Introduction
Recently, many studies have focused on investigating
the role of non-coding RNAs (ncRNAs), RNA sequences that are not
translated into protein. Not only are the functions of ncRNAs only
partially understood, but the clinical or biological significance
of most ncRNAs has not yet been determined. Among ncRNAs, long
ncRNAs, consisting of over 200 bases have been reported to
associate with DNA-binding proteins, such as chromatin modifying
complexes, and certain ncRNAs can epigenetically regulate the
expression of multiple genes through this mechanism (1–3).
Hox transcript antisense intergenic RNA
(HOTAIR) is a long ncRNA that was identified from a custom
tilling array of the HOXC locus (12q13.13) (4). HOTAIR forms a complex with the
polycomb-repressive complex 2 (PRC2), composed of EZH2, SUZ12 and
EED, to trimethylate histone H3 at lysine 27 (H3K27me3), thereby
inhibiting HOXD gene expression. Thus, HOTAIR
epigenetically regulates the expression of HOXD, a gene
located on a separate chromosome. Recent studies have demonstrated
the clinical significance of HOTAIR in surgical solid
malignancies. Gupta et al found that patients with high
HOTAIR expression in primary lesions had a poorer prognosis
for both overall- and metastases-free survival than those with low
HOTAIR expression in breast cancer (4). Moreover, we reported the clinical
significance of HOTAIR in patients with advanced colorectal
cancer (CRC), where HOTAIR was more highly expressed in
cancerous tissues than in noncancerous tissues (5). High HOTAIR expression
correlated well with the presence of liver metastases, and was
significantly associated with poorer prognoses. In addition, we
found that HOTAIR expression was associated with a
genome-wide reprogramming of PRC2 (SUZ12, EZH2 and H3K27me3) by
gene set enrichment analysis using cDNA array data from a pilot
study (5). Since HOTAIR
expression has clinical significance in both breast and colorectal
cancers, we expect that this ncRNA may also play an important role
in other intractable malignancies.
Hepatocellular carcinoma (HCC) is one of the most
common cancers in the world. In Japan, HCC is an intractable tumor,
causing approximately 30,000 deaths per year and representing the
third leading cause of death from malignant neoplasms in men, and
the fifth leading cause of death from malignant neoplasms in women
(6,7). The major causes of HCC are viral
infections, alcohol and tobacco use. While efforts have been made
to identify appropriate prognostic markers for HCC (8–12),
including primary tumor size, elevated AFP levels, and gene
expression markers in the primary tumor, these method have not
proven adequate to predict the prognosis of all HCC patients. In
addition, few studies describing the expression of ncRNAs and their
clinical significance in HCC have been published. In the present
study, we determined the clinicopathological significance of
HOTAIR expression in HCC patients, and investigated whether
HOTAIR is a useful prognostic indicator in HCC patients.
Materials and methods
Clinical tissue samples
A total of 64 patients with HCC who underwent
surgery at Beppu Hospital were enrolled in this study. The resected
tumor and paired non-tumor tissue specimens were immediately frozen
in liquid nitrogen and kept at −80<C until analysis. Frozen
tissue specimens were homogenized in guanidinium thiocyanate, and
total RNAs were obtained by ultracentrifugation through a cesium
chloride cushion. Written informed consent was obtained from all
patients. All patients were closely followed after surgery at
regular one-month intervals.
RNA preparation, reverse transcription
and quantitative real-time PCR
Total RNA from frozen HCC samples (n=64) was
extracted using Isogen (Nippon Gene Co., Ltd.) following the
manufacturer’s protocol. As previously reported, cDNAs from all
samples were synthesized from 8.0 μg of total RNA (13). HOTAIR levels were quantified
using a LightCycler 480 Probes Master kit (Roche Applied Science)
following the manufacturer’s protocol with the following specific
HOTAIR primers: (forward, 5′-CAGTGGGGAACTCTGACTCG-3′ and
reverse, 5′-GTGCCTGGTGCTCTCTTACC-3′). HOTAIR levels were
normalized to GAPDH (forward, 5′-GTCAACGGATTTGG TCTGTATT-3′ and
reverse, 5′-AGTCTTCTGGGTGGCAGT GAT-3′).
Cell lines
HepG2 cells, human liver cancer cells, were provided
by the Cell Resource Center for Biomedical Research, Institute of
Development, Aging and Cancer, Tohoku University, Japan. All cell
lines were maintained in Dulbecco’s modified Eagle’s media (DMEM)
supplemented with 10% fetal calf serum and antibiotics. We cultured
the cells at 37°C in a humidified atmosphere of 5% CO2
and 95% air.
HOTAIR expression lentiviral vector
To generate a HOTAIR expression lentiviral
vector, we amplified full-length human HOTAIR by PCR using
MCF7 cDNA. Lentiviruses were produced by transient transfection of
HEK293T cells with pCMV-VSV-G-RSV-Rev, pCAG-HIVgp, and either
CSII-CMV-HOTAIR or CSII-CMV-MCS (empty) plasmid DNAs
(5′-XhoI and 3′-NotI sites) using Lipofectamine 2000
(Invitrogen), following the manufacturer’s protocol. Forty-eight
hours after cotransfection, the lentivirus-containing supernatant
was collected and passed through a 0.45-μm filter. The titer of the
lentivirus vector in filtered supernatants was estimated by
measuring the concentration of HIV p24 gag antigen with an ELISA
kit (Perkin-Elmer Life Science).
Cell proliferation assay
Cell proliferation was assessed by
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl tetrazolium bromide (MTT)
assay. In brief, we plated HepG2 cells infected with either the
human HOTAIR full-length lentiviral vector or the empty
lentiviral vector, in 96-well tissue culture plates at a density of
5.0×103 cells per well. At different time points (24,
72, and 120 h after plating, representing the 0-, 48-, and 96-h
time points respectively), 10 μl MTT (5 mg/ml in phosphate-buffered
saline) was added to each well, and plates were incubated for an
additional 4 h at 37°C. The colored formazan product was then
dissolved in 100 μl DMSO. We then evaluated mitochondrial activity,
reflecting cellular growth and viability, by measuring the optical
density at a test wavelength of 570–650 nm using a microplate
reader (Bio-Rad, Japan); results were expressed as optimal density
per milligram protein (OD/mg protein).
Statistical analysis
The significance of differences between two groups
was estimated using the Student’s t-test and the χ2
test. Overall survival curves were plotted according to the
Kaplan-Meier method, with the log-rank test applied for comparison.
Variables with a p-value of <0.05 by univariate analysis were
used in subsequent multivariate analysis on the basis of the Cox
proportional hazards model. All differences were considered
statistically significant when p<0.05. Statistical analyses were
conducted using JMP 5 software (SAS Institute).
Gene set enrichment analysis (GSEA) of
HCC in the GEO database
For GSEA (14), we
used GSE27462 in the GEO database; this was a dataset for RNA
expression profiles collected using a genome tiling array in 10 HCC
samples (15). HOTAIR
expression was treated as a binary variable divided into low or
high expression according to medians. For functional gene sets for
GSEA, we used gene sets arising from global occupancy of H3K27me3,
and EZH2, a PRC2 subunit induced by HOTAIR overexpression in
MDA-MB-231 breast cancer cells (4).
As a metric for ranking genes in GSEA, the difference between the
means of samples with low and high HOTAIR expression was
used, and other parameters were set to default values.
Results
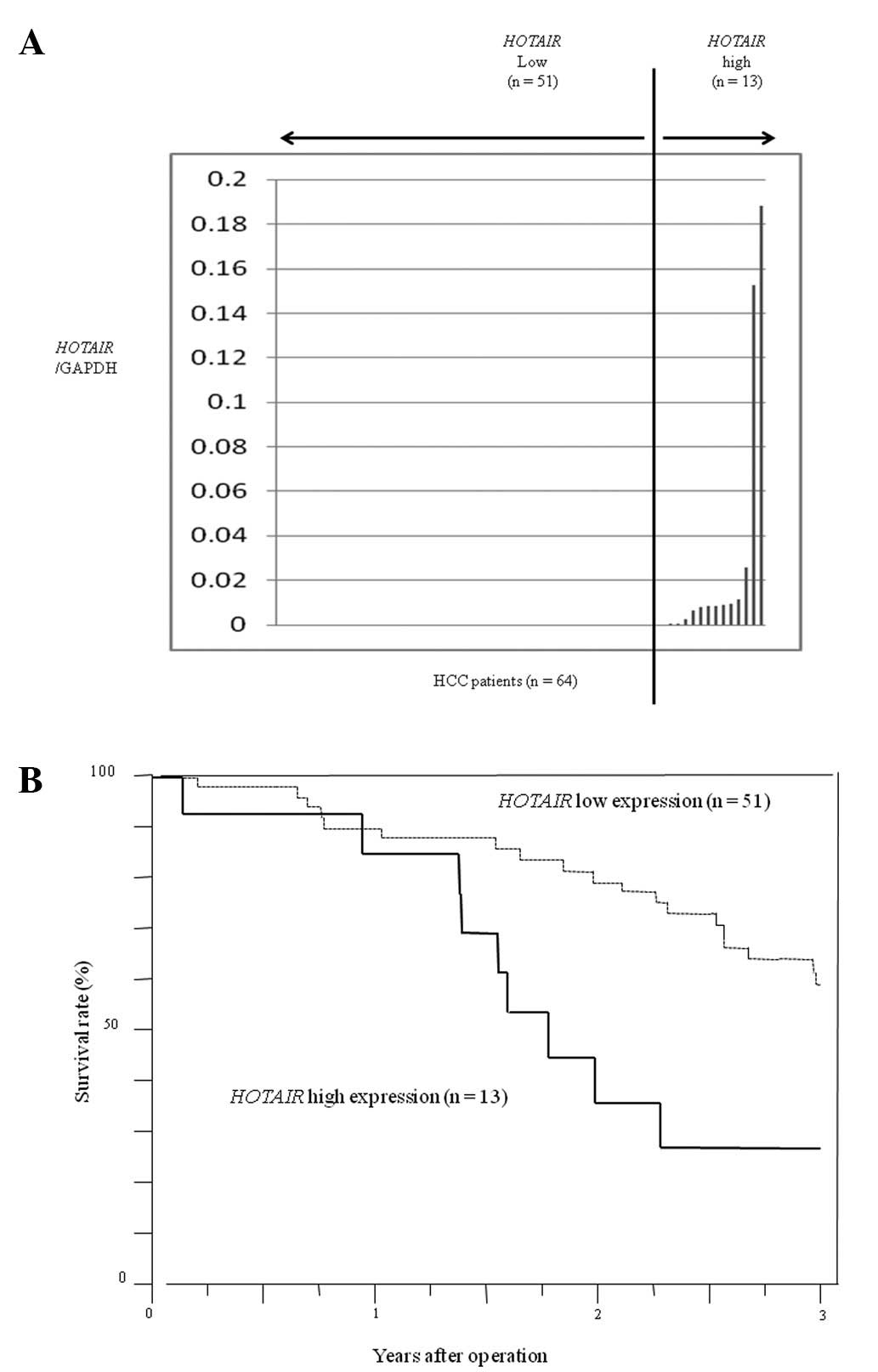
We first evaluated HOTAIR expression in primary
tumors from HCC patients (n=64) by quantitative real-time PCR. From
these data, we divided the 64 patients with HCC into a
HOTAIR high expression group (n=13) and a low expression
group (n=51) (Fig. 1A). In
consideration of clinical applications, we set cut-off values as
the upper limit of normal samples, and we found a
HOTAIR/GAPDH ratio of 0.027 in HCC samples. Patients with
high HOTAIR expression had a significantly poorer prognosis
with regard to overall survival and a significantly larger tumor
size than those with low HOTAIR expression (p<0.01)
(Fig. 1B). Clinicopathological
factors were analyzed between groups (Table I), and no significant differences in
other clinicopathological factors or in recurrence-free survival
were noted between the high and low expression groups (data not
shown).
 | Table IHOTAIR expression and
clinicopathological features of the HCC patients. |
Table I
HOTAIR expression and
clinicopathological features of the HCC patients.
| Features | HOTAIR high
expression (n=13) | HOTAIR low
expression (n=51) | p-value |
|---|
| Age (years) | 61.3±2.8 | 67.2±1.4 | 0.06 |
| Gender |
| Male | 9 (69.2%) | 34 (66.7%) | 0.86 |
| Female | 4 (30.8%) | 17 (33.3%) | |
| Virus |
| HBV(+) | 4 (30.8%) | 9 (17.7%) | 0.38 |
| HCV(+) | 9 (69.2%) | 38 (74.5%) | |
| non-B, non-C | 0 (0.0%) | 4 (7.8%) | |
| Child-Pugh |
| A | 11 (84.6%) | 44 (86.2%) | 0.88 |
| B | 2 (15.4%) | 6 (11.8%) | |
| C | 0 (0.0%) | 1 (2.0%) | |
| Tumor size
(cm) | 5.7±0.8 | 3.2±0.4 | 0.009a |
| fc |
| (+) | 4 (30.8%) | 15 (29.4%) | 0.92 |
| (−) | 9 (69.2%) | 36 (70.6%) | |
| fc-inf |
| (+) | 7 (53.9%) | 22 (43.1%) | 0.49 |
| (−) | 6 (46.1%) | 29 (56.9%) | |
| vp |
| 0 | 5 (38.5%) | 24 (47.1%) | 0.61 |
| 1 | 7 (53.9%) | 20 (39.2%) | |
| 2 | 1 (7.6%) | 7 (13.7%) | |
| vv |
| 0 | 12 (92.3%) | 49 (96.1%) | 0.59 |
| 1 | 1 (7.7%) | 2 (3.9%) | |
| b |
| 0 | 13 (100%) | 50 (98.0%) | 0.61 |
| 1 | 0 (0%) | 1 (2.0%) | |
| im |
| 0 | 12 (92.3%) | 45 (88.2%) | 0.85 |
| 1 | 1 (7.7%) | 5 (9.8%) | |
| 2 | 0 (0.0%) | 1 (2.0%) | |
| Tumor number |
| Single | 9 (69.2%) | 36 (70.6%) | 0.92 |
| Multiple | 4 (30.8%) | 15 (29.4%) | |
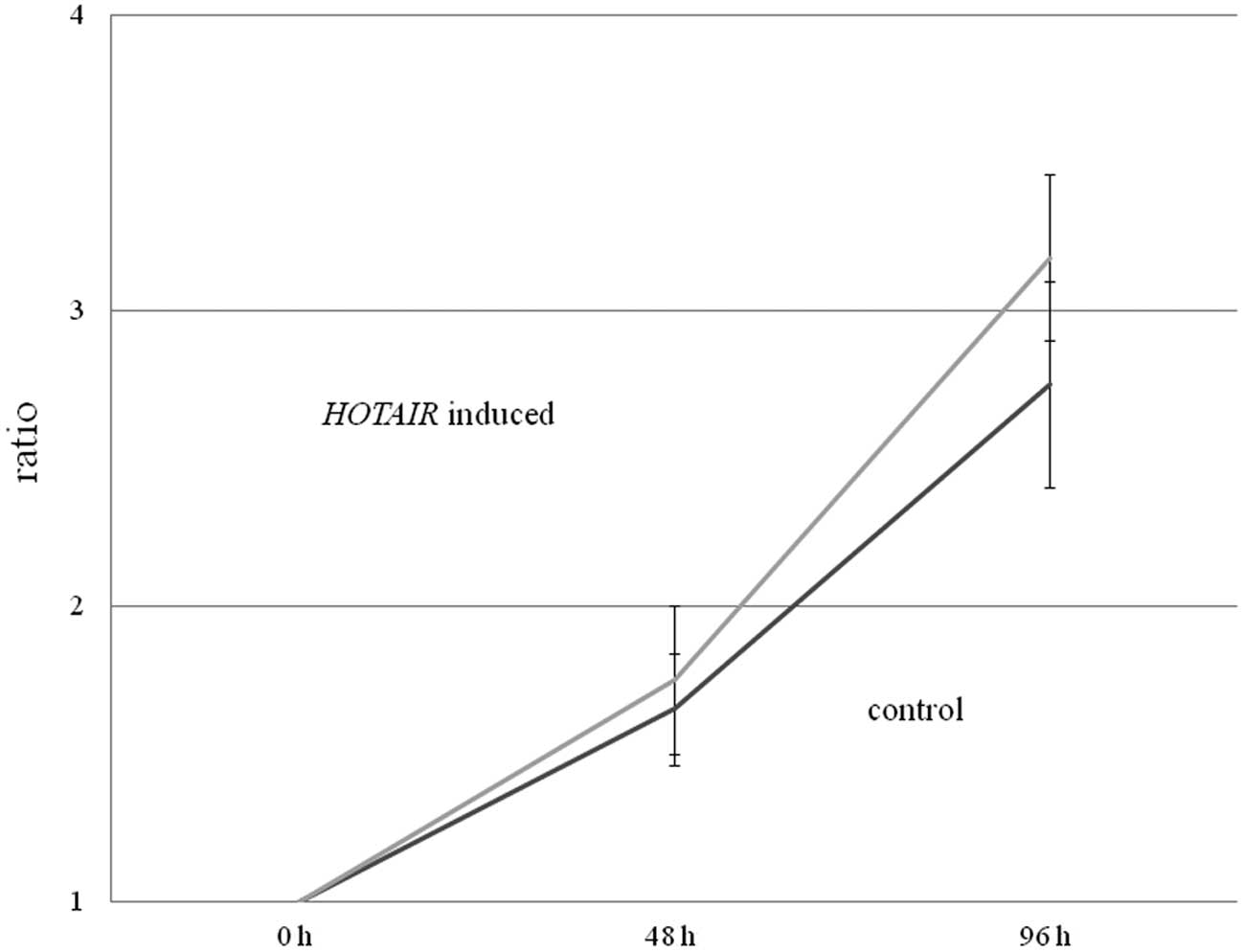
Subsequently, we examined whether HOTAIR
overexpression in HepG2 cells induced more rapid cell growth using
an MTT assay. While we observed no significant differences between
normal HepG2 cells (control) and HepG2 cells overexpressing
HOTAIR, there was a tendency for the
HOTAIR-expressing HepG2 cells to grow more rapidly than the
control HepG2 cells (p=0.10) (Fig.
2).
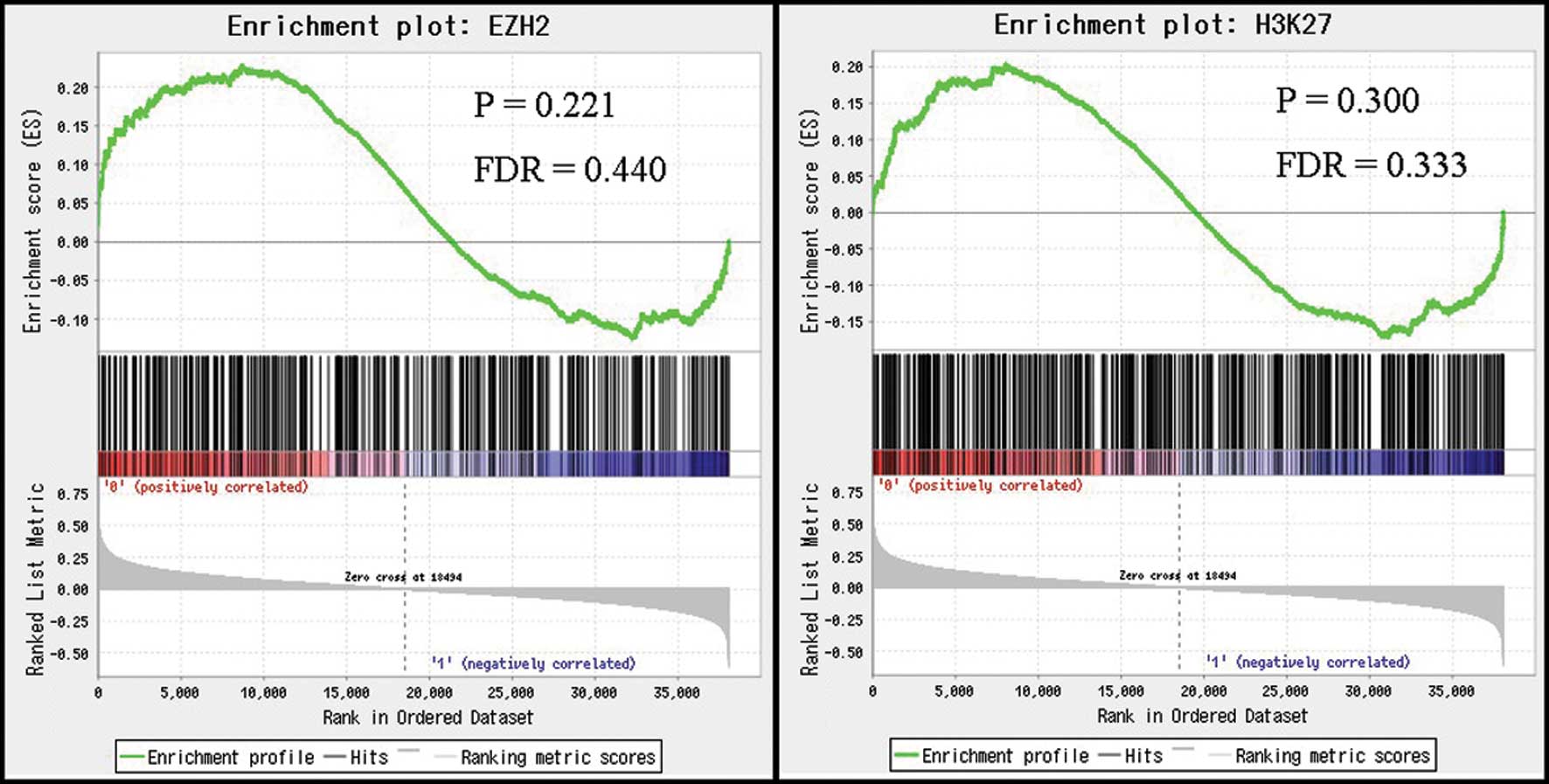
We next investigated the relationship between
HOTAIR expression and gene signatures after HOTAIR-induced
H3K27me3 and EZH2 occupancy using GSEA. Based on the results of
GSEA, there were no significant correlations between HOTAIR
expression levels in HCC and the expression levels of signatures
induced by H3K27me3 and EZH2 occupancy (Fig. 3). These results suggest that
HOTAIR expression in HCC may not induce genome-wide
retargeting of PRC2, unlike that noted in breast and colorectal
cancers.
Discussion
In the present study, we found that high
HOTAIR expression in HCC primary tumors was associated with
a poor prognosis. As for clinicopathological factors, there was a
significant association between HOTAIR expression and tumor
size. Additionally, we found that introduction of human
HOTAIR in liver cancer cells resulted in a tendency for more
rapid proliferation, compared with the control cells. We speculate
that HOTAIR may promote cell growth, explaining the
significant association between HOTAIR expression and tumor
size in HCC, and leading to poorer prognoses in HCC patients with
high HOTAIR expression.
Recently, researchers have been searching for novel
biomarkers to predict tumor recurrence in patients who have
undergone liver transplantation (16). In the present study, we showed that
high HOTAIR expression in primary HCC could be a useful
marker for predicting HCC recurrence since HCC cases can be clearly
categorized into high and low expression groups.
In previous studies, including ours, HOTAIR
expression was shown to interact with multiple genes in cooperation
with PRC2 (5), suggesting an
important role for HOTAIR in tumor growth and tumorigenicity
of breast and colorectal cancers. In addition, Yang et al
reported that long ncRNAs, including HOTAIR, are expressed
at high levels in HCC tumors in vivo(15). HOTAIR and other long ncRNAs,
such as XIST, and other unidentified ncRNAs, may play significant
roles in tumor growth in cooperation with PRC2 and histone
modification genes in HCC (17–19).
In the present study, however, GSEA results showed that the
expression profiles generally induced by H3K27me3 and EZH2, a PRC2
element, were not significantly associated with HOTAIR
expression in HCC. Therefore, this ncRNA may cooperate with other
molecules or in different pathways in HCC, rather than functional
with PRC2 complex proteins, and further research is required to
fully elucidate these mechanisms.
In conclusion, certain HCC patients express the long
ncRNA HOTAIR, and HCC patients with high HOTAIR
expression exhibits poorer prognoses than those with low
HOTAIR expression. HOTAIR expression in primary HCC
tumors may be a prognostic marker in HCC; however, the clinical
significance of HOTAIR expression in HCC appears to be less
important than that in breast and colorectal cancers, including
parameters such as histological grade, tumor depth and lymph node
metastasis. Our data demonstrate the need for further study to
support the use of ncRNAs as new clinical indicators of poor
prognosis in HCC.
Acknowledgements
The authors thank T. Shimooka, and M. Kasagi for
their technical assistance and H. Miyoshi (RIKEN BioResource
Center) for providing the lentiviral vector plasmid DNA. This study
was supported in part by the following grants and foundations:
CREST, Japan Science and Technology Agency (JST); Japan Society for
the Promotion of Science (JSPS) Grant-in-Aid for Scientific
Research (grant nos. 20390360, 20591547, 20790960, 21591644,
21791295, 21791297, 215921014 and 21679006); the Funding Program
for Next Generation World-Leading Researchers (LS094); NEDO (New
Energy and Industrial Technology Development Organization)
Technological Development for Chromosome Analysis; and Grant-in-Aid
from the Tokyo Biochemical Research Foundation.
References
|
1
|
Ponting CP, Oliver PL and Reik W:
Evolution and functions of long noncoding RNAs. Cell. 136:629–641.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Rinn JL, Kertesz M, Wang JK, et al:
Functional demarcation of active and silent chromatin domains in
human HOX loci by noncoding RNAs. Cell. 129:1311–1323. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Khalil AM, Guttman M, Huarte M, et al:
Many human large intergenic noncoding RNAs associate with
chromatin-modifying complexes and affect gene expression. Proc Natl
Acad Sci USA. 106:11667–11672. 2009. View Article : Google Scholar
|
|
4
|
Gupta RA, Shah N, Wang KC, et al: Long
non-coding RNA HOTAIR reprograms chromatin state to promote cancer
metastasis. Nature. 464:1071–1076. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Kogo R, Shimamura T, Mimori K, et al: Long
noncoding RNA HOTAIR regulates polycomb-dependent chromatin
modification and is associated with poor prognosis in colorectal
cancers. Cancer Res. 71:6320–6326. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Kudo M, Izumi N, Kokudo N, et al:
Management of hepatocellular carcinoma in Japan: Consensus-Based
Clinical Practice Guidelines proposed by the Japan Society of
Hepatology (JSH) 2010 updated version. Dig Dis. 29:339–364. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Umemura T, Ichijo T, Yoshizawa K, Tanaka E
and Kiyosawa K: Epidemiology of hepatocellular carcinoma in Japan.
J Gastroenterol. 44(Suppl 19): 102–107. 2009. View Article : Google Scholar
|
|
8
|
Ammerpohl O, Pratschke J, Schafmayer C, et
al: Distinct DNA methylation patterns in cirrhotic liver and
hepatocellular carcinoma. Int J Cancer. 130:1319–1328. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
McKnight R, Nassar A, Cohen C and Siddiqui
MT: Arginase-1: a novel immunohistochemical marker of
hepatocellular differentiation in fine needle aspiration cytology.
Cancer Cytopathol. 120:223–229. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Pan K, Liang XT, Zhang HK, et al:
Characterization of BIN1 as a potential tumor suppressor and
prognostic marker in hepatocellular carcinoma. Mol Med. 18:507–518.
2012.PubMed/NCBI
|
|
11
|
Andrisani OM, Studach L and Merle P: Gene
signatures in hepatocellular carcinoma (HCC). Semin Cancer Biol.
21:4–9. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Qiu J, Huang P, Liu Q, et al:
Identification of MACC1 as a novel prognostic marker in
hepatocellular carcinoma. J Transl Med. 9:1662011. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Inoue H, Mori M, Honda M, et al: The
expression of tumor-rejection antigen ‘MAGE’ genes in human gastric
carcinoma. Gastroenterology. 109:1522–1525. 1995.
|
|
14
|
Subramanian A, Tamayo P, Mootha VK, et al:
Gene set enrichment analysis: a knowledge-based approach for
interpreting genome-wide expression profiles. Proc Natl Acad Sci
USA. 102:15545–15550. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Yang F, Zhang L, Huo XS, et al: Long
noncoding RNA high expression in hepatocellular carcinoma
facilitates tumor growth through enhancer of zeste homolog 2 in
humans. Hepatology. 54:1679–1689. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Yang Z, Zhou L, Wu LM, et al:
Overexpression of long non-coding RNA HOTAIR predicts tumor
recurrence in hepatocellular carcinoma patients following liver
transplantation. Ann Surg Oncol. 18:1243–1250. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Huarte M, Guttman M, Feldser D, et al: A
large intergenic noncoding RNA induced by p53 mediates global gene
repression in the p53 response. Cell. 142:409–419. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Braconi C, Valeri N, Kogure T, et al:
Expression and functional role of a transcribed noncoding RNA with
an ultraconserved element in hepatocellular carcinoma. Proc Natl
Acad Sci USA. 108:786–791. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Tsai MC, Manor O, Wan Y, et al: Long
noncoding RNA as modular scaffold of histone modification
complexes. Science. 329:689–693. 2010. View Article : Google Scholar : PubMed/NCBI
|