Introduction
Cervical cancer is one of the most commonly
occurring cancer types in females worldwide (1–3). Each
year, new cases of cervical cancer are detected in ~528,000 women,
and the worldwide fatality toll of cervical cancer is ~275,000
(3–5). High-risk human papillomavirus (HPV)
infection is the primary cause of cervical carcinogenesis (6). More than 200 different HPV genotypes
have been identified, there are >10% differences in the L1
nucleotide sequence between the genotypes (7). A minimum of 13 high-risk HPV genotypes
(HPV16, 18, 31, 33, 35, 39, 45, 51, 52, 56, 58, 59, and 68)
(8) are recognized as the causative
agents of cervical cancer and numerous other types of cancer
(9). HPV16 and HPV18 are the two
most prevalent genotypes in cervical cancer (CC) (10). Although the high prevalence of
HPV16/18 in CC is common throughout the world, the distribution of
other high-risk HPV types in the remaining fraction of CC reveals
region-specific variations (11).
Notably, in East Asian countries (including Japan, South Korea,
Taiwan and China) HPV52 infection is more prevalent compared with
European, North American and African regions (12–15). The
present study has focused on this type of HPV. The HPV E2 protein
controls replication, transcription and viral genome partitioning
during the viral infectious life cycle (16). The HPV E2 protein is a key
transcriptional regulator of the E6 and E7 genes (17). HPV DNA sequences are typically
integrated into the host cell genome, and cervical cancer
progression is significantly associated with integration of the
viral genome (18–20). Integrated and episomal HPV may be
confirmed using a polymerase chain reaction (PCR)-based protocol
for the amplification of papillomavirus oncogene transcripts
(APOT), which was developed by Klaes et al (21). This protocol is based on the
hypothesis that HPV transcripts derived from the integrated HPV
genome represent suitable molecular markers for a cervical lesion
at risk of progression to carcinoma. However, there are limited
available data concerning HPV52 status and integration
patterns.
The aim of the present study was to assess HPV52
infection and integration into the host cell genome in exfoliated
cervical cells. Briefly, cervical cytology specimens were collected
and genomic DNA and RNA were extracted from each sample. For every
specimen HPV DNA was detected using polymerase chain reaction (PCR)
amplification with the MY09/11 primers (22), HPV52 was detected using HPV52
type-specific primers. The viral load and integration state were
determined for HPV52 positive specimens by quantitative (q)PCR
using HPV52 E2, E6 and β-actin primers, the number of copies of
HPV52 was calculated using the formula: (E6 copy/β-actin copy) ×2,
and the HPV52 integration state was determined by the ratio of E2
to E6 copy number. The integration of HPV in the host chromosome
integration site may be accurately located by detecting
transcription of the poly (A) tail (21,23,24).
cDNA was synthesized by reverse transcription using an RNA
template, and a (dT)17-p3 as primer; PCR amplification
was conducted using cDNA as a template and p1-HPV52 E7 and p3 as
primers; nest PCR was conducted using the product as a template and
p2-HPV52 E7 and (dT)17-p3 as primers; the PCR product
underwent sequence analysis. The integration sites were determined
by sequence alignment of the HPV and human chromosome sequence.
Materials and methods
Specimen collection
Cervical cytology specimens from 468 female patients
were collected from the Affiliated Hospital of North China
University of Science and Technology (Tangshan, China) between
October 2012 and June 2014. These specimens were collected through
a sterile swab scraped in a clockwise rotation three times in the
cervix from every patient, the cervical exfoliation cells were
subsequently washed down from the swab using physiological saline
solution and collected. The age of the patients ranged from 26–60
years, and the mean age was 41.5±5 years. The patients had no
medical history of cervical diseases prior to this diagnosis and
were not menstruating or pregnant. The inclusion criteria for all
cases was a conventional cervical cytology diagnosis; exclusion
criteria were a diagnosis and treatment for cervical
intraepithelial neoplasia, cervical cancer and hysterectomy prior
to the study. The present study was approved by the Ethics
Committee of North China University of Science and Technology, and
every specimen donor provided informed consent. These specimens
were stored at −80°C.
Reagents
The Ex Taq Polymerase kit and pMD-18T vector kit
were provided by Takara Biotechnology Co., Ltd., Dalian, China.
EvaGreen was provided by Biotium, Inc., Freemont, CA, USA. AmpliTaq
Gold DNA Polymerase kit was provided by Applied Biosystems; Thermo
Fisher Scientific, Inc., Waltham, MA, USA. The RNeasy Mini kit was
provided by Qiagen GmbH, Hilden, Germany. Goldview I nuclear
staining dye and M-MLV Reverse Transcriptase kit were provided by
BioTeke Corporation, Beijing, China.
Media
Dulbecco's modified Eagle medium (DMEM) with
antibiotics (100 U/ml penicillin-streptomycin), PBS, fetal bovine
serum, trypsin, lysogeny broth and other media used in the present
study were provided by BioTeke Corporation.
Plasmid and cell DNA
The plasmid including the complete HPV52 genome, the
plasmid encorporating part of the human β-actin gene, 293 and HeLa
cell DNA, and E. coli DH5α competent cells, were obtained from the
National Institute for Viral Disease Control and Prevention,
Chinese Center for Disease Control and Prevention (Beijing,
China).
Cell culture conditions
The HeLa and 293 cells (National Institute for Viral
Disease Control and Prevention, Chinese Center for Disease Control
and Prevention) were cultivated using 10 ml media (DMEM with 100
U/ml penicillin-streptomycin and 10% fetal bovine serum) in a 10 cm
diameter petri dish and maintained at 37°C in a humidified
atmosphere with 5% CO2. The fresh cell media was changed
every 2–3 days.
Pathological cytology
The cervical cytology specimens were diagnosed by
pathological cytology according to the 2001 Bethesda system (TBS)
(25,26). According to the TBS, the cytological
diagnosis of abnormal cervical squamous cells is divided into the
following categories: Atypical squamous epithelial cells (ASC) of
undetermined significance; ASC-cannot exclude high-grade squamous
epithelial lesion (HSIL); low-grade squamous intraepithelial lesion
(LSIL); and HSIL. Sample smear preparation and staining were
performed using BD PrepStain™ Slide Processor (BD Biosciences, San
Jose, CA, USA) according to the manufacturer's protocol, and the
results were observed using an optical microscope.
Primer design and synthesis
The primers used for HPV52 E2 and E6 were designed
according to the HPV52 gene sequence in GenBank (X74481.1;
http://www.ncbi.nlm.nih.gov/nuccore/X74481) and were
synthesized with the β-actin primer by Sangon Biotech Shanghai Co.,
Ltd. (Shanghai, China). The primer sequences for HPV52 E2, HPV52 E6
and β-actin are presented in Table
I.
 | Table I.Primers used in the present
study. |
Table I.
Primers used in the present
study.
| Primer | Sequence
(5′-3′) | Position | Amplicon size
(bp) |
|---|
| HPV52 E2
primers |
|
|
|
| HPV52
E2-1 |
GGAAAACGATGGAGTCGATAC | 2735–2755 | 176 |
| HPV52
E2-2 |
GTGGCCTATATGAGTTATTC | 2891–2910 |
|
| HPV52
E2-3 |
CGATGCAAAGCAATATTGTG | 3261–3280 | 235 |
| HPV52
E2-4 |
GTACTTGGTGTTTCTGGAGTC | 3475–3395 |
|
| HPV52
E2-5 |
GCACCTATAATACACCTAAAAGG | 3604–3626 | 245 |
| HPV52
E2-6 |
CACAATGACATGACACCTTG | 3829–3848 |
|
| HPV52 E6
primers |
|
|
|
| HPV52
E6-1 |
GTTTGAGGATCCAGCAACAC | 104–123 | 369 |
| HPV52
E6-2 |
CGCTTGTTTGCATTAACATG | 453–472 |
|
| β-actin
primers |
|
|
|
|
Forward |
CACCCACACTGTGCCCATCT | 550–560 | 289 |
|
Reverse |
GAACCGCTCATTGCCAATGG | 819–838 |
|
Detection of specimen quality
Genomic DNA was extracted from each exfoliated
cervical cell sample using the phenol-chloroform method (27,28). The
quality of the sample DNA was determined by PCR amplification which
was conducted using a Takara Ex Taq kit in a 20 µl reaction mixture
[containing 1X Ex Taq Buffer (MgCl2 free), 2.5 mM
MgCl2, 0.2 mM dNTPs, 1 unit Ex Taq DNA polymerase, 5
pmol β-actin primers, 40 ng sample DNA template], the 293 cell DNA
served as a positive control and sterile water served as a negative
control. The thermocycling profile used was as follows: 95°C for 5
min; followed by 31 cycles of 95°C for 30 sec, 55°C for 30 sec and
72°C for 30 sec; followed by a final extension at 72°C for 5 min;
and storage at 4°C.
Detection of HPV DNA
HPV DNA was detected in each exfoliated cervical
cell sample using PCR as described above with the MY09/11 primers
(MY09: 5′-CGTCCMARRGGAWACTGATC-3′; MY11:
5′-GCMCAGGGWCATAAYAATGG-3′; (amplicon size, 450 bp) for the HPV L1
gene as described previously (22).
The PCR products were resolved on a 1% agarose gel and visualized
using Goldview I nuclear staining dye. The MY09/11 primers are able
to detect >20 HPV types, including HPV52. In the present
experiment, HPV infection-free 293 cell DNA was used as a negative
control, and HeLa cell DNA and HPV52 plasmid DNA were used as
positive controls. HeLa cell DNA was used as a HPV52 type-specific
control and HPV52 plasmid DNA was used as a positive control to
detect HPV52 E6 using HPV52 type-specific primers. PCR was
conducted in a 20-µl reaction mixture containing 1X Ex Taq Buffer
(MgCl2-free), 2.5 mM MgCl2, 0.2 mM dNTPs, 1
unit Ex Taq DNA polymerase, 5 pmol of each primer from the primer
pair described above (MY09/MY11 or HPV52 E6 primers) and 40 ng
sample DNA or control DNA. The thermocycling profile used was as
follows: 95°C for 5 min; followed by 31 cycles of 95°C for 30 sec,
55°C for 30 sec and 72°C for 30 sec; followed by a final extension
at 72°C for 5 min; and a holding temperature of 4°C. The PCR
products were resolved on a 1.0% agarose gel and observed with a UV
transilluminator.
qPCR analysis
The viral load and integration state were determined
using qPCR with HPV52 E2/E6/β-actin primers. Briefly, the number of
copies of HPV52 was calculated using the formula: (E6 copy/β-actin
copy) ×2, and the HPV52 integration state was determined by the
ratio of E2 to E6 copy number. The reactions were performed in a
25-µl reaction mixture and standard curves were generated with
quantified DNA standards (a 10-fold dilution series of full length
HPV52 and β-actin plasmids). The PCR mixtures consisted of 1X
AmpliTaq gold buffer (MgCl2-free), 2.5 mmol/l
MgCl2, 0.2 mmol/l dNTPs, 2 pmol/l primers, 20X EvaGreen
(1.25 µl), 0.25 µl RX (stabilizer), 2 µl AmpliTaq Gold DNA
polymerase, 0.4 µl DNA template (80 ng/µl) and double distilled
water to a final volume of 25 µl. The initial denaturation was
performed at 95°C for 5 min; followed by 5 cycles of 95°C for 1
min, 55°C for 1 min and 72°C for 1 min; followed by 35 cycles at
95°C for 30 sec, 55°C for 50 sec and 72°C for 50 sec; and a final
extension at 72°C for 5 min. The threshold cycle (Cq) (29,30) was
determined by the CFX96 Touch Real-time PCR system (Bio-Rad
Laboratories, Inc., Hercules, CA, USA), with the baseline set
automatically to standard deviations above the background
fluorescence generated in the first five cycles. For each specimen,
identical amounts of DNA were analyzed for E6 and E2 and compared
with expression of the internal reference gene β-actin. The
specificity of each amplification was confirmed by checking the
dissociation curve against the expected melting temperature of the
amplification product.
Reverse transcription (RT)
RNA was extracted from the patient samples that were
identified to have integrated HPV52 using the RNeasy Mini kit. RT
was performed using Oligo (dT)17-p3 (primer:
5′-GACTCGAGTCGACATCGATTTTTTTTTTTTTTTTT) to produce double-stranded
DNA, which was subsequently amplified using PCR. The components of
the RT reaction were as follows: RNase-free H2O (9.2
µl), 5X first-strand buffer (BioTeke Corporation; 4 µl), 0.2 mmol/l
dNTPs (1 µl), 40 units RNase inhibitor (1 µl), 10 pmol
(dT)17-p3 (oligonucleotide primer,
GACTCGAGTCGACATCGATTTTTTTTTTTTTTTTT-3′; 1 µl), 200 units
SuperScript reverse transcriptase (BioTeke Corporation; 1 µl), and
total RNA (0.5–1 µg) in a total volume of 20 µl. RNA was reverse
transcribed by heating at 42°C for 50 min, and the reaction was
terminated by heating at 70°C for 15 min. The cDNA samples were
stored at 4°C.
To confirm that HPV52 E7 mRNA was detectable in all
of the samples and to ensure that the isolated mRNA was of
sufficient integrity for PCR amplification, mRNA corresponding to
the human reference gene GAPDH was analyzed by RT-PCR. The RT
reactions were performed using a Power M-MLV Reverse Transcriptase
kit (BioTeke Corporation) according to the manufacturer's protocol
and the cDNA was stored at 4°C. The PCR was conducted using a
Takara Ex Taq Polymerase kit in a 20 µl reaction mixture containing
1X Ex buffer (MgCl2-free), 2.5 mM MgCl2, 0.2
mM dNTPs, 1 unit Ex Taq DNA polymerase, 5 pmol each GAPDH primer
(forward, 5′-CATCACCATCTTCCAGGA-3′ and reverse,
5′-GTCTACCACCCTATTGCA-3′) and 2 µl cDNA as a template. The
thermocycling profile used was as follows: 95°C for 5 min; followed
by 31 cycles of 95°C for 30 sec, 52°C for 30 sec and 72°C for 30
sec; followed by a final extension at 72°C for 5 min; and storage
at 4°C.
Detection of viral-cell fusion
transcripts using nested PCR
The generation of viral-cellular fusion transcripts
encompasses the E6-E7-E1 sequences at their 5′-ends and cellular
sequences at their 3′-ends. APOT analysis used an
oligo(dT)17-primer coupled to a linker sequence (dT)17-p3, the RT
of all of the mRNAs was initiated by binding to their poly(A) tail,
and integrate-derived HPV oncogene transcripts were amplified by
nested PCR reactions as described previously (21,23,24). The
cDNA was synthesized by RT using RNA as a template and (dT)17-p3 as
a primer. Nested PCR amplification was then performed. The first
PCR reactions were performed using cDNA as a template, and p1-HPV52
E7 and p3 as primers. The second PCR reactions were conducted using
first PCR products as template, and p2-HPV52 E7 and (dT)17-p3 as
primers. First PCR reactions were conducted with 10 pmol/l primers
(forward, p1-HPV52 E7, 5′-CTACGGCTATGCATTCATAGC-3′; and reverse,
p3, 5′-GACTCGAGTCGACATCG-3′), 1X Ex buffer, 2.5 mmol/l
MgCl2, 0.2 mmol/l dNTPs, 1 unit Ex Taq DNA polymerase,
and 2 µl cDNA in a total volume of 25 µl; following thermocycling
conditions: 95°C for 5 min; followed by 30 cycles of denaturation
at 95°C for 1 min, annealing at 56°C for 1 min and elongation at
72°C for 3 min; followed by a final extension at 72°C for 5 min.
Then, second PCR was performed using the first PCR product (5 µl)
as a template, and using the p2-HPV52 E7 forward primer
(5′-CGTACTCTACAGCAAATGCTGTTG-3′) and the (dT)17-p3
reverse primer, with the exception that annealing was performed at
67°C. The positions of the two primers were 751–771 for p1-HPV52
and 787–810 for p2-HPV52. To control for false-positives, a
negative control (293 cell DNA template) was included in each set
of amplifications. Electrophoresis was performed using a 1.2%
agarose gel with Goldview I nuclear staining dye.
Cloning and sequence analysis
To further confirm specific HPV52 oncogene
transcription in all of the samples, the final nested PCR products
were cloned into a pMD-18T vector according to a previous study
(31), except the temperature of the
target segment ligation to the pMD-18T vector was changed to room
temperature for 1 h. Then Beijing Rui Bo Xing ke Biological
Technology (Beijing, China) was commisioned to sequence the
products. The results were analyzed using the National Center for
Biotechnology Information BLAST (blast.ncbi.nlm.nih.gov/Blast.cgi?PROGRAM=blastn&PAGE
_TYPE=BlastSearch&LINK_LOC=blasthome), and HPV52
integration sites in human chromosomes were determined for every
integration specimen.
Results
Determination of specimen DNA
quality
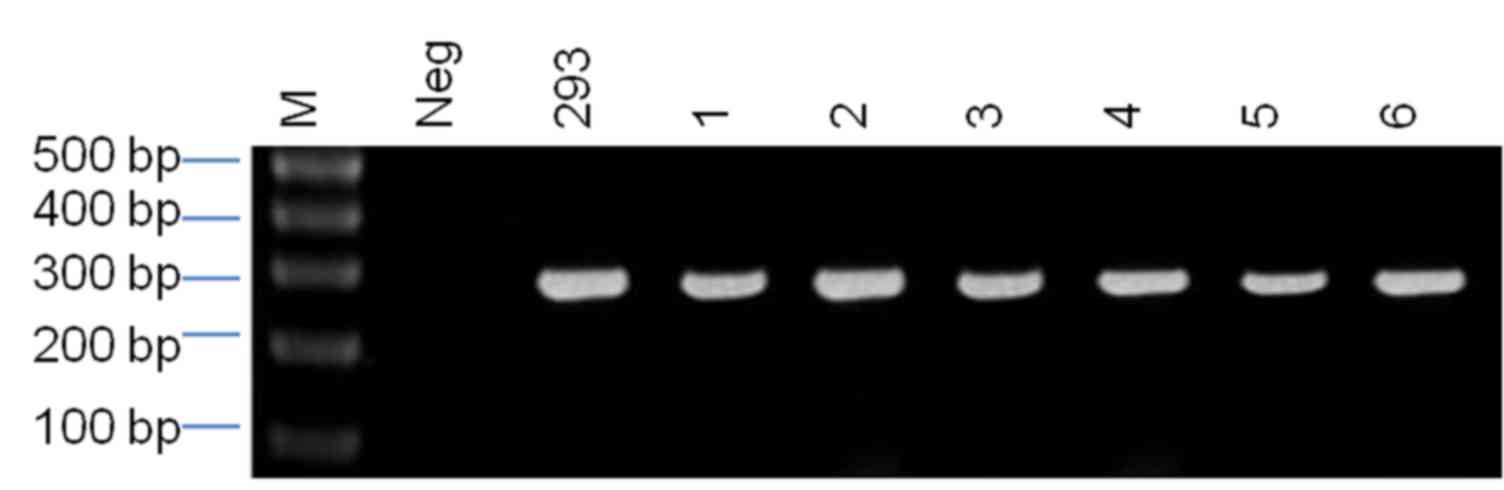
The 289-bp β-actin product size was confirmed by
agarose gel electrophoresis, and PCR amplification of β-actin in
the DNA specimens and the positive control was observed to be
similar (Fig. 1). The results
suggest that all of the DNA samples were of satisfactory quality
for subsequent experiments.
Detection of HPV DNA in the
samples
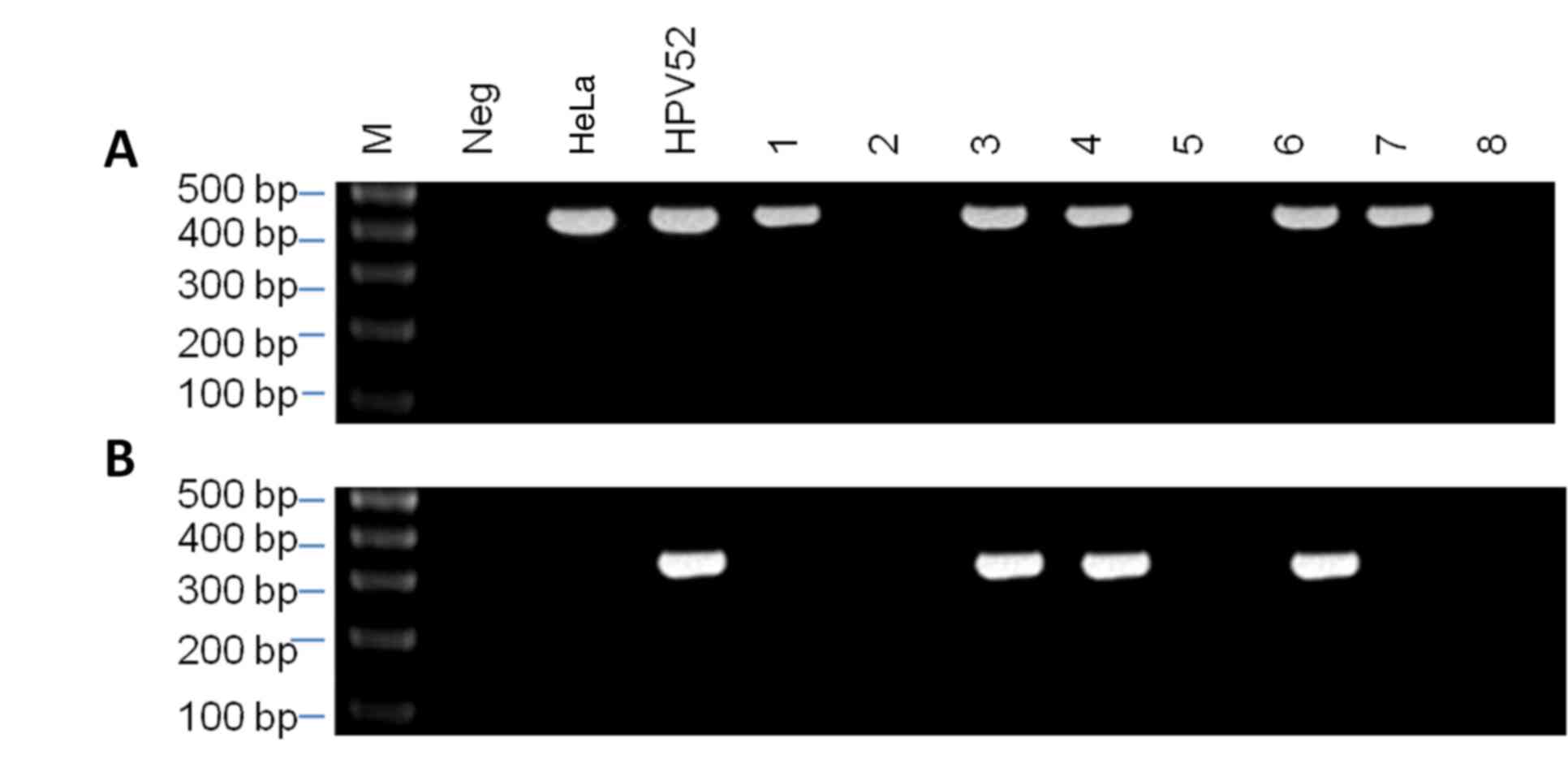
In total, 52 HPV-positive samples were identified
from among the 468 samples by PCR using the MY09/11 general primer
set, which generated a 450-bp product (Fig. 2A). Among them, 13 cases were found to
be HPV52-positive using the HPV52 E6 type-specific primer set,
which generated a 369-bp product (Fig.
2B).
Association between cervical lesions
and HPV52 infection
The 13 HPV52-positive samples were subjected to
cytological diagnosis, which indicated that these 13 samples
included 1 case of ASC (Fig. 3A), 5
cases of LSIL (Fig. 3B) and 7 cases
of HSIL (Fig. 3C).
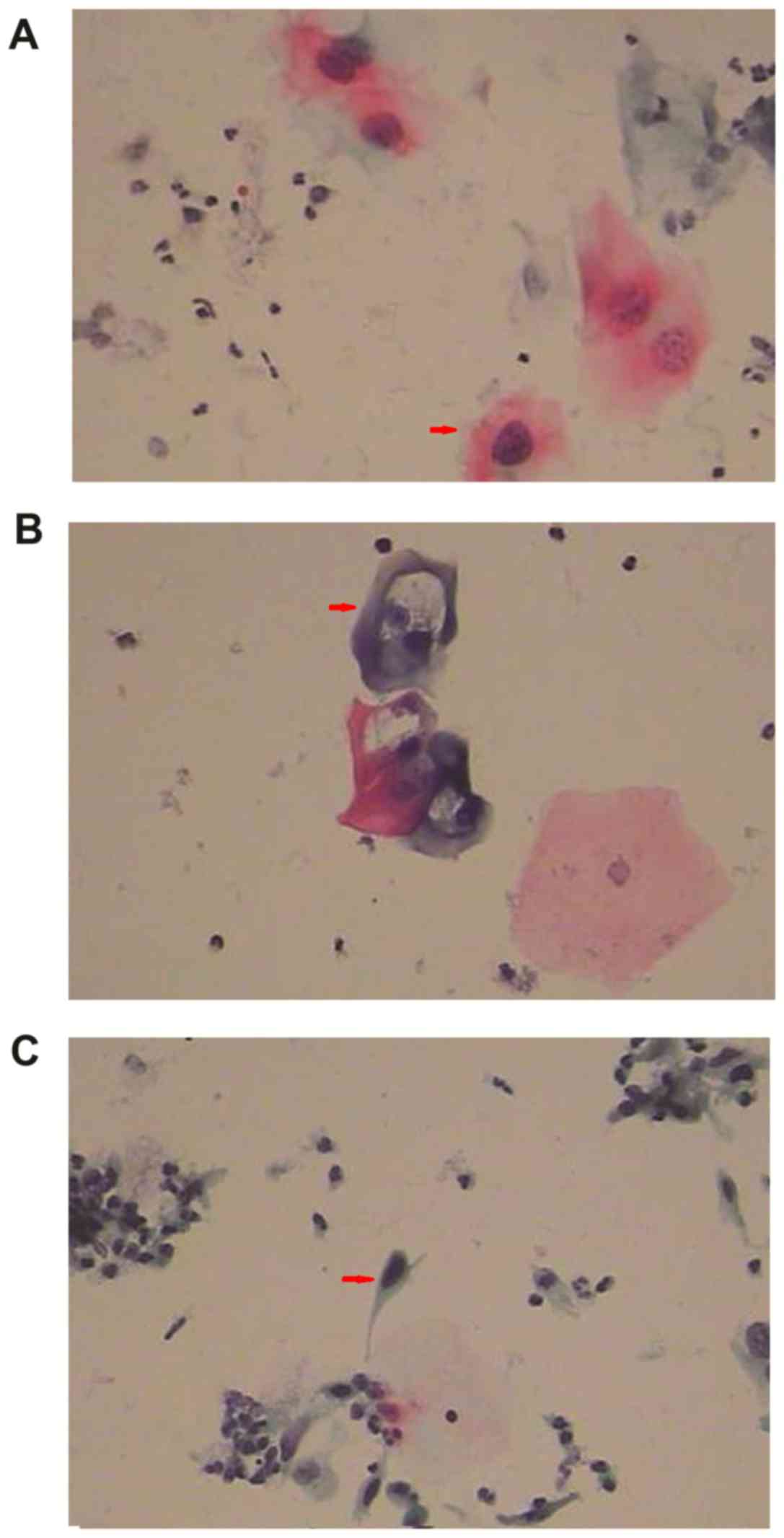 | Figure 3.Cytological diagnosis of cervical
lesions. (A) Atypical squamous cells, (B) low-grade squamous
intraepithelial lesions and (C) high-grade squamous epithelial
lesions. (A) The arrowhead indicates the cell volume increased, the
cytoplasm was rich, the nucleus was slightly enlarged and nuclear
polarity was lacking in the atypical squamous cells. (B) The
arrowhead indicates koilocyte was visible near the edge and in the
middle of the field of view, with an atypical concave nuclear
space, an irregular nuclear membrane, a visible dual-core, a free
halo around the nucleus and a stiff surrounding halo, with an iron
wire mesh-like shape. (C) The arrowhead indicates the cell was
alone with a larger nucleus, the cytoplasm exhibited keratosis and
the cytoplasmic area was decreased. Magnification, ×400. |
Analysis of HPV52 infection and viral
load
The standard curve and copy number of β-actin, E2
and E6 for every sample were automatically generated and determined
by the qPCR instrumentation based on six 10-fold dilutions of
β-actin, E2 and E6 standard plasmids. The HPV52 viral load of each
sample was calculated (Table II).
No clear association of lesion pathology with viral load was
observed.
 | Table II.Distribution of viral load and
integration state of HPV52. |
Table II.
Distribution of viral load and
integration state of HPV52.
|
|
|
| HPV52 integration
state, n (%) |
|---|
|
|
|
|
|
|---|
| Cytodiagnosis | Viral load (copy
no./cell) | No. of cases | Episomal | Mixeda | Integrated |
|---|
| ASC | 23.26 | 1 | 1 (100.0) | 0 | 0 |
| LSIL | 9.88–189.25 | 5 | 0 | 4 (80.0) | 1 (20.0) |
| HSIL | 19.52–98.52 | 7 | 0 | 5 (71.4) | 2 (28.6) |
| Total | 9.88–189.25 | 13 | 1 (7.7) | 9 (69.2) | 3 (23.1) |
The HPV52 integration status was determined based on
the E2/E6 ratio. Among the 13 HPV52-positive samples, 1 case was
episomal, and the E2 gene was not destroyed. A total of 3 cases
were fully integrated, with E2/E6=0. The remaining 9 cases were a
mix of integrated and episomal, with an E2/E6 ratio >0 and
<1. The results are presented in Table II. As the pathology of the lesions
advanced from ACS to LSIL and HSIL, the proportion of episomal
cases decreased and that of integrated (episomal + integrated)
cases increased. These results indicate that the pathology of the
cervical lesions is associated with HPV52 infection and viral DNA
integration into the host genome.
Location of HPV52 integration in the
host chromosome
RNA was extracted from the HPV52-integrated samples.
A product of ~600 bp was generated according to nested PCR using
the E7-specific primers with the template cDNA obtained by RT with
oligo (dT)17-p3. In 3 samples, the HPV52 integration
site was in human chromosome 2, 5 or 8; this was determined by
sequencing the ~600 bp product and cloning it into the pMD-18
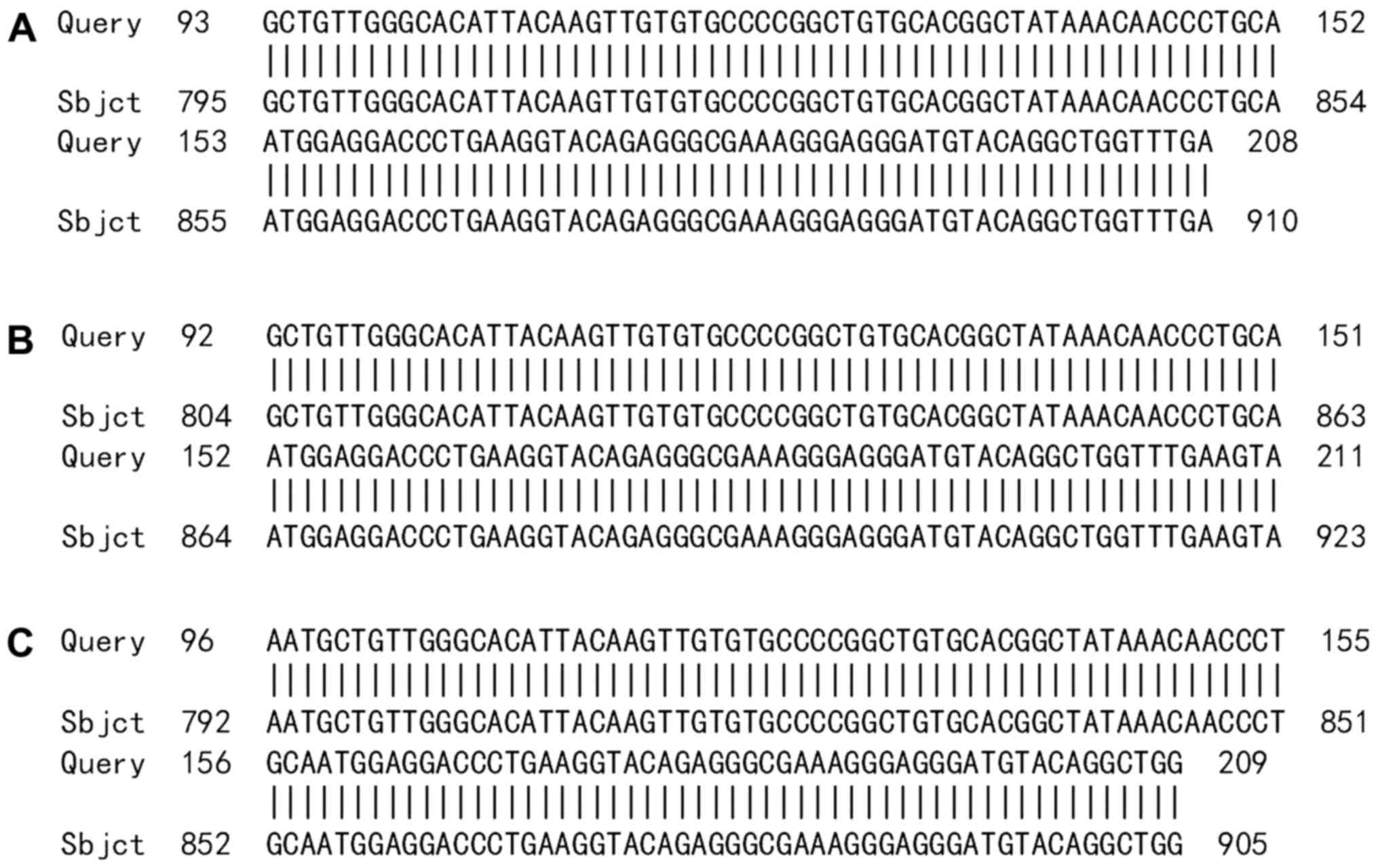
vector. The sequencing results indicated that the HPV52 E7 PCR
product that was amplified using the p2-HPV52 primer was part of
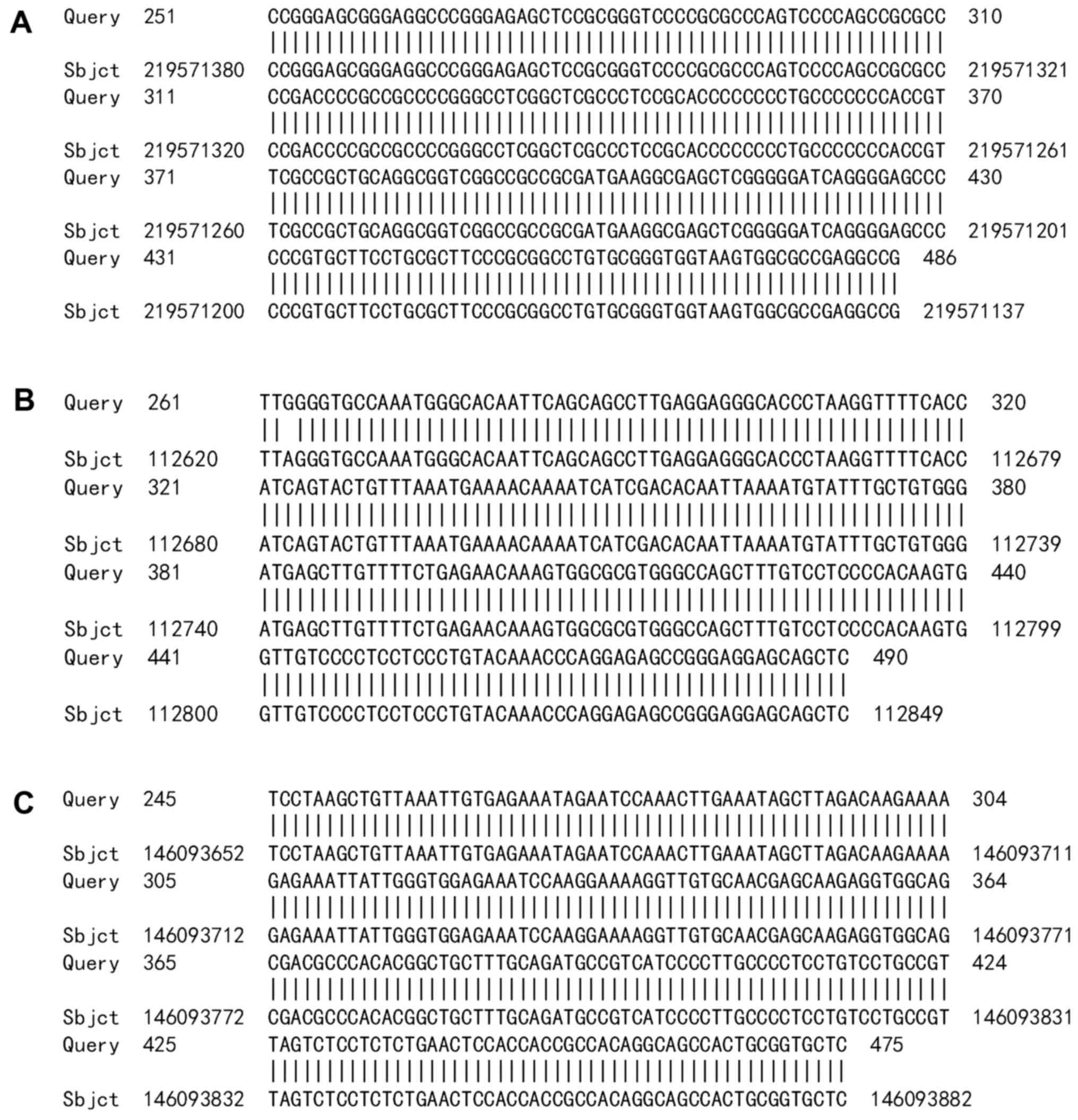
the HPV52 E7-E1 sequence (Fig. 4).
Another section of the amplified sequence was similar to human
chromosome 2 (Fig. 5A), 5 (Fig. 5B) or 8 (Fig. 5C).
Discussion
Persistent high-risk HPV infection, with HPV types
including HPV16, 18 and 31, has been identified as the major
causative agent of cervical cancer (32). HPV52 infection is an important factor
for cervical, anal and oropharyngeal carcinoma (33–36).
Viral load is recommended as a precancer biomarker, but this has
been indicated only for HPV16 infection (37,38).
qPCR is regarded as the standard method for viral load
quantification (39). Although viral
load has been recommended as a precancer biomarker for HPV16
infection (37–39), there is no consistent evidence that
viral load is a useful marker of prevalent disease or disease
progression (40–42). In the present study, the results
revealed that viral load did not have a clear association with
cervical lesion pathology. Multiple E1-L1/E6E7 (including E1/E6E7
and L1/E6E7) ratio analysis has been demonstrated to be more
sensitive than E2/E6E7 ratio analysis in the detection of HPV
integration status, and may be used to select candidates for
cervical biopsy and reduce the incidence of cervical cancer
(43).
The present study identified 13 HPV52-positive
patients by the analysis of 468 exfoliated cervical cell samples.
Among them, 1 was purely episomal, 3 were purely integrated, and 9
were mixed (episomal and integrated). The results of 3 integrated
HPV52 samples indicated that the integration of high-risk HPV may
have an important role in cervical cancer. Furthermore, viral
integration may occur in the E6 and E7 genes or downstream in the
E1 or E2 area, causing gene inactivation (18,44,45).
Integration of the viral genome is typically caused by an HPV E2
open reading frame that is broken or missing; in such cases,
certain areas of the DNA fragment are lost, and the E7 or E6 gene
is integrated into the host cell, which promotes and maintains the
integration of the viral genome and the interaction with host genes
(46).
Cytological analysis of the 13 HPV18-positive
specimens according to the TBS indicated that there was 1 case of
ASC, 5 cases of LSIL and 7 cases of HSIL. As the lesion stage
advanced from ASC to LSIL and HSIL, the proportion of episomal
samples decreased, and that of integrated samples increased for the
HPV52-infected patients. The results indicate that pathological
changes are associated with integration status, and the 3
HPV52-positive samples integrated into human chromosome 2, 5 or 8.
This is consistent with previous results, which have suggested that
integration of the viral genome increases the severity of the
disease, at least as indicated by biomarker studies of
HPV16-infected cervical lesions and precancerous lesions (38,32,47).
Therefore, a persistent infection following integration of the
viral genome into human host cell chromosomes may be key for cell
transformation and cancer development.
HPV52 integration into the host genome may be a key
factor in cervical lesions, therefore, patients at high risk for
cervical lesions may potentially be identified by screening for
HPV52 infection and integration.
Acknowledgements
The present study was supported by the project of
Science and Technology for Overseas Scholars in Hebei Province
(grant no. CY201620), the Project of Hebei Education Department
(grant no. ZD2016003), the Project of North China University
Science and Technology (grant no. SP201506) and Beijing Natural
Science Foundation (grant no. 5162003).
References
|
1
|
Oh JK and Weiderpass E: Infection and
cancer: Global distribution and burden of diseases. Ann Glob
Health. 80:384–392. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Arbyn M, Castellsagué X, de Sanjosé S,
Bruni L, Saraiya M, Bray F and Ferlay J: Worldwide burden of
cervical cancer in 2008. Ann Oncol. 22:2675–2686. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Vaccarella S, Laversanne M, Ferlay J and
Bray F: Cervical cancer in Africa, Latin America and the Caribbean
and Asia: Regional inequalities and changing trends. Int J Cancer.
Jul 22–2017.(Epub ahead of print). View Article : Google Scholar
|
|
4
|
Nayir T, Okyay RA, Nazlican E, Yesilyurt
H, Akbaba M, Ilhan B and Kemik A: Cervical cancer screening in an
early diagnosis and screening center in mersin, Turkey. Asian Pac J
Cancer Prev. 16:6909–6912. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Torre LA, Bray F, Siegel RL, Ferlay J,
Lortet-Tieulent J and Jemal A: Global cancer statistics, 2012. CA
Cancer J Clin. 65:87–108. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
zur Hansen H: Papillomaviruses and cancer:
From basic studies to clinical application. Nat Rev Cancer.
2:342–350. 2002. View
Article : Google Scholar : PubMed/NCBI
|
|
7
|
Bzhalava D, Eklund C and Dillner J:
International standardization and classification of human
papillomavirus types. Virology. 476:341–344. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Arbyn M, Tommasino M, Depuydt C and
Dillner J: Are 20 human papillomavirus types causing cervical
cancer? J Pathol. 234:431–435. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Doorbar J, Egawa N, Griffin H, Kranjec C
and Murakami I: Human papillomavirus molecular biology and disease
association. Rev Med Virol. 25 Suppl 1:S2–S23. 2015. View Article : Google Scholar
|
|
10
|
de Sanjose S, Quint WG, Alemany L, Geraets
DT, Klaustermeier JE, Lloveras B, Tous S, Felix A, Bravo LE, Shin
HR, et al: Human papillomavirus genotype attribution in invasive
cervical cancer: A retrospective cross-sectional worldwide study.
Lancet Oncol. 11:1048–1056. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Smith JS, Lindsay L, Hoots B, Keys J,
Franceschi S, Winer R and Clifford GM: Human papillomavirus type
distribution in invasive cervical cancer and high-grade cervical
lesions: A meta-analysis update. Int J Cancer. 121:621–632. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Onuki M, Matsumoto K, Satoh T, Oki A,
Okada S, Minaguchi T, Ochi H, Nakao S, Someya K, Yamada N, et al:
Human papillomavirus infections among Japanese women: Age-related
prevalence and type-specific risk for cervical cancer. Cancer Sci.
100:1312–1316. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Li N, Franceschi S, Howell-Jones R,
Snijders PJ and Clifford GM: Human papillomavirus type distribution
in 30,848 invasive cervical cancers worldwide: Variation by
geographical region, histological type and year of publication. Int
J Cancer. 128:927–935. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Azuma Y, Kusumoto-Matsuo R, Takeuchi F,
Uenoyama A, Kondo K, Tsunoda H, Nagasaka K, Kawana K, Morisada T,
Iwata T, et al: Human papillomavirus genotype distribution in
cervical intraepithelial neoplasia grade 2/3 and invasive cervical
cancer in Japanese women. Jpn J Clin Oncol. 44:910–917. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Chan PK, Ho WC, Chan MC, Wong MC, Yeung
AC, Chor JS and Hui M: Meta-analysis on prevalence and attribution
of human papillomavirus types 52 and 58 in cervical neoplasia
worldwide. PLoS One. 9:e1075732014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Graham SV: Human Papillomavirus E2
Protein: Linking replication, transcription and RNA processing. J
Virol. 90:8384–8388. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Chaiwongkot A, Vinokurova S, Pientong C,
Ekalaksananan T, Kongyingyoes B, Kleebkaow P, Chumworathayi B,
Patarapadungkit N, Reuschenbach M and von Knebel Doeberitz M:
Differential methylation of E2 binding sites in episomal and
integrated HPV 16 genomes in preinvasive and invasive cervical
lesions. Int J Cancer. 132:2087–2094. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Alp Avcı G: Genomic organization and
proteins of human papillomavirus. Mikrobiyol Bul. 46:507–515.
2012.(In Turkish). PubMed/NCBI
|
|
19
|
Akagi K, Li J, Broutian TR, Padilla-Nash
H, Xiao W, Jiang B, Rocco JW, Teknos TN, Kumar B, Wangsa D, et al:
Genome-wide analysis of HPV integration in human cancers reveals
recurrent, focal genomic instability. Genome Res. 24:185–199. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Raybould R, Fiander A, Wilkinson GW and
Hibbitts S: HPV integration detection in CaSki and SiHa using
detection of integrated papillomavirus sequences and
restriction-site PCR. J Virol Methods. 206:51–54. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Klaes R, Woerner SM, Ridder R, Wentzensen
N, Duerst M, Schneider A, Lotz B, Melsheimer P and von Knebel
Doeberitz M: Dectection of High-Risk cervical intraepithelial
neoplasia and cervical cancer by amplification of transcripts
derived from integrated papillomavirus oncogenes. Cancer Res.
59:6132–6136. 1999.PubMed/NCBI
|
|
22
|
Karlsen F, Kalantari M, Jenkins A,
Pettersen E, Kristensen G, Holm R, Johansson B and Hagmar B: Use of
Multiple PCR Sets for Optimal Detection of Human papillomavirus. J
Clin Microbiol. 34:2095–2100. 1996.PubMed/NCBI
|
|
23
|
Hillemanns P and Wang XL: Integration of
HPV-16 and HPV-18 DNA in vulvar intraepithelial neoplasia. Gynecol
Oncol. 100:276–282. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Klimov E, Vinokourova S, Moisjak E,
Rakhmanaliev E, Kobseva V, Laimins L, Kisseljov F and Sulimova G:
Human papilloma viruses and cervical tumours: Mapping of
integration sites and analysis of adjacent cellular sequences. BMC
Cancer. 2:242002. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Ok Atılgan A, Tepeoğlu M, Haberal AN,
Durukan E, Kuşcu E and Haberal M: Papanicolaou smear findings in
solid-organ transplant recipients compared with normal subjects
according to the Bethesda 2001 system. Exp Clin Transplant. 13
Suppl 1:S219–S222. 2015.
|
|
26
|
Al-Kadri HM, Kamal M, Bamuhair SS, Omair
AA and Bamefleh HS: Prevalence and characteristics of abnormal
Papanicolaou smear in Central Saudi Arabia. Saudi Med J.
36:117–122. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Psifidi A, Dovas CI and Banos G: A
comparison of six methods for genomic DNA extraction suitable for
PCR-based genotypingapplications using ovine milk samples. Mol Cell
Probes. 24:93–98. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Psifidi A, Dovas CI, Bramis G, Lazou T,
Russel CL, Arsenos G and Banos G: Comparison of eleven methods for
genomic DNA extraction suitable for large-scale whole-genome
genotyping and long-Term DNA banking using blood samples. PLoS One.
10:e01159602015. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Kim ST, Chun JW, Park G and Koh JW:
Comparative quantification of plasma TDRD7 mRNA in cataract
patients by real-time polymerase chain reaction. Korean J
Ophthalmol. 28:343–350. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Yuan B, Li XY, Zhu T, Yuan L, Hu JP, Chen
J, Gao W and Ren WZ: Antibody study in canine distemper virus
nucleocapsid protein gene-immunized mice. Genet Mol Res.
14:3098–3105. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Das P, Thomas A, Kannan S, Deodhar K,
Shrivastava SK, Mahantshetty U and Mulherkar R: Human
papillomavirus (HPV) genome status & cervical cancer outcome-A
retrospective study. Indian J Med Res. 142:525–532. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Onyekwuluje JM, Steinau M, Swan DC and
Unger ER: A real-time PCR assay for HPV52 detection and viral load
quantification. Clin Lab. 58:61–66. 2012.PubMed/NCBI
|
|
34
|
Ardhaoui M, Ennaifer E, Letaief H,
Salsabil R, Lassili T, Chahed K, Bougatef S, Bahrini A, El Fehri E,
Ouerhani K, et al: Prevalence, genotype distribution and risk
factors for cervical human papillomavirus infection in the grand
tunis region, Tunisia. PLoS One. 11:e01574322016. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Salehi-Vaziri M, Sadeghi F,
Bokharaei-Salim F, Younesi S, Alinaghi S, Monavari SH and Keyvani
H: The prevalence and genotype distribution of human papillomavirus
in the genital tract of males in Iran. Jundishapur J Microbiol.
8:e219122015. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Ewaisha R, Meshay I, Resnik J, Katchman BA
and Anderson KS: Programmable protein arrays for immunoprofiling
HPV-associated cancers. Proteomics. 16:1215–1224. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Castle PE, Schiffman M, Scott DR, Sherman
ME, Glass AG, Rush BB, Schussler JE, Wacholder S and Lorincz AT:
Semiquantitative human papillomavirus type 16 viral load and the
prospective risk of cervicalprecancer and cancer. Cancer Epidemiol
Biomarkers Prev. 14:1311–1314. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Shukla S, Mahata S, Shishodia G, Pande S,
Verma G, Hedau S, Bhambhani S, Kumari A, Batra S, Basir SF, et al:
Physical state & copy number of high risk human papillomavirus
type 16 DNA in progression of cervical cancer. Indian J Med Res.
139:531–543. 2014.PubMed/NCBI
|
|
39
|
Wentzensen N, Gravitt PE, Long R,
Schiffman M, Dunn ST, Carreon JD, Allen RA, Gunja M, Zuna RE,
Sherman ME, et al: Human papillomavirus load measured by Linear
Array correlates with quantitative PCR in cervical cytology
specimens. J Clin Microbiol. 50:1564–1570. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Theelen W, Reijans M, Simons G, Ramaekers
FC, Speel EJ and Hopman AH: A new multiparameter assay to assess
HPV 16/18, viral load and physical status together with gain of
telomerase genes in HPV-related cancers. Int J Cancer. 126:959–975.
2010.PubMed/NCBI
|
|
41
|
Jacobs MV, Walboomers JM, van Beek J,
Voorhorst FJ, Verheijen RH, Meijer CJ, van den Brule AJ,
Helmerhorst TJ and Snijders PJ: A quantitative polymerase chain
reaction-enzyme immunoassay for accurate measurements of human
papillomavirus type 16 DNA levels in cervical scrapings. Br J
Cancer. 81:114–121. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Constandinou-Williams C, Collins SI,
Roberts S, Young LS, Woodman CB and Murray PG: Is human
papillomavirus viral load a clinically useful predictive marker? A
longitudinal study. Cancer Epidemiol Biomarkers Prev. 19:832–837.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Zhang R, He YF, Chen M, Chen CM, Zhu QJ,
Lu H, Wei ZH, Li F, Zhang XX, Xu CJ and Yu L: Diagnosis of 25
genotypes of human papillomaviruses for their physical statuses in
cervical precancerous/cancerous lesions: a comparison of E2/E6E7
ratio-based vs. Multiple E1-L1/E6E7 ratio-based detection
techniques. J Transl Med. 12:2822014. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Kadaja M, Sumerina A, Verst T, Ojarand M,
Ustav E and Ustav M: Genomic instability of the host cell induced
by the human papillomavirus replication machinery. EMBO J.
26:2180–2191. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Johansson C, Somberg M, Li X, Winquist
Backström E, Fay J, Ryan F, Pim D, Banks L and Schwartz S: HPV-16
E2 contributes to induction of HPV-16 late gene expression by
inhibiting early polyadenylation. EMBO J. 31:3212–3227. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Al-Shabanah OA, Hafez MM, Hassan ZK,
Sayed-Ahmed MM, Abozeed WN, Al-Rejaie SS and Alsheikh AA: Human
papillomavirus genotyping and integration in ovarian cancer Saudi
patients. Virol J. 10:3432013. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Das P, Thomas A, Mahantshetty U,
Shrivastava SK, Deodhar K and Mulherkar R: HPV genotyping and site
of viral integration in cervical cancers in Indian women. PLoS One.
7:e410122012. View Article : Google Scholar : PubMed/NCBI
|