Introduction
Bladder cancer is theninth most frequently diagnosed
cancer globally, with an estimated 429,000 new cases and 165,000
cancer-related deaths worldwide in 2012 (1,2). Notably,
a significant difference in the incidence rate between sexes
exists; men always suffer from a higher incidence than women due to
their addiction with smoking (3,4). Most
bladder cancers (75–80%) that do not involve the muscular wall of
the bladder usually obtain favorable clinical results through
endoscopic treatment (4). In
addition, radical cystectomy combined with abdominal lymph node
dissection is the gold standard for the treatment of invasive
bladder cancer. However, due to the difficulty of this surgery and
the large number of postoperative complications, it is not suitable
for all patients (5). Therefore, it
is necessary to study the diagnosis and pathogenesis of bladder
cancer. Exploring potential targets for bladder cancer treatment is
urgently needed to further improve the therapeutic effects on
bladder cancer.
In recent years, growing evidence has revealed that
long non-coding RNAs act as biomarkers for diagnosis and prognosis
of cancer patients, thus providing new therapeutic targets for
cancer treatment (6–8). Numerous lncRNAs are aberrantly expressed
in various tumors, and a number of lncRNAs, such as MALAT1 in lung
cancer, are stable in body fluids and could be detected in the
plasma and urine of patients with cancer (9–11).
Increasing evidence has demonstrated that lncRNAs participate in
bladder cancer formation, progression, and metastasis (12,13). Wei
et al have reported thatlncRNA MBNL1-AS1 suppressed cell
proliferation and enhanced cell apoptosis via targeting of the
microRNA (miR)-135a-5p/PHLPP2/FOXO1 axis in bladder cancer
(14). Among these lncRNAs,
LINC00858, located in 10q23.1, has been reported in three types of
tumors, including lung cancer, osteosarcoma and colorectal cancer
(15). In these malignancies,
LINC00858 has been revealed to induce tumor progression and
metastasis. However, the role of LINC00858 in the regulation of
bladder cancer remains unknown.
In the present study, the expression level of
LINC00858 was detectedin bladder cancer tissues and cells. Then,
LINC00858 was knocked down by RNAi technology, and abilities
including cell proliferation, migration and invasion were
investigated. Futhermore, the mechanism underlying LINC00858 in
bladder cancer was studied.
Materials and methods
Human samples and ethical
approval
Tumor tissues and paired adjacent normal tissues
were obtained from 60 bladder cancer patients (28 females and 32
males) from June 2011 to July 2019 who underwent surgery at Jiangxi
Cancer Hospital (Nanchang, China). The inclusion criteria was as
follows: Age, from 28 to 61 years; histologically confirmed
non-muscular invasive urothelial carcinoma of the bladder. The
exclusion criteria was as follows: Patients with other concurrent
uropoiesis reproductive system tumors; patients with severe
cardiovascular and cerebrovascular disease such as heart failure;
known immunodeficiency; various mental disorders. The present study
was approved by the Ethics Committee of Jiangxi Cancer Hospital
(approval no. 2019014). Written informed consent was obtained from
all participants. All the tissue samples were collected and frozen
in liquid nitrogen, and then stored at −80°C for further use. The
clinicopathological data of the patients are presented in Table SI.
Cell culture
Three human bladder cancer cell lines,T24, J82 and
5637 and a normal human bladder epithelial cell line SV-HUC-1, were
purchased from the American Type Culture Collection (ATCC). All
cell lines were cultured in RPMI-1640 medium supplemented with 10%
fetal bovine serum (both from Invitrogen; Thermo Fisher Scientific,
Inc.) at 37°C with 5% CO2.
Cell transfection
The bladder cancer cell lines 5637 and T24 were
respectively transfected with 0.2 µg LINC00858 overexpression
vector, 50 pmol LINC00858 short hairpin (sh)RNA, 50 pmol negative
control (NC) shRNA, 50 pmol CTGF shRNA, 50 pmol NC mimic (a
non-targeting miR-scramble), 50 pmol miR-3064-5p mimic, 50 pmol
miR-3064-5p inhibitor, 50 pmol miR-NC inhibitor and 0.2 µg empty
vector was used as a control for the overexpression experiments
(all from Shanghai GenePharma Co., Ltd. The sequences were:
sh-LINC00858 (5′-GCGACATTAATGGGAATGA-3′); sh-CTGF (forward,
5′-CACCGCACCAGAATGTATATTAATTCAAGAGATTAATATACATTCTGGTGCTTTTTTG-3′
and reverse,
5′-GATCCAAAAAAGCACCAGAATGTATATTAATCTCTTGAATTAATATACATTCTGGTGC-3′);
sh-RNA NC (5′-GTTCTCCGAACGTGTCACGT-3′); miR-3064-5p mimic
(5′-TTACTGGCTGTTGTGGTGTGC-3′); NC mimic (forward,
5′-UUCUCCGAACGUGUCACGUTT-3′ and reverse,
5′-ACGUGACACGUUCGGAGAATT-3′); miR-3064-5p inhibitor
(5′-UUGCACACCACAACAGCCAGA-3′); and miR-NC inhibitor
(5′-CAGUACUUUUGUGUAGUACAA-3′). Lipofectamine 2000 (Invitrogen;
Thermo Fisher Scientific, Inc.) was used for transfection according
to the manufacturer's protocols. After transfection for 6 h at
37°C, fresh culture medium was added and cells were cultured for 48
h before futher experiments. Real-time PCR and green fluorescence
microscopy were used to assess the transfection efficiency.
Bioinformatics analysis
To predict the interaction site of miR-3064-5p with
LINC00858, starBase v2.0 (http://starbase.sysu.edu.cn/) (16) was employed according to the online
instructions. In addition, the binding site between miR-3064-5p and
CTGF was predicted by searching starBase v2.0 online.
CCK-8 and colony formation assays
Cell proliferation was detected by CCK-8 and colony
formation assays. The CCK-8 assay was performed according to the
manufacturer's instructions (Beyotime Biotechnology). In brief,
5,000 cells were seeded into a 96-well plate and cultured for 24,
48 and 72 h. Then 5 µl of CCK-8 reagent was added into each well
and cultured for another 1 h at 37°C before detection on a
microplate reader. The optical density (OD) values were then
measured at 450 nm. For the colony formation assay, 500 cells were
seeded into a 6-well plate and cultured for ~14 days, and then the
colonies (>5 cells per colony) were stained with 0.5% crystal
violet at room temperature for 10 min and counted with a
stereomicroscope (magnification, ×40; Leica MZ8; Leica Microsystems
GmbH).
Wound healing assay
Approximately 20,000 cells were seeded into a
24-well plate and cultured at 37°C for 24 h. The scratch wounds
were created in 5637 and T24 cells by 10-µl pipette tips. The cells
were washed 3 times with PBS, to remove the marked cells, and
serum-free medium was added for culture. Representative images were
captured after 48 h using a microscope (magnification, ×200; Leica
MZ8; Leica Microsystems GmbH).
Transwell chamber assay
Approximately 20,000 cells in serum-free medium were
seeded in the upper Transwell chamber (8 µm; Cell Biolabs, Inc.)
with a Matrigel-coated membrane (pre-coated at 4°C for 12 h) for
invasion assays or in chambers not coated with Matrigel for
migration assays, while the lower chamber was filled with complete
medium as a chemoattractant. After 24 h at 37°C, the cells on the
upper part of the filters were removed with a cotton swab. The
migrated and invasive cells were counted in 5 different fields
(magnification, ×200) under a light microscope (Nikon Corporation)
after staining with 0.5% crystal violet at 37°C for 15 min.
Dual luciferase reporter assay
To assess the relationship between CTGF and
miR-3064-5p (or LINC00585), the online tool LncTar (http://www.cuilab.cn/lnctar) (17) was used to predict the binding site
between CTGF 3′UTR (or LINC00585) and miR-3064-5p. Then, a
dual-luciferase reporter assay was carried out. First, CTGF 3′-UTR
was cloned into the downstream of the psiCheck2 vector (Promega
Corporation) to generate the wild-type CTGF 3′-UTR luciferase
reporter vector; a mutant CTGF 3′-UTR luciferase reporter vector
was generated by mutating the predicted miR-3064-5p binding site
within the CTGF 3′-UTR. These two reporter vectors were
cotransfected by Lipofectamine 2000 (Invitrogen; Thermo Fisher
Scientific, Inc.) with 50 pmol of miR-3064-5p mimics (Shanghai
GenePharma Co., Ltd.) into 5637 and T24 cells. Then, 48 h later,
the cells were harvested and underwent a dual-luciferase reporter
assay (Promega Corporation) to evaluate firefly and Renilla
the luciferase activities. Renilla luciferase activity
served as a normalization control.
RNA immunoprecipitation (RIP)
assay
The Immunoprecipitation Kit (cat. no. 17-10085; EMD
Millipore) was used for an RNA immunoprecipitation (RIP) assay
according to the manufacturer's instructions. Briefly, the lysed
cells (5×106) were incubated in 100 µl of RIP buffer
solution (cat. no. KT102-01; GZSCBio Co. Ltd.), and the magnetic
beads (included in the kit) were labeled with anti-Ago2 or IgG
antibodies (included in the kit) at 4°C for 2 h. The abundance of
LINC00858, miR-3064-5p, and CTGF were verified by RT-qPCR.
Determination of reactive oxygen
species (ROS)
Approximately 1×105 cells were seeded
into a 6-well plate and cultured at 37°C for 24 h. Then CellROX™
Green reagent (ThermoFisher Scientific, Inc.) was added at a 1/500
volume of medium and cultured for 1 h at room temperature. After
being washed with PBS buffer (Gibco; Thermo Fisher Scientific,
Inc.), cells were monitored under a fluorescence microscope
(Olympus BX43;Olympus Corporation) and the ROS content was
calculated according to the control.
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
Total RNA from tissues or target cells was extracted
using TRIzol reagent (Invitrogen; Thermo Fisher Scientific, Inc.),
following the manufacturer's procedures. A NanoDrop-1000 (Thermo
Fisher Scientific, Inc.) was used to determine concentrations and
for quality control purposes. Complementary DNA was synthesized
from extracted RNA (5 µg) using the Prime Script® RT
Reagent kit with gDNA Eraser (Invitrogen; Thermo Fisher Scientific,
Inc.) in accordance with the recommended protocol (samples and
reagent mix were incubated at 37°C for 15 min and 85°C for 5 sec,
and finally stored at 4°C). qPCR was performed using
SYBR®Premix Ex Taq™ II (Invitrogen; Thermo Fisher
Scientific, Inc.). An ABI 7500 real-time PCR detection system
(Applied Biosystems; Thermo Fisher Scientific, Inc.) was used for
detection. The qPCR thermocycling conditions were as follows: The
initial denaturation was first performed at 95°C for 2 min followed
by denaturation at 95°C for 15 sec and annealing and extension at
60°C for 30 sec. Then, denaturation, annealing and extension were
repeated for 40 cycles. After amplification, data were collected
and processed by the comparative cycle threshold method. U6 and
GAPDH expression levels were used as internal references for miRNA
and mRNA expression detection, respectively. Finally, the data were
processed using the 2−ΔΔCq relative expression method
(18). The PCR amplification products
were quantified by the FastStart Universal SYBR Green Master (Roche
Diagnostics) and normalized to GAPDH or U6.
Western blotting
Total protein was extracted from cultured cells
using RIPA buffer (Beyotime Biotechnology) containing protease and
phosphotase inhibitorcocktails. A BCA protein assay kit (Beyotime
Biotechnology) was used to determine protein concentrations
according to the manufacturer's instructions. Approximately 10 µg
of proteins were separated by 12% SDS-PAGE and transferred to PVDF
(EMD Millipore) membranes. The membranes were blocked with 5%
non-fat milk at room temperature for 1 hand then incubated with
primary antibodies overnight at 4°C, and subsequently incubated
with HRP-conjugated secondary antibodies (cat. no. A0216; dilution
1:5,000; Beyotime Biotechnology) at room temperature for 1 h.
Proteins were visualized using ECL Western blotting detection
reagent (EMD Millipore). Immunoreactive bands were quantified using
ImageJ (1.8.0; NIH). The following primary antibodies were used:
Anti-Cox-2 (product code ab179800; 1:1,000), anti-MMP2 (product
code ab92536; 1:2,000), anti-MMP9 (product code ab76003; 1:2,000),
anti-CTGF (product code ab209780; 1:1,000) and β-actin (product
code ab8226; 1:1,000) (all from Abcam).
Statistical analysis
All data are presented as the mean ± standard
deviation and all experiments were repeated three times.
Statistical significance between normal and tumor tissues was
analyzed using a paired t-test, and other comparisons between two
groups were analyzed by unpaired t-test. One-way ANOVA followed by
Tukey's post hoc test was performed for multiple comparisons using
SPSS version 21.0 software (IBM Corp.). Pearson's correlation
analysis was used to analyze the relationship between LINC00858 and
miR-3064-5p. P<0.05 was considered to indicate a statistically
significant difference.
Results
LINC00858 is increased in bladder
cancer tissues and the cancer cell lines
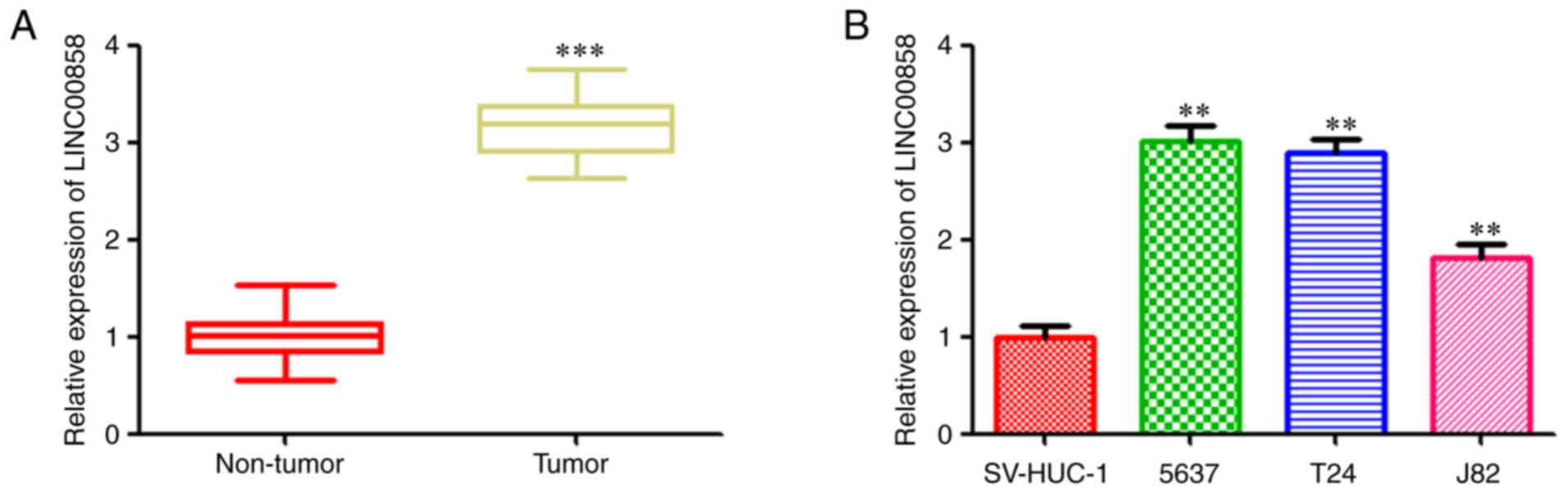
As revealed in Fig.
1A, RT-qPCR analysis revealed that the RNA expression level of
LINC00858 was increased in the bladder cancer tissues compared with
that in the paired adjacent normal tissues. Consistently with the
findings in tissues, LINC00858 was revealed to be upregualted in
three bladder cancer cell lines (5673, T24 and J82) compared with
the normal bladder cells (Fig. 1B).
All these data indicated a positive relationship between LINC00858
and bladder cancer progression.
Knockdown of LINC00858 suppresses
bladder cancer cell proliferation, migration and invasion
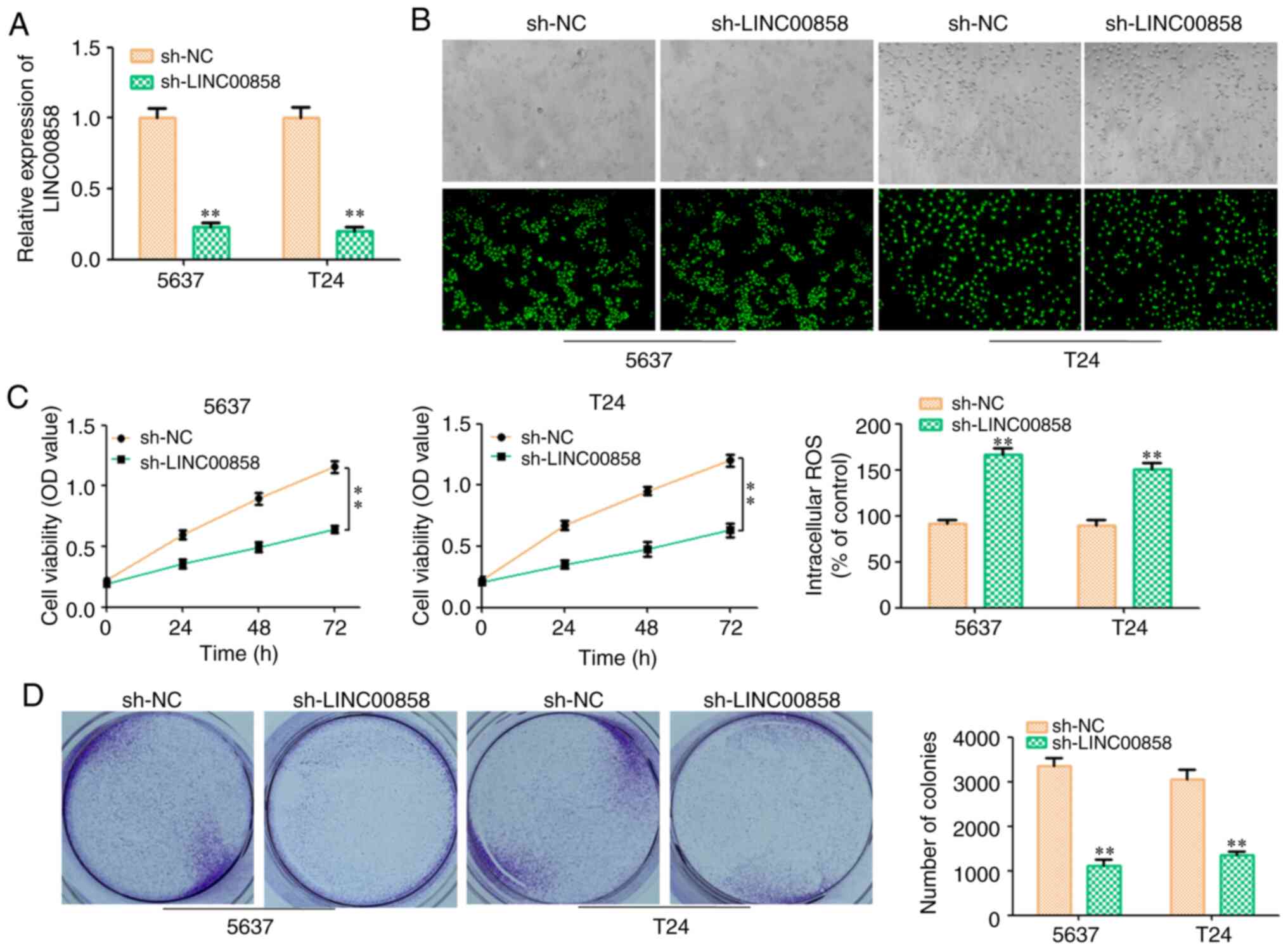
To investigate the role of LINC00858 in bladder
cancer, the specific shRNA targeting LINC00858 was used to knock
down the expression level of LINC00858 both in 5637 and T24 bladder
cancer cells. The transfection efficiency is presented in Fig. 2A and B. CCK-8 and colony formation
assays revealed that knockdown of LINC00858 inhibited the
proliferation of 5637 and T24 cells (Fig.
2C and D). It has been previously reported that the cell
apoptosis of bladder cancer was induced by reactive oxygen species
(ROS) (19). In the present study, it
was revealed that the content of ROS was increased in the
sh-LINC00858 group (Fig. 2C).
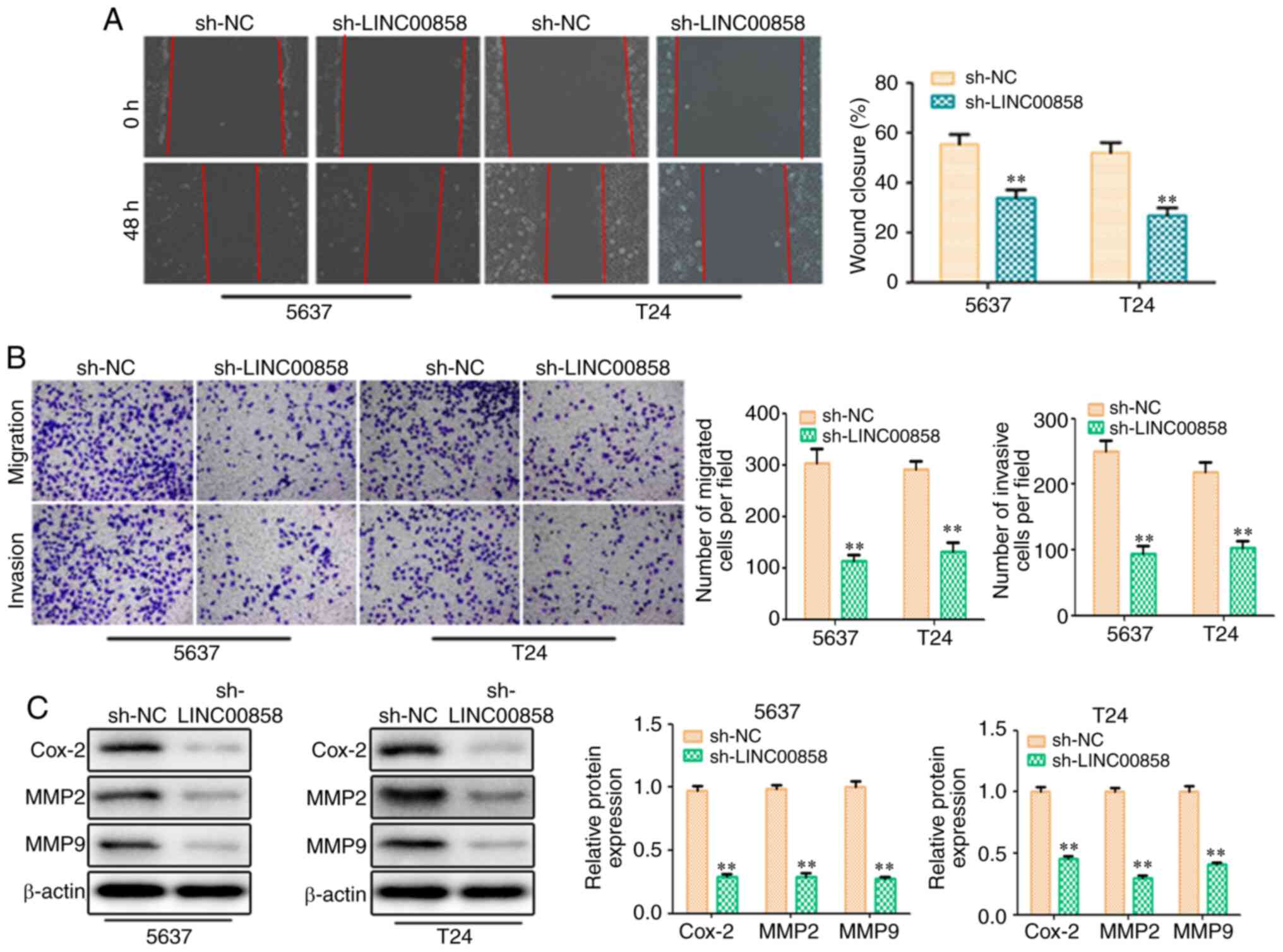
Furthermore, Transwell migration and invasion assays were conducted
to determine whether LINC00858 is involved in the regulation of
bladder cell migration and invasion. In the present study,
knockdown of LINC00858 significantly inhibited bladder cancer cell
migration and invasion as evidenced by the wound healing and
Transwell chamber assays (Fig. 3A and
B). At the molecular level, the expression levels of
migration/invasion-associated proteins, including Cox-2, MMP2 and
MMP9, were markedly decreased in response to LINC00858 knockdown
(Fig. 3C). These findings indicated
that LIN00858 may exert an oncogenic action on the aggressiveness
of bladder cells in vitro.
LINC00858 acts as a sponge for
miR-3064-5p in bladder cancer cells
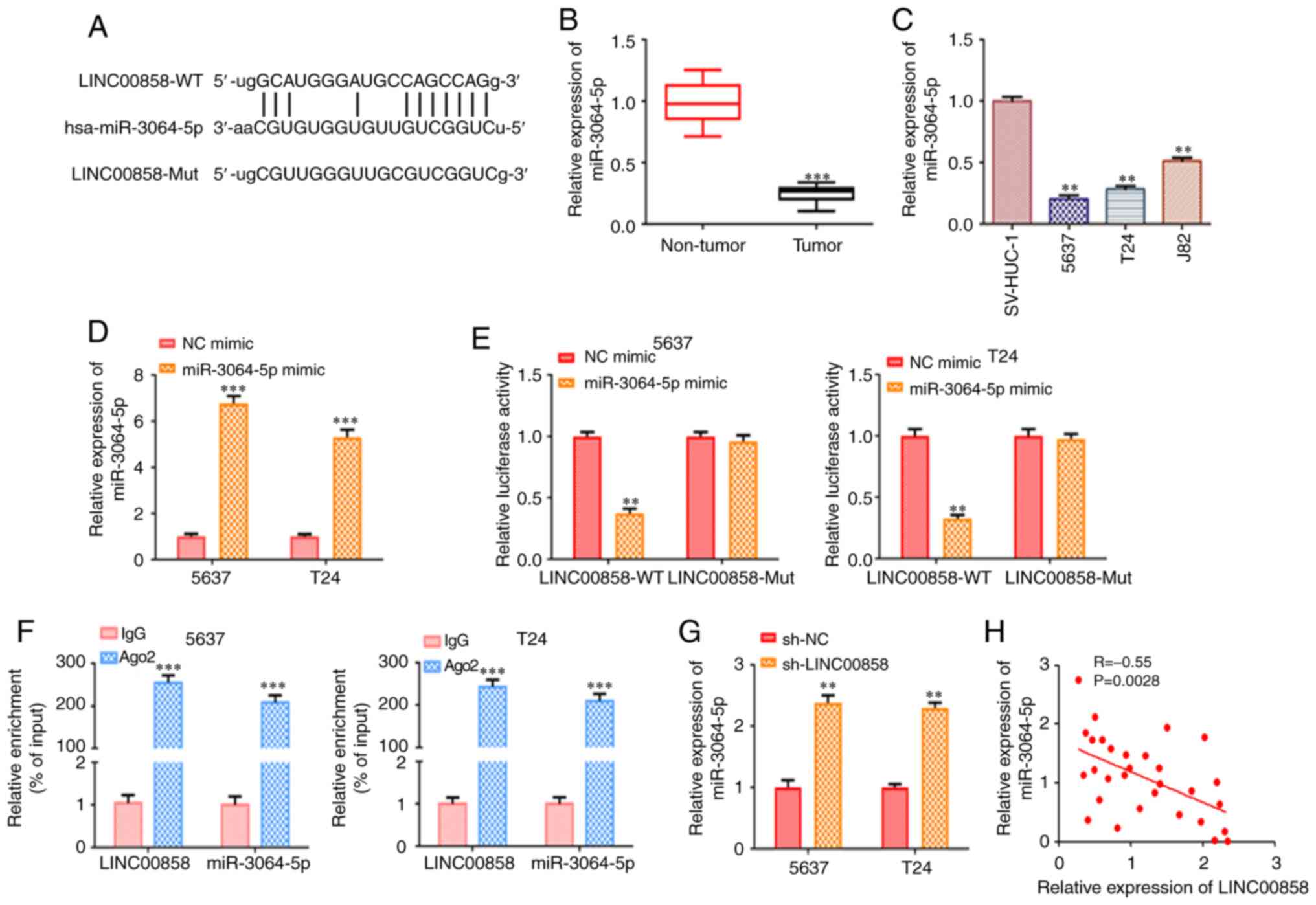
As revealed in Fig.
4A, bioinformatics analysis using starBase v2.0 indicated that
miR-3064-5p was a potential downstream target of LINC00858. RT-qPCR
analysis revealed that miR-3064-5p was expressed at a low level in
bladder cancer tissues and cells compared with normal samples and
cells (Fig. 4B and C). A previous
study has revealed that lncRNAs may function as ceRNAs by sponging
miRNAs (20).
To validate true functional binding between
LINC00858 and miR-3064-5p, luciferase reporter assays were
performed. The mimics of miR-3064-5p were transfected into 5637 and
T24 cells for miR-3064-5p overexpression, as confirmed by RT-qPCR
(Fig. 4D). The luciferase reporter
assay results revealed that miR-3064-5p mimics transfected into
5637 and T24 cells significantly reduced the activity of the
luciferase reporter, while no significant alterations were observed
in the luciferase activity of mutant LINC00858 (Fig. 4E). Notably, RIP assays revealed that
the expression levels of LINC00858 and miR-3064-5p in Ago2 RIP were
significantly higher than those in IgG RIP, suggesting the direct
binding of these two RNAs (Fig. 4F).
Moreover, based on the results of the RT-qPCR assays, the relative
levels of miR-3064-5p were notably increased due to silencing of
LINC00858 in 5637 and T24 cells (Fig.
4G). In addition, Pearson's correlation analysis indicated that
miR-3064-5p was negatively correlated with LINC00858 (Fig. 4H).
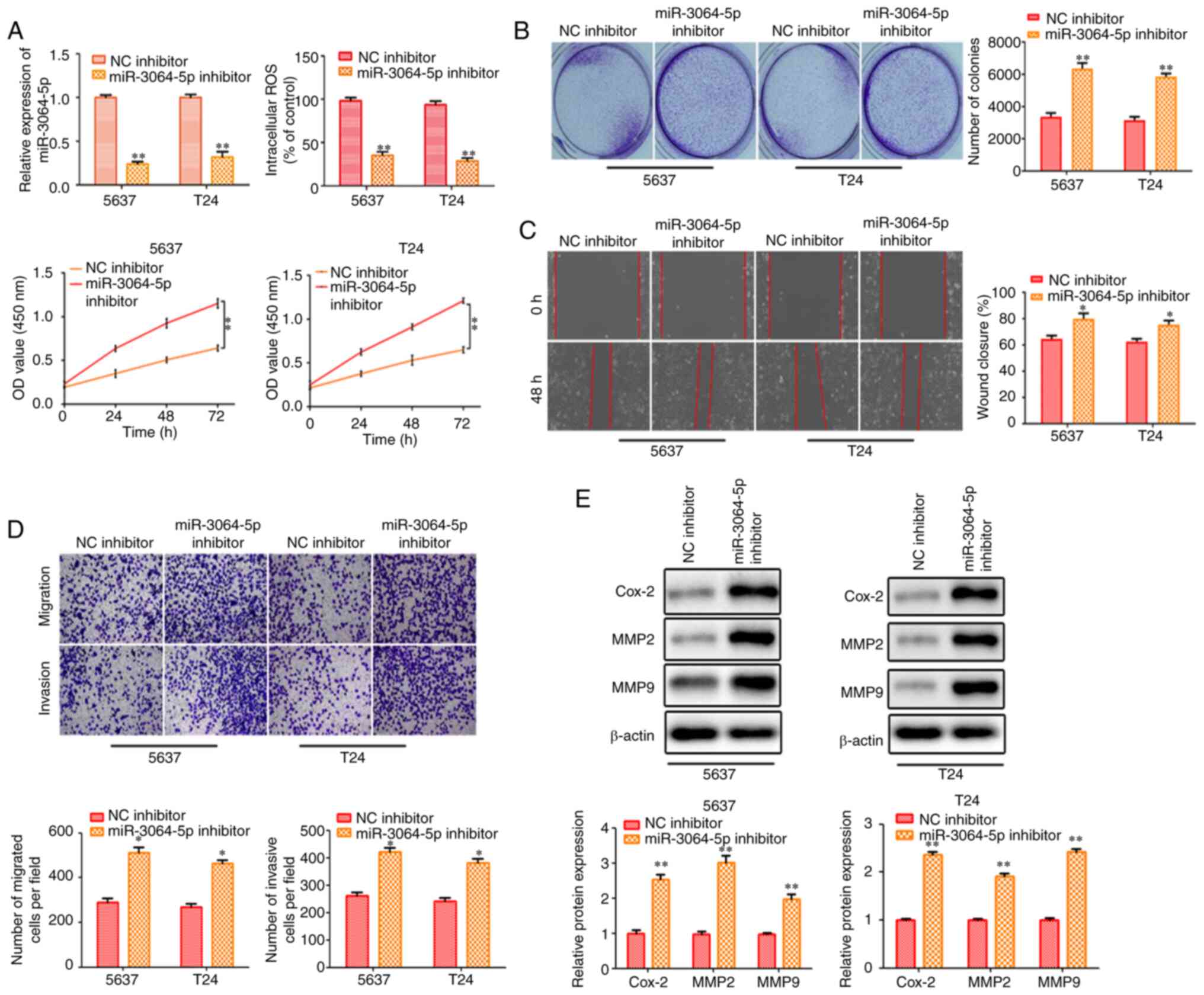
Inhibition of miR-3064-5p contributes
to cell proliferation, migration and invasion of bladder cancer
cells
The aforementioned data clearly demonstrated that
LINC00858 directly interacted with miR-3064-5p. Thus, it was
further investigated whether the LINC00858/miR-3064-5p axis
modulated bladder cancer. 5637 and T24 cells were co-transfected
with miR-3064-5p inhibitor and sh-LINC00858 (Fig. 5A). As revealed in Fig. 5A, the cell viability decreased by
LINC00858 knockdown was rescued by the miR-3064-5p inhibitor, while
increased ROS contents were reversed by miR-3064-5p inhibitor.
miR-3064-5p inhibitor also rescued the effect of LINC00858
knockdown on bladder cancer cells as revealed inthe colony
formation assay (Fig. 5B).
Furthermore, the wound healing assay and the Transwell assays
revealed that the migration and invasion of bladder cancer cells
inhibited by sh-LINC00858 were increased by miR-3064-5p inhibitor.
Cox-2, MMP2 and MMP9 protein levels that were significantly
decreased by LINC00858 knockdown, were increased by miR-3064-5p
inhibition. The effects of sh-LINC00858 on migration and invasion
were partially attenuated by miR-3064-5p inhibitor (Fig. 5C-E).
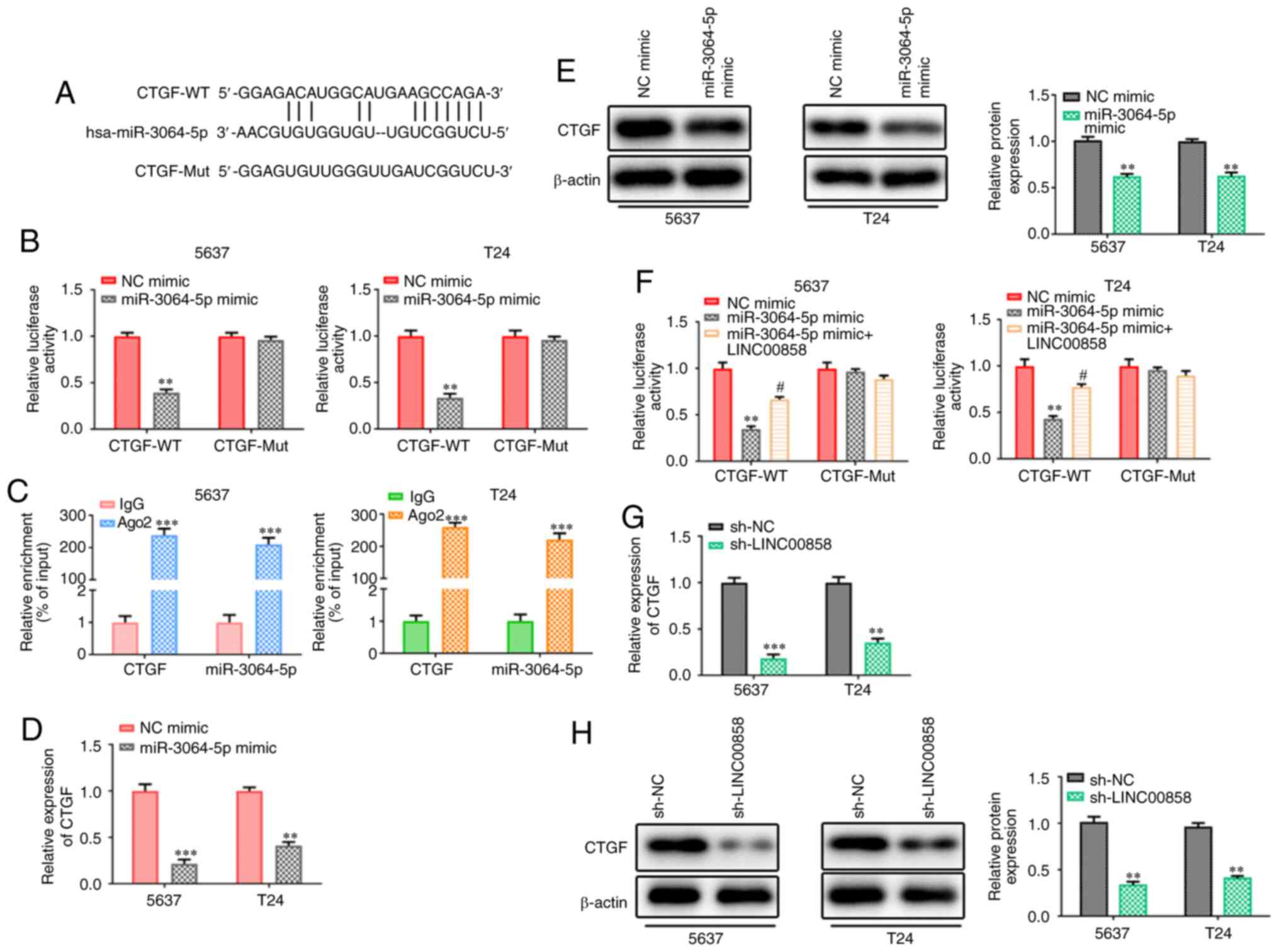
LINC00858 modulates CTGF via
competitive binding with miR-3064-5p
Using starBase v2.0, it was determined that CTGF was
a downstream target gene of miR-3064-5p and the potential binding
sequences are presented in Fig. 6A.
Reporter gene assays in 5637 and T24 cells revealed that
overexpression of miR-3064-5p led to a notable decrease in the
transcriptional activity of CTGF 3′-UTR promoter constructs, while
no significant changes were observed in the CTGF-Mut group
(Fig. 6B). RIP assays suggested that
miR-3064-5p binded to CTGF (Fig. 6C).
Similar results of CTGF expression pattern at the mRNA and protein
levels were observed and confirmed by RT-qPCR and western blot
analyses (Fig. 6D and E). Notably,
the results revealed that the expression levels of CTGF were
markedly impeded by miR-3064-5p overexpression. In addition, it was
revealed that the miR-3064-5p-induced inhibitory effects on CTGF
transcription could be partially restored by overexpression of
LINC00858, indicating that LINC00858 competes with CTGF for
miR-3064-5p (Fig. 6F). Finally,
RT-qPCR and western blotting revealed that the expression levels of
CTGF were also markedly impeded by knockdown of LINC00858 (Fig. 6G and H).
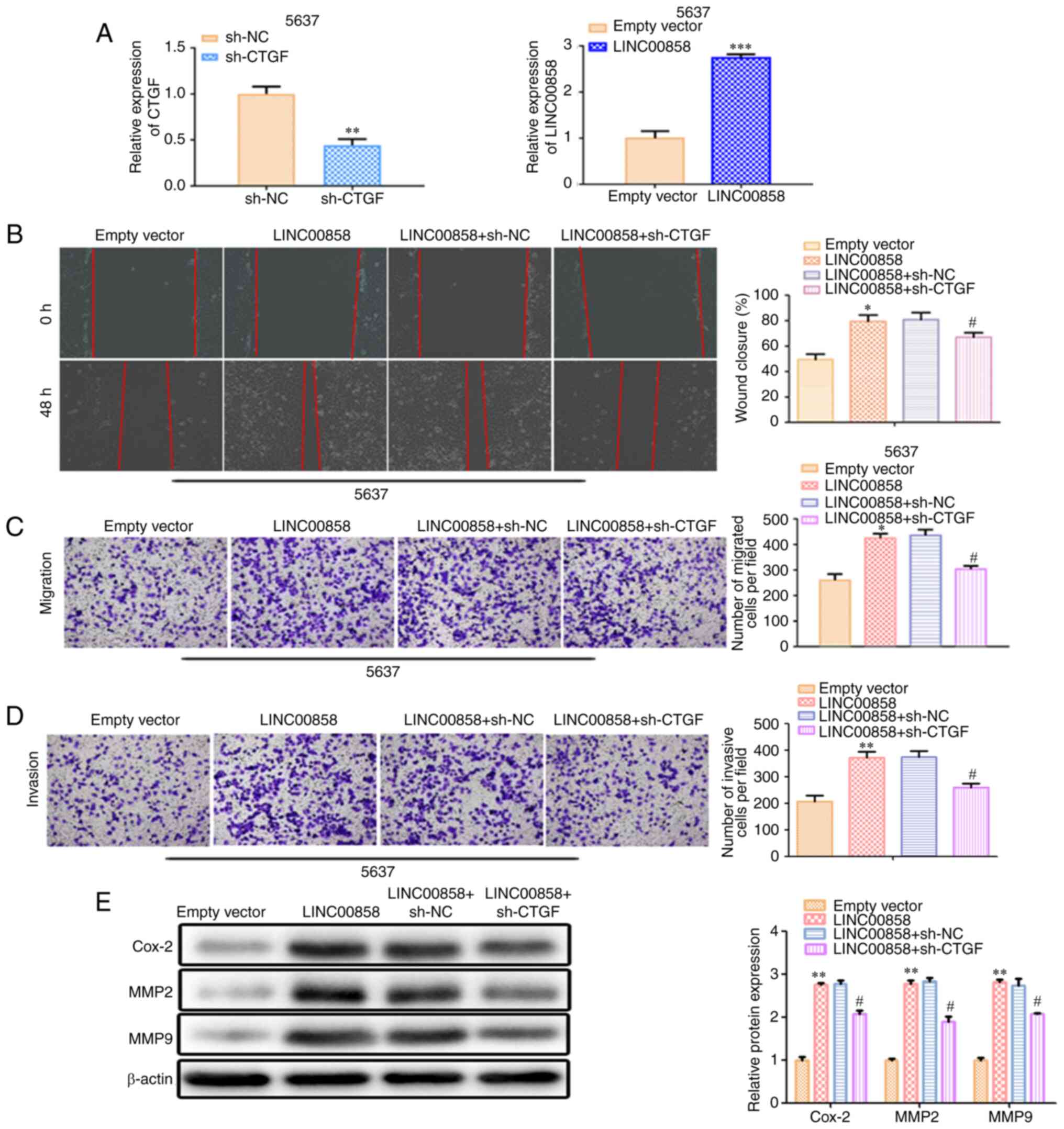
LINC00858/miR-3064-5p/CTGF axis in
bladder cancer
To further study the role of
LINC00858/miR-3064-5p/CTGF axis in bladder cancer, rescue assays
were performed. As revealed in Fig.
7A, the transfection efficiency in 5637 was confirmed by
RT-qPCR. Wound healing analysis and Transwell analysis revealed
that overexpression of LINC00858 promoted cell migration and
invasion, while knockdown of CTGF inhibitedcell migration and
invasion (Fig. 7B-D). Moreover,
western blot analysis revealed that overexpression of LINC00858
resulted in the increased expression levels of Cox-2, MMP2 and MMP9
and the effects of LINC00858 overexpression were counteracted by
silencing of CTGF (Fig. 7E).
Discussion
With the advancement of RNA-Seq technologies,
lncRNAs are closely associated with bladder cancer (21). Previous studies have revealed that
LINC00858, a novel lncRNA, was upregulated in several cancers and
functioned as a tumor promoter in colorectal cancer, non-small cell
lung cancer and osteosarcoma (8,15,22–24).
LINC00858 promoted cell proliferation, migration and invasion by
acting as a ceRNA of microRNA in the aforementioned cancers. In the
present study, it was revealed that LINC00858 was aberrantly highly
expressed in bladder cancer tissues and cells. Mechanism studies
revealed that LINC00858 could increase the expression level of CTGF
in bladder cancer cells by combining with miR-3064-5p. For the
first time, to the best of our knowledge, direct evidence was
provided that LINC00858 acts as an oncogene in bladder cancer and
promotes the growth and metastasis of bladder cancer cells.
In addition, lncRNAs have also been revealed to act
as diagnostic and prognostic biomarkersin bladder cancer (25). For instance, Shan et al have
reported that lncRNA NEAT1 promoted bladder progression by
regulating miR-410-mediated HMGB1 (23). Liu and Wu demonstrated that
lncRNA NNT-AS1 enhanced bladder cancer cell growth by targeting the
miR-1301-3p/PODXL axis and activating the Wnt pathway (24), and in 2019, Fang et al revealed
that DLX6-AS1 promoted cell growth and invasiveness in bladder
cancer by modulating the miR-223-HSP90B1 axis (25).
Functionally, some lncRNAs contain miRNA-binding
elements and act as ceRNAs, suppressing miRNA activities. In the
present study, it was revealed using a bioinformatics database that
miR-3064-5p may be a target of LINC00858. miR-3064-5p was revealed
to be downregulated in cancer tissues and cells in the present
study. A recent study indicated that miR-3064-5p served as a tumor
suppressor in gastric cancer (26).
Herein, for the first time, to the best of our knowledge, it was
revealed that miR-3064-5p reversed the effects of LINC00858 on
bladder cancer cells.
CTGF, encoded within chromosomal 6q23.2, has been
reported to be a potential oncogene in cancer (27–29). CTGF
played a crucial role for osteolytic bone metastasis both by
enhancing invasiveness of tumor cells and producing RANKL for
osteoclastogenesis (30). In
addition, CTGF has been confirmed as a potential prognostic marker
for medulloblastoma (31). With
regard to the present research, it was revealed that CTGF was a
direct target of miR-3064-5p. Moreover, it was also revealed that
the expression level of CTGF was significantly decreased by
knockdown of LINC00858. Knockdown of CTGF reversed the effects of
LINC00858 overexpression on bladder cancer cells. Therefore, CTGF
mediated the oncogenic role of LINC00858 in the development of
bladder cancer.
In summary, the present study demonstrated that
knockdown of LINC00858 suppressed the progression andmetastasis of
bladder cancer cells in vitro. In terms of the mechanism,
LINC00858 functioned as a ceRNA by regulating the expression level
of CTGF by sponging miR-3064-5p.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
All data generated or analysed during this study are
included in this published article.
Authors' contributions
JH, WMZ and XHT conceived the study and revised the
manuscript. QMH and CH conducted all the experiments and drafted
the manuscript. QW and GXW interpreted and analyzed the data. All
authors reviewed, read, and approved this manuscript and agree to
be accountable for all aspects of the research in ensuring that the
accuracy or integrity of any part of the work are appropriately
investigated and resolved.
Ethics approval and consent to
participate
The present study was approved by the Ethics
Committee of Jiangxi Cancer Hospital (approval no. 2019014).
Written informed consent was obtained from all participants.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Ferlay J, Soerjomataram I, Dikshit R, Eser
S, Mathers C, Rebelo M, Parkin DM, Forman D and Bray F: Cancer
incidence and mortality worldwide: Sources, methods and major
patterns in GLOBOCAN 2012. Int J Cancer. 136:E359–E386. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Hedegaard J, Lamy P, Nordentoft I, Algaba
F, Høyer S, Ulhøi BP, Vang S, Reinert T, Hermann GG, Mogensen K, et
al: Comprehensive transcriptional analysis of early-stage
urothelial carcinoma. Cancer Cell. 30:27–42. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Antoni S, Ferlay J, Soerjomataram I, Znaor
A, Jemal A and Bray F: Bladder cancer incidence and mortality: A
global overview and recent trends. Eur Urol. 71:96–108. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Renganathan A and Felley-Bosco E: Long
noncoding RNAs in cancer and therapeutic potential. Adv Exp Med
Biol. 1008:199–222. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Lenis AT, Lec PM, Chamie K and Mshs MD:
Bladder cancer: A review. JAMA. 324:1980–1991. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Luo D, Deng B, Weng M, Luo Z and Nie X: A
prognostic 4-lncRNA expression signature for lung squamous cell
carcinoma. Artif Cells Nanomed Biotechnol. 46:1207–1214. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Yang G, Lu X and Yuan L: lncRNA: A link
between RNA and cancer. Biochim Biophys Acta. 1839:1097–1109. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Sha QK, Chen L, Xi JZ and Song H: Long
non-coding RNA LINC00858 promotes cells proliferation, migration
and invasion by acting as a ceRNA of miR-22-3p in colorectal
cancer. Artif Cells Nanomed Biotechnol. 47:1057–1066. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Shi T, Gao G and Cao Y: Long noncoding
RNAs as novel biomarkers have a promising future in cancer
diagnostics. Dis Markers. 2016:90851952016. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Toiyama Y, Okugawa Y and Goel A: DNA
methylation and microRNA biomarkers for noninvasive detection of
gastric and colorectal cancer. Biochem Biophys Res Commun.
455:43–57. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Bhan A, Soleimani M and Mandal SS: Long
noncoding RNA and cancer: A new paradigm. Cancer Res. 77:3965–3981.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Martens-Uzunova ES, Böttcher R, Croce CM,
Jenster G, Visakorpi T and Calin GA: Long noncoding RNA in
prostate, bladder, and kidney cancer. Eur Urol. 65:1140–1151. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Zhang X, Gejman R, Mahta A, Zhong Y, Rice
KA, Zhou Y, Cheunsuchon P, Louis DN and Klibanski A: Maternally
expressed gene 3, an imprinted noncoding RNA gene, is associated
with meningioma pathogenesis and progression. Cancer Res.
70:2350–2358. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Wei X, Yang X, Wang B, Yang Y, Fang Z, Yi
C, Shi L and Song D: lncRNA MBNL1-AS1 represses cell proliferation
and enhances cell apoptosis via targeting miR-135a-5p/PHLPP2/FOXO1
axis in bladder cancer. Cancer Med. 9:724–736. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Xue M, Shi D, Xu G and Wang W: The long
noncoding RNA linc00858 promotes progress of lung cancer through
miR-3182/MMP2 axis. Artif Cells Nanomed Biotechnol. 47:2091–2097.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Li JH, Liu S, Zhou H, Qu LH and Yang JH:
starBase v2.0: Decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA
interaction networks from large-scale CLIP-Seq data. Nucleic Acids
Res. 42((Database Issue)): D92–D97. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Li J, Ma W, Zeng P, Wang J, Geng B, Yang J
and Cui Q: LncTar: A tool for predicting the RNA targets of long
noncoding RNAs. Brief Bioinform. 16:806–812. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Takeuchi H, Taoka R, Mmeje CO, Jinesh GG,
Safe S and Kamat AM: CDODA-Me decreases specificity protein
transcription factors and induces apoptosis in bladder cancer cells
through induction of reactive oxygen species. Urol Oncol.
34:337.e11–e18. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Tay Y, Rinn J and Pandolfi PP: The
multilayered complexity of ceRNA crosstalk and competition. Nature.
505:344–352. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Zhu W, Liu H, Wang X, Lu J and Yang W:
Long noncoding RNAs in bladder cancer prognosis: A meta-analysis.
Pathol Res Pract. 215:1524292019. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Quan J, Pan X, Zhao L, Li Z, Dai K, Yan F,
Liu S, Ma H and Lai Y: lncRNA as a diagnostic and prognostic
biomarker in bladder cancer: A systematic review and meta-analysis.
Onco Targets Ther. 11:6415–6424. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Shan G, Tang T, Xia Y and Qian HJ: Long
non-coding RNA NEAT1 promotes bladder progression through
regulating miR-410 mediated HMGB1. Biomed Pharmacother.
121:1092482020. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Liu Y and Wu G: NNT-AS1 enhances bladder
cancer cell growth by targeting miR-1301-3p/PODXL axis and
activating Wnt pathway. Neurourol Urodyn. 39:547–557. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Fang C, Xu L, He W, Dai J and Sun F: Long
noncoding RNA DLX6-AS1 promotes cell growth and invasiveness in
bladder cancer via modulating the miR-223-HSP90B1 axis. Cell Cycle.
18:3288–3299. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Sun X, Zhang X, Zhai H, Zhang D and Ma S:
A circular RNA derived from COL6A3 functions as a ceRNA in gastric
cancer development. Biochem Biophys Res Commun. 515:16–23. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Yang L, Hou J, Cui XH, Suo LN and Lv YW:
miR-133b regulates the expression of CTGF in epithelial-mesenchymal
transition of ovarian cancer. Eur Rev Med Pharmacol Sci.
21:5602–5609. 2017.PubMed/NCBI
|
|
28
|
Alam KJ, Mo JS, Han SH, Park WC, Kim HS,
Yun KJ and Chae SC: MicroRNA 375 regulates proliferation and
migration of colon cancer cells by suppressing the CTGF-EGFR
signaling pathway. Int J Cancer. 141:1614–1629. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Lun W, Wu X, Deng Q and Zhi F: miR-218
regulates epithelial-mesenchymal transition and angiogenesis in
colorectal cancer via targeting CTGF. Cancer Cell Int. 18:832018.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Kim B, Kim H, Jung S, Moon A, Noh DY, Lee
ZH, Kim HJ and Kim HH: A CTGF-RUNX2-RANKL axis in breast and
prostate cancer cells promotes tumor progression in bone. J Bone
Miner Res. 35:155–166. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Cruzeiro GAV, Lira RCP, de Almeida
Magalhães T, Scrideli CA, Valera ET, Baumgartner M and Tone LG:
CTGF expression is indicative of better survival rates in patients
with medulloblastoma. Cancer Gene Ther. 27:378–382. 2020.
View Article : Google Scholar : PubMed/NCBI
|