Introduction
Keloids are benign fibrotic tumors of the dermis
that form during a prolonged wound healing process (1). Keloids exhibit aggressive dermal growth
beyond the boundaries of the original margins of wounds, causing
pain, pruritis and contractures. They are a cosmetic and
psychological burden to patients (2).
The effect of one type of glucocorticoid,
hydrocortisone (HC), on keloids has previously been investigated.
HC is known to alter the chemistry and morphology of connective
tissue cells and to hinder the production of intercellular
substances, including collagenous fiber and glycosaminoglycan
(3). It may also decrease the
maximum density of keloid-derived fibroblasts (4), increase proline transport (5), lower prolyl hydroxylase activity,
reduce the rate of collagen synthesis (6), enhance apoptosis rates (7) and diminish hyaluronan accumulation
(8).
Microarray studies identify a broad spectrum of
differentially regulated genes in biological samples under genomic
sequencing, cloning, cDNA or PCR approaches (9). In 2008, Smith et al (10) conducted a bioinformatic analysis on
RNA obtained from fibroblasts cultured from normal scars and
keloids grown in the absence and presence of HC. The results
indicated that there was elevated expression of a number of
insulin-like growth factor (IGF)-binding and IGF-binding related
proteins, in addition to decreased expression of a set of Wnt
pathway inhibitors and numerous interleukin (IL)-1-inducible genes.
Furthermore, it was observed that IGF binding protein (IGFBP)-3 and
connective tissue growth factor (CTGF) were associated with the
increase of fibroblast and collagen deposition and were
overexpressed in keloid fibroblasts only in the presence of HC,
suggesting that glucocorticoid resistance of HC is involved in the
pathogenesis of keloids formation (10). However, the adverse effects of HC
during its treatment on keloids and the drugs that may potentially
be used to weaken or reverse these adverse effects, remain
unknown.
The present study used the microarray data collected
by Smith et al (10) to
identify differentially expressed genes (DEGs) in fibroblasts
cultured from keloids treated with or without HC. A Gene Ontology
(GO) enrichment analysis was performed on DEGs, which were
potentially associated with the positive efficacy of HC, and a
pathway enrichment analysis for DEGs that may be associated with
adverse effects of HC was completed. Furthermore, the adverse
effects of HC were analyzed and small molecule drugs that may
reduce the occurrence of adverse effects were screened from the
connectivity map (CMAP) database. The aim of the present study was
to elucidate the molecular mechanisms of HC treatment on keloids
and provide novel information for the clinical treatment of this
disease.
Materials and methods
Affymetrix microarray data
The gene expression profile data of GSE7890
(10) was downloaded from the Gene
Expression Omnibus (GEO; http://www.ncbi.nlm.nih.gov/geo/) database, which was
based on the platform of GPL570 [HG-U133_Plus_2] Affymetrix Human
Genome U133 Plus 2.0 Array (Affymetrix Inc., Santa Clara, CA, USA).
A total of 19 primary cultures of dermal fibroblast samples were
included in this dataset, including 4 samples from normal scar
tissue and 5 samples from keloids treated with 1.5 µM HC; as well
as 5 samples from normal scar and 5 samples from keloids that did
not undergo HC treatment.
Affymetrix CEL files and the probe annotation files
were downloaded and the gene expression data of all samples were
preprocessed via background correction, quantile normalization and
probe summarization using the Affy software (version 1.30.0)
package of Bioconductor (available at http://www.bioconductor.org/packages/2.8/bioc/html/affy.html)
(11).
DEGs screening
Linear Models for Microarray Data package (version
3.22.7) (12) of Bioconductor
(available at http://www.bioconductor.org/packages/3.0/bioc/html/limma.html)
was used to identify genes that were differentially expressed in
dermal fibroblasts from keloids and those from normal scars. The
raw P-value was adjusted into False Discovery Rate (FDR) by the
Bonferroni method (13) in a
multtest package (version 2.20.0) of Bioconductor (http://www.bioconductor.org/packages/2.14/bioc/html/multtest.html).
The |log2FC (fold change)|>1 and FDR <0.05 were
selected as the cut-off criteria.
Functional classification of DEGs
GO functional enrichment analysis of DEGs was
performed to interpret their biological significance, via the
Database for Annotation, Visualization and Integrated Discovery
(http://david.abcc.ncifcrf.gov/)
(14). P<0.01 was used as the
cut-off criterion.
Pathway enrichment analysis of DEGs was performed
using a Kegg Orthology Based Annotation System, version 2.0
(http://kobas.cbi.pku.edu.cn) (15). P<0.05 was used as the cut-off
criterion.
Identification of potential diseases
associated with DEGs
Potential diseases associated with the DEG genesets
A and C were identified using the WEB-based GEne SeT AnaLysis
Toolkit (WebGestalt; www.webgestalt.org) (16) with a cut-off criterion of
P<0.05.
Prediction of small molecule
drugs
CMAP database (www.connectivitymap.org/cmap) (17) was used to predict candidate small
molecule drugs targeting DEGs genesets A and C. An enrichment score
between 0.9–1 was selected as the cut-off criterion. The closer to
−1 the enrichment score was, the stronger the effect of the drug on
the disease.
Results
Identification of DEGs
A total of 192 DEGs with HC treatment (50
upregulated and 142 downregulated) and 67 DEGs without HC treatment
(15 upregulated and 52 downregulated) were screened from dermal
fibroblasts in keloids and compared with normal scar tissue.
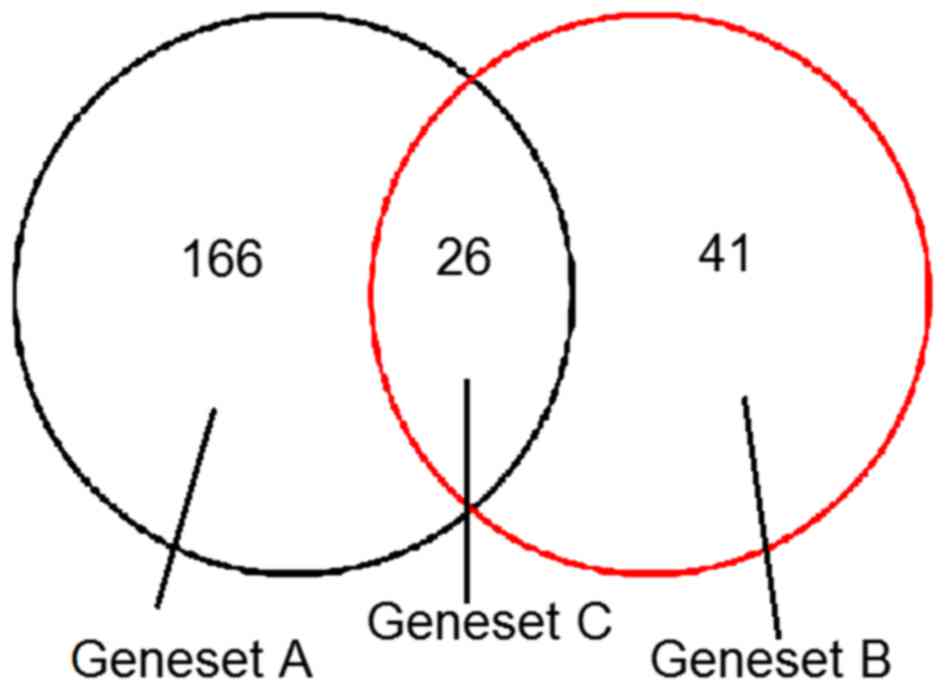
Among them, 166 DEGs were only present in the dermal
fibroblasts from keloids treated with HC (geneset A), 41 DEGs were
only present in dermal fibroblasts from keloids with no HC
treatment (geneset B) and 26 DEGs were common in dermal fibroblasts
from keloids with and without HC treatment (geneset C; Fig. 1).
Functional annotation of DEG
To investigate the potential molecular functions in
keloid fibroblasts affected by HC, a GO functional enrichment
analysis of the DEGs in geneset B, which were potentially
associated with the positive efficacy of HC, was performed.
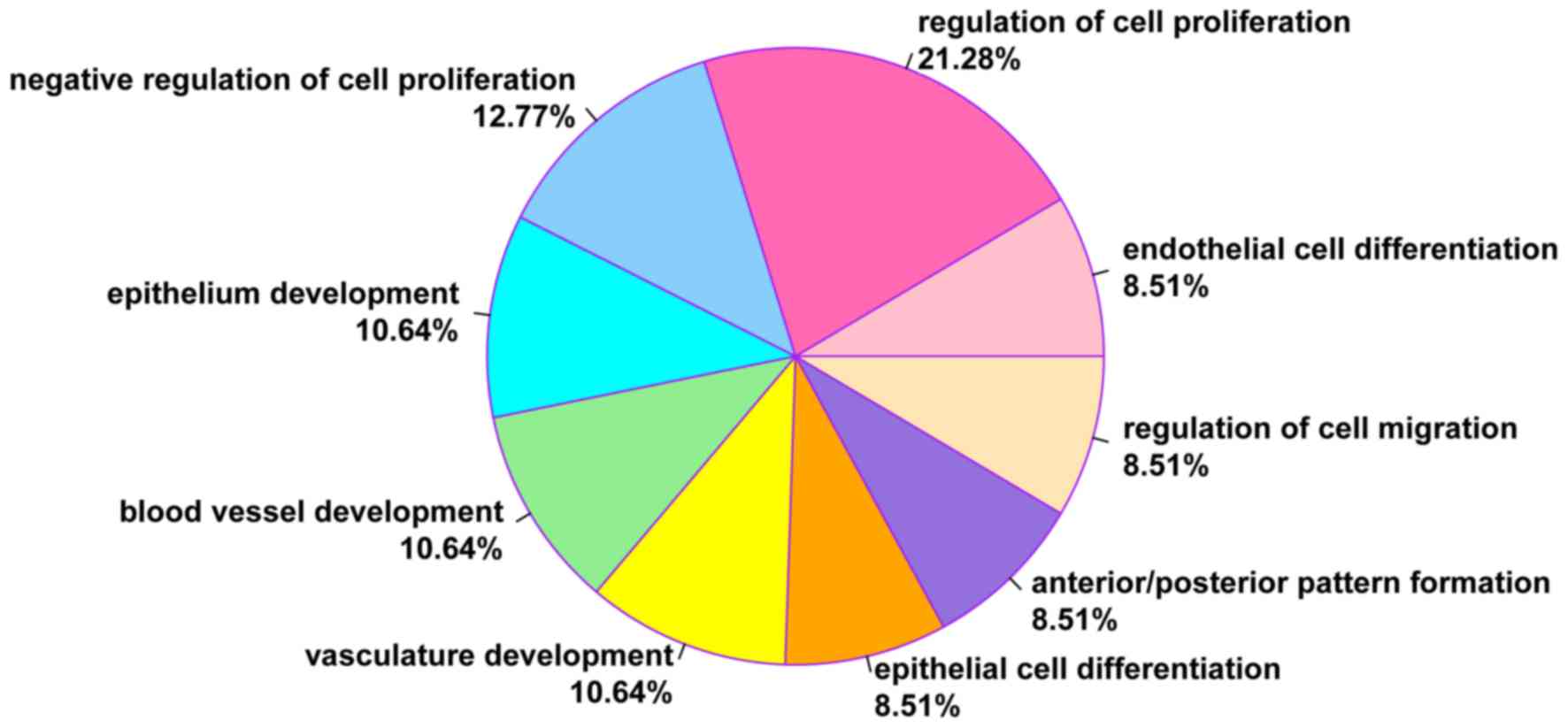
According to the GO functional enrichment analysis, DEGs in geneset
B were significantly enriched in 9 GO terms (all P<0.01),
including endothelial cell differentiation (COL18A1,
HOXB5 and JAG1) and regulation of cell proliferation
(DLC1, COL18A1 and JAG1; Table I). Furthermore, 21.28% of DEGs were
associated with the regulation of cell proliferation and 8.51% of
DEGs with endothelial cell differentiation (Fig. 2).
 | Table I.Enriched GO terms for differentially
expressed genes in geneset B. |
Table I.
Enriched GO terms for differentially
expressed genes in geneset B.
| Term | Description | Count | P-value | Genes |
|---|
| GO:0045446 | Endothelial cell
differentiation | 4 |
P<0.001a | COL18A1,
HOXB5, JAG1, NR2F2 |
| GO:0042127 | Regulation of cell
proliferation | 10 |
P<0.001a | DLC1,
COL18A1, NCK2, TBX3, NKX3-1,
PTN, JAG1, PPAP2A, CLEC11A,
IGFBP5 |
| GO:0008285 | Negative regulation
of cell proliferation | 6 |
0.001937a | DLC1,
COL18A1, NCK2, NKX3-1, PPAP2A,
IGFBP5 |
| GO:0060429 | Epithelium
development | 5 |
0.002412a | DLC1,
COL18A1, HOXB5, JAG1, NR2F2 |
| GO:0001568 | Blood vessel
development | 5 |
0.003177a | COL18A1,
TBX3, MMP19, JAG1, NR2F2 |
| GO:0001944 | Vasculature
development | 5 |
0.003465a | COL18A1,
TBX3, MMP19, JAG1, NR2F2 |
| GO:0030855 | Epithelial cell
differentiation | 4 |
0.004833a | COL18A1,
HOXB5, JAG1, NR2F2 |
| GO:0009952 | Anterior/posterior
pattern formation | 4 |
0.005134a | TBX3,
HOXB5, NKX3-1, NR2F2 |
| GO:0030334 | Regulation of cell
migration | 4 |
0.008627a | DLC1,
COL18A1, JAG1, IGFBP5 |
To identify the potential dysregulation pathways
associated with HC treatment and those that were unaffected by HC,
a pathway enrichment analysis of the DEGs in geneset A
(specifically, novel DEGs following HC treatment) and genes in
geneset C (namely, genes that were not affected by HC), was
performed. Based on this pathway enrichment analysis, the DEGs in
genesets A and C were significantly enriched in two pathways: Cell
cycle (CCNB1, MAD2L1 and BUB1) and p53
signaling pathway (CCNB1, RRM2 and PERP;
Table II; P<0.05).
 | Table II.Results of pathway enrichment
analysis for differentially expressed genes in genesets A and
C. |
Table II.
Results of pathway enrichment
analysis for differentially expressed genes in genesets A and
C.
| Term | Description | Count | P-value | Genes |
|---|
| hsa04110 | Cell cycle | 9 |
4.29E-05a | CCNB1,
MAD2L1, CCNB2, DBF4, BUB1, TTK,
CDC20, CDC25C, CCNA2 |
| hsa04115 | p53 signaling
pathway | 5 |
0.005598a | CCNB1,
CCNB2, RRM2, PERP, PTEN |
Potential diseases associated with
DEGs in genesets A and C
To investigate the adverse effects of HC during its
treatment on keloids, potential diseases associated with DEGs in
genesets A and C were identified. It was determined that the newly
added DEGs in dermal fibroblasts from keloids following HC
treatment and genes that were not altered by HC were associated
with two diseases: Bone loss (IL1R1, HSPA2 and
COL1A1) and osteoarthritis (IL1R1, IGFBP7 and
COL1A1; Table III).
 | Table III.Potential diseases associated with
differentially expressed genes in genesets A and C. |
Table III.
Potential diseases associated with
differentially expressed genes in genesets A and C.
| Disease | P-value | Genes |
|---|
| Bone loss |
0.002593a | IL1R1,
HSPA2, COL1A1 |
| Osteoarthritis |
0.033811a | IL1R1,
IGFBP7, COL1A1, CALM1 |
Small molecule drugs targeting DEGs in
genesets A and C
To further investigate potential drugs that are able
to weaken or eliminate the adverse effects of HC, small molecule
drugs targeting DEGs in genesets A and C were identified. A total
of 9 small molecule drugs were selected from the CMAP database,
including acemetacin, scriptaid and alsterpaullone. Among them,
acemetacin has a negative score and the lowest P-value (P=0.00002),
indicating it had the strongest effect on the adverse effects of HC
on gene expression (Table IV).
 | Table IV.Small molecule drugs potentially used
to treat the diseases associated with differentially expressed
genes in genesets A and C. |
Table IV.
Small molecule drugs potentially used
to treat the diseases associated with differentially expressed
genes in genesets A and C.
| Small molecule
drug | Enrichment | P-value |
|---|
| Acemetacin | −0.925 |
0.00002a |
| Scriptaid | 0.912 |
0.00142a |
| Alsterpaullone | 0.923 |
0.00094a |
| Mycophenolic
acid | 0.957 | 0.0001a |
| MG-262 | 0.962 |
0.00008a |
| Trifluridine | 0.975 | <0.001 |
| Camptothecin | 0.975 |
0.00004a |
| Ciclopirox | 0.987 | <0.001 |
| MS-275 | 0.998 |
0.00002a |
Discussion
Keloids are benign fibrotic tumors of the dermis and
are a cosmetic and psychological burden to patients (2). In the present study, 192 and 67 DEGs
were screened from dermal fibroblasts in keloids with and without
HC treatment, respectively, and compared with gene expression in
normal scar tissue. According to the GO functional enrichment
analysis for DEGs only present in dermal fibroblasts from keloids
without HC treatment, namely those expressed normally following HC
treatment, a set of genes (COL18A1 and JAG1) were
significantly associated with endothelial cell differentiation and
regulation of cell proliferation (P<0.001).
COL18A1 encodes the alpha chain of type XVIII
collagen, which is an inhibitor of angiogenesis (18). It has demonstrated that collagen
XVIII levels are downregulated in keloid patients compared with
normal controls (19). Inhibition of
collagen synthesis was observed following the addition of higher
concentrations of HC (20).
JAG1 is one of Notch ligands from the Notch family (21). Increased expression of JAG1
and Notch receptors has been observed in keloid fibroblasts and the
inhibition of Notch signaling via JAG1 knockdown led to the
inhibition of proliferation, migration and invasion properties of
keloid fibroblasts (22).
Furthermore, the endothelial cells within blood vessels in keloids
are somewhat rounded and projected into the lumen of the vessel
(23); this has been speculated as
critical to facilitate the development and maintenance of keloids
(24). Vascular endothelial growth
factor, which promotes angiogenesis and enhances endothelial cell
survival (25), is abundantly
produced in keloids (26,27). As DEGs in geneset B were not present
in the samples following HC treatment, these genes were considered
to be potentially associated with the positive efficacy of HC.
Therefore, HC may serve a key role in the treatment of keloids by
targeting genes associated with endothelial cell differentiation
and regulation of cell proliferation (COL18A1 and
JAG1).
The pathway enrichment analysis for the DEGs in
genesets A and C revealed that certain DEGs (CCNB1 and
CCNB2) were significantly enriched in the cell cycle and p53
signaling pathways (P<0.05). Keloids are characterized by
aggressive dermal growth beyond the boundaries of the original
margins of a wound (2), which is
dependent on the ectopic cell cycle. A previous study demonstrated
that mRNAs associated with cell cycle suppression, such as cyclin
B1, were detected in the bottom region of keloids (28). Furthermore, CCNB1 gene expression is
reduced by low-dose 5-fluorouracil in treated keloid fibroblasts
(29). The p53 tumor suppressor acts
as a transcription factor and has a central function in controlling
apoptosis (30). p53 levels are
higher in keloids compared with normal scar tissues (31). Therefore, the current study
demonstrated that CCNB1 and CCNB2 are involved in the
cell cycle and p53 signaling pathways may be essential in the
development of keloids. However, the current results indicated that
the expression of these genes was altered following HC treatment,
and did not change in the samples without HC treatment. Thus, it is
speculated that the alterations of CCNB1 and CCNB2
may contribute to keloids via dysregulation of cell cycle and p53
signaling pathways.
A number of diseases, such as bone loss, were
predicted to be associated with DEGs in keloid fibroblasts treated
with HC (IL1R1 and COL1A1). Bone loss is one of the
most devastating adverse effects of glucocorticoids as it results
in the inhibition of calcium transport and the impairment of
osteoblast function (32). A
previous study has reported that long-term treatment of secondary
hypocortisolism with a high replacement dose of hydrocortisone (30
mg/day) induces bone loss (33).
IL1R1 encodes the type I interleukin 1 receptor. It has been
demonstrated that IL-1 production may accelerate bone loss in
postmenopausal women (34). The
interleukin-1 receptor antagonist (IL-1ra) may decrease bone loss
and bone resorption in ovariectomized rats (35) and it has been reported that a gene
polymorphism of IL-1ra is associated with bone mineral density and
osteoporosis in postmenopausal women (36). Additionally, it has been demonstrated
that COL1A1 (encoding collagen, type I, alpha 1) Sp1 alleles
are associated with a modest reduction in bone mineral density and
a markedly increased risk of osteoporotic fracture, particularly
vertebral fracture (37).
Previously, it was demonstrated that the crosslinking process of
collagen is important in bone strength, osteogenesis imperfecta and
osteoporosis (38). Therefore, the
present study hypothesized that HC may contribute to bone loss
during treatment with keloids, through certain genes including
IL1R1 and COL1A1.
Although HC has been successfully used to treat
keloids, it also induces certain adverse effects, such as
arthralgia (39). In the present
study, 9 small molecule drugs were identified by the CMAP database
and acemetacin was predicted to be the most effective drug at
reducing the adverse effects of HC. It has previously been
demonstrated that acemetacin may treat arthralgia effectively
(40). Therefore, combining
acemetacin with HC may be a novel therapeutic method of treating
keloids, thus reducing the adverse effects of HC treatment.
Despite the aforementioned results, there were a
number of limitations in the present study. Confirmation by
experiment of these predictions is necessary. Future in vivo
studies are required to detect the expression of genes associated
with endothelial cell differentiation and regulation of cell
proliferation. Furthermore, the efficacy of acemetacin-HC
combination treatment compared with HC treatment alone requires
evaluation.
In conclusion, the present study suggests that HC
treats keloids effectively by targeting COL18A1 and
JAG1, which are associated with endothelial cell
differentiation. However, the cell cycle and p53 signaling pathways
in keloids do not return to normal following treatment with HC. A
number of adverse effects such as bone loss, which involves
IL1R1 and COL1A1, may occur during the therapeutic
process of HC treatment. Additionally, it was predicted that
acemetacin may be used in combination with HC to treat keloids,
although further studies are required to demonstrate its efficacy.
These findings may contribute to the understanding of mechanisms
involved in HC treatment on keloids and provide candidates for
subsequent validation and further study.
References
|
1
|
Marneros AG, Norris JE, Watanabe S,
Reichenberger E and Olsen BR: Genome scans provide evidence for
keloid susceptibility loci on chromosomes 2q23 and 7p11. J Invest
Dermatol. 122:1126–1132. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Fong CY, Biswas A, Subramanian A,
Srinivasan A, Choolani M and Bongso A: Human keloid cell
characterization and inhibition of growth with human Wharton's
jelly stem cell extracts. J Cell Biochem. 115:826–838. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Asboe-Hansen G, Brodthagen H and Zachariae
L: Treatment of keloids with topical injections of hydrocortisone
acetate. AMA Arch Derm. 73:162–165. 1956. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Russell JD, Russell SB and Trupin KM:
Differential effects of hydrocortisone on both growth and collagen
metabolism of human fibroblasts from normal and keloid tissue. J
Cell Physiol. 97:221–229. 1978. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Russell SB, Russell JD and Trupin JS:
Alteration of amino acid transport by hydrocortisone. Different
effects in human fibroblasts derived from normal skin and keloid. J
Biol Chem. 257:9525–9531. 1982.PubMed/NCBI
|
|
6
|
Trupin JS, Russell SB and Russell JD:
Variation in prolyl hydroxylase activity of keloid-derived and
normal human fibroblasts in response to hydrocortisone and ascorbic
acid. Coll Relat Res. 3:13–23. 1983. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Ladin DA, Hou Z, Patel D, McPhail M, Olson
JC, Saed GM and Fivenson DP: p53 and apoptosis alterations in
keloids and keloid fibroblasts. Wound Repair Regen. 6:28–37. 1998.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Meyer LJ, Russell SB, Russell JD, Trupin
JS, Egbert BM, Shuster S and Stern R: Reduced hyaluronan in keloid
tissue and cultured keloid fibroblasts. J Investig Dermatol.
114:953–959. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Mantripragada KK, Buckley PG, de Ståhl
Diaz T and Dumanski JP: Genomic microarrays in the spotlight.
Trends Genet. 20:87–94. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Smith JC, Boone BE, Opalenik SR, Williams
SM and Russell SB: Gene profiling of keloid fibroblasts shows
altered expression in multiple fibrosis-associated pathways. J
Invest Dermatol. 128:1298–1310. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Seo J and Hoffman EP: Probe set
algorithms: Is there a rational best bet? BMC Bioinformatics.
7:3952006. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Smyth GK: Linear models and empirical
bayes methods for assessing differential expression in microarray
experiments. Stat Appl Genet Mol Biol. 3:Article32004. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Bland JM and Altman DG: Multiple
significance tests: The Bonferroni method. BMJ. 310:1701995.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Huang da W, Sherman BT and Lempicki RA:
Systematic and integrative analysis of large gene lists using DAVID
bioinformatics resources. Nat Protoc. 4:44–57. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Xie C, Mao X, Huang J, Ding Y, Wu J, Dong
S, Kong L, Gao G, Li CY and Wei L: KOBAS 2.0: A web server for
annotation and identification of enriched pathways and diseases.
Nucleic Acids Res. 39:W316–W322. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Wang J, Duncan D, Shi Z and Zhang B:
WEB-based GEne SeT AnaLysis Toolkit (WebGestalt): Update 2013.
Nucleic Acids Res. 41:W77–W83. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Lamb J: The Connectivity Map: A new tool
for biomedical research. Nat Rev Cancer. 7:54–60. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Pufe T, Petersen WJ, Miosge N, Goldring
MB, Mentlein R, Varoga DJ and Tillmann BN: Endostatin/collagen
XVIII-an inhibitor of angiogenesis-is expressed in cartilage and
fibrocartilage. Matrix Biol. 23:267–276. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Mogili NS, Krishnaswamy VR, Jayaraman M,
Rajaram R, Venkatraman A and Korrapati PS: Altered angiogenic
balance in keloids: A key to therapeutic intervention. Transl Res.
159:182–189. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Deshpande M, Papp S, Schaffer L and
Pouyani T: Hydrocortisone and triiodothyronine regulate hyaluronate
synthesis in a tissue-engineered human dermal equivalent through
independent pathways. J Biosci Bioeng. 119:226–236. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Bolós V, Grego-Bessa J and de la Pompa JL:
Notch signaling in development and cancer. Endocr Rev. 28:339–363.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Syed F and Bayat A: Notch signaling
pathway in keloid disease: Enhanced fibroblast activity in a
Jagged-1 peptide-dependent manner in lesional vs. Extralesional
fibroblasts. Wound Repair Regen. 20:688–706. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Ehrlich HP, Desmoulière A, Diegelmann RF,
Cohen IK, Compton CC, Garner WL, Kapanci Y and Gabbiani G:
Morphological and immunochemical differences between keloid and
hypertrophic scar. Am J Pathol. 145:105–113. 1994.PubMed/NCBI
|
|
24
|
Kischer CW, Thies AC and Chvapil M:
Perivascular myofibroblasts and microvascular occlusion in
hypertrophic scars and keloids. Hum Pathol. 13:819–824. 1982.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Nör JE, Christensen J, Mooney DJ and
Polverini PJ: Vascular endothelial growth factor (VEGF)-mediated
angiogenesis is associated with enhanced endothelial cell survival
and induction of Bcl-2 expression. Am J Pathol. 154:375–384. 1999.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Gira AK, Brown LF, Washington CV, Cohen C
and Arbiser JL: Keloids demonstrate high-level epidermal expression
of vascular endothelial growth factor. J Am Acad Dermatol.
50:850–853. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Fujiwara M, Muragaki Y and Ooshima A:
Upregulation of transforming growth factor-beta1 and vascular
endothelial growth factor in cultured keloid fibroblasts: Relevance
to angiogenic activity. Arch Dermatol Res. 297:161–169. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Javad F, Marriage F, Bayat A and Day PJ:
Perturbation of cell cycle expression in keloid fibroblast.
Skinmed. 10:152–159. 2012.PubMed/NCBI
|
|
29
|
Huang L, Wong Y, Cai Y, Lung I, Leung C
and Burd A: Low-dose 5-fluorouracil induces cell cycle G2 arrest
and apoptosis in keloid fibroblasts. Br J Dermatol. 163:1181–1185.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Lane DP: The regulation of p53 function:
Steiner Award Lecture. Int J Cancer. 57:623–627. 1994. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Tanaka A, Hatoko M, Tada H, Iioka H,
Niitsuma K and Miyagawa S: Expression of p53 family in scars. J
Dermatol Sci. 34:17–24. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Lukert BP and Raisz LG:
Glucocorticoid-induced osteoporosis. Rheum Dis Clin North Am.
20:629–650. 1994.PubMed/NCBI
|
|
33
|
Wichers M, Springer W, Bidlingmaier F and
Klingmüller D: The influence of hydrocortisone substitution on the
quality of life and parameters of bone metabolism in patients with
secondary hypocortisolism. Clin Endocrinol (Oxf). 50:759–765. 1999.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Langdahl BL, Løkke E, Carstens M, Stenkjær
LL and Eriksen EF: Osteoporotic Fractures Are Associated with an
86-Base Pair Repeat Polymorphism in the Interleukin-1-Receptor
Antagonist Gene But Not with Polymorphisms in the Interleukin-1beta
Gene. J Bone Miner Res. 15:402–414. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Kimble RB, Vannice JL, Bloedow DC,
Thompson RC, Hopfer W, Kung VT, Brownfield C and Pacifici R:
Interleukin-1 receptor antagonist decreases bone loss and bone
resorption in ovariectomized rats. J Clin Invest. 93:1959–1967.
1994. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Chen HY, Chen WC, Wu MC, Tsai FJ and Lin
CC: Interleukin-1beta and interleukin-1 receptor antagonist gene
polymorphism in postmenopausal women: Correlation to bone mineral
density and susceptibility to osteoporosis. Maturitas. 44:49–54.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Mann V and Ralston SH: Meta-analysis of
COL1A1 Sp1 polymorphism in relation to bone mineral density and
osteoporotic fracture. Bone. 32:711–717. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Viguet-Carrin S, Garnero P and Delmas P:
The role of collagen in bone strength. Osteoporos Int. 17:319–336.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Sun Y, Sun Q, Fan C, Shen J, Zhao W, Guo
Y, Su T, Wang W, Ning G and Bian L: Diagnosis and therapy for
Cushing's disease with negative dynamic MRI finding: A
single-centre experience. Clin Endocrinol (Oxf). 76:868–876. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Bori Segura G, Rubio Torres y Gutierrez A,
Herrera Gómez LE and Olguín Uribe J: Efficacy and tolerability of
acemetacin, a non-steroidal anti-inflammatory drug, in Mexican
patients: Result of the ETAPAM Study. Proc West Pharmacol Soc.
45:pp. 104–107. 2002, PubMed/NCBI
|