|
1
|
Schlaeger C, Longerich T, Schiller C,
Bewerunge P, Mehrabi A, Toedt G, Kleeff J, Ehemann V, Eils R,
Lichter P, et al: Etiology-dependent molecular mechanisms in human
hepatocarcinogenesis. Hepatology. 47:511–520. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
El-Serag HB and Davila JA: Is
fibrolamellar carcinoma different from hepatocellular carcinoma? A
US population-based study. Hepatology. 39:798–803. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Mavros MN, Mayo SC, Hyder O and Pawlik TM:
A systematic review: Treatment and prognosis of patients with
fibrolamellar hepatocellular carcinoma. J Am Coll Surg.
215:820–830. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Kakar S, Chen X, Ho C, Burgart LJ, Sahai
V, Dachrut S, Yabes A, Jain D and Ferrell LD: Chromosomal changes
in fibrolamellar hepatocellular carcinoma detected by array
comparative genomic hybridization. Mod Pathol. 22:134–141. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Craig JR, Peters RL, Edmondson HA and
Omata M: Fibrolamellar carcinoma of the liver: A tumor of
adolescents and young adults with distinctive clinico-pathologic
features. Cancer. 46:372–379. 1980. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Kakar S, Burgart LJ, Batts KP, Garcia J,
Jain D and Ferrell LD: Clinicopathologic features and survival in
fibrolamellar carcinoma: Comparison with conventional
hepatocellular carcinoma with and without cirrhosis. Mod Pathol.
18:1417–1423. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Edmondson HA: Differential diagnosis of
tumors and tumor-like lesions of liver in infancy and childhood.
AMA J Dis Child. 91:168–186. 1956.PubMed/NCBI
|
|
8
|
Sanyal AJ, Yoon SK and Lencioni R: The
etiology of hepatocellular carcinoma and consequences for
treatment. Oncologist. 15 (Suppl 4):S14–S22. 2010. View Article : Google Scholar
|
|
9
|
Lafaro KJ and Pawlik TM: Fibrolamellar
hepatocellular carcinoma: Current clinical perspectives. J
Hepatocell Carcinoma. 2:151–157. 2015.PubMed/NCBI
|
|
10
|
Lane DP: Cancer. p53, guardian of the
genome. Nature. 358:15–16. 1992. View
Article : Google Scholar : PubMed/NCBI
|
|
11
|
Efeyan A and Serrano M: p53: Guardian of
the genome and policeman of the oncogenes. Cell Cycle. 6:1006–1010.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Selivanova G: Wild type p53 reactivation:
From lab bench to clinic. FEBS Lett. 588:2628–2638. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Shiloh Y: ATM: Ready, set, go. Cell Cycle.
2:116–117. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Shiloh Y: ATM and related protein kinases:
Safeguarding genome integrity. Nat Rev Cancer. 3:155–168. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Banin S, Moyal L, Shieh S, Taya Y,
Anderson CW, Chessa L, Smorodinsky NI, Prives C, Reiss Y, Shiloh Y
and Ziv Y: Enhanced phosphorylation of p53 by ATM in response to
DNA damage. Science. 281:1674–1677. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Polyak K, Xia Y, Zweier JL, Kinzler KW and
Vogelstein B: A model for p53-induced apoptosis. Nature.
389:300–305. 1997. View
Article : Google Scholar : PubMed/NCBI
|
|
17
|
Livingstone LR, White A, Sprouse J,
Livanos E, Jacks T and Tlsty TD: Altered cell cycle arrest and gene
amplification potential accompany loss of wild-type p53. Cell.
70:923–935. 1992. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Sengupta S and Harris CC: p53: Traffic cop
at the crossroads of DNA repair and recombination. Nat Rev Mol Cell
Biol. 6:44–55. 2005. View
Article : Google Scholar : PubMed/NCBI
|
|
19
|
Marine JC: MDM2 and MDMX in cancer and
development. Curr Top Dev Biol. 94:45–75. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Marine JC and Jochemsen AG: Mdmx as an
essential regulator of p53 activity. Biochem Biophys Res Commun.
331:750–760. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Haupt Y, Maya R, Kazaz A and Oren M: Mdm2
promotes the rapid degradation of p53. Nature. 387:296–299. 1997.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Shvarts A, Bazuine M, Dekker P, Ramos YF,
Steegenga WT, Merckx G, van Ham RC, van der Houven van Oordt W, van
der Eb AJ and Jochemsen AG: Isolation and identification of the
human homolog of a new p53-binding protein, Mdmx. Genomics.
43:34–42. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Shvarts A, Steegenga WT, Riteco N, van
Laar T, Dekker P, Bazuine M, van Ham RC, van der Houven van Oordt
W, Hateboer G, van der Eb AJ and Jochemsen AG: MDMX: A novel
p53-binding protein with some functional properties of MDM2. EMBO
J. 15:5349–5357. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Leroy B, Anderson M and Soussi T: TP53
mutations in human cancer: Database reassessment and prospects for
the next decade. Hum Mutat. 35:672–688. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Li L, Tan Y, Chen X, Xu Z, Yang S, Ren F,
Guo H, Wang X, Chen Y, Li G and Wang H: MDM4 overexpressed in acute
myeloid leukemia patients with complex karyotype and wild-type
TP53. PLoS One. 9:e1130882014. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Quintás-Cardama A, Hu C, Qutub A, Qiu YH,
Zhang X, Post SM, Zhang N, Coombes K and Kornblau SM: p53 pathway
dysfunction is highly prevalent in acute myeloid leukemia
independent of TP53 mutational status. Leukemia. 31:1296–1305.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Li Q and Lozano G: Molecular pathways:
Targeting Mdm2 and Mdm4 in cancer therapy. Clin Cancer Res.
19:34–41. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Gembarska A, Luciani F, Fedele C, Russell
EA, Dewaele M, Villar S, Zwolinska A, Haupt S, de Lange J, Yip D,
et al: MDM4 is a key therapeutic target in cutaneous melanoma. Nat
Med. 18:1239–1247. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Haupt S, Buckley D, Pang JM, Panimaya J,
Paul PJ, Gamell C, Takano EA, Lee YY, Hiddingh S, Rogers TM, et al:
Targeting Mdmx to treat breast cancers with wild-type p53. Cell
Death Dis. 6:e18212015. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Kan Z, Zheng H, Liu X, Li S, Barber TD,
Gong Z, Gao H, Hao K, Willard MD, Xu J, et al: Whole-genome
sequencing identifies recurrent mutations in hepatocellular
carcinoma. Genome Res. 23:1422–1433. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Kunst C, Haderer M, Heckel S, Schlosser S
and Müller M: The p53 family in hepatocellular carcinoma. Transl
Cancer Res. 5:632–638. 2016. View Article : Google Scholar
|
|
32
|
Ward SC and Waxman S: Fibrolamellar
carcinoma: A review with focus on genetics and comparison to other
malignant primary liver tumors. Semin Liver Dis. 31:61–70. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Sorenson EC, Khanin R, Bamboat ZM, Cavnar
MJ, Kim TS, Sadot E, Zeng S, Greer JB, Seifert AM, Cohen NA, et al:
Genome and transcriptome profiling of fibrolamellar hepatocellular
carcinoma demonstrates p53 and IGF2BP1 dysregulation. PLoS One.
12:e01765622017. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Cancer Genome Atlas Research Network.
Electronic address, . wheeler@bcm.edu; Cancer Genome AtlasResearch
Network: Comprehensive and integrative genomic characterization of
hepatocellular carcinoma. Cell. 169:1327–1341.e23. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Arai Y, Honda S, Haruta M, Kasai F,
Fujiwara Y, Ohshima J, Sasaki F, Nakagawara A, Horie H, Yamaoka H,
et al: Genome-wide analysis of allelic imbalances reveals 4q
deletions as a poor prognostic factor and MDM4 amplification at
1q32.1 in hepatoblastoma. Genes Chromosomes Cancer. 49:596–609.
2010.PubMed/NCBI
|
|
36
|
LaQuaglia MJ, Grijalva JL, Mueller KA,
Perez-Atayde AR, Kim HB, Sadri-Vakili G and Vakili K: YAP
subcellular localization and Hippo pathway transcriptome analysis
in pediatric hepatocellular carcinoma. Sci Rep. 6:302382016.
View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Fedchenko N and Reifenrath J: Different
approaches for interpretation and reporting of immunohistochemistry
analysis results in the bone tissue-a review. Diagn Pathol.
9:2212014. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Hemming AW, Langer B, Sheiner P, Greig PD
and Taylor BR: Aggressive surgical management of fibrolamellar
hepatocellular carcinoma. J Gastrointest Surg. 1:342–346. 1997.
View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Stevens WR, Johnson CD, Stephens DH and
Nagorney DM: Fibrolamellar hepatocellular carcinoma: Stage at
presentation and results of aggressive surgical management. AJR Am
J Roentgenol. 164:1153–1158. 1995. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
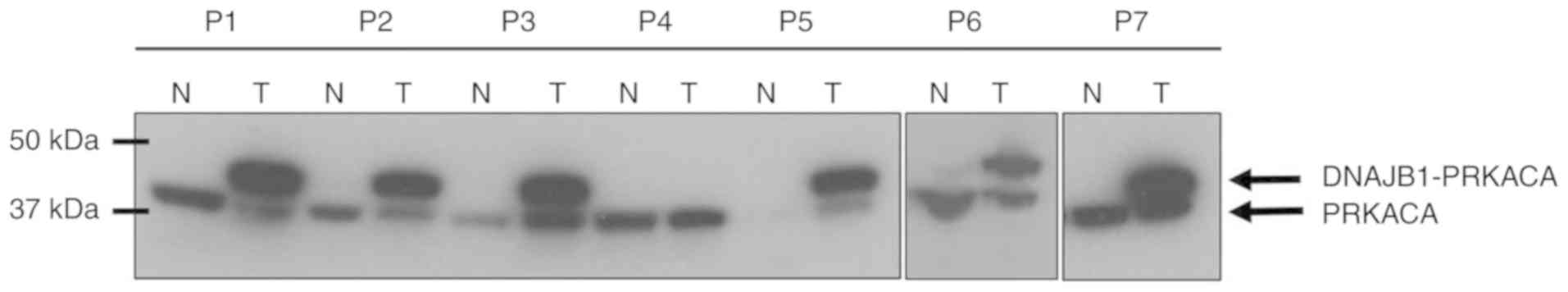
Honeyman JN, Simon EP, Robine N,
Chiaroni-Clarke R, Darcy DG, LimI I, Gleason CE, Murphy JM,
Rosenberg BR, Teegan L, et al: Detection of a recurrent
DNAJB1-PRKACA chimeric transcript in fibrolamellar hepatocellular
carcinoma. Science. 343:1010–1014. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Graham RP, Jin L, Knutson DL, Kloft-Nelson
SM, Greipp PT, Waldburger N, Roessler S, Longerich T, Roberts LR,
Oliveira AM, et al: DNAJB1-PRKACA is specific for fibrolamellar
carcinoma. Mod Pathol. 28:822–829. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Turenne GA, Paul P, Laflair L and Price
BD: Activation of p53 transcriptional activity requires ATM's
kinase domain and multiple N-terminal serine residues of p53.
Oncogene. 20:5100–5110. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Haupt S, Vijayakumaran R, Miranda PJ,
Burgess A, Lim E and Haupt Y: The role of MDM2 and MDM4 in breast
cancer development and prevention. J Mol Cell Biol. 9:53–61. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Zhang H, Hu L, Qiu W, Deng T, Zhang Y,
Bergholz J and Xiao ZX: MDMX exerts its oncogenic activity via
suppression of retinoblastoma protein. Oncogene. 34:5560–5569.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Carneiro A, Isinger A, Karlsson A,
Johansson J, Jönsson G, Bendahl PO, Falkenback D, Halvarsson B and
Nilbert M: Prognostic impact of array-based genomic profiles in
esophageal squamous cell cancer. BMC Cancer. 8:982008. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Tomasini MD, Wang Y, Karamafrooz A, Li G,
Beuming T, Gao J, Taylor SS, Veglia G and Simon SM: Conformational
LANDSCAPE of the PRKACA-DNAJB1 chimeric kinase, the driver for
fibrolamellar hepatocellular carcinoma. Sci Rep. 8:7202018.
View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Petitjean A, Mathe E, Kato S, Ishioka C,
Tavtigian SV, Hainaut P and Olivier M: Impact of mutant p53
functional properties on TP53 mutation patterns and tumor
phenotype: Lessons from recent developments in the IARC TP53
database. Hum Mutat. 28:622–629. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Olivier M, Hollstein M and Hainaut P: TP53
mutations in human cancers: Origins, consequences, and clinical
use. Cold Spring Harb Perspect Biol. 2:a0010082010. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Lawrence MS, Stojanov P, Polak P, Kryukov
GV, Cibulskis K, Sivachenko A, Carter SL, Stewart C, Mermel CH,
Roberts SA, et al: Mutational heterogeneity in cancer and the
search for new cancer-associated genes. Nature. 499:214–218. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Xu L, Hazard FK, Zmoos AF, Jahchan N,
Chaib H, Garfin PM, Rangaswami A, Snyder MP and Sage J: Genomic
analysis of fibrolamellar hepatocellular carcinoma. Hum Mol Genet.
24:50–63. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Finlay CA, Hinds PW and Levine AJ: The p53
proto-oncogene can act as a suppressor of transformation. Cell.
57:1083–1093. 1989. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Harper JW, Adami GR, Wei N, Keyomarsi K
and Elledge SJ: The p21 Cdk-interacting protein Cip1 is a potent
inhibitor of G1 cyclin-dependent kinases. Cell. 75:805–816. 1993.
View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Kastenhuber ER and Lowe SW: Putting p53 in
context. Cell. 170:1062–1078. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Mills KD: Tumor suppression: Putting p53
in context. Cell Cycle. 12:3461–3462. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
O'Brate A and Giannakakou P: The
importance of p53 location: Nuclear or cytoplasmic zip code? Drug
Resist Updat. 6:313–322. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Sembritzki O, Hagel C, Lamszus K, Deppert
W and Bohn W: Cytoplasmic localization of wild-type p53 in
glioblastomas correlates with expression of vimentin and glial
fibrillary acidic protein. Neuro Oncol. 4:171–178. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Green DR and Kroemer G: Cytoplasmic
functions of the tumour suppressor p53. Nature. 458:1127–1130.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Moll UM, Riou G and Levine AJ: Two
distinct mechanisms alter p53 in breast cancer: Mutation and
nuclear exclusion. Proc Natl Acad Sci USA. 89:7262–7266. 1992.
View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Bosari S, Viale G, Roncalli M, Graziani D,
Borsani G, Lee AK and Coggi G: p53 gene mutations, p53 protein
accumulation and compartmentalization in colorectal adenocarcinoma.
Am J Pathol. 147:790–798. 1995.PubMed/NCBI
|
|
60
|
Vici P, Di Benedetto A, Ercolani C,
Pizzuti L, Di Lauro L, Sergi D, Sperati F, Terrenato I, Dattilo R,
Botti C, et al: Predictive significance of DNA damage and repair
biomarkers in triple-negative breast cancer patients treated with
neoadjuvant chemotherapy: An exploratory analysis. Oncotarget.
6:42773–42780. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Shen X, Zuo X, Zhang W, Bai Y, Qin X and
Hou N: MiR-370 promotes apoptosis in colon cancer by directly
targeting MDM4. Oncol Lett. 15:1673–1679. 2018.PubMed/NCBI
|
|
62
|
Han H, Wang L, Xu J and Wang A: miR-128
induces pancreas cancer cell apoptosis by targeting MDM4. Exp Ther
Med. 15:5017–5022. 2018.PubMed/NCBI
|
|
63
|
Zache N, Lambert JM, Rökaeus N, Shen J,
Hainaut P, Bergman J, Wiman KG and Bykov VJ: Mutant p53 targeting
by the low molecular weight compound STIMA-1. Mol Oncol. 2:70–80.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Bykov VJ, Issaeva N, Shilov A, Hultcrantz
M, Pugacheva E, Chumakov P, Bergman J, Wiman KG and Selivanova G:
Restoration of the tumor suppressor function to mutant p53 by a
low-molecular-weight compound. Nat Med. 8:282–288. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Hiraki M, Hwang SY, Cao S, Ramadhar TR,
Byun S, Yoon KW, Lee JH, Chu K, Gurkar AU, Kolev V, et al:
Small-molecule reactivation of mutant p53 to wild-type-like p53
through the p53-Hsp40 regulatory axis. Chem Biol. 22:1206–1216.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Bossi G and Sacchi A: Restoration of
wild-type p53 function in human cancer: Relevance for tumor
therapy. Head Neck. 29:272–284. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Stad R, Little NA, Xirodimas DP, Frenk R,
van der Eb AJ, Lane DP, Saville MK and Jochemsen AG: Mdmx
stabilizes p53 and Mdm2 via two distinct mechanisms. EMBO Rep.
2:1029–1034. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Garcia D, Warr MR, Martins CP, Brown
Swigart L, Passegué E and Evan GI: Validation of MdmX as a
therapeutic target for reactivating p53 in tumors. Genes Dev.
25:1746–1757. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Danovi D, Meulmeester E, Pasini D,
Migliorini D, Capra M, Frenk R, de Graaf P, Francoz S, Gasparini P,
Gobbi A, et al: Amplification of Mdmx (or Mdm4) directly
contributes to tumor formation by inhibiting p53 tumor suppressor
activity. Mol Cell Biol. 24:5835–5843. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Wang H, Ma X, Ren S, Buolamwini JK and Yan
C: A small-molecule inhibitor of MDMX activates p53 and induces
apoptosis. Mol Cancer Ther. 10:69–79. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Reed D, Shen Y, Shelat AA, Arnold LA,
Ferreira AM, Zhu F, Mills N, Smithson DC, Regni CA, Bashford D, et
al: Identification and characterization of the first small molecule
inhibitor of MDMX. J Biol Chem. 285:10786–10796. 2010. View Article : Google Scholar : PubMed/NCBI
|