Introduction
Acute myeloid leukemia (AML) is a common
hematological malignancy caused by the clonal expansion of
undifferentiated myeloid progenitors (1,2).
Globally, AML accounts for ~1.5% of all new cases of cancer and
remains one of the most lethal hematological malignancies, with a
5-year survival rate of <30%, The resulting hematopoietic
failure represents a primary cause of patient mortality (3). Although chemotherapy, radiotherapy and
hematopoietic stem cell transplantation are the current mainstays
of treatment, the frequent development of multidrug resistance
leads to limited efficacy and poor prognosis (4). Therefore, the identification of novel,
naturally derived active compounds with distinct mechanisms of
action, low toxicity and high efficacy remains a key direction in
AML drug development.
Natural products and their derivatives have been a
vital source of antitumor agents (5,6).
Diterpenoids derived from plants of the genus Isodon
suzhouens have gained increasing attention due to their notable
antitumor activities (7–9). Family members such as glaucocalyxin A
(GLA) and B have been confirmed to induce apoptosis and autophagy
in various tumor models by modulating pathways including PI3K/AKT.
There are preliminary reports on the pro-apoptotic mechanisms of
GLA in AML (10–13). However, the pharmacological
mechanism of action of its structural congener, glaucocalyxin D
(GLD), which demonstrates potent in vitro activity against
chronic myeloid leukemia K562 cells (IC50=1.60 µmol)
(14), remains largely unexplored
in the context of AML. Specifically, whether GLD exerts its effects
through unique targets and signaling pathways is unknown and the
characteristics of the types of cell death and cell cycle arrest
GLD induces require systematic elucidation. This lack of
mechanistic insight severely hinders the evaluation of its clinical
translation potential.
Network pharmacology provides a powerful
methodological framework in elucidating the complex interactions
between compounds and diseases at a systems level by integrating
systems biology and network analysis to establish multi-level
associations among compounds, targets and disease states (15,16).
The present study aimed to systematically investigate the anti-AML
activity of GLD and elucidate its underlying molecular mechanisms
through an integrated approach combining network pharmacology,
molecular docking and in vitro experimental validation.
Materials and methods
Cell lines and reagents
HEL and K562 cell lines were provided by the State
Key Laboratory of Discovery and Utilization of Functional
Components in Traditional Chinese Medicine, Natural Products
Research Center of Guizhou Province. RPMI-1640 culture medium was
purchased from Gibco (cat. no. 11875093; Thermo Fisher Scientific,
Inc.) and fetal bovine serum was supplied by Biological Industries
(cat. no. 04-007-1A; Sartorius AG). Furthermore,
penicillin-streptomycin solution (100X), MTT reagent and
phosphate-buffered saline (PBS) were acquired from Beijing Solarbio
Science & Technology Co., Ltd. (cat. nos. P1400; IM0280;
P1020). The present study also used the Annexin V-FITC/PI apoptosis
detection kit (cat. no. C1383M; Beyotime Biotechnology) and the
cell cycle analysis kit (cat. no. C1052; Beyotime
Biotechnology).
Isolation and characterization of
GLD
GLD was isolated from Isodon suzhouensis aerial
parts. Dried material (1.00×108 mg) was extracted with
95% ethanol and partitioned between water and ethyl acetate to
yield the ethyl acetate fraction (3.60×106 mg). This
fraction was subjected to silica gel column chromatography (300–400
mesh; Qingdao Haiyang Chemical Co., Ltd., Qingdao, China) using
stepwise gradient elution with petroleum ether-acetone (80:1→1:1,
v/v). Fraction 5 was further separated on silica gel with petroleum
ether-acetone (15:1→1:1, v/v). Subfraction Fr.5.2 was purified by
D-101 macroporous resin (Cangzhou Bon Adsorber Technology Co.,
Ltd., Cangzhou, China) with ethanol-water (30→70%, v/v) to yield
GLD (40 mg) (14). Purity (>98%)
was determined by analytical HPLC using a Waters 1525 EF system
(Waters Corporation, Milford, MA, USA) on a YMC C18 column (4.6×250
mm, 5 µm; YMC Co., Ltd., Kyoto, Japan) at 30°C. The mobile phase
consisted of water and methanol (55:45, isocratic; no gradient was
used) at a flow rate of 1.0 ml/min. The injection volume was 10 µl,
and detection was performed at 239 nm using a ZF-1 UV analyzer
(Shanghai Baoshan Guncun Electro-optical Instrument Factory,
Shanghai, China) (Fig. S1). No
internal standard was used. The structure was confirmed by NMR and
MS. NMR spectra were recorded on a Varian INOVA 600 MHz
spectrometer (Varian, Inc., Palo Alto, CA, USA) at 600 MHz for ¹H
and 150 MHz for ¹3C in CDCl3 with
tetramethylsilane (TMS) as the internal standard. HR-MS was
performed on a Waters Autospec Premier P776 mass spectrometer
(Waters Corporation, Milford, MA, USA). The NMR and MS data were
consistent with the literature (Figs.
S2 and S3).
Cell culture
HEL and K562 cells were cultured in RPMI-1640 medium
supplemented with 10% fetal bovine serum and 1%
penicillin-streptomycin at 37°C in a humidified atmosphere with 5%
CO2.
Network pharmacology analysis
Obtaining the intersection genes of GLD and AML
potential molecular targets of GLD were identified by screening the
PharmMapper (http://lilab.ecust.edu.cn/pharmmapper/) and
SwissTargetPrediction (http://www.swisstargetprediction.ch/) databases.
Targets associated with AML were retrieved by querying the
GeneCards (https://www.genecards.org/), Online
Mendelian Inheritance in Man (OMIM) (http://www.omim.org) and Therapeutic Target Database
(TTD) (https://db.idrblab.net/ttd) using the
key word ‘acute myeloid leukemia’. Standardized gene nomenclature
was ensured by referencing the UniProt database (https://www.uniprot.org/). Subsequently, overlapping
genes between GLD targets and AML-associated genes were determined
using Venny 2.1.0 (Spanish National Biotechnology Centre, Madrid,
Spain; http://bioinfogp.cnb.csic.es/tools/venny/).
Construction of protein interaction
networks and core target screening
The identified drug-disease intersection targets
were input into the Search Tool for the Retrieval of Interacting
Genes/Proteins (STRING) database (https://string-db.org/), with the target species set
to Homo sapiens and a minimum interaction score threshold of
0.4. Peripheral-free targets were excluded to generate a network
interaction diagram and a tab-separated values (TSV) file
illustrating target interactions. This TSV file was then imported
into Cytoscape (version 0.1. http://apps.cytoscape.org/apps/cytohubba) to construct
the protein-protein interaction (PPI) network diagram. The
‘cytoHubba’ and ‘molecular complex detection MCODE (version 1.2;
http://apps.cytoscape.org/apps/mcode)
plugins were used for network topology analysis, producing the most
significant core target network diagram and the clustered network
relationship diagram, respectively.
GO and KEGG analyses
The intersecting genes were imported into the
Database for Annotation, Visualization and Integrated Discovery
(DAVID) database (version 6.8; http://david.ncifcrf.gov/). The present study selected
Homo sapiens for Gene Ontology (GO) functional enrichment
analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway
enrichment analysis to identify key signaling pathways. Data
analysis and visualization were conducted using the
MicroBioinformatics online platform (version 1.0; Shanghai
BioGenius Biotechnology Co., Ltd.).
Molecular docking
The top 10 core targets of GLD, ranked by their
binding affinity, were subjected to molecular docking analysis. The
three-dimensional structures of the target proteins were retrieved
from the Protein Data Bank (PDB) database (https://www.rcsb.org/). The target proteins were
processed by removing ligands and water molecules, followed by
hydrogenation using ‘AutoDockTools’ (version 1.5.6; Molecular
Graphics Laboratory; Scripps Research Institute). Both the protein
receptors and ligands were then converted into PDB, partial charge
and atom type format. Molecular docking was performed using
AutoDock Vina (version 1.2.0; Molecular Graphics Laboratory;
Scripps Research Institute) software and the model with the lowest
free energy was selected for analysis. The results were visualized
using PyMOL (version 2.5; Schrödinger, Inc.) software. Each
interaction was assigned a docking score, with lower scores
indicating improved docking quality.
Cell viability assay
The MTT method was used to evaluate cell viability.
Briefly, cells were seeded into 96-well plates at a density of
5×103 cells per well. After 24 h, the cells were treated
with different concentrations of GLD (0.625, 1.25, 2.5, 5 and 10
µmol/l) dissolved in DMSO (0.1% v/v) for 24, 48 and 72 h, and
doxorubicin was used as a positive control. Following treatment,
MTT reagent was added for 4 h, the formazan crystals were dissolved
in DMSO (≥99.9%), and absorbance was measured at 490 nm using an
ELx808 microplate reader (Agilent Technologies, Inc) Cell viability
was calculated relative to untreated controls.
Detection of apoptosis by Hoechst
33258 staining
To detect cell apoptosis, the cells were seeded into
6-well plates at a density of 3×105 cells per well.
After treatment with GLD for 48 h, cells were collected, washed
with PBS and fixed with 4% paraformaldehyde at room temperature for
15 min. Following fixation, cells were washed twice with PBS and
stained with Hoechst 33258 (5 µg/ml, 0.5% v/v) for 10 min at room
temperature in the dark. After washing to remove excess stain,
cells were resuspended in a small volume of PBS. A drop of the cell
suspension was mounted with anti-fade mounting medium, covered with
a coverslip and immediately observed under a fluorescence
microscope. Apoptotic cells were identified by the presence of
condensed and fragmented nuclei.
Apoptosis detection and cell cycle
analysis
For the detection of apoptosis and cell cycle
analysis, cells were seeded in 6-well plates at a density of
3×105 cells per well. Using DMSO (0.1% v/v) as the
control group, cells were treated with different concentrations of
GLD for 48 h. Subsequently, cells were collected and stained using
the Annexin V-FITC/PI apoptosis detection kit (cat. no. C1062M;
Beyotime Biotechnology) and the cell cycle analysis kit (cat. no.
C1052M; Beyotime Biotechnology) for flow cytometry (BD
Biosciences). Staining was performed at room temperature for 15 min
in the dark, according to the manufacturer's instructions. No
intracellular cytokine staining was performed. The collected data
were analyzed using FlowJo software (version 10.10; BD
Biosciences). For apoptosis analysis, cells were gated on forward
scatter/side scatter to exclude debris and the percentage of
Annexin V-FITC-positive and PI-positive cells was determined. For
cell cycle analysis, single cells were gated, and the proportions
of cells in G1, S and G2M phases were
calculated using the Dean-Jett-Fox model (17).
Western blotting
After treatment with GLD for 48 h, cells were
harvested and washed with PBS. Total proteins were extracted using
radioimmunoprecipitation assay lysis buffer (cat. no. P0013B;
Beyotime Biotechnology; RIPA:PMSF=100:1)) supplemented with a
protease inhibitor cocktail (cat. no. P1005; Beyotime
Biotechnology). Cell lysis was performed on ice, followed by brief
sonication (20 kHz; 4°C; 10 sec on, 10 sec off for 2 cycles). The
lysates were centrifuged at 12,000 × g for 15 min at 4°C to collect
the supernatant. Protein concentration was determined using the BCA
Protein Assay Kit (cat. no. P0012; Beyotime Biotechnology)
according to the manufacturer's instructions. Equal amounts of
protein (30 µg per lane) were mixed with 5X loading buffer,
denatured at 95°C for 10 min and separated by 10% sodium dodecyl
sulfate-polyacrylamide gel electrophoresis. The separated proteins
were then transferred onto polyvinylidene fluoride membranes. The
membranes were blocked with 5% bovine serum albumin in
Tris-buffered saline with 0.1% Tween 20 for 2 h at room temperature
and subsequently incubated overnight at 4°C with the following
primary antibodies (all diluted 1:1,000): Anti-phosphorylated
(p)-JAK2 (Tyr1007/1008; cat. no. 3771; Cell Signaling Technology,
Inc.), anti-JAK2 (cat. no. 17670-1-AP; Proteintech Group Inc.),
anti-p-STAT3 (Tyr705; cat. no. 9145; Cell Signaling Technology,
Inc.), anti-STAT3 (cat. no. 10253-2-AP; Proteintech Group Inc.),
anti-Bcl-2 (cat. no. 12789-1-AP; Proteintech Group Inc.), anti-Bax
(cat. no. 50599-2-Ig; Proteintech Group Inc.), anti-cyclin B1 (cat.
no. 55004-1-AP; Proteintech Group Inc.), anti-p-CDK1 (Tyr15; cat.
no. 4539; Cell Signaling Technology, Inc.) and anti-β-actin
(anti-β-actin (cat. no. 66009-1-Ig; Proteintech Group Inc.) and
anti-GAPDH (cat. no. 60004-1-Ig; Proteintech Group Inc.). After
washing, the membranes were incubated with horseradish
peroxidase-conjugated goat anti-rabbit IgG secondary antibody
(1:50,000; cat. no. SA00001-2; Proteintech Group Inc.) for 2 h at
room temperature. Protein bands were visualized using an enhanced
chemiluminescence (ECL) detection kit (cat. no. P0018FS; Beyotime
Biotechnology) and the band intensities were semi-quantified using
ImageJ software (version 1.54; National Institutes of Health). The
expression levels of target proteins were normalized to β-actin or
GAPDH.
Statistical analysis
Statistical analyses were performed using GraphPad
Prism software (version 9.5.1; Dotmatics). Data from at least three
independent experiments are presented as mean ± standard deviation.
Comparisons among multiple groups were conducted using one-way
analysis of variance, followed by Tukey's honest significant
difference post hoc test for multiple comparisons. P<0.05 was
considered to indicate a statistically significant difference.
Results
Network pharmacology
Active components and potential targets of GLD in
AML were investigated in the present study. In total, 74 potential
targets associated with GLD were retrieved from the PharmMapper and
SwissTarget prediction databases. Concurrently, 5,576 targets were
identified using the term AML in the OMIM, GeneCards and TTD
databases (Fig. 1A). The Venny
2.1.0 online tool (https://bioinfogp.cnb.csic.es/tools/venny/) was used
to identify the 55 overlapping targets between the 74
GLD-associated targets and the 5,576 AML-associated targets
(Fig. 1B).
 | Figure 1.Potential AML-associated targets of
GLD. (A) Venn diagram of AML-associated target interactions. The
numbers of targets retrieved from the OMIM, GeneCards and TTD
databases are shown: OMIM, 92; GeneCards, 5,446; TTD, 16. (B) Venn
diagram of GLD and AML target intersections. A total of 74
GLD-associated targets and 5,576 AML-associated targets were
analyzed, yielding 55 overlapping targets. (C) Construction of the
GLD and AML gene-protein interaction network. The network was
generated using the STRING database and visualized in Cytoscape.
(D) Screening of core targets for GLD treatment of AML using the
cytoHubba plugin. (E) Screening of core targets using the MCODE
plugin. (F) Combined core target network diagram. AML, acute
myeloid leukemia; GLD, glaucocalyxin D; OMIM, Online Mendelian
Inheritance in Man; TTD, Therapeutic Target Database. |
Construction of the PPI interaction
network and identification of core targets
The 55 intersecting genes were input into the STRING
database and a PPI network was constructed using the Cytoscape
software (Fig. 1C). Subsequently,
the core regions of the network were analyzed using ‘cytoHubba’ and
‘MCODE’ plugins, ultimately yielding three core topological
networks (Fig. 1D-F). These
networks encompassed 10 key targets: STAT3, nuclear factor-κ B
subunit 1 (NFKB1), heat shock protein 90 kDa α, class B, member 1
(HSP90AB1), hypoxia-inducible factor-1α (HIF1A), mTOR, STAT1,
MAPK1, toll-like receptor 4 (TLR4), growth factor receptor-bound
protein 2 (GRB2) and protein tyrosine phosphatase non-receptor type
11 (PTPN11).
GO and KEGG enrichment analyses reveal
the potential multi-target mechanism of GLD in AML
To elucidate the biological functions and pathways
associated with the 55 overlapping targets, GO and KEGG enrichment
analyses were performed. The GO analysis indicated that these
targets are significantly involved in key oncogenic processes,
including the ‘positive regulation of phosphatidylinositol
3-kinase/protein kinase B signal transduction’, ‘positive
regulation of cell migration’ and ‘positive regulation of
angiogenesis’ (biological process). They are primarily located in
the ‘nucleoplasm’ and ‘cytoplasm’ (cellular component) and possess
molecular functions such as ‘protein kinase binding’ and ‘histone
H3Y41 kinase activity’ (molecular function) (Fig. 2A). These enriched terms collectively
suggested that GLD may interfere with AML progression by broadly
impacting signal transduction, cell motility and epigenetic
regulation. KEGG pathway enrichment analysis of the predicted
targets revealed 20 key signaling pathways potentially modulated by
GLD (Table I). Pathways closely
associated with cancer and immunoinflammatory responses, such as
‘proteoglycans in cancer’, ‘JAK-STAT signaling pathway’ and
‘chemical carcinogenesis-receptor activation’ were among the most
significantly enriched. Notably, 9.5 targets were specifically
enriched in the ‘JAK-STAT signaling pathway’, strongly suggesting
its central role in the anti-AML activity of GLD (Fig. 2B). Subsequent molecular docking
analysis demonstrated that GLD could form stable binding
conformations with core proteins JAK2/STAT3 pathway, including JAK2
and STAT3, providing direct computational evidence for further
experimental validation.
 | Table I.Annotation analysis of the top 20
KEGG signaling pathways. |
Table I.
Annotation analysis of the top 20
KEGG signaling pathways.
| Pathway | P-value | Count | Targets |
|---|
| Chemical
carcinogenesis-receptor activation |
4.08×10−9 | 13 | STAT1, STAT3, TLR4
and NFKB1 |
| Human
cytomegalovirus infection |
7.20×10−9 | 13 | HSP90AB1, STAT3,
PRKCA, PIK3R1, CYP3A4, mTOR, NFKB1, AR, KLF5, Bcl-6, RPS6KA1, MAPK1
and GRB2 |
| Chemokine signaling
pathway |
1.92×10−7 | 11 | ZAP70, PTK2B,
MAPK1, PTPN11, GRB2, PRKCA and PIK3R1 |
| Proteoglycans in
cancer |
3.22×10−7 | 11 | CHUK, SERPINE1,
MAPK1, PIK3R1, TLR4 and NFKB1 |
| JAK-STAT signaling
pathway |
6.87×10−6 | 9 | MAPK1, GRB2, PRKCA,
PIK3R1 and mTOR |
| Measles |
1.89×10−5 | 8 | MAPK1, GRB2, PRKCA
and PIK3R1 |
| Osteoclast
differentiation |
2.28×10−5 | 8 | CHUK, CCNE1, STAT1,
CDK4, STAT3, PIK3R1, TLR4 and NFKB1 |
| Natural killer
cell-mediated cytotoxicity |
7.10×10−5 | 7 | CCR1, PDGFRA, CHUK,
ITGB3, STAT3, PRKCA, PIK3R1, mTOR, NFKB1, CDK4, PTK2B, MAPK1 and
GRB2 |
| Growth hormone
synthesis, secretion and action |
8.97×10−5 | 7 | STAT1, STAT3,
MAPK1, GRB2, PRKCA, PIK3R1 and mTOR |
| Chagas disease |
3.84×10−4 | 6 | CCR1, ITK, CHUK,
STAT1, STAT3, PRKCD, PTK2B, MAPK1, GRB2, PIK3R1 and NFKB1 |
| Toll-like receptor
signaling pathway |
4.99×10−4 | 6 | CHUK, STAT1, ITGB3,
MAPK1, GRB2, PIK3R1, MAP3K14 and NFKB1 |
|
Progesterone-mediated oocyte
maturation |
5.43×10−4 | 6 | CHUK, STAT1, MAPK1,
PIK3R1, TLR4, NFKB1 |
| Leukocyte
transendothelial migration |
6.64×10−4 | 6 | CHUK, MAPK1, GRB2,
PIK3R1 and NFKB1 |
| ErbB signaling
pathway |
1.80×10−3 | 5 | HSP90AB1, RPS6KA1,
CDK1, MAPK1, PIK3R1 and CDC25C |
| B cell receptor
signaling pathway |
2.22×10−3 | 5 | ITGB1, ITGB3,
STAT3, MAPK1, PTPN11, GRB2, PRKCA, PIK3R1, HIF1A, TLR4 and
mTOR |
| Th1 and Th2 cell
differentiation |
2.40×10−3 | 5 | ZAP70, CHUK, STAT1,
MAPK1 and NFKB1 |
| GnRH signaling
pathway |
2.40×10−3 | 5 | PRKCD, PTK2B,
MAPK1, GRB2 and PRKCA |
| Inflammatory bowel
disease |
7.45×10−3 | 4 | PDGFRB, PDGFRA,
STAT1, STAT3, PTPN11, GRB2, PIK3R1, PTPN2 and mTOR |
| FcεRI signaling
pathway |
8.43×10−3 | 4 | CCNE1, CDK4,
SERPINE1 and CDK1 |
| p53 signaling
pathway |
1.06×10−2 | 4 | ITGB1, ITK, PTK2B,
PTPN11, PRKCA and PIK3R1 |
Molecular docking validates the direct
binding of GLD to key signaling targets
To structurally validate the predictions from
network pharmacology, molecular docking was performed to assess the
binding of GLD to core targets. The results demonstrated strong
binding affinities (ΔG <-5.0 kcal/mol) between GLD and several
pivotal targets (Table II). Of
note, GLD exhibited high affinity for STAT3 (ΔG=−6.3 kcal/mol), the
central transcription factor within the significantly enriched
‘JAK-STAT signaling pathway’, corroborating the KEGG enrichment
analysis. Docking poses suggested that GLD binds to the Src
homology 2 domain of STAT3, a key site for phosphotyrosine binding
and dimerization, implying a potential mechanism for direct
interference with its activation. Notable binding was also observed
for other relevant targets, including MAPK1 (−6.8 kcal/mol) and
STAT1 (−6.6 kcal/mol). Visualization of the optimal binding
conformations (Fig. 3A-J) revealed
stabilizing interactions mediated by hydrogen bonds and hydrophobic
forces. These findings provided notable computational structural
evidence that GLD can directly engage with key effectors like STAT3
and potentially modulate the associated signaling networks.
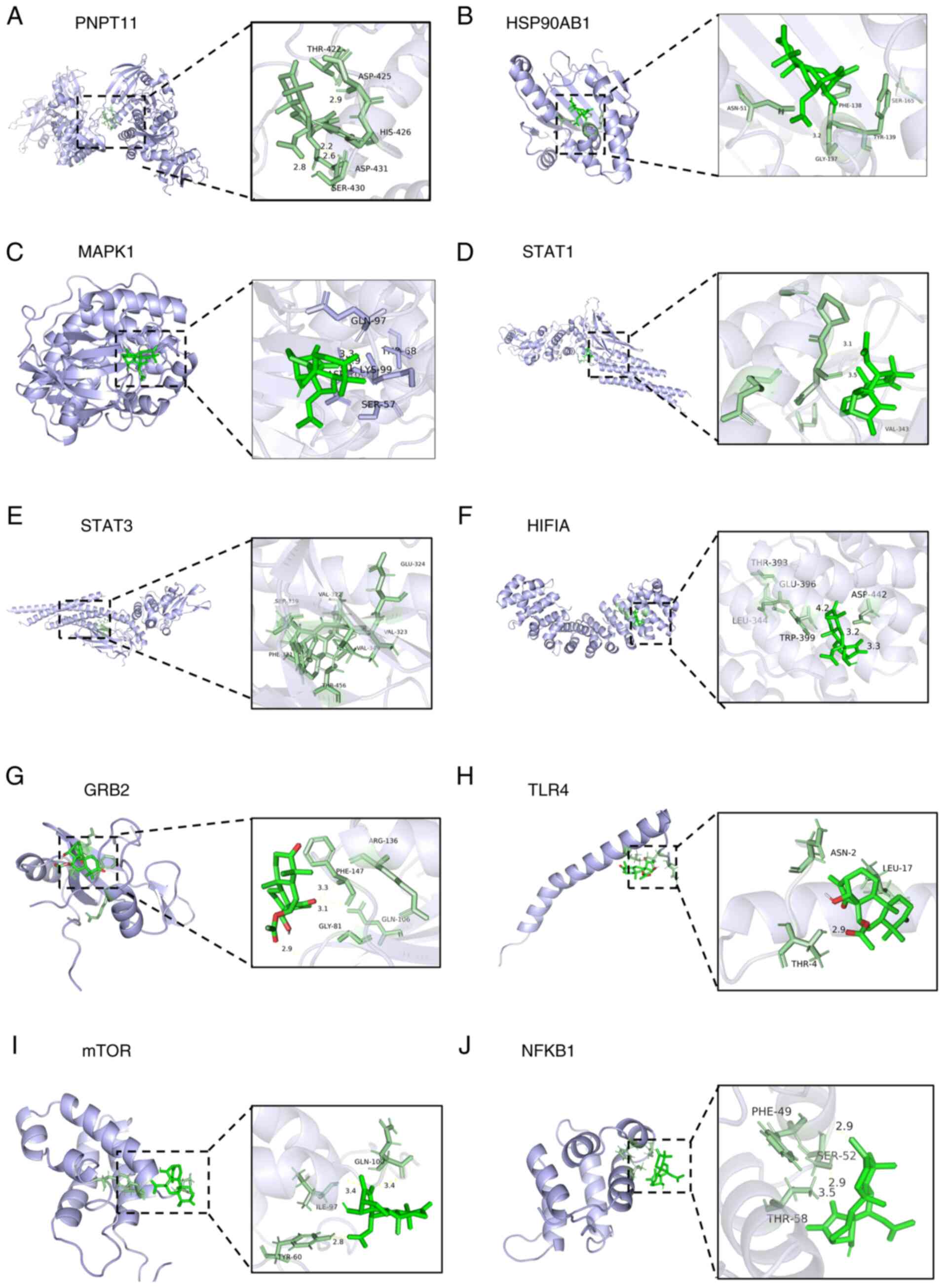 | Figure 3.Docking results of glaucocalyxin D
with 10 core target molecules. (A) PNPT11, (B) HSP90AB1, (C) MAPK1,
(D) STAT1, (E) STAT3, (F) HIF1A, (G) GRB2, (H) TLR4, (I) mTOR, (J)
NFKB1. Binding energies (kcal/mol) and detailed interactions
(hydrogen bonds, hydrophobic interactions, salt bridges,
π-stacking) for each target are presented in Table II. In the docking analysis, ‘-’
indicates that no corresponding interaction was detected. |
 | Table II.Binding energy in molecular docking
between glaucocalyxin D and core targets. |
Table II.
Binding energy in molecular docking
between glaucocalyxin D and core targets.
| Target symbol | PDB ID | Binding energy,
kcal/mol | Hydrogen bonds | Hydrophobic
interactions | Salt bridges | π-stacking |
|---|
| PTPN11 | Q06124 | −7.7 | LYS-2.22 | THR-3.17 | LYS-4.49 and
LYS-5.40 | LYS-4.49 and
LYS-5.40 |
| HSP90AB1 | P08238 | −7.1 | ILE-2.06 and
ILE-2.60 | PHE-3.99, ILE-3.67
and GLN-3.03 | LYS-5.26 | LYS-5.26 |
| MAPK1 | P28482 | −6.8 | ALA-2.65, GLU-2.95
and GLU-2.35 | ARG-3.98, ALA-3.66
and ALA-3.88 | - | - |
| STAT1 | P42224 | −6.6 | TYR-3.26, LYS-2.81
and ARG-2.46 | - | LYS-3.74 and
ARG-4.83 | - |
| STAT3 | P40763 | −6.3 | ARG-2.89 | GLN-3.75, PRO-3.95
and ARG-3.82 | - | - |
| HIF1A | Q309Z6 | −5.8 | - | - | - | - |
| GRB2 | P62993 | −5.6 | - | - | - | - |
| TLR4 | O00206 | −5.3 | - | - | - | - |
| mTOR | P42345 | −4.9 | - | - | - | - |
| NFKB1 | P19838 | −4.5 | - | - | - | - |
GLD inhibits the proliferation of AML
cells
To evaluate the anti-proliferative effect of GLD
(chemical structure presented in Fig.
4A) on AML cell viability was assessed using the MTT assay. GLD
treatment markedly inhibited the growth of HEL and K562 cells in a
concentration- and time-dependent manner (Fig. 4C and D). The half-maximal inhibitory
concentration (IC50) for HEL cells was determined to be
3.039±0.692 µmol/l at 24 h, 0.808±0.166 µmol/l at 48 h and
0.385±0.011 µmol/l at 72 h. For K562 cells, the IC50
values were 5.165±0.768 µmol/l, 1.479±0.049 µmol/l and 0.388±0.010
µmol/l at 24, 48 and 72 h, respectively (Fig. 4B). The progressive leftward shift of
the dose-response curves with prolonged exposure (Fig. 4C and D) visually represents a strong
time-dependent effect, which is quantitatively corroborated by the
marked decrease in IC50 values over time (Fig. 4B). Notably, HEL cells exhibited
consistently lower IC50 values and left-shifted
inhibition curves compared with K562 cells at both 24 and 48 h,
indicating greater sensitivity to GLD. This differential
sensitivity aligns with the distinct molecular background of the
two cell lines; HEL cells harbor the activating JAK2 V617F
mutation, which confers constitutive JAK-STAT pathway activation
and may render them more susceptible to agents targeting this
signaling axis. Furthermore, the IC50 of GLD against
K562 cells at 48 h is similar to that reported for the classic
chemotherapeutic agent cytarabine (~1.95 µmol/l in the same cell
line) (18), indicating potent
anti-proliferative activity of GLD in vitro. Morphological
assessment of nuclear changes was performed using Hoechst 33258
staining. Compared with control cells, GLD-treated cells exhibited
intensely stained, condensed nuclei, a morphological hallmark often
associated with DNA damage and apoptosis (Fig. 4E).
GLD induces apoptosis in AML cell
lines
To further investigate the effects of GLD on the
survival of AML cell lines, flow cytometry was performed in the
present study. AML cell lines were treated with GLD at
concentrations of 0.25, 0.5 and 1 µmol/l for 48 h, followed by
Annexin V-FITC/PI staining and flow cytometry analysis (Fig. 5A). Compared with the control group,
the apoptosis rates in HEL and K562 cells treated with 0.25, 0.5
and 1 µmol/l GLD exhibited a progressive significant increase in
association with the increasing concentrations of GLD (Fig. 5B and C).
GLD blocks AML cells in the
G2/M phase
To further investigate the effects of GLD on the AML
cell cycle, the present study observed a progressive significant
increase in the proportion of cells in the G2/M phase
with increasing concentrations of GLD. This finding indicated that
GLD induces cell cycle arrest specifically at the G2/M
phase (Fig. 5D-F).
GLD exerts its anti-AML effects by
targeting the JAK-STAT signaling pathway, leading to the induction
of apoptosis and G2/M phase arrest
To experimentally validate the mechanism predicted
by computational analysis, the present study examined the effect of
GLD on the JAK-STAT axis and its functional consequences. Western
blotting analysis revealed that GLD treatment significantly
inhibited the phosphorylation of JAK2 and its key downstream
effector, STAT3, in AML cells in a dose-dependent manner,
demonstrating an effective suppression of this pathway at the
upstream level. This suppression elicited coordinated pro-apoptotic
and cell cycle-disruptive responses. Specifically, GLD
significantly downregulated the anti-apoptotic protein Bcl-2 and
significantly upregulated the pro-apoptotic protein Bax, thereby
shifting the cellular balance toward apoptosis. Concurrently, GLD
significantly reduced the protein levels of cyclin B1 and its
catalytic partner CDK1, the key driver complex for the
G2/M phase transition, which explains the observed cell
cycle arrest (Fig. 6A-D).
Collectively, these results outlined a coherent mechanistic
pathway: GLD inhibits the JAK2-STAT3 signaling axis, which in turn
orchestrates mitochondrial apoptosis and G2/M phase cell
cycle blockade in AML cells.
 | Figure 6.GLD exerts its anti-AML effects by
targeting the JAK-STAT signaling pathway. (A) HEL cells were
treated with the indicated concentrations of GLD (0, 0.25, 0.5 and
1 µmol/l) for 48 h. The protein expression levels of Bax, Bcl-2,
CDK1, cyclin B1, JAK2, p-JAK2, STAT3 and p-STAT3 were detected by
western blotting. β-actin and GAPDH were used as loading controls.
(B) K562 cells were treated with GLD (0, 0.25, 0.5 and 1 µmol/l)
for 48 h and the protein expression levels were detected as
described in (A). (C) Quantification of the protein bands presented
in (A) and (B). The relative protein expression levels were
normalized to the loading controls, and p-JAK2 and p-STAT3 were
further normalized to total JAK2 and STAT3, respectively. Data are
presented as the mean ± SD (n=3). *P<0.05, **P<0.01 and
***P<0.001 vs. the control group. p, phosphorylated; AML, acute
myeloid leukemia; GLD, glaucocalyxin D. |
Discussion
AML is an aggressive hematological malignancy
associated with a relapse rate of ~50% ≤1 year of achieving
complete remission and therapeutic resistance, highlighting an
urgent need for novel agents with distinct mechanisms of action
(19). Natural products serve as a
valuable source of such candidates due to their structural
diversity and multi-target potential (20,21).
Although diterpenoids such as GLA and GLB have demonstrated
anticancer activities, the specific mechanism by which GLD exerts
its effects against AML remains insufficiently explored,
representing a key knowledge gap. The present study is the first to
employ an integrated strategy combining network pharmacology,
molecular docking and experimental validation to systematically
elucidate the anti-AML mechanism of GLD, to the best of our
knowledge. The core finding of the present study revealed that GLD
exerts potent anti-leukemic effects primarily by inhibiting the
JAK-STAT signaling pathway, thereby inducing mitochondrial
apoptosis and G2/M phase cell cycle arrest.
Network pharmacology analysis identified 55
potential overlapping targets between GLD and AML. Construction of
a PPI network highlighted core targets including STAT3, STAT1,
NFKB1, HSP90AB1, HIF1A and mTOR, suggesting that GLD may act
through a multi-target, multi-pathway network. To functionally
annotate these targets, GO and KEGG enrichment analyses were
performed. GO analysis indicated extensive involvement in
fundamental oncogenic processes, whereas KEGG analysis identified
the ‘JAK-STAT signaling pathway’ as one of the most significantly
enriched pathways. Notably, key target genes within this pathway,
such as JAK2, STAT3 and Bcl-2, were specifically identified,
providing precise hypotheses for subsequent molecular validation.
Molecular docking further supported the structural feasibility of
GLD binding to the core target STAT3. Due to the well-documented,
pivotal role of the JAK-STAT signaling axis in AML pathogenesis
(22–25), this pathway was prioritized for
definitive experimental validation in the present study.
Based on these predictions, in vitro
experiments confirmed that GLD effectively inhibits AML cell
proliferation. Using HEL and K562 cells for mechanistic
exploration, MTT assays demonstrated that GLD significantly and
dose-dependently suppressed proliferation in a time- and
concentration-dependent manner. Notably, the anti-proliferative
activity of GLD was similar to that of the clinical
chemotherapeutic agent doxorubicin, employed as a positive control.
This finding suggested the potential of GLD as a lead compound with
efficacy rivaling current standard therapy. Furthermore, the
multi-target profile inferred from network pharmacology may
potentially offer a theoretical advantage in mitigating the drug
resistance frequently encountered with single-target agents.
To validate the mechanistic basis of anti-AML action
by GLD, the present study focused on key proteins within the
JAK-STAT signaling pathway and cell-cycle regulators. The JAK-STAT
pathway participates in diverse biological processes, including
cell proliferation, differentiation, apoptosis and immune
regulation (26,27). Dysregulation of this pathway is
implicated in various cancer types, including hematological
malignancies (for example, acute myeloid leukemia or
myeloproliferative neoplasms) and solid tumors (for example, breast
cancer or lung cancer), as well as autoimmune disorders such as
rheumatoid arthritis, systemic lupus erythematosus and inflammatory
bowel disease (24,25,28–31).
Previous research indicated that constitutive STAT3 activation can
suppress erythroid differentiation via upregulation of PU.1,
thereby exacerbating erythroleukemia progression (32). By contrast, inhibitors of JAK2 or
STAT3 phosphorylation can ameliorate anemia by restoring erythroid
cell development (33). In the
present study, western blotting analysis confirmed that GLD
inhibited the phosphorylation of both JAK2 and STAT3 in a
dose-dependent manner, as demonstrated by the reduced levels of
phosphorylated p-JAK2 and p-STAT3 Apoptosis and cell-cycle arrest
are two common indicators in antitumor research (34,35).
Regarding apoptosis regulation, GLD significantly upregulated the
pro-apoptotic protein Bax while downregulating the anti-apoptotic
protein Bcl-2, representing a key molecular switch for initiating
the mitochondrial-dependent intrinsic apoptotic pathway and
mechanistically explaining GLD-induced apoptosis. In cell-cycle
regulation, western blotting analysis demonstrated that GLD
markedly downregulated the protein levels of cyclin B1 and its
catalytic partner CDK1, core regulators of the G2/M
checkpoint. This suppression directly led to G2/M phase
arrest, which was fully consistent with flow cytometry results.
The present study findings underscored the
therapeutic potential of GLD as a novel natural inhibitor of the
JAK-STAT pathway, a key driver in AML. The implied multi-targeted
potential may confer an advantage in modulating complex disease
networks. However, certain limitations of the present study should
be acknowledged. The validation was conducted primarily in
established cell lines. Confirmation of the efficacy of GLD in
primary patient-derived AML cells and relevant in vivo
models is key to evaluating its translational potential and safety
profile. Furthermore, to address potential pharmacokinetic
challenges and enhance its therapeutic index, future research could
explore advanced drug-delivery strategies. As highlighted by recent
advances in nanocarrier technology, exemplified by
antibody-functionalized lipid nanocarriers for targeted cancer
therapy, encapsulating bioactive compounds such as GLD into
optimized nano-formulations could markedly improve its
bioavailability, targeting specificity and reduce off-target
toxicity, thereby facilitating clinical translation (36,37).
In conclusion, to the best of our knowledge, the
present integrated study provides the first systematic evidence
that GLD inhibits AML cell proliferation by suppressing the
JAK-STAT pathway, which orchestrates downstream apoptotic and
cell-cycle-disruptive events. These findings not only elucidate a
coherent molecular mechanism for GLD but also potentially
establishes GLD as a promising lead compound targeting the JAK-STAT
axis for AML therapy in the future.
In summary, the present study systematically
elucidated the mechanism by which GLD exerts anti-AML effects in
vitro. GLD acts by inhibiting the JAK-STAT signaling pathway
and coordinately regulating key downstream executioners of
apoptosis and the cell cycle (Bax/Bcl-2 and cyclin B1/CDK1). These
findings identify GLD as a multi-targeted natural compound that
inhibits the JAK-STAT signaling pathway, supporting its potential
as a lead molecule for developing therapeutic strategies against
AML subtypes with aberrant JAK-STAT signaling. Future research
should aim to overcome current limitations using in vivo
validation and formulation optimization to advance its translation
into preclinical development.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
The present study was supported by grants from the National
Natural Science Foundation of China (grant no. 82460687), the
Natural Science Foundation of China (grant no. 82460838), the
Department of Science and Technology of Guizhou Province [grant no.
QKZC(2024)YB068], the Science and Technology Foundation Project of
Guizhou Provincial Health Commission (grant no. GZWJK2024-241), the
Organization Department of Guizhou Provincial Committee [grant no.
QKHPTRC-GCC(2022)034-2] and the Anshun City Science and Technology
Plan Project [grant no. ASKSS(2024)01].
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
LC wrote the main manuscript and completed the main
experiments under the supervision of CY and HL. LC and CY conceived
and designed the study. HL contributed to the interpretation of
data and critically revised the manuscript. WY isolated the
compound and performed the structural characterization; WZ assisted
in compound isolation. LC and CY confirm the authenticity of all
the raw data. All authors reviewed and approved the final
manuscript and agree to be accountable for all aspects of the
work.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Schratz KE and Armanios M: Cancer and
myeloid clonal evolution in the short telomere syndromes. Curr Opin
Genet Dev. 60:112–118. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Cai SF and Levine RL: Genetic and
epigenetic determinants of AML pathogenesis. Semin Hematol.
56:84–89. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Weinberg OK, Sohani AR, Bhargava P and
Nardi V: Diagnostic work-up of acute myeloid leukemia. Am J
Hematol. 92:317–321. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Long L, Assaraf YG, Lei ZN, Peng H, Yang
L, Chen ZS and Ren S: Genetic biomarkers of drug resistance: A
compass of prognosis and targeted therapy in acute myeloid
leukemia. Drug Resist Updat. 52:1007032020. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Yang GX, Ma GL, Li H, Huang T, Xiong J and
Hu JF: Advanced natural products chemistry research in China
between 2015 and 2017. Chin J Nat Med. 16:881–906. 2018.PubMed/NCBI
|
|
6
|
Mansoori B, Mohammadi A, Amin Doustvandi
M, Mohammadnejad F, Kamari F, Gjerstorff MF, Baradaran B and
Hamblin MR: Photodynamic therapy for cancer: Role of natural
products. Photodiagnosis Photodyn Ther. 26:395–404. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Gao C, Yu S and Liang Q: New research
progress on Rabdosia japonica var. glaucocalyx. J Mudanjiang Med
Univ. 32:54–56. 2011.(In Chinese).
|
|
8
|
Su Y, Cui J, Shi W, Wang X and Zhou Y:
Research progress on Chinese herbal medicine isodon. Asia Pac
Tradit Med. 7:155–161. 2011.(In Chinese).
|
|
9
|
Dong R, Gao H and Liu Z: Research progress
on the biological activities of diterpenoids from isodon
(lamiaceae). China Pharm. 21:651–653. 2010.(In Chinese).
|
|
10
|
Lin W, Xie J, Xu N, Huang L, Xu A, Li H,
Li C, Gao Y, Watanabe M, Liu C and Huang P: Glaucocalyxin A induces
G2/M cell cycle arrest and apoptosis through the PI3K/Akt pathway
in human bladder cancer cells. Int J Biol Sci. 14:418–426. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Hou X, Xu G, Wang Z, Zhan X, Li H, Li R,
Shi W, Wang C, Chen Y, Ai Y, et al: Glaucocalyxin A alleviates
LPS-mediated septic shock and inflammation via inhibiting NLRP3
inflammasome activation. Int Immunopharmacol. 81:1062712020.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Gan P, Zhang L, Chen Y, Zhang Y, Zhang F,
Zhou X, Zhang X, Gao B, Zhen X, Zhang J and Zheng LT:
Anti-inflammatory effects of glaucocalyxin B in microglia cells. J
Pharmacol Sci. 128:35–46. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Pan Y, Bai J, Shen F, Sun L, He Q and Su
B: Glaucocalyxin B induces apoptosis and autophagy in human
cervical cancer cells. Mol Med Rep. 14:1751–1755. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Wei MQ, Chen L, Zhang WQ, Sun M, Wang JJ,
Zhan JP, Wang CL and Yan C: Chemical constituents from Isodon
suzhouensis and their anti-tumor activity. Nat Prod Res Deve.
37:262–270+261. 2024.(In Chinese).
|
|
15
|
Zhao L, Zhang H, Li N, Chen J, Xu H, Wang
Y and Liang Q: Network pharmacology, a promising approach to reveal
the pharmacology mechanism of Chinese medicine formula. J
Ethnopharmacol. 309:1163062023. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Yuan Z, Pan Y, Leng T, Chu Y, Zhang H, Ma
J and Ma X: Progress and prospects of research ideas and methods in
the network pharmacology of traditional Chinese medicine. J Pharm
Pharm Sci. 25:218–226. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Dean PN and Jett JH: Mathematical analysis
of DNA distributions derived from flow microfluorometry. J Cell
Biol. 60:523–527. 1974. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Zhu XJ, Li YM, Gu JY, Guo M and Fei J:
Enhanced Chemosensitivty of Leukemic Cells to Cytarabine by
Targeted Suppression of miRNA-21. Life Sci Res. 15:317–322.
2011.(In Chinese).
|
|
19
|
Wang ES, Montesinos P, Foran J, Erba H,
Rodríguez-Arbolí E, Fedorov K, Heiblig M, Heidel FH, Altman JK,
Baer MR, et al: Ziftomenib in relapsed or refractory NPM1-mutated
AML. J Clin Oncol. 43:3381–3390. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Atanasov AG, Waltenberger B,
Pferschy-Wenzig EM, Linder T, Wawrosch C, Uhrin P, Temml V, Wang L,
Schwaiger S, Heiss EH, et al: Discovery and resupply of
pharmacologically active plant-derived natural products: A review.
Biotechnol Adv. 33:1582–1614. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Aung TN, Qu Z, Kortschak RD and Adelson
DL: Understanding the effectiveness of natural compound mixtures in
cancer through their molecular mode of action. Int J Mol Sci.
18:6562017. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Hirose R, Miura T, Sha R, Shinkai Y,
Tanaka-Kagawa T and Kumagai Y: A method for detecting covalent
modification of sensor proteins associated with
1,4-naphthoquinone-induced activation of electrophilic signal
transduction pathways. J Toxicol Sci. 37:891–898. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Furtek SL, Backos DS, Matheson CJ and
Reigan P: Strategies and approaches of targeting STAT3 for cancer
treatment. ACS Chem Biol. 11:308–318. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Fasouli ES and Katsantoni E: JAK-STAT in
early hematopoiesis and leukemia. Front Cell Dev Biol.
9:6693632021. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
How J, Garcia JS and Mullally A: Biology
and therapeutic targeting of molecular mechanisms in MPNs. Blood.
141:1922–1933. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Jaśkiewicz A, Domoradzki T and Pająk B:
Targeting the JAK2/STAT3 pathway-can we compare it to the two faces
of the god janus? Int J Mol Sci. 21:82612020. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Agashe RP, Lippman SM and Kurzrock R: JAK:
Not just another kinase. Mol Cancer Ther. 21:1757–1764. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Qin JJ, Yan L, Zhang J and Zhang WD: STAT3
as a potential therapeutic target in triple negative breast cancer:
A systematic review. J Exp Clin Cancer Res. 38:1952019. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Gao SP, Mark KG, Leslie K, Pao W, Motoi N,
Gerald WL, Travis WD, Bornmann W, Veach D, Clarkson B and Bromberg
JF: Mutations in the EGFR kinase domain mediate STAT3 activation
via IL-6 production in human lung adenocarcinomas. J Clin Invest.
117:3846–3856. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Banerjee S, Biehl A, Gadina M, Hasni S and
Schwartz DM: JAK-STAT signaling as a target for inflammatory and
autoimmune diseases: Current and future prospects. Drugs.
77:521–546. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Salas A, Hernandez-Rocha C, Duijvestein M,
Faubion W, McGovern D, Vermeire S, Vetrano S and Vande Casteele N:
JAK-STAT pathway targeting for the treatment of inflammatory bowel
disease. Nat Rev Gastroenterol Hepatol. 17:323–337. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Hegde S, Ni S, He S, Yoon D, Feng GS,
Watowich SS, Paulson RF and Hankey PA: Stat3 promotes the
development of erythroleukemia by inducing Pu.1 expression and
inhibiting erythroid differentiation. Oncogene. 28:3349–3359. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Chen C, Lu M, Lin S and Qin W: The nuclear
gene rpl18 regulates erythroid maturation via JAK2-STAT3 signaling
in zebrafish model of Diamond-Blackfan anemia. Cell Death Dis.
11:1352020. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Fuchs Y: The therapeutic promise of
apoptosis. Science. 363:1050–1051. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Talib WH: Melatonin and cancer hallmarks.
Molecules. 23:5182018. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Nabih NW, Hassan HAFM, Preis E, Schaefer
J, Babker A, Abbas AM, Amin MU, Bakowsky U and Fahmy SA:
Antibody-functionalized lipid nanocarriers for RNA-based cancer
gene therapy: Advances and challenges in targeted delivery.
Nanoscale Adv. 7:5905–5931. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Wafik Nabih N, Nafie MS, Babker A, Hassan
HAFM and Fahmy SA: Recent advances in nano vehicles encapsulating
cinnamic acid and its derivatives as promising anticancer agents.
RSC Adv. 15:20815–20847. 2025. View Article : Google Scholar : PubMed/NCBI
|
















![Enrichment analysis. (A) GO
biological function analysis; (B) KEGG pathway enrichment analysis:
Top 20 significantly enriched pathways [-log10 (P-value)]. GO, Gene
Ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes.](/article_images/ol/31/6/ol-31-06-15622-g01.jpg)



