Introduction
Screening of human tissues for cancer biomarkers is
an important task in cancer diagnosis and treatment, which is
hindered by the complexity of the sample systems studied. A less
complex system such as urine is a preferred medium to screen for
protein or peptide biomarkers due to the non-invasive sampling of
patients, ease of sampling and the unrestricted quantities
obtainable. Urine is relatively stable in terms of protein/peptide
composition and fragmentation compared with other bodily fluids
such as serum, where proteolytic degradation by endogenous
proteases has been shown to occur during or after sample collection
(1).
Several investigations have been published
describing the urinary peptidome and proteome (as well as biomarker
discoveries for several diseases) using methodologies ranging from
traditional 2D gel electrophoresis alone (2), or coupled with mass spectrometry
(2-DE-MS) (3),
immunohistochemistry (4), liquid
chromatography mass spectrometry (LC-MS) (5), and surface enhanced laser desorption
ionisation-time of flight mass spectrometry (SELDI-TOF-MS)
(6–9).
The proteomic screening of urine for potential
cancer markers has shown several proteins to be differentially
present in ovarian cancer (10).
Bladder cancer biomarkers constitute a different non-overlapping
set of molecules (11–13), as do potential biomarkers for upper
gastrointestinal cancers (9). An
improvement in the reliability of diagnostic tests is to employ
more than one biomarker synchronously (9,14).
For example, one previous study employed an antibody-based array of
810 different antibodies to define peptide patterns in urine
associated with cancer (15). A
different approach was used successfully in recent years, combining
urinary mass spectroscopy with protein/peptide pattern analysis to
identify kidney disease (16).
There is a clear need to collect and cross-correlate
the wealth of data published in the scientific literature.
Currently, there are a number of urinary databases available. The
majority consist of lists of identified proteins derived from
tryptic digests analysed by liquid chromatography tandem mass
spectrometry (LC-MS/MS), such as the Max-Planck Unified Proteome
Database (MAPU) (17) and
Sys-BodyFluid (18). More
recently, a urinary database combining chromatographic
reverse-phase retention times and m/z values has been established
(19).
However, there is no database available which
integrates all of the data. In order to fill this gap, we have
assembled datasets from 100 urinary proteomic studies in our novel
proteomic database termed the Large Scale Screening Resource
(LSSR). LSSR is accessible and downloadable through the Proteomic
Analysis DataBase (PADB) portal at www.PADB.org.
In this study, we explore the possibility of
discovering novel cancer-associated molecular markers in human
urine by subtractive analysis using a novel dataset of the human
cancer urinary proteome [derived from patients with upper
gastrointestinal (GI) cancer] and comparing it to non-cancer
urinary datasets.
Materials and methods
Materials
Tris/Tricine peptide gels, gel-running buffers, CM
and IMAC resins, and chromatography buffers were from Bio-Rad
(Hemel Hempstead, UK). All other chemicals were obtained from
Sigma-Aldrich (Gillingham, UK).
Sample collection
Urine samples were obtained from upper GI cancer
patients (n=41) and non-cancer controls (n=21) as described
previously (9). Summary
participant demographics are shown in Table I. Participant age ranged between 21
and 84 (control group), and 43 and 82 (cancer group). Random
morning urine samples were collected over a time period of 2 years.
Cancer urine samples were collected prior to surgery if the patient
was being considered for resection. All procedures were approved by
the local research ethics committee, and written informed consent
was obtained. The study conformed to the standards set by the
Declaration of Helsinki. All urine samples were kept at −40°C for
short-term or −80°C for long-term storage.
 | Table IDemographics of the study cohort. |
Table I
Demographics of the study cohort.
| Cancer (n=41) | Control (n=21) | Entire cohort
(n=62) |
|---|
| Age (years) | 64 (9.5) | 62.1 (23.5) | 63.4 (15.6) |
| Male (M:F) | 26:15 | 17:04 | 43:19 |
| Primary tumor
origin | | | |
| Pancreas | 15 | N/A | |
| Oesophagus | 9 | | |
| OGJ | 7 | | |
| Stomach | 5 | | |
| Duodenum | 1 | | |
| Unknown | 4 | | |
| Histology | | | |
| Adenocarcinoma | 34 | N/A | |
| Squamous
carcinoma | 3 | | |
| Unknown | 4 | | |
Chromatographic enrichment of urine
proteins and peptides, and sample preparation
Aliquots of 0.5 ml from individual cancer or control
urine samples was added to either 30 μl CM10 (n=33 cancer urines,
n=8 control urines) or 30 μl IMAC30 (Cu2+-chelated)
(n=21 cancer urines, n=19 control urines) spin column resin
(Bio-Rad) and 0.75 ml binding buffer (either 0.1 M
NaH3C2O2 pH 4.0 for CM resin, or
0.1 M NaHPO4 pH 7.0 including 0.5 M NaCl for IMAC30
resin) and incubated for 1 h at room temperature under constant
agitation. Sample and resin combinations were chosen based on
independent analyses using peak stratification by SELDI mass
spectrometry (9). Unbound material
was removed and the resin washed four times with 0.3 ml binding
buffer. Bound material was separated by electrophoresis on a 16.5%
Tris-Tricine gel (Bio-Rad), and gel bands in the region of 2–10 kDa
were excised after Coomassie staining (BioSafe Coomassie; Bio-Rad).
The molecular mass range of 2–10 kDa was selected since many
urinary proteins are derived from proteolytic processing and
urinary shedding as described (20). Additionally, we previously observed
potential urinary cancer markers in this mass range (9).
LC-MS/MS mass spectrometry
Proteins and peptides from gel bands were digested
in situ with trypsin. The resulting peptides were eluted
with acetonitrile (ACN), and analysed by LC-MS/ MS (21). The LC-MS system consisted of an
Agilent 1200 Series HPLC (Agilent Technologies, Yarnton, UK) with a
Kasil sealed fused silica pre-column (Next Advance, New York, NY,
USA) packed to a length of ~3 cm with Pursuit C18, 5 μm particle
size (Varian, Crawley, UK) and PicoTip Emitter analytical column PF
360-75-15-N-5 (New Objective, Woburn, MA, USA) packed to a length
of ~20 cm with Pursuit C18, 5 μm particle size (Varian). The column
was equilibrated with solvent A (0.1% formic acid in 2.5%
acetonitrile) and eluted with a linear gradient from 0 to 10% over
6 to 8 min; from 8 to 60% over 8 to 35 min; from 60 to 100% over 35
to 40 min; solvent B (0.1% formic acid, 0.025% TFA in 90%
acetonitrile) over 45 min at a flow rate of 5 μl/min. The LTQ mass
spectrometer (Thermo Scientific, Epsom, UK) was fitted with a
NanoLC ESI source. Data-dependent acquisition was controlled by
XCalibur software. Fragmentation spectra were then processed by
XCalibur and BioWorks software (Thermo Fisher Scientific,
Loughborough, UK) and submitted to the Mascot search engine (Matrix
Science, London, UK) using UniProt/SwissProt (release May 2011,
Homo sapiens, 18055 sequences) as the reference database.
Mascot search parameters were: enzyme specificity trypsin, maximum
missed cleavage 1, fixed modifications cysteine
carbamidomethylation, variable modification methionine oxidation,
precursor mass tolerance +/−3 kDa, fragment ion mass tolerance
+/−0.4 kDa. Only Mascot hits with a false discovery rate (FDR)
≤0.05 were taken into consideration.
Meta-analysis and subtractive data
analysis
Proteins with at least two peptide matches were
analysed further by comparing molecules that were only observed in
urine samples from cancer patients with a database consisting of
proteins found by other studies in urine, blood and kidney. This
database was assembled from 136 publications, listing 146
tissue-specific datasets. The blood datasets covered plasma, serum
and erythrocytes; the kidney studies were derived from analyses of
cortex, medulla, epithelium, glomerulus, inner medullary collecting
duct, mesangium, parenchyma, peroxisomal membrane, peroxisome,
basolateral membrane vesicles, brush border membrane vesicles,
urothelial mucosa and whole kidney; and urine datasets described
either the whole or exosomal proteomes. All entries were then
matched to the UniProt database, followed by clustering to
individual (unique) entries by annotating splice and variant
entries to common parent molecules and ultimately assigning each
unique cluster an in-house specific accession number. Additionally,
all proteins mapping to immunoglobulins were clustered into one
generic cluster, as well as all proteins belonging to the Major
Histocompatibility Complex (MHC). Merging and subtraction analysis
was done using software written in-house. We also manually added
our own functional classification tags to each molecular cluster,
based on known properties of each molecule, giving an abridged view
of proteome compositions.
Results
Urine samples were extracted from 21 healthy
non-cancer controls and 41 patients with upper GI cancer (n=41)
(Table I). Of the 41 cancer
patients, staging investigations demonstrated that at least 29
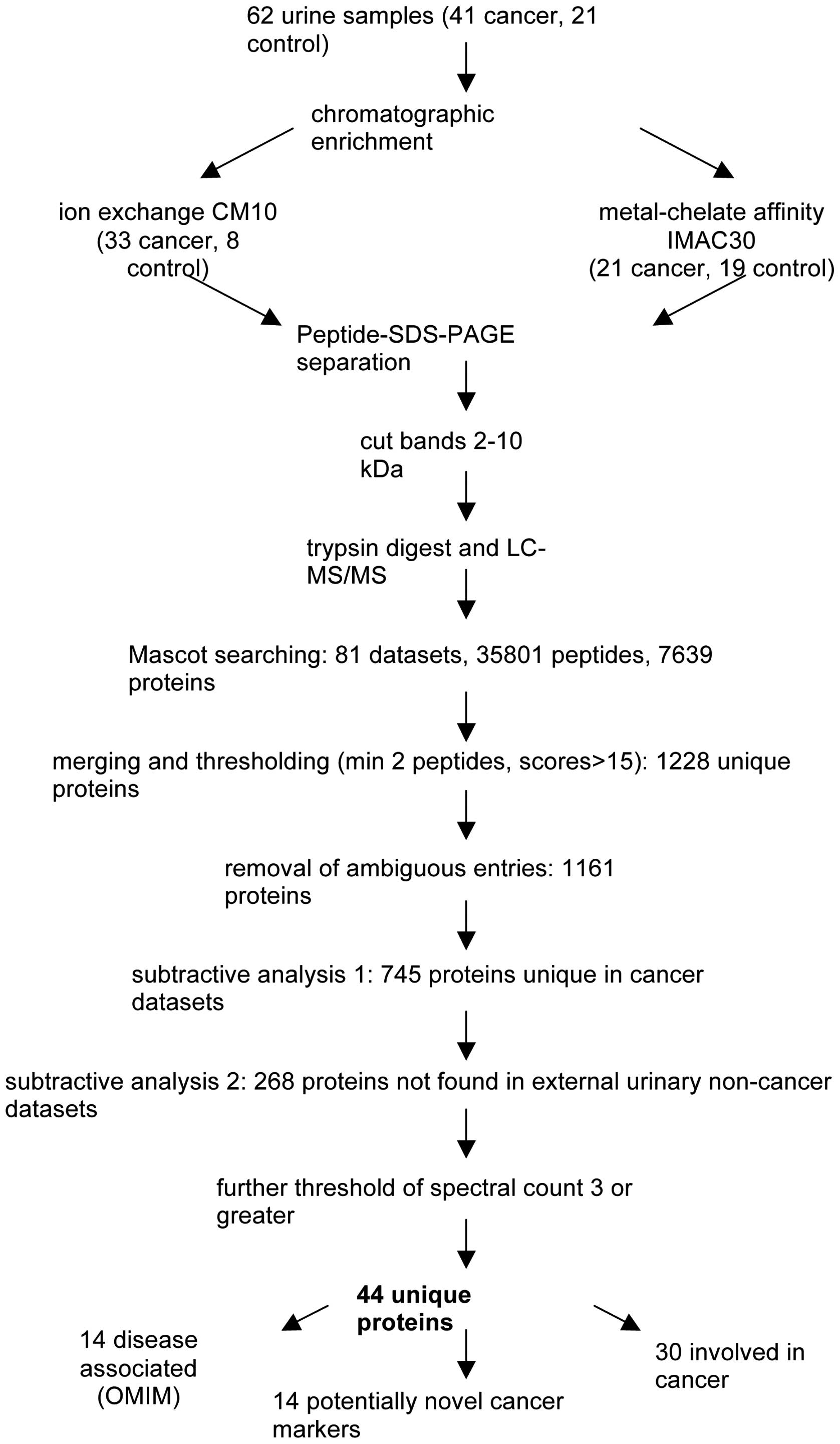
(70.7%) had nodal or metastatic disease. We analysed all 62 urine
samples by LC-MS/MS in the region of 2–10 kDa by chromatographic
enrichment using either CM10, IMAC30, or both resin types
individually, resulting in a total of 81 chromatographic
enrichments, followed by gel analysis, tryptic digestion and mass
spectrometry. All molecular weight regions cut from gels were
identical in at least three samples from each cohort group, thus
also allowing comparison of identified molecules on a gel-region by
gel-region basis After data extraction by Mascot searching
(resulting in 35,801 peptides covering 7,639 proteins) and applying
discovery criteria of a FDR ≤0.05 and a minimal Mascot score of 13,
the resulting 81 datasets were further analysed by merging all
protein lists. This yielded 1,228 unique non-redundant entries
(data not shown). Additionally, all molecules relating to either
immunoglobulins or MHC were also merged into two individual
clusters since members of these two families are well known to show
a great degree of hypervariability, and therefore they may skew any
analysis towards single entries from those classes, since they are
not expected to show any duplications across the datasets analysed
in this study. The final list consisted of 1,161 molecular
clusters. Furthermore, we re-classified all molecules in the
datasets available by manually annotating every protein with a
single molecular property or functionality tag as listed in the
legend of Fig. 1. The properties
or functionalities were assigned based on known properties of each
individual protein, either from original publications or derived
from database annotations, such as enzyme nomenclatures, sequence
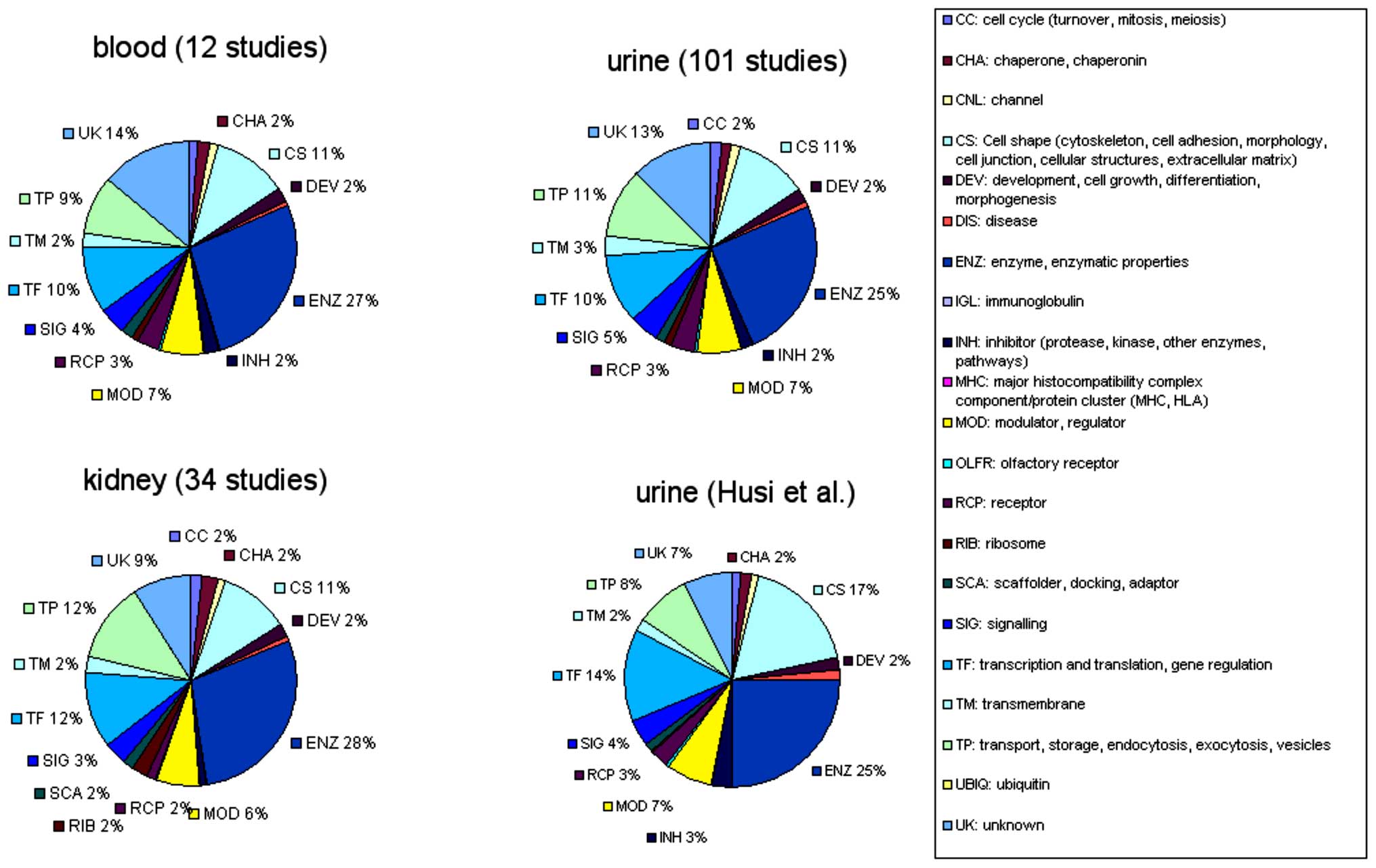
homologies and domain analysis. The compositional analysis of the
merged datasets of blood, urine and kidney proteomes, as well as
our urinary dataset is shown in Fig.
2. It was clear that all merged datasets consist of ~25%
enzymes, 10% cell-shape molecules, 10% transcriptional or
translational elements and 10% transport molecules. However, our
novel dataset appeared to contain more cell-shape and
transcriptional/translational proteins and less transport
molecules, which may reflect an association with disease, rather
than a general breakdown of cellular components.
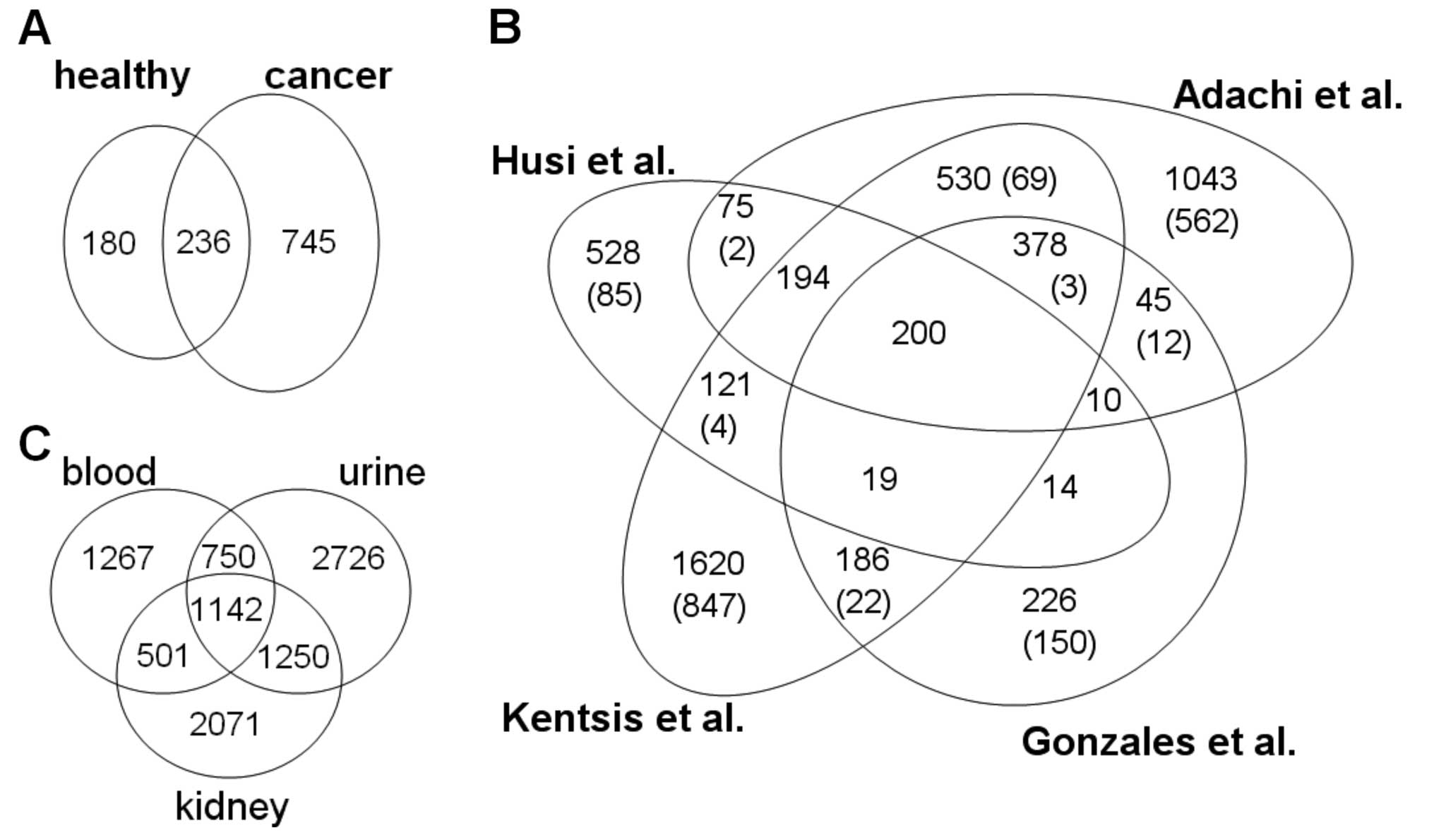
The 1,161 molecules were then split into groups
depending on whether they were observed in cancer urine samples, or
urine from healthy individuals (Fig.
3A). The 745 proteins only found in cancer urine samples were
then tagged and the entire dataset compared to data of 31,743
unmerged entries derived from 146 tissue-specific datasets from 137
publications (data not shown). This external data consisted of
9,707 merged entries, covering proteomic studies from urine, kidney
and blood (Table II). A
comparative analysis of our dataset with the three largest urinary
proteome profiling datasets showed a 46% overlap of our data with
the dataset from Kentsis et al (22), a 41% overlap with the study by
Adachi et al (23), and a
21% with the urinary exosome dataset from Gonzales et al
(24) (Fig. 3B). A global comparison between
proteomes from urine, kidney and blood (Fig. 3C) demonstrated a slightly larger
overlap of the urinary proteome with the kidney proteome than the
blood proteome.
 | Table IINumber of entries listed in the LSSR
database for analysed samples derived from blood, urine and
kidney. |
Table II
Number of entries listed in the LSSR
database for analysed samples derived from blood, urine and
kidney.
| No. of entries
prior to merge | Merged entries | No. of studies |
|---|
| Urine | 13,635 | 5,868 | 101 |
| Blood | 4,433 | 3,660 | 12 |
| Kidney | 13,675 | 4,964 | 34 |
We then performed subtractive analysis on our
urinary proteome data by eliminating any potential cancer candidate
molecule if it was found in any of the urinary datasets unrelated
to cancer. This reduced dataset of 268 proteins (data not shown)
was further condensed by removing any entries which did not have a
spectral count of at least two, resulting in 44 proteins, of which
24 were found uniquely in our study (in comparison to all other
datasets), and 20 which were also found in the other tissues
(Table III). All 44 of these
proteins were then analysed by searching the Online Mendelian
Inheritance in Man (OMIM) database for publications where these
molecules were reported to be directly associated with human
disease or cancer. Fourteen proteins were annotated in OMIM to be
causative for a disease, and 30 were known to be involved in
cancer.
 | Table IIIList of potential cancer candidate
markers from human urine. |
Table III
List of potential cancer candidate
markers from human urine.
| Peptide count | Spectral count | Gene | Protein | OMIM disease | PADB
classification | Tissue | Molecular
function | Cancer type | PubMed (cancer
association) |
|---|
| Only detected in
cancer patient urine, high confidence dataset |
| 10 | 11 | POTEKP | Putative
β-actin-like protein 3 | | CS: Cell shape | Urine | Actin filament
de-/re-polymerization | Hepatocellular
carcinoma | 16824795 |
| 194 | 3 | DCP1A | mRNA-decapping
enzyme 1A | | ENZ: enzyme,
enzymatic properties | Urine | Transcriptional
co-activator | Gastric cancer | 23932921 |
| 93 | 3 | NAV1 | Neuron navigator
1 | | DEV:
development | Urine | neuronal
migration | | |
| 80 | 3 | ZFYVE20 | Rabenosyn-5 | | TP: transport,
storage, endocytosis, exocytosis, vesicles | Urine | endosomal
transport | | |
| 77 | 3 | PLA1A | Phospholipase A1
member A | | ENZ: enzyme,
enzymatic properties | Urine | Lipid
metabolism | Prostate
cancer | 22904677 |
| 5 | 3 | GLB1L |
β-galactosidase-1-like protein | | ENZ: enzyme,
enzymatic properties | Urine | Glycosyl hydrolase,
carbohydrate metabolism | | |
| 32 | 2 | COX4I2 | Cytochrome c
oxidase subunit 4 isoform 2, mitochondrial | Exocrine pancreatic
insufficiency, dyserythropoietic anemia, calvarial
hyperostosis | TP: transport,
storage, endocytosis, exocytosis, vesicles | Urine | Mitochondrial
electron transport | General (Warburg
effect) | 22320183 |
| 20 | 2 | SOS2 | Son of sevenless
homolog 2 | | MOD: modulator,
regulator | Urine | Guanine-nucleotide
releasing factor | | |
| 20 | 2 | GALNT6 | Polypeptide
N-acetylgalactosa minyltransferase 6 | | ENZ: enzyme,
enzymatic properties | Urine | Post-translational
protein O-linked glycosylation | Breast cancer | 20215525 |
| 17 | 2 | CCDC88C | Protein Daple | Autosomal recessive
nonsyndromic hydrocephalus HYC1 | SIG: signaling | Urine | Negative regulator
of canonical Wnt signaling pathway | Breast cancer | 23593120 |
| 15 | 2 | TTI1 | TEL2-interacting
protein 1 homolog | | MOD: modulator,
regulator | Urine | Regulator of DNA
damage response | Multiple
myeloma | 23263282 |
| 13 | 2 | RPGRIP1 | X-linked
retinitis-pigmentosa GTPase regulator interacting protein 1 | Leber congenital
amaurosis 6 | CS: Cell shape | Urine | Sensory
transduction | | |
| 13 | 2 | GBP4 | Guanylate-binding
protein 4 | | DEV:
development | Urine | GTP hydrolysis | | |
| 11 | 2 | MTTP | Microsomal
triglyceride transfer protein large subunit | Aβ
lipoproteinemia | TP: transport,
storage, endocytosis, exocytosis, vesicles | Urine | Lipid transport,
plasma lipoprotein secretion | Small intestinal
cancer | 12630961 |
| 11 | 2 | ERBB2 | Receptor
tyrosine-protein kinase erbB-2 | Glioma
susceptibility 1; ovarian cancer; lung cancer; gastric cancer | ENZ: enzyme,
enzymatic properties | Urine | Protein tyrosine
kinase involved in transcriptional regulation | Multiple | 22014070 |
| 9 | 2 | PLEKHG2 | Pleckstrin homology
domain-containing family G member 2 | | MOD: modulator,
regulator | Urine | Guanine-nucleotide
releasing factor | Pancreatic
cancer | 24041470 |
| 7 | 2 | POLA2 | DNA polymerase α
subunit B | | TF: transcription
and translation | Urine | DNA replication and
cell proliferation | Melanoma | 24987109 |
| 6 | 2 | GPSM2 | G-protein-signaling
modulator 2 | Deafness, autosomal
recessive 82 | CC: cell cycle
(turnover, mitosis, meiosis) | Urine | G-protein coupled
receptor signaling pathway, spindle pole orientation | Breast cancer | 20589935 |
| 6 | 2 | GDNF | Glial cell
line-derived neurotrophic factor | Central
hypoventilation syndrome; Hirschsprung disease, susceptibility to,
3 | SIG: signaling | Urine | neurotrophic
factor | Pancreatic
cancer | 20960036 |
| 5 | 2 | DDHD2 | Phospholipase
DDHD2 | | ENZ: enzyme,
enzymatic properties | Urine | Lipid degradation
and metabolism | Breast cancer | 20940404 |
| 4 | 2 | TEAD2 | Transcriptional
enhancer factor TEF-4 | | TF: transcription
and translation | Urine | Transcription
regulation | Prostate
cancer | 19478945 |
| 3 | 2 | SARM1 | Sterile α and TIR
motif-containing protein 1 | | MOD: modulator,
regulator | Urine | Regulator of
Toll-like receptor signaling pathway | Colorectal
cancer | 20426761 |
| 3 | 2 | DOK7 | Downstream of
tyrosine kinase 7 | Myasthenia,
limb-girdle | SIG: signaling | Urine | Neuromuscular
synaptogenesis | Breast cancer | 23054610 |
| 2 | 2 | ZDHHC6 | Probable
palmitoyltransferase ZDHHC6 | | ENZ: enzyme,
enzymatic properties | Urine | Protein
palmitoylation | | |
| Detected in urine
from cancer patients and other tissues, high confidence
dataset |
| 9 | 5 | HIST3H3 | Histone H3.1t | | TF: transcription
and translation | Kidney, urine | Transcription
regulation, DNA repair, DNA replication | | |
| 25 | 4 | HIST1H1E | Histone H1.4 | | TF: transcription
and translation | Kidney, urine | Regulator of gene
transcription | Endometrial cancer
cells | 23682076 |
| 3 | 4 | BOLA2 | BolA-like protein
2 | | UK: unknown | Kidney, urine | Redox control | Liver cancer | 22653869 |
| 116 | 3 | NAV2 | Neuron navigator
2 | | ENZ: enzyme,
enzymatic properties | Blood, urine | Neuronal
development | Colorectal
carcinoma | 22810696 |
| 25 | 3 | MLL3 | Histone-lysine
N-methyltransferase MLL3 | | ENZ: enzyme,
enzymatic properties | Blood, urine | Transcriptional
coactivation | Colorectal
cancer | 21853109 |
| 39 | 2 | FMR1 | Fragile X mental
retardation 1 protein | Fragile x
tremor/ataxia syndrome; fragile x mental retardation syndrome;
premature ovarian failure 1 | TF: transcription
and translation | Kidney, urine | Translation
repressor | Hepatocellular
carcinoma | 17786358 |
| 37 | 2 | TJP1 | Tight junction
protein ZO-1 | | CS: cell shape | Kidney, urine | Tight junction
assembly, cell migration | Non-small cell lung
cancer | 24294375 |
| 35 | 2 | PCDH17 |
Protocadherin-17 | | CS: cell shape | Blood, urine | Calcium-dependent
cell-adhesion protein | Laryngeal squamous
cell carcinoma | 21213369 |
| 30 | 2 | NSUN5 | Putative
methyltransferase NSUN5 | Williams-Beuren
syndrome | ENZ: enzyme,
enzymatic properties | Blood, urine | Methyl-transferase,
embryonic development | | |
| 13 | 2 | ST14 | Suppressor of
tumorigenicity 14 protein | Ichthyosis with
hypotrichosis, autosomal recessive | ENZ: enzyme,
enzymatic properties | Kidney, urine | Degradation of
extracellular matrix | Breast cancer | 20716618 |
| 13 | 2 | SON | Bax antagonist
selected in saccharomyces 1 | | TF: transcription
and translation | Kidney, urine | Splicing
cofactor | | |
| 9 | 2 | CPB1 | Carboxypeptidase
B | | ENZ: enzyme,
enzymatic properties | Blood, urine | Carboxypeptidase,
protein degradation | Pancreatic
cancer | 1688389 |
| 6 | 2 | HBG2 | Hemoglobin subunit
γ-2 | Cyanosis transient
neonatal | TP: transport,
storage, endocytosis, exocytosis, vesicles | Blood, kidney,
urine | Oxygen
transport | | |
| 6 | 2 | AKAP2 | A-kinase anchor
protein 2 | | SCA: scaffolder,
docking, adaptor | Kidney, urine | Protein kinase
A-anchoring protein | Ovarian cancer | 19123201 |
| 5 | 2 | DYNLL2 | Dynein light chain
2, cytoplasmic | | TP: transport,
storage, endocytosis, exocytosis, vesicles | Blood, kidney,
urine | Microtubule-based
transport | | |
| 5 | 2 | NCOR1 | Nuclear receptor
corepressor 1 | | TF: transcription
and translation | Blood, urine | Transcriptional
repressor | Prostate
cancer | 20466759 |
| 5 | 2 | MKX | Homeobox protein
Mohawk | | TF: transcription
and translation | Kidney, urine | Morphogenetic
regulator of cell adhesion | | |
| 4 | 2 | ACTN2 | α-actinin-2 | Cardiomyopathy,
dilated, 1aa | CS: cell shape | Blood, kidney,
urine |
Actin-anchoring | Metastatic
pancreatic endocrine neoplasm | 15448002 |
| 4 | 2 | CORO1A | Coronin-1A | Immunodeficiency
8 | CS: cell shape | Blood, kidney,
urine | Crucial component
of cytoskeletal modulation | Breast cancer | 21489049 |
| 2 | 2 | CELF5 | CUGBP Elav-like
family member 5 | | TF: transcription
and translation | Kidney, urine | Regulation of
pre-mRNA alternative splicing | | |
Discussion
Proteomic large-scale analysis of tissues to define
a cancer state can be time- and resource-consuming, especially in
light of an unknown end-point. Therefore, it could be helpful to
compare a novel dataset with known data in order to establish
whether potential disease markers are observable, and thereby
analyse a simplified dataset for the disease in question. This
approach does not address the issue of quantitative comparisons,
but it is rather a qualitative approach. However, the resulting
list of potential candidate molecules will have a specificity of
100%. Here, we test this hypothesis by applying a subtractive
analysis method in conjunction with large-scale meta-analysis of
urinary datasets to screen for potential novel cancer markers
observable in human urine.
An initial comparison of functional profiles of
urine, blood and kidney proteomes showed no major discernible
difference between those datasets. This finding, in itself is not
surprising, since it is expected that these systems should reflect
an overall similar composition through a combination of immediate
environment and source. Blood, containing a substantial amount of
cells, is also expected to show a reasonably uniform functional
composition profile compared with other tissues e.g. kidney. Our
novel urinary dataset, having an expected bias towards an aberrant
functional profile due to overexpressed molecules associated with
disease, contains more molecules involved in cellular contacts,
morphology and cytoskeletal aspects, as well as
transcriptional/translational components, which may be directly
linked to abnormal and uncontrolled cellular growth.
Comparison of our dataset with known non-cancer
urinary proteomes yielded a set of only 44 molecules specific for
our cancer data, of which 68% are already known to be involved in
cancer. The functional profile of those 44 proteins in comparison
to the merged urinary proteome profile showed mainly an enrichment
of developmental proteins (5%), signaling molecules (7%) and, most
strikingly, transcriptional/ translational proteins (20%). The
known cancer-associated molecules described have been suggested to
be involved in hepatocellular carcinoma [κ actin (POTEKP) (25); BolA-like protein 2 (BOLA2)
(26); fragile X mental
retardation 1 protein (FMR1) (27)]; mammary carcinogenesis [polypeptide
N-acetylgalactosaminyltrans ferase 6 (GALNT6) (28); protein Daple (CCDC88C) (29); G-protein-signaling modulator 2
(GPSM2) (30);
phospho-lipase DDHD2 (DDHD2) (31); downstream of tyrosine kinase 7
(DOK7) (32); suppressor of
tumorigenicity 14 protein (ST14) (33); coronin-1A (CORO1A) (34)], lung cancer [tight junction protein
ZO-1 (TJP1) (35)], prostate
cancer [phos-pholipase A1 member A (PLA1A) (36); transcriptional enhancer factor
TEF-4 (TEAD2) (37);
nuclear receptor corepressor 1 (NCOR1) (38)], ovarian cancer [A-kinase anchor
protein 2 (AKAP2) (39)],
colorectal cancer [sterile α and TIR motif-containing protein 1
(SARM1) (40); neuron navigator 2
(NAV2) (41); histone-lysine
N-methyltransferase MLL3 (MLL3) (42)], pancreatic cancer [pleckstrin
homology domain-containing family G member 2 (PLEKHG2) (43); glial cell line-derived neurotrophic
factor (GDNF) (44);
carboxypeptidase B (CPB1) (45); α-actinin-2 (ACTN2) (46)], gastric cancer [mRNA-decapping
enzyme 1A (DCP1A) (47), a
co-activator in TGF-β signaling (48)], melanoma [DNA polymerase α subunit
B (POLA2) (49)], multiple myeloma
[TEL2-interacting protein 1 homolog (TTI1) (50)], endometrial cancer cells [Histone
H1.4 (HIST1H1E (51)], laryngeal
squamous cell carcinoma [protocadherin-17 (PCDH17) (52)], and adenocarcinoma [microsomal
triglyceride transfer protein large subunit (MTTP) (53)]. Additionally, the latter protein
was also described to be a pivotal element in the cancer-associated
muscle-wasting disease cachexia (54). Some of these proteins may be
differentially regulated across a range of different cancer types
and may therefore represent key cancer markers. For example,
receptor tyrosine-protein kinase erbB-2 (ERBB2) has been described
to be a marker for various cancer types, such as gastroesophageal
(55), breast (56), lung (57), gallbladder (58) and pancreatic cancer (59), as well as uterine serous
adenocarcinoma (60), and others.
Another known protein to be involved in cancer progression is the
mitochondrial cytochrome c oxidase subunit 4 isoform 2
(COX4I2), which is part of the Warburg effect, where cancer cells
show higher propensity to produce lactate independent of oxygen
presence or absence (61).
Of the proteins not previously described in
association with cancer, transcription factor Bax antagonists
selected in Saccharomyces 1 (SON), homeobox protein Mohawk (MKX)
and CUGBP Elav-like family member 5 (CELF5) may represent other
potential lead candidates in cancer stratification. Other important
markers may include developmental molecules, such as
guanylate-binding protein 4 GBP4, which is a negative regulator of
virus-triggered cellular responses (62) and is involved in GTP hydrolysis, or
neuron navigator NAV1, which has been reported to be a neuronal
guidance molecule (63). However,
its role in cancer or outside the neuronal environment remains to
be elucidated.
In conclusion, we have demonstrated that a
subtractive analysis of proteomic datasets can yield a number of
potential diagnostic cancer targets in human urine. Further
specific screening of urine, based on our findings, using, for
example, an antibody-based approach, will establish whether our
potential markers are associated with a general cancer status, or
if they are specific for a defined cancer type such as pancreatic
or esophageal cancer. Additionally, since the data in our database
can easily be expanded to contain further datasets, there are
other, as yet undefined diseases, which can be addressed by
establishing and comparing a relatively small disease-specific
dataset. This approach also has the advantage of rapid turnover and
increased cost-effectiveness relating to large-scale analyses of
tissue and cell proteomes for the discovery of novel molecular
markers. In this regard, we are encouraging researchers to submit
their published datasets to be incorporated in the LSSR database.
All data will be freely available through the PADB portal at
www.PADB.org.
Acknowledgements
We thank C.A. Greig, N.A. Stephens and H. Wackerhage
for patient recruitment. Funding of this study was provided by the
University of Edinburgh.
References
|
1
|
Good DM, Thongboonkerd V, Novak J,
Bascands JL, Schanstra JP, Coon JJ, Dominiczak A and Mischak H:
Body fluid proteomics for biomarker discovery: Lessons from the
past hold the key to success in the future. J Proteome Res.
6:4549–4555. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Marshall T and Williams K: Two-dimensional
electrophoresis of human urinary proteins following concentration
by dye precipitation. Electrophoresis. 17:1265–1272. 1996.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Pieper R, Gatlin CL, McGrath AM, Makusky
AJ, Mondal M, Seonarain M, Field E, Schatz CR, Estock MA, Ahmed N,
et al: Characterization of the human urinary proteome: A method for
high-resolution display of urinary proteins on two-dimensional
electrophoresis gels with a yield of nearly 1400 distinct protein
spots. Proteomics. 4:1159–1174. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Büeler MR, Wiederkehr F and Vonderschmitt
DJ: Electrophoretic, chromatographic and immunological studies of
human urinary proteins. Electrophoresis. 16:124–134. 1995.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Spahr CS, Davis MT, McGinley MD, Robinson
JH, Bures EJ, Beierle J, Mort J, Courchesne PL, Chen K, Wahl RC, et
al: Towards defining the urinary proteome using liquid
chromatography-tandem mass spectrometry. I Profiling an
unfractionated tryptic digest. Proteomics. 1:93–107. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Cadieux PA, Beiko DT, Watterson JD, Burton
JP, Howard JC, Knudsen BE, Gan BS, McCormick JK, Chambers AF,
Denstedt JD, et al: Surface-enhanced laser
desorption/ionization-time of flight-mass spectrometry
(SELDI-TOF-MS): A new proteomic urinary test for patients with
urolithiasis. J Clin Lab Anal. 18:170–175. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Roelofsen H, Alvarez-Llamas G, Schepers M,
Landman K and Vonk RJ: Proteomics profiling of urine with surface
enhanced laser desorption/ionization time of flight mass
spectrometry. Proteome Sci. 5:22007. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Vanhoutte KJ, Laarakkers C, Marchiori E,
Pickkers P, Wetzels JF, Willems JL, van den Heuvel LP, Russel FG
and Masereeuw R: Biomarker discovery with SELDI-TOF MS in human
urine associated with early renal injury: Evaluation with
computational analytical tools. Nephrol Dial Transplant.
22:2932–2943. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Husi H, Stephens N, Cronshaw A, MacDonald
A, Gallagher I, Greig C, Fearon KC and Ross JA: Proteomic analysis
of urinary upper gastrointestinal cancer markers. Proteomics Clin
Appl. 5:289–299. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Petri AL, Simonsen AH, Yip TT, Hogdall E,
Fung ET, Lundvall L and Hogdall C: Three new potential ovarian
cancer biomarkers detected in human urine with equalizer bead
technology. Acta Obstet Gynecol Scand. 88:18–26. 2009. View Article : Google Scholar
|
|
11
|
Tsui KH, Tang P, Lin CY, Chang PL, Chang
CH and Yung BY: Bikunin loss in urine as useful marker for bladder
carcinoma. J Urol. 183:339–344. 2010. View Article : Google Scholar
|
|
12
|
Chen YT, Chen CL, Chen HW, Chung T, Wu CC,
Chen CD, Hsu CW, Chen MC, Tsui KH, Chang PL, et al: Discovery of
novel bladder cancer biomarkers by comparative urine proteomics
using iTRAQ technology. J Proteome Res. 9:5803–5815. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Tan LB, Chen KT, Yuan YC, Liao PC and Guo
HR: Identification of urine PLK2 as a marker of bladder tumors by
proteomic analysis. World J Urol. 28:117–122. 2010. View Article : Google Scholar
|
|
14
|
Xue A, Scarlett CJ, Chung L, Butturini G,
Scarpa A, Gandy R, Wilson SR, Baxter RC and Smith RC: Discovery of
serum biomarkers for pancreatic adenocarcinoma using proteomic
analysis. Br J Cancer. 103:391–400. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Schröder C, Jacob A, Tonack S, Radon TP,
Sill M, Zucknick M, Rüffer S, Costello E, Neoptolemos JP,
Crnogorac-Jurcevic T, et al: Dual-color proteomic profiling of
complex samples with a microarray of 810 cancer-related antibodies.
Mol Cell Proteomics. 9:1271–1280. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Good DM, Zürbig P, Argilés A, Bauer HW,
Behrens G, Coon JJ, Dakna M, Decramer S, Delles C, Dominiczak AF,
et al: Naturally occurring human urinary peptides for use in
diagnosis of chronic kidney disease. Mol Cell Proteomics.
9:2424–2437. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Zhang Y, Zhang Y, Adachi J, Olsen JV, Shi
R, de Souza G, Pasini E, Foster LJ, Macek B, Zougman A, et al:
MAPU: Max-Planck Unified database of organellar, cellular, tissue
and body fluid proteomes. Nucleic Acids Res. 35:D771–D779. 2007.
View Article : Google Scholar :
|
|
18
|
Li SJ, Peng M, Li H, Liu BS, Wang C, Wu
JR, Li YX and Zeng R: Sys-BodyFluid: A systematical database for
human body fluid proteome research. Nucleic Acids Res.
37:D907–D912. 2009. View Article : Google Scholar :
|
|
19
|
Agron IA, Avtonomov DM, Kononikhin AS,
Popov IA, Moshkovskii SA and Nikolaev EN: Accurate mass tag
retention time database for urine proteome analysis by
chromatography-mass spectrometry. Biochemistry (Mosc). 75:636–641.
2010. View Article : Google Scholar
|
|
20
|
Carson JM, Okamura K, Wakashin H, McFann
K, Dobrinskikh E, Kopp JB and Blaine J: Podocytes degrade
endocytosed albumin primarily in lysosomes. PLoS One. 9:e997712014.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Collins MO, Yu L and Choudhary JS:
Analysis protein complexes by 1D-SDS-PAGE and tandem mass
spectrometry. Protocol Exchange. 2008. View Article : Google Scholar
|
|
22
|
Kentsis A, Monigatti F, Dorff K, Campagne
F, Bachur R and Steen H: Urine proteomics for profiling of human
disease using high accuracy mass spectrometry. Proteomics Clin
Appl. 3:1052–1061. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Adachi J, Kumar C, Zhang Y, Olsen JV and
Mann M: The human urinary proteome contains more than 1500
proteins, including a large proportion of membrane proteins. Genome
Biol. 7:R802006. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Gonzales PA, Pisitkun T, Hoffert JD,
Tchapyjnikov D, Star RA, Kleta R, Wang NS and Knepper MA:
Large-scale proteomics and phosphoproteomics of urinary exosomes. J
Am Soc Nephrol. 20:363–379. 2009. View Article : Google Scholar :
|
|
25
|
Chang KW, Yang PY, Lai HY, Yeh TS, Chen TC
and Yeh CT: Identification of a novel actin isoform in
hepatocellular carcinoma. Hepatol Res. 36:33–39. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Hunecke D, Spanel R, Länger F, Nam SW and
Borlak J: MYC-regulated genes involved in liver cell dysplasia
identified in a transgenic model of liver cancer. J Pathol.
228:520–533. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Liu Y, Zhu X, Zhu J, Liao S, Tang Q, Liu
K, Guan X, Zhang J and Feng Z: Identification of differential
expression of genes in hepatocellular carcinoma by suppression
subtractive hybridization combined cDNA microarray. Oncol Rep.
18:943–951. 2007.PubMed/NCBI
|
|
28
|
Park JH, Nishidate T, Kijima K, Ohashi T,
Takegawa K, Fujikane T, Hirata K, Nakamura Y and Katagiri T:
Critical roles of mucin 1 glycosylation by transactivated
polypeptide N-acetylgalactosaminyltransferase 6 in mammary
carcinogenesis. Cancer Res. 70:2759–2769. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Long J, Zhang B, Signorello LB, Cai Q,
Deming-Halverson S, Shrubsole MJ, Sanderson M, Dennis J,
Michailidou K, Easton DF, et al: Evaluating genome-wide association
study-identified breast cancer risk variants in African-American
women. PLoS One. 8:e583502013. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Fukukawa C, Ueda K, Nishidate T, Katagiri
T and Nakamura Y: Critical roles of LGN/GPSM2 phosphorylation by
PBK/TOPK in cell division of breast cancer cells. Genes Chromosomes
Cancer. 49:861–872. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Yang ZQ, Liu G, Bollig-Fischer A, Giroux
CN and Ethier SP: Transforming properties of 8p11–12 amplified
genes in human breast cancer. Cancer Res. 70:8487–8497. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Heyn H, Carmona FJ, Gomez A, Ferreira HJ,
Bell JT, Sayols S, Ward K, Stefansson OA, Moran S, Sandoval J, et
al: DNA methylation profiling in breast cancer discordant identical
twins identifies DOK7 as novel epigenetic biomarker.
Carcinogenesis. 34:102–108. 2013. View Article : Google Scholar :
|
|
33
|
Kauppinen JM, Kosma VM, Soini Y, Sironen
R, Nissinen M, Nykopp TK, Kärjä V, Eskelinen M, Kataja V and
Mannermaa A: ST14 gene variant and decreased matriptase protein
expression predict poor breast cancer survival. Cancer Epidemiol
Biomarkers Prev. 19:2133–2142. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Hattori N, Okochi-Takada E, Kikuyama M,
Wakabayashi M, Yamashita S and Ushijima T: Methylation silencing of
angio-poietin-like 4 in rat and human mammary carcinomas. Cancer
Sci. 102:1337–1343. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Ni S, Xu L, Huang J, Feng J, Zhu H, Wang G
and Wang X: Increased ZO-1 expression predicts valuable prognosis
in non-small cell lung cancer. Int J Clin Exp Pathol. 6:2887–2895.
2013.PubMed/NCBI
|
|
36
|
Paulo P, Ribeiro FR, Santos J, Mesquita D,
Almeida M, Barros-Silva JD, Itkonen H, Henrique R, Jerónimo C,
Sveen A, et al: Molecular subtyping of primary prostate cancer
reveals specific and shared target genes of different ETS
rearrangements. Neoplasia. 14:600–611. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Blum R, Gupta R, Burger PE, Ontiveros CS,
Salm SN, Xiong X, Kamb A, Wesche H, Marshall L, Cutler G, et al:
Molecular signatures of prostate stem cells reveal novel signaling
pathways and provide insights into prostate cancer. PLoS One.
4:e57222009. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Battaglia S, Maguire O, Thorne JL, Hornung
LB, Doig CL, Liu S, Sucheston LE, Bianchi A, Khanim FL, Gommersall
LM, et al: Elevated NCOR1 disrupts PPARalpha/gamma signaling in
prostate cancer and forms a targetable epigenetic lesion.
Carcinogenesis. 31:1650–1660. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Quinn MC, Filali-Mouhim A, Provencher DM,
Mes-Masson AM and Tonin PN: Reprogramming of the transcriptome in a
novel chromosome 3 transfer tumor suppressor ovarian cancer cell
line model affected molecular networks that are characteristic of
ovarian cancer. Mol Carcinog. 48:648–661. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Quyun C, Ye Z, Lin SC and Lin B: Recent
patents and advances in genomic biomarker discovery for colorectal
cancers. Recent Pat DNA Gene Seq. 4:86–93. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Cancer Genome Atlas Network. Comprehensive
molecular characterization of human colon and rectal cancer.
Nature. 487:330–337. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Watanabe Y, Castoro RJ, Kim HS, North B,
Oikawa R, Hiraishi T, Ahmed SS, Chung W, Cho MY, Toyota M, et al:
Frequent alteration of MLL3 frameshift mutations in microsatellite
deficient colorectal cancer. PLoS One. 6:e233202011. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Shain AH, Salari K, Giacomini CP and
Pollack JR: Integrative genomic and functional profiling of the
pancreatic cancer genome. BMC Genomics. 14:6242013. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Liu H, Ma Q and Li J: High glucose
promotes cell proliferation and enhances GDNF and RET expression in
pancreatic cancer cells. Mol Cell Biochem. 347:95–101. 2011.
View Article : Google Scholar
|
|
45
|
Fernstad R, Pousette A, Carlström K and
Sköldefors H: A novel assay for pancreatic cellular damage: IV.
Serum concentrations of pancreas-specific protein (PASP) in acute
pancreatitis and other abdominal diseases. Pancreas. 5:42–49. 1990.
View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Hansel DE, Rahman A, House M, Ashfaq R,
Berg K, Yeo CJ and Maitra A: Met proto-oncogene and insulin-like
growth factor binding protein 3 overexpression correlates with
metastatic ability in well-differentiated pancreatic endocrine
neoplasms. Clin Cancer Res. 10:6152–6158. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Iio A, Takagi T, Miki K, Naoe T, Nakayama
A and Akao Y: DDX6 post-transcriptionally down-regulates
miR-143/145 expression through host gene NCR143/145 in cancer
cells. Biochim Biophys Acta. 1829:1102–1110. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Bai RY, Koester C, Ouyang T, Hahn SA,
Hammerschmidt M, Peschel C and Duyster J: SMIF, a Smad4-interacting
protein that functions as a co-activator in TGFbeta signaling. Nat
Cell Biol. 4:181–190. 2002. View
Article : Google Scholar : PubMed/NCBI
|
|
49
|
Lu YC, Yao X, Crystal JS, Li YF, El-Gamil
M, Gross C, Davis L, Dudley ME, Yang JC, Samuels Y, et al:
Efficient identification of mutated cancer antigens recognized by T
cells associated with durable tumor regressions. Clin Cancer Res.
20:3401–3410. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Fernández-Sáiz V, Targosz BS, Lemeer S,
Eichner R, Langer C, Bullinger L, Reiter C, Slotta-Huspenina J,
Schroeder S, Knorn AM, et al: SCFFbxo9 and CK2 direct the cellular
response to growth factor withdrawal via Tel2/Tti1 degradation and
promote survival in multiple myeloma. Nat Cell Biol. 15:72–81.
2013. View Article : Google Scholar
|
|
51
|
Lee LR, Teng PN, Nguyen H, Hood BL,
Kavandi L, Wang G, Turbov JM, Thaete LG, Hamilton CA, Maxwell GL,
et al: Progesterone enhances calcitriol antitumor activity by
upregulating vitamin D receptor expression and promoting apoptosis
in endometrial cancer cells. Cancer Prev Res (Phila). 6:731–743.
2013. View Article : Google Scholar
|
|
52
|
Giefing M, Zemke N, Brauze D,
Kostrzewska-Poczekaj M, Luczak M, Szaumkessel M, Pelinska K,
Kiwerska K, Tönnies H, Grenman R, et al: High resolution ArrayCGH
and expression profiling identifies PTPRD and PCDH17/PCH68 as tumor
suppressor gene candidates in laryngeal squamous cell carcinoma.
Genes Chromosomes Cancer. 50:154–166. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Al-Shali K, Wang J, Rosen F and Hegele RA:
Ileal adenocarcinoma in a mild phenotype of abetalipoproteinemia.
Clin Genet. 63:135–138. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Silvério R, Laviano A, Rossi Fanelli F and
Seelaender M: L-Carnitine induces recovery of liver lipid
metabolism in cancer cachexia. Amino Acids. 42:1783–1792. 2012.
View Article : Google Scholar
|
|
55
|
Hjortland GO, Meza-Zepeda LA, Beiske K,
Ree AH, Tveito S, Hoifodt H, Bohler PJ, Hole KH, Myklebost O,
Fodstad O, et al: Genome wide single cell analysis of chemotherapy
resistant metastatic cells in a case of gastroesophageal
adenocarcinoma. BMC Cancer. 11:4552011. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Adachi R, Horiuchi S, Sakurazawa Y,
Hasegawa T, Sato K and Sakamaki T: ErbB2 down-regulates
microRNA-205 in breast cancer. Biochem Biophys Res Commun.
411:804–808. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Janku F, Garrido-Laguna I, Petruzelka LB,
Stewart DJ and Kurzrock R: Novel therapeutic targets in non-small
cell lung cancer. J Thorac Oncol. 6:1601–1612. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Goldin RD and Roa JC: Gallbladder cancer:
A morphological and molecular update. Histopathology. 55:218–229.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Komoto M, Nakata B, Amano R, Yamada N,
Yashiro M, Ohira M, Wakasa K and Hirakawa K: HER2 overexpression
correlates with survival after curative resection of pancreatic
cancer. Cancer Sci. 100:1243–1247. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Elsahwi KS and Santin AD: erbB2
overexpression in uterine serous cancer: A molecular target for
trastuzumab therapy. Obstet Gynecol Int. 2011:1282952011.
View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Mazzio EA, Boukli N, Rivera N and Soliman
KF: Pericellular pH homeostasis is a primary function of the
Warburg effect: Inversion of metabolic systems to control lactate
steady state in tumor cells. Cancer Sci. 103:422–432. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Hu Y, Wang J, Yang B, Zheng N, Qin M, Ji
Y, Lin G, Tian L, Wu X, Wu L, et al: Guanylate binding protein 4
negatively regulates virus-induced type I IFN and antiviral
response by targeting IFN regulatory factor 7. J Immunol.
187:6456–6462. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Maes T, Barceló A and Buesa C: Neuron
navigator: A human gene family with homology to unc-53, a cell
guidance gene from Caenorhabditis elegans. Genomics. 80:21–30.
2002. View Article : Google Scholar : PubMed/NCBI
|