intersection of the core-enriched genes from the GSEA of the
TCGA-LUAD and -LUSC cohorts produced 49 candidates (Fig. 7A). The independent overlap of
BUD23-correlated genes from the same two cohorts yielded 76 genes
(Fig. 7B). Finally, the
intersection of the 49 genes derived from the focused GSEA set with
the 79 BUD23-correlated genes yielded four high-confidence targets:
TATA-box binding protein associated factor 6 (TAF6), RNA polymerase
II subunit J (POLR2J), replication factor C subunit 2 (RFC2) and
vacuolar protein sorting-associated protein 37D (VPS37D) (Fig. 7C). RT-qPCR validation confirmed that
POLR2J expression is significantly attenuated following BUD23
knockdown in A549 cells and that only POLR2J expression exhibited
the expected change (Fig. 7D). As
DNA-repair processes are tightly associated with cell-cycle
progression, the impact of BUD23 knockdown on the cell cycle of
NSCLC cells was examined. The results revealed that BUD23 knockdown
decreased the S- and G2-phase fractions in both A549 and H1299
cells (Fig. 7E and F).
Collectively, these data suggest that POLR2J may be a
BUD23-regulated effector involved in DNA repair in NSCLC.
 | Figure 7.BUD23 knockdown significantly
downregulates POLR2J expression in non-small cell lung cancer. (A)
Venn diagram showing the intersection between LUAD GSEA core
enrichment genes (key genes driving Hallmark DNA repair pathway
enrichment) and LUSC GSEA core enrichment genes within the Hallmark
DNA repair gene set, identifying 49 common genes. (B) Venn diagram
showing the intersection between BUD23-correlated genes
(correlation index >0.3) derived from the TCGA-LUAD cohort and
BUD23-correlated genes (correlation index >0.3) derived from the
TCGA-LUSC cohort via multi-gene correlation analysis, yielding 79
overlapping genes. (C) Venn diagram showing the secondary
intersection between the 49 overlapping GSEA core enrichment genes
(from panel A) and the 79 overlapping BUD23-correlated genes (from
panel B), identifying 4 shared common genes: TAF6, POLR2J, RFC2 and
VPS37D. (D) Validation of TAF6, POLR2J, RFC2 and VPS37D mRNA
expression following BUD23 knockdown in A549 cells via reverse
transcription-quantitative PCR. Cell-cycle analysis in (E) A549 and
(F) H1299 cells following BUD23 knockdown. ***P<0.001 vs. Con.
POLR2J, RNA polymerase II subunit J; LUAD, lung adenocarcinoma;
GSEA, gene set enrichment analysis; LUSC, lung squamous cell
carcinoma; TCGA, The Cancer Genome Atlas; TAF6, TATA-box binding
protein associated factor 6; RFC2, replication factor C subunit 2;
VPS37D, vacuolar protein sorting-associated protein 37D; Con,
control; Si1, small interfering RNA targeting BUD23. |
Discussion
Lung cancer is the leading cause of cancer-related
mortality worldwide. Therefore, elucidation of the molecular
mechanisms underlying the development and progression of NSCLC is
important to improve the prognosis of patients with this disease.
In the present study, multi-omics profiling revealed pronounced
BUD23 upregulation in NSCLC, which strongly predicts inferior
patient outcomes. Integrative immunogenomic analysis further
indicated that high BUD23 expression levels are associated with
altered tumor immune infiltration. Mechanistically, GSEA, KEGG and
single-cell pathway enrichment convergently implicated BUD23 in
DNA-repair and cell-cycle networks that drive NSCLC progression.
Functional experiments revealed that BUD23 depletion attenuated the
proliferation and migration of NSCLC cells, and POLR2J was
identified as a direct transcriptional target whose expression
levels were markedly reduced following BUD23 knockdown. These
findings provide a strong basis for future research into the role
of BUD23 in the development and progression of NSCLC.
Previous studies have suggested that BUD23 is an
oncogene, intimately associated with tumorigenesis and tumor
progression, and with upregulated expression in various cancers,
including breast cancer (12),
myeloma (11), colorectal cancer
(40) and hepatocellular carcinoma
(41). Similarly, the differential
analysis performed in the current study revealed upregulated BUD23
expression in NSCLC. Consistent findings were observed across
multiple omics datasets through comparative analyses of tumor and
normal tissues. Therefore, it can be concluded that BUD23 is
upregulated in NSCLC. Further analysis revealed that elevated BUD23
expression is significantly associated with shorter OS, FPS and
PPS, and an advanced clinical stage. These findings indicate that
in NSCLC, patients with elevated BUD23 levels are likely to have
worse clinical outcomes. Subgroup analysis revealed no
statistically significant association between BUD23 expression and
OS in patients with stage N0 NSCLC, although a trend towards
shorter OS was observed in patients with high BUD23 expression. By
contrast, patients with stage N1 or N2 NSCLC and high BUD23
expression exhibited a significantly shorter OS than those with
these stages and low BUD23 expression. This may be attributed to
BUD23 having a stronger association with advanced disease or the
presence of additional confounding factors in patients with stage
N0 NSCLC that could have influenced the results. These findings are
consistent with a previous study, which found that high BUD23
expression predicted a shorter OS in patients with colorectal
cancer (40). Similarly, a study of
glioblastoma demonstrated that elevated BUD23 expression is
associated with a poorer prognosis (10).
Multiple algorithms, including TIMER, EPIC,
MCPcounter and QUANTISEQ, were employed in the present study to
investigate the relationship between BUD23 and immune infiltration
in NSCLC. The analysis consistently revealed a correlation between
BUD23 and immune cell infiltration in NSCLC. Specifically, multiple
algorithms indicated a negative correlation between BUD23 and the
infiltration of B cells and CD8+ T cells. Previous
studies have shown that tumor-infiltrating B cells (TIL-Bs) play an
important role in tumor immunology; specifically, TIL-Bs promote
antitumor immunity by a unique antigen presentation pathway, which
enhances T-cell activation, and TIL-B infiltration is positively
associated with prognosis in multiple types of cancer (42,43).
In the present study, high BUD23 expression was negatively
correlated with B-cell infiltration, and associated with a poorer
prognosis in patients with NSCLC. This suggests that BUD23 might
influence patient prognosis by modulating B-cell immune
infiltration. The in vitro experiments showed that in NSCLC
cells, the knockdown of BUD23 enhanced the cytotoxic activity of
Jurkat T cells. These in vitro data imply a potential
association between BUD23 and tumor immune-related crosstalk.
Similarly, previous research has established that CD8+ T
cells selectively kill tumor cells and play a crucial role in tumor
immunotherapy (44). The negative
correlation observed between BUD23 expression and CD8+
T-cell infiltration suggest that patients with high BUD23
expression may exhibit diminished antitumor immune activity. The
ESTIMATE algorithm is a method for calculating tumor purity. The
present study revealed that BUD23 shows a negative correlation with
stromal and immune scores in NSCLC, hinting that high BUD23
expression may be related to elevated tumor purity and decreased
non-tumor cell infiltration, which could be implicated in the
malignant progression of NSCLC. Collectively, these results imply a
potential association between BUD23 expression and immune
infiltration status within the NSCLC tumor microenvironment, which
may partly account for the unfavorable prognosis in patients with
NSCLC with high BUD23 expression.
The potential regulatory mechanisms by which BUD23
modulates the malignant progression of NSCLC were explored in the
present study using various enrichment analyses. The results
suggest that BUD23 might contribute to NSCLC progression by
regulating DNA repair and cell cycle pathways, the latter being
central to cell proliferation. A previous study has demonstrated
that BUD23 promotes glioblastoma cell proliferation through the
increased phosphorylation of AKT and increased expression of cyclin
D1 and β-catenin (10). Notably,
another study observed no significant change in BUD23 expression
levels in HeLa cells subjected to UV irradiation (45). In the present study, in vitro
data demonstrate that BUD23 contributes to the proliferative and
migratory capacities of NSCLC cells. Knockdown of BUD23 affected
the cell cycle of NSCLC cells but not their apoptosis, suggesting
that BUD23 does not affect apoptosis signaling and mainly affects
pathways associated with cell viability and migration. While the
precise underlying molecular mechanisms remain to be clarified, the
present enrichment analysis, cell cycle assays and preliminary
downstream gene expression data collectively provide partial clues
supporting that BUD23 may regulate cell cycle progression via the
activation of cell cycle checkpoints and alterations in cell cycle
regulatory proteins. Other potential mechanisms, including
metabolic reprogramming and intercellular crosstalk, represent
purely speculative hypotheses without direct experimental support
from the current study and warrant further in-depth
investigation.
POLR2J emerged as a potential downstream target of
BUD23 based on intersection analysis and RT-qPCR validation.
Previous studies have implicated POLR2J as an oncogenic factor in
multiple malignancies, including glioma (46), NSCLC (47), breast cancer (48), reproductive-system tumors (49) and colorectal carcinoma (50), where its upregulation has been
associated with aggressive tumor behavior and adverse clinical
outcomes. Therefore, these findings suggest that the BUD23-mediated
control of POLR2J may be a previously unrecognized signaling axis
influencing NSCLC progression. Nevertheless, further in-depth
mechanistic analyses are urgently required to fully verify and
characterize this novel regulatory axis.
However, several limitations of the present study
should be acknowledged. First, although publicly available datasets
and online analytic platforms enabled efficient data processing,
they may introduce potential biases stemming from heterogeneous
data collection protocols and patient baseline characteristics,
while limiting in-depth mechanistic exploration. However,
consistent results were obtained from multiple online databases and
diverse data processing methods, which supports the reliability of
the conclusions. Second, although analyses using multiple datasets
and methods demonstrated an association of BUD23 expression with OS
and immune infiltration in patients with NSCLC, BUD23 was not
evaluated as a definitive prognostic predictor. Third, the
mechanistic role of BUD23 in modulating immune infiltration, and
its potential relevance to the response to checkpoint inhibitor
therapy remains to be elucidated. However, it is planned to perform
an immunohistochemical analysis of BUD23 expression and relevant
immune markers in clinical lung cancer specimens to examine their
correlations. Additionally, human NSCLC xenograft models coupled
with immunohistochemical staining will be used to explore the
relationship between BUD23 expression and solid tumor infiltration.
Extensive in vitro and in vivo experiments, as well
as additional clinical data, are necessary to validate the findings
of the present study.
In summary, the collective findings of integrative
multi-omics analyses in the present study reveal that BUD23
expression is markedly upregulated in NSCLC and its high expression
portends a poor prognosis. Immunogenomic analyses revealed that
elevated BUD23 expression is associated with alterations in immune
cell infiltration levels. Functional enrichment analyses, including
GSEA, KEGG and single-cell approaches, consistently implicate ‘DNA
repair’ and ‘cell cycle’ pathways as processes associated with
BUD23 expression. In vitro experiments demonstrated that
siRNA-mediated BUD23 knockdown significantly attenuates NSCLC cell
proliferation and migration, and POLR2J is a downstream target, the
expression of which is markedly downregulated upon BUD23 depletion.
These data provide convergent evidence that the present study
indicates that the BUD23-POLR2J axis could act as a relevant
regulatory module in NSCLC progression, highlighting BUD23 as a
candidate prognostic biomarker and a promising therapeutic target
for further investigation.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
YT, JM and JZ performed the data analysis,
experiments and figure preparation. JL conceived the study and
participated in its design. JM and YT wrote the manuscript. YT and
JL confirm the authenticity of all the raw data. All authors read
and approved the final version of the manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Bray F, Laversanne M, Sung H, Ferlay J,
Siegel RL, Soerjomataram I and Jemal A: Global cancer statistics
2022: GLOBOCAN estimates of incidence and mortality worldwide for
36 cancers in 185 countries. CA Cancer J Clin. 74:229–263.
2024.PubMed/NCBI
|
|
2
|
Marx A, Chan JK, Coindre JM, Detterbeck F,
Girard N, Harris NL, Jaffe ES, Kurrer MO, Marom EM, Moreira AL, et
al: The 2015 World Health Organization classification of tumors of
the thymus: Continuity and changes. J Thorac Oncol. 10:1383–1395.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Adams SJ, Stone E, Baldwin DR,
Vliegenthart R, Lee P and Fintelmann FJ: Lung cancer screening.
Lancet. 401:390–408. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Asamura H, Nishimura KK, Giroux DJ,
Chansky K, Hoering A, Rusch V and Rami-Porta R; Members of the
IASLC Staging, Prognostic Factors Committee of the Advisory Boards,
Participating Institutions, : IASLC lung cancer staging project:
The new database to inform revisions in the ninth edition of the
TNM classification of lung cancer. J Thorac Oncol. 18:564–575.
2023. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Thai AA, Solomon BJ, Sequist LV, Gainor JF
and Heist RS: Lung cancer. Lancet. 398:535–554. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Ferrari M and Stagi S: Oxidative stress in
down and williams-beuren syndromes: An overview. Molecules.
26:31392021. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Zorbas C, Nicolas E, Wacheul L, Huvelle E,
Heurgué-Hamard V and Lafontaine DL: The human 18S rRNA base
methyltransferases DIMT1L and WBSCR22-TRMT112 but not rRNA
modification are required for ribosome biogenesis. Mol Biol Cell.
26:2080–2095. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Petrossian TC and Clarke SG: Uncovering
the human methyltransferasome. Mol Cell Proteomics. 10:M110.000976.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Õunap K, Käsper L, Kurg A and Kurg R: The
human WBSCR22 protein is involved in the biogenesis of the 40S
ribosomal subunits in mammalian cells. PLoS One. 8:e756862013.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Chi Y, Liang Z, Guo Y, Chen D, Lu L, Lin
J, Qiu S, Wang X, Qiu E, Lin F, et al: WBSCR22 confers cell
survival and predicts poor prognosis in glioma. Brain Res Bull.
161:1–12. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Tiedemann RE, Zhu YX, Schmidt J, Shi CX,
Sereduk C, Yin H, Mousses S and Stewart AK: Identification of
molecular vulnerabilities in human multiple myeloma cells by RNA
interference lethality screening of the druggable genome. Cancer
Res. 72:757–768. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Nakazawa Y, Arai H and Fujita N: The novel
metastasis promoter Merm1/Wbscr22 enhances tumor cell survival in
the vasculature by suppressing Zac1/p53-dependent apoptosis. Cancer
Res. 71:1146–1155. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Khan AA, Huang H, Zhao Y, Li H, Pan R,
Wang S and Liu X: WBSCR22 and TRMT112 synergistically suppress cell
proliferation, invasion and tumorigenesis in pancreatic cancer via
transcriptional regulation of ISG15. Int J Oncol. 60:242022.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Yan D, Zheng X, Tu L, Jia J, Li Q, Cheng L
and Wang X: Knockdown of Merm1/Wbscr22 attenuates sensitivity of
H460 non-small cell lung cancer cells to SN-38 and 5-FU without
alteration to p53 expression levels. Mol Med Rep. 11:295–302. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Gillette MA, Satpathy S, Cao S,
Dhanasekaran S, Vasaikar S, Krug K, Petralia F, Li Y, Liang WW,
Reva B, et al: A02 Proteogenomic characterization reveals
therapeutic vulnerabilities in lung adenocarcinoma. J Thorac Oncol.
15 (Suppl):S122020. View Article : Google Scholar
|
|
16
|
Satpathy S, Krug K, Jean Beltran PM,
Savage SR, Petralia F, Kumar-Sinha C, Dou Y, Reva B, Kane MH,
Avanessian SC, et al: A proteogenomic portrait of lung squamous
cell carcinoma. Cell. 184:4348–4371.e40. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Rousseaux S, Debernardi A, Jacquiau B,
Vitte AL, Vesin A, Nagy-Mignotte H, Moro-Sibilot D, Brichon PY,
Lantuejoul S, Hainaut P, et al: Ectopic activation of germline and
placental genes identifies aggressive metastasis-prone lung
cancers. Sci Transl Med. 5:186ra662013. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Hou J, Aerts J, den Hamer B, van Ijcken W,
den Bakker M, Riegman P, van der Leest C, van der Spek P, Foekens
JA, Hoogsteden HC, et al: Gene expression-based classification of
non-small cell lung carcinomas and survival prediction. PLoS One.
5:e103122010. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Zhang Y, Foreman O, Wigle DA, Kosari F,
Vasmatzis G, Salisbury JL, van Deursen J and Galardy PJ: USP44
regulates centrosome positioning to prevent aneuploidy and suppress
tumorigenesis. J Clin Invest. 122:4362–4374. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Kim IJ, Quigley D, To MD, Pham P, Lin K,
Jo B, Jen KY, Raz D, Kim J, Mao JH, et al: Rewiring of human lung
cell lineage and mitotic networks in lung adenocarcinomas. Nat
Commun. 4:17012013. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Seo JS, Ju YS, Lee WC, Shin JY, Lee JK,
Bleazard T, Lee J, Jung YJ, Kim JO, Shin JY, et al: The
transcriptional landscape and mutational profile of lung
adenocarcinoma. Genome Res. 22:2109–2119. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Girard L, Rodriguez-Canales J, Behrens C,
Thompson DM, Botros IW, Tang H, Xie Y, Rekhtman N, Travis WD,
Wistuba II, et al: An expression signature as an aid to the
histologic classification of non-small cell lung cancer. Clin
Cancer Res. 22:4880–4889. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Chen CH, Lai JM, Chou TY, Chen CY, Su LJ,
Lee YC, Cheng TS, Hong YR, Chou CK, Whang-Peng J, et al: VEGFA
upregulates FLJ10540 and modulates migration and invasion of lung
cancer via PI3K/AKT pathway. PLoS One. 4:e50522009. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Wei TY, Juan CC, Hisa JY, Su LJ, Lee YC,
Chou HY, Chen JM, Wu YC, Chiu SC, Hsu CP, et al: Protein arginine
methyltransferase 5 is a potential oncoprotein that upregulates G1
cyclins/cyclin-dependent kinases and the phosphoinositide
3-kinase/AKT signaling cascade. Cancer Sci. 103:1640–1650. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Robles AI, Arai E, Mathé EA, Okayama H,
Schetter AJ, Brown D, Petersen D, Bowman ED, Noro R, Welsh JA, et
al: An integrated prognostic classifier for stage I lung
adenocarcinoma based on mRNA, microRNA, and DNA methylation
biomarkers. J Thorac Oncol. 10:1037–1048. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Sun Z, Wang L, Eckloff BW, Deng B, Wang Y,
Wampfler JA, Jang J, Wieben ED, Jen J, You M and Yang P: Conserved
recurrent gene mutations correlate with pathway deregulation and
clinical outcomes of lung adenocarcinoma in never-smokers. BMC Med
Genomics. 7:322014. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Li J, Wang B, Li X and Zhu Y: Estimation
of hub genes and infiltrating immune cells in non-smoking females
with lung adenocarcinoma by integrated bioinformatic analysis. Med
Sci Monitor. 26:e9226802020. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Okayama H, Kohno T, Ishii Y, Shimada Y,
Shiraishi K, Iwakawa R, Furuta K, Tsuta K, Shibata T, Yamamoto S,
et al: Identification of genes upregulated in ALK-positive and
EGFR/KRAS/ALK-negative lung adenocarcinomas. Cancer Res.
72:100–111. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Győrffy B, Surowiak P, Budczies J and
Lánczky A: Online survival analysis software to assess the
prognostic value of biomarkers using transcriptomic data in
non-small-cell lung cancer. PLoS One. 8:e822412013. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Li T, Fan J, Wang B, Traugh N, Chen Q, Liu
JS, Li B and Liu XS: TIMER: A web server for comprehensive analysis
of tumor-infiltrating immune cells. Cancer Res. 77:e108–e110. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Racle J, de Jonge K, Baumgaertner P,
Speiser DE and Gfeller D: Simultaneous enumeration of cancer and
immune cell types from bulk tumor gene expression data. Elife.
6:e264762017. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Becht E, Giraldo NA, Lacroix L, Buttard B,
Elarouci N, Petitprez F, Selves J, Laurent-Puig P, Sautès-Fridman
C, Fridman WH and de Reyniès A: Estimating the population abundance
of tissue-infiltrating immune and stromal cell populations using
gene expression. Genome Biol. 17:2182016. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Finotello F, Mayer C, Plattner C,
Laschober G, Rieder D, Hackl H, Krogsdam A, Loncova Z, Posch W,
Wilflingseder D, et al: Molecular and pharmacological modulators of
the tumor immune contexture revealed by deconvolution of RNA-seq
data. Genome Med. 11:342019. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Yoshihara K, Shahmoradgoli M, Martínez E,
Vegesna R, Kim H, Torres-Garcia W, Treviño V, Shen H, Laird PW,
Levine DA, et al: Inferring tumour purity and stromal and immune
cell admixture from expression data. Nat Commun. 4:26122013.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Charoentong P, Finotello F, Angelova M,
Mayer C, Efremova M, Rieder D, Hackl H and Trajanoski Z: Pan-cancer
immunogenomic analyses reveal genotype-immunophenotype
relationships and predictors of response to checkpoint blockade.
Cell Rep. 18:248–262. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Chandrashekar DS, Karthikeyan SK, Korla
PK, Patel H, Shovon AR, Athar M, Netto GJ, Qin ZS, Kumar S, Manne
U, et al: UALCAN: An update to the integrated cancer data analysis
platform. Neoplasia. 25:18–27. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Yuan H, Yan M, Zhang G, Liu W, Deng C,
Liao G, Xu L, Luo T, Yan H, Long Z, et al: CancerSEA: A cancer
single-cell state atlas. Nucl Acids Res. 47(D1): D900–D908. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Aliazis K, Christofides A, Shah R, Yeo YY,
Jiang S, Charest A and Boussiotis VA: The tumor microenvironment's
role in the response to immune checkpoint blockade. Nat Cancer.
6:924–937. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Yan D, Tu L, Yuan H, Fang J, Cheng L,
Zheng X and Wang X: WBSCR22 confers oxaliplatin resistance in human
colorectal cancer. Sci Rep. 7:154432017. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Stefanska B, Cheishvili D, Suderman M,
Arakelian A, Huang J, Hallett M, Han ZG, Al-Mahtab M, Akbar SM,
Khan WA, et al: Genome-wide study of hypomethylated and induced
genes in patients with liver cancer unravels novel anticancer
targets. Clin Cancer Res. 20:3118–3132. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Petitprez F, de Reyniès A, Keung EZ, Chen
TW, Sun CM, Calderaro J, Jeng YM, Hsiao LP, Lacroix L, Bougoüin A,
et al: B cells are associated with survival and immunotherapy
response in sarcoma. Nature. 577:556–560. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Helmink BA, Reddy SM, Gao J, Zhang S,
Basar R, Thakur R, Yizhak K, Sade-Feldman M, Blando J, Han G, et
al: B cells and tertiary lymphoid structures promote immunotherapy
response. Nature. 577:549–555. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Philip M, Fairchild L, Sun L, Viale A,
Merghoub T, Wolchok JD, Leslie CS and Schietinger A: Chromatin
state dynamics underlying CD8 T cell differentiation and
dysfunction in cancer. Blood. 128:8612016. View Article : Google Scholar
|
|
45
|
Stixová L, Tichý V and Bártová E:
RNA-related DNA damage and repair: The role of N7-methylguanosine
in the cell nucleus exposed to UV light. Heliyon. 10:e255992024.
View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Zheng XL, Li ZD, Luo KZ, Li YL, Liu YH,
Shen SY, Shen FY, Li WY, Chen GQ, Zhang C and Zeng LH: POLR2J
expression promotes glioblastoma malignancy by regulating oxidative
stress and the STAT3 signaling pathway. Am J Cancer Res.
14:2037–2054. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Campbell JM, Lockwood WW, Buys TP, Chari
R, Coe BP, Lam S and Lam WL: Integrative genomic and gene
expression analysis of chromosome 7 identified novel oncogene loci
in non-small cell lung cancer. Genome. 51:1032–1039. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Nourbakhsh M, Saksager A, Tom N, Chen XS,
Colaprico A, Olsen C, Tiberti M and Papaleo E: A workflow to study
mechanistic indicators for driver gene prediction with moonlight.
Brief Bioinform. 24:bbad2742023. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Yao L, Cong R, Ji C, Zhou X, Luan J, Meng
X and Song N: RNA-binding proteins play an important role in the
prognosis of patients with testicular germ cell tumor. Front Genet.
12:6102912021. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Costales-Carrera A, Fernández-Barral A,
Bustamante-Madrid P, Domínguez O, Guerra-Pastrián L, Cantero R, Del
Peso L, Burgos A, Barbáchano A and Muñoz A: Comparative study of
organoids from patient-derived normal and tumor colon and rectal
tissue. Cancers (Basel). 12:23022020. View Article : Google Scholar : PubMed/NCBI
|