Introduction
Prostate cancer is the second leading cause of
cancer-related mortality in males in the UK, with 9,900 deaths each
year, and it accounts for 13% of cancer-related deaths in males.
Approximately 85% of these cases involve men over 70 years of age.
The mortality rate for prostate cancer peaked in the early 1990s
and has now fallen to approximately 25 per 100,000 individuals at
risk. In the UK, the survival rates have been improving, and the
5-year relative survival rate was 60% in the period 1993 to 1995
(1).
Metastasis is a complex multi-step process by which
primary tumour cells invade adjacent tissue, enter the systemic
circulation (intravasate), translocate through the vasculature,
arrest in distant capillaries, extravasate into the surrounding
tissue parenchyma and finally proliferate from microscopic growths
(micrometastases) into macroscopic secondary tumours. Metastases
can be located in various organs and in different regions of the
same organ. The organ microenvironment modifies the response of
metastatic tumour cells to therapy and alters the effectiveness of
anticancer agents in destroying the tumour cells without producing
undesirable toxic effects. The major obstacle to treating
metastasis is the biological heterogeneity of primary neoplasms and
metastases. By the time of diagnosis, cancers contain multiple
genetically unstable cell populations with diverse karyotypes,
growth rates, cell-surface properties, antigenicities,
immunogenicities, marker enzymes, sensitivity to various cytotoxic
drugs and abilities to invade and produce metastasis (2).
Angiogenesis plays a key role in the pathogenesis of
a variety of disorders, including cancer, proliferative
retinopathies and rheumatoid arthritis. Accumulating evidence
indicates that, for most tumours, the switch to an angiogenic
phenotype depends upon the outcome of a balance between angiogenic
stimuli and angiogenic inhibitors, both of which may be produced by
tumour cells and perhaps by certain host cells (3). Growth and motility factors play an
essential role in migration processes at various levels of the
metastatic cascade. These factors include metastasis activators and
suppressors, which act in autocrine or paracrine manners through
special receptors that mediate different signals through tyrosine
phosphorylation (4). However,
metastasis suppressors may inhibit metastasis at any step of the
metastatic cascade without blocking tumourigenicity (5).
Metastasis tumour suppressor-1 (MTSS1), also known
as Missing in Metastasis (MIM), was originally identified as a
tumour suppressor since it is expressed in non-metastatic, but
absent from metastatic, bladder cancer cells (6,7). It
is expressed during development in muscles, kidneys and the liver
(8,9). The MTSS1 gene encodes a 5.3-kb mRNA,
and it is a polypeptide of 356 amino acids with homology to the
Wiscott-Aldrich Syndrome protein family (6). The MIM protein and MIM-B, a much
longer 759-amino acid protein whose C-terminus is identical to the
356 amino acids encoded by the MIM gene (10), are cytoplasmic in
location and have multidomain and scaffolding function (9).
MIM-B induces actin-rich protrusions resembling
microspikes and lamellipodia at the plasma membrane and promotes
disassembly of actin stress fibres. The actin cytoskeleton plays a
key role in regulating essential cellular processes, such as
endocytosis, cell migration, cytokinesis and various morphogenetic
processes. In addition, MTSS1 enhances Arp2/3-mediated actin
polymerization through interactions with cortactin (11). It is thus involved in cell motility
and morphogenesis, and studies suggest that further analysis of
MTSS1 expression or inactivation in tissue samples may define a new
candidate for use as a marker for primary tumours or metastasis
(12). MTSS1 is also a member of
the sonic hedgehog signalling pathway, which interacts and
modulates Gli responses during cell growth and carcinogenesis
(8).
This study aimed to assess MTSS1 expression levels
in prostate cancer cell lines in order to reveal any changes in
cell properties and to clarify the cellular function of MTSS1 in
these cancer cells. In the present study, we analysed MTSS1 through
a series of expression and inactivation studies to clarify the
function of MTSS1 in prostate cancer cells.
Materials and methods
Cell lines and culture
The cells were routinely cultured with Dulbecco’s
modified Eagle’s medium (DMEM) (PAA Laboratories Ltd., Somerset,
UK) supplemented with 10% fetal calf serum (PAA Laboratories Ltd.),
penicillin and streptomycin. This study used human prostate cancer
(DU-145, PC-3 and CA-HPV10) and human breast cancer (ZR-751 and
MDA-MB-231) cell lines. The cell lines were purchased from the
American Type Culture Collection (ATCC, Rockville, MD, USA) and
were stored at 37°C in 5% CO2 and under 95%
humidity.
RNA preparation and reverse
transcription-polymerase chain reaction
Total cellular RNA was isolated from the homogenized
human cell lines using Total RNA Isolation reagent (Advanced
Biotechnologies Ltd., Epsom, Surrey, UK). RNA concentration and
quality were determined through spectrophotometric measurement (WPA
UV 1101; Biotech Photometer, Cambridge, UK). cDNA was generated
using 200 ng of each RNA sample and a transcription kit (Sigma,
Poole, Dorset, UK) to reverse transcribe the RNA samples into cDNA.
DNA quality was verified using GAPDH. MTSS1 mRNA levels were
assessed using MTSS1 primers (sense, TCAAGAACAGATGGAAGAATGG;
antisense, TGCGGTAGCGGTAATGTG). PCR was carried out using a T-Cy
Thermocycler (Creacon Technologies Ltd., The Netherlands) and
REDTaq® ReadyMix™ PCR reaction mix (Sigma). The PCR
conditions consisted of a 40-sec denaturation step (95°C), a 2-min
annealing step (58°C) and a 3-min elongation step (72°C);
elongation took place over 36 cycles. The PCR products were next
loaded onto a 0.8% agarose gel and electrophoretically separated
prior to being stained with ethidium bromide and visualised under
UV light.
Quantitative real-time polymerase chain
reaction
Quantitative real-time polymerase chain reaction
(QPCR) is a technology that provides a broad dynamic range for
detecting specific gene sequences with high sensitivity. To
quantify the level of MTSS1 transcripts in the prostate cancer cell
lines, the iCycler IQ system (BioRad, Camberley, UK) was used. cDNA
samples were examined for MTSS1 expression using the MTSS1 QPCR
primers (sense, ATATCCCAGGATGCCTTC; antisense,
ACTGAACCTGACCGTACACGGTTCTCGCTTCTCTTT). The QPCR technique utilized
the Amplifluor System™ (Intergen Inc., UK) and Q-PCR master mix
(ABgene, Surrey, UK), in conjunction with a universal probe
(UniPrimer™). The real-time QPCR conditions were: 95°C for 15 min,
followed by 60 cycles at 95°C for 20 sec, 55°C for 30 sec and 72°C
for 20 sec.
Generation of ribozyme transgenes and
MTSS1 knockdown cells
MTSS1 expression levels were reduced in the DU-145
prostate cancer cell line using a ribozyme system. Briefly,
ribozyme transgenes that specifically cleave MTSS1 mRNA were
designed based on the predicted secondary structure of MTSS1. These
ribozymes were then generated using touchdown PCR and subsequently
cloned into the pEF6/V5-His-TOPO vector and amplified in
Escherichia coli. Plasmids were purified and verified for
correct size and orientation of the ribozymes, and electroporated
into the DU-145 prostate cancer cell line. A closed
pEF6/V5/His-TOPO plasmid (containing no ribozyme sequence) was also
electroporated into the same cell line to create a control group.
After selection using blasticidin, the unaltered wild-type cells
were termed DU-145 WT, the wild-type cells containing closed
plasmid only were termed DU-145 PEF and the wild-type cells
containing plasmid with a ribozyme sequence were termed DU-145
MTSSIKD.
Generation of MTSS1-overexpressing cell
lines
The full sequence of MTSS1 was amplified from cDNA
using the standard PCR procedure and a master mix with a
proofreading enzyme (sense primer, ATGGAGGCTGTGATTGAG; antisense,
CTAAGAAAAGCGAGGGG). This MTSS1 sequence was then T-A cloned into
the pEF6/V5/His-TOPO vector (Invitrogen, Paisley, UK) and then
electroporated into the PC-3 prostate cancer cell line with the aim
of enhancing MTSS1 expression in a cell line that does not normally
express it. Multiple clones were used, assessed and sequenced. The
PC-3 cells thus prepared and expressing MTSS1 were referred to in
the study as PC-3 MTSS1Exp. The control group of cells contained
the same plasmid vector (minus the MTSS1 sequence) and was termed
PC-3 PEF.
Confirmation of MTSS1 overexpression and
knockdown by Western blotting
MTSS1 protein expression was assessed in the human
prostate cancer cell line lysates through standard SDS-PAGE and
Western blot analysis. The cells were grown to confluence in a
75-cm2 tissue culture flask before being detached using
a cell scraper and pelleted. The cell pellet was then lysed in HCMF
buffer with 0.5% SDS, 1% Triton X-100, 2 mM CaCl2, 100
μg/ml phenylmethylsulfonyl fluoride, 1 mg/ ml leupeptin, 1
mg/ml aprotinin and 10 mM sodium orthovana-date on a rotor wheel
for 1 h and spun at 13,000 × g for 15 min to remove insolubles. The
lysed protein was then quantified using the Bio-Rad DC Protein
Assay kit (Bio-Rad Laboratories, CA, USA), and the samples were
normalized to a standard final concentration of 1.5 mg/ml following
addition of Sample Buffer, Laemmli 2X concentrate (Sigma) in a 1:1
ratio. The samples were then boiled for 5 min before being loaded
into a 10% polyacrylamide gel. Following electrophoresis, proteins
were blotted onto a Hybond-C Extra nitrocellulose membrane
(Amersham Biosciences UK Ltd., Bucks, UK), blocked in 10% milk and
subjected to specific antibody probing. The anti-MTSS1 antibody
(SC-98376, Santa Cruz Biotechnologies Inc., Santa Cruz, CA, USA)
was used to probe for MTSS1 at a concentration of 1:500 and
anti-GAPDH (Santa Cruz Biotechnologies Inc.) at a concentration of
1:500 for GAPDH. Probing with the primary antibody was followed by
probing with peroxidase-conjugated anti-rabbit (MTSS1) or
anti-rabbit (GAPDH) antibody (Sigma) at a 1:1,000 concentration.
The Supersignal West Dura Extended Duration substrate
chemiluminescent system (Perbio Science UK Ltd., Cramlington, UK)
was then used to visualize the protein bands. GAPDH expression was
used as an internal control (Santa Cruz Biotechnologies, Santa
Cruz, CA, USA). Protein expression was assessed and quantified
using Uvitech analysis software (Uvitech, Cambridge, UK).
Tumour cell growth assay
The effects of the modification of MTSS1 expression
on prostate cancer cell growth rates were assessed using an in
vitro growth assay. The cells were seeded in triplicate into
96-well plates at a density of 3,000 cells/well. Plates were then
incubated for 1, 3 and 5 days. Cell density was recorded after 1, 3
and 5 days by fixing cells in 4% formaldehyde, washing and staining
with 0.5% (w/v) crystal violet. Following this, the crystal violet
stain was extracted with 10% (v/v) acetic acid before reading the
absorbance on a Bio-Tek ELx800 multiplate reader (Bio-Tek
Instruments Inc., VT, USA).
Cell adhesion assay
The adhesive properties of the MTSS1-modified cells
to an artificial basement membrane were quantified using the in
vitro Matrigel adhesion assay. DU-145 WT, DU-145 PEF control
and DU-145 MTSS1KD cells and PC-3 WT, PC-3 PEF control and PC-3
MTSS1Exp prostate cancer cells were seeded at a density of
50,000/well (in triplicate) into a 96-well plate that had been
previously coated with 5 μg of Matrigel artificial basement
membrane. The cells were then incubated for 45 min to allow them to
adhere before being subjected to vigorous washing (x4) in BSS to
remove the non-adherent cells. The adherent cells were then fixed
in 4% formaldehyde (v/v) and stained with 0.5% (w/v) crystal
violet. The number of stained adherent cells was counted in several
random fields (≤40 objective magnifications).
Wounding assay
The migratory properties of PC-3 and DU-145 cells
were assessed to determine the impact of the forced expression or
knockout of MTSS1 on the invasive nature of these prostate cancer
cells. Cells at a density of 106 were incubated in 10 ml
of growth medium containing 100 μl of Cytodex-2 beads (GE
Healthcare, Cardiff, UK) for 3 h to allow the cells to adhere to
the beads. The beads were then washed to remove non-adherent or
dead cells and resuspended in 5 ml of medium. The cell/bead complex
was then incubated in a 96-well plate overnight. Following
incubation, the cells were washed with BSS, and cells that had
migrated from the Cytodex-2 beads and adhered to the base of the
well were fixed in 4% formaldehyde (v/v), stained with 0.5% crystal
violet (w/v) and counted under x40 objective magnification. At
least five random fields were counted per well, and five duplicate
wells were set up per sample. The entire experimental procedure was
repeated three independent times.
Statistical analysis
All in vitro experimentation was repeated at
least three independent times. The results were assessed using a
two-sample, two-tailed t-test and the Minitab 14 statistical
package. The values presented represent the mean ± SEM, and p≤0.05
was considered statistically significant. In vivo data were
analysed using a non-parametric Mann-Whitney test, as the data did
not follow a normal distribution.
Results
MTSS1 expression in human cancer cell
lines and creation of prostate cancer cell sublines with
differential patterns of MTSS1 expression
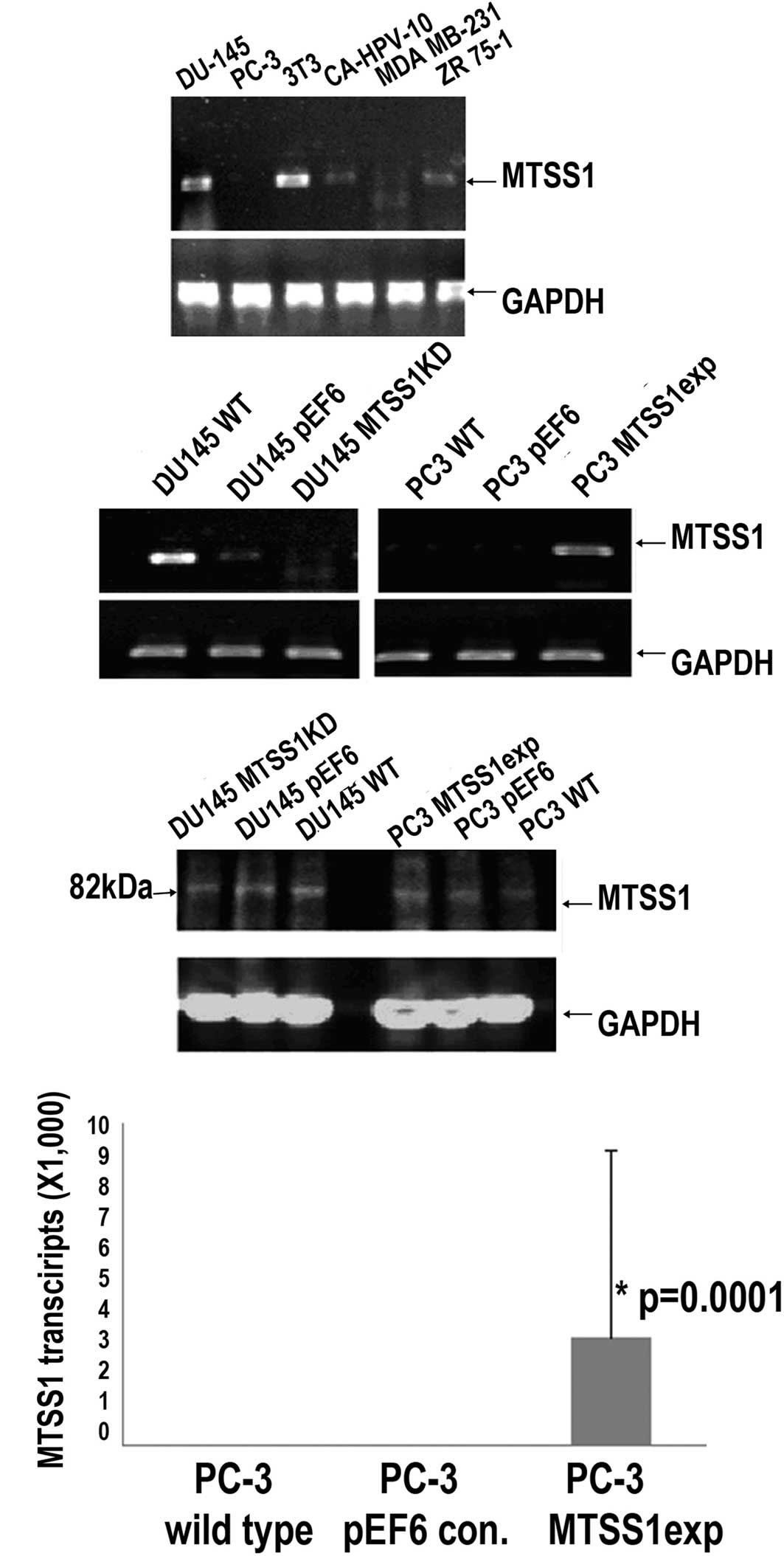
MTSS1 was found to be highly expressed in the DU-145
and 3T3 prostate cancer cells and at moderate levels in the
CA-HPV-10 prostate and ZR 75-1 breast cancer cells, while the
highly invasive prostate cancer cell line PC-3 showed little
expression (Fig. 1A). The highly
invasive PC-3 prostate cancer cell line was transfected with the
MTSS1 expression vector. Following selection, an
MTSS1-overexpressing subline was established (Fig. 1B–D). The DU-145 prostate cancer
cell line was also an appropriate candidate for knockdown as the
low-invasive cell line expressed levels of MTSS1. Anti-MTSS1
trangenes were used to knockdown MTSS1 expression in the DU-145
cells, followed by the establishment of a new subline which
expressed a low level of MTSS1 (Fig.
1B and C).
Regulation of MTSS1 expression affects
the rate of cell growth of prostate cancer cells
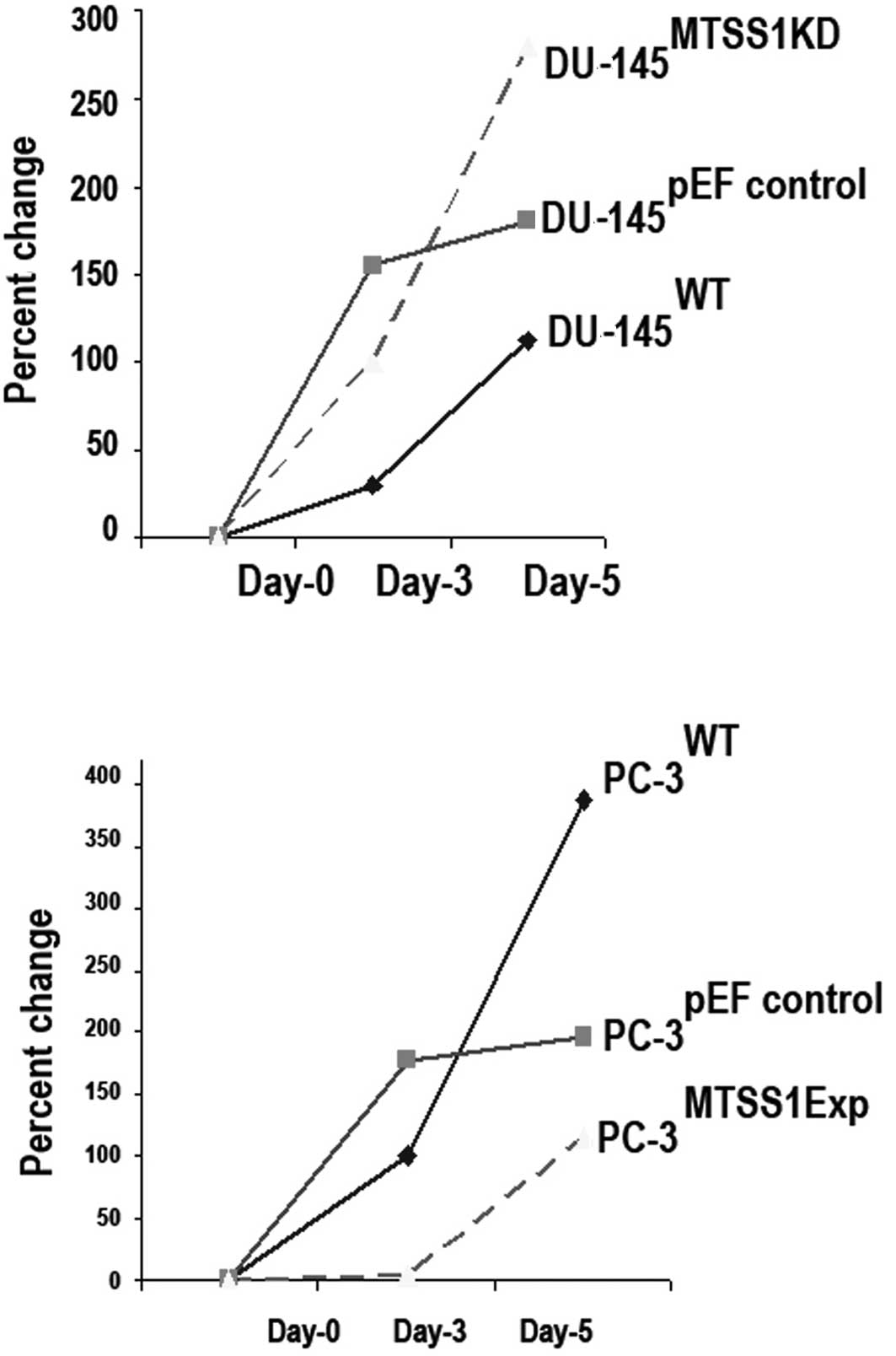
Knockdown of MTSS1 expression increases prostate
cancer cell growth. This study found that the levels of MTSS1
protein had an effect on the growth rate of cells. The effects of
the suppression of MTSS1 expression on the growth of DU-145 cells
was examined following a 5-day incubation period using an in
vitro cell growth assay. There was a significant difference in
the cell growth rate between the wild-type (DU-145 WT) and
MTSS1-suppressed (DU-145 MTSSIKD) cells. Suppression of MTSS1
expression was found to increase the cell growth rate (DU-145
MTSSIKD compared to DU-145 WT cells; p=0.001) (Fig. 2A).
Enhanced MTSS1 expression suppresses prostate
cancer cell growth. The growth capacity of the prostate cancer
cells following MTSS1 overexpression was examined and compared to
the wild-type cells using an in vitro cell growth assay for
PC-3 cell lines. A significant reduction in cell growth after 5
days following enhancement of MTSS1 expression was noted (PC-3
MTSS1Exp when compared to PC-3 WT cells; p=0.002) (Fig. 2B).
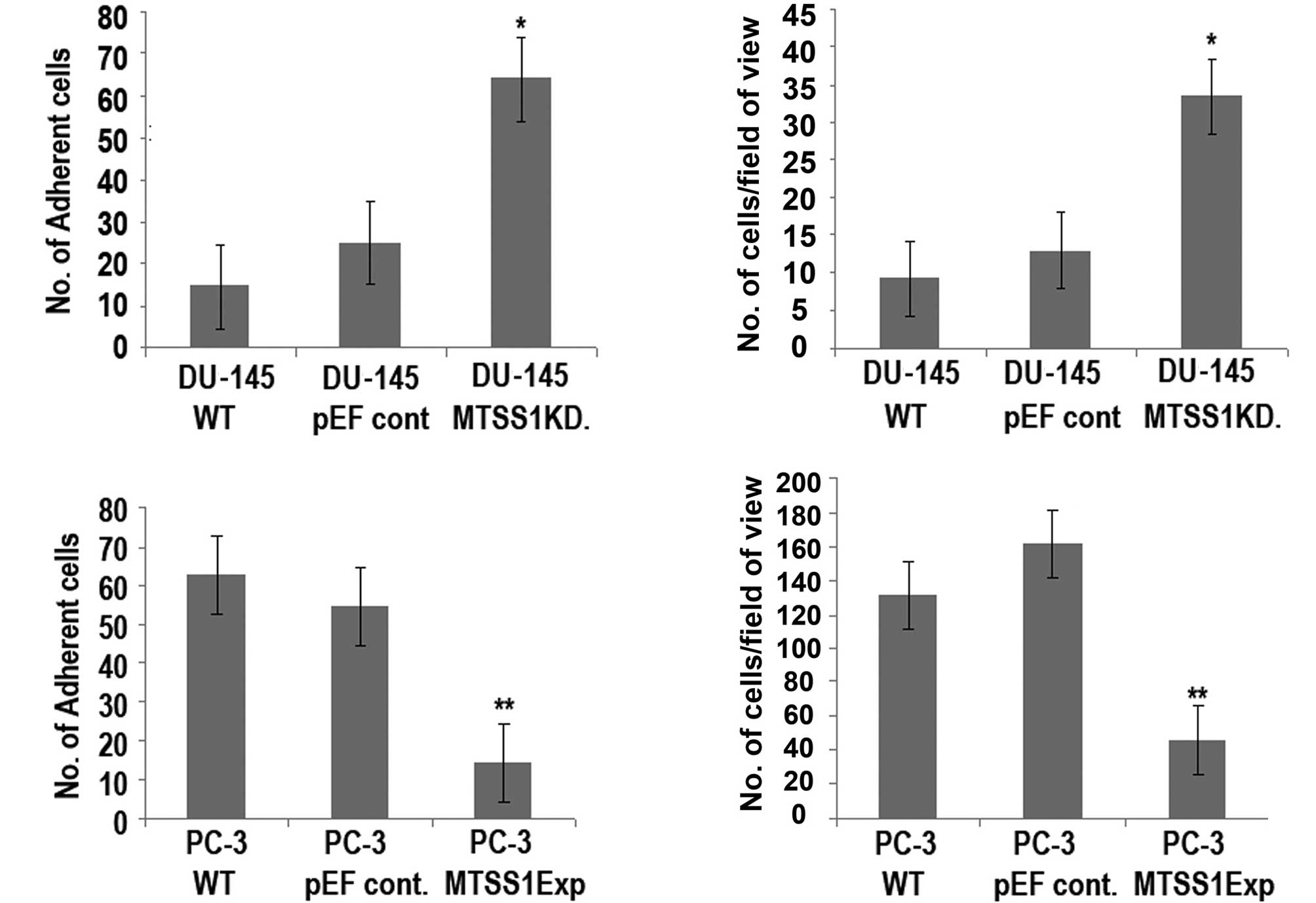
MTSS1 influences the adhesive ability of
prostate cancer cell lines
To identify and examine the adhesive nature of the
prostate cancer cells to attach to a basement membrane, an adhesion
assay was carried out (Fig. 3).
The results obtained show a significant increase (DU-145 MTSSIKD
compared to DU-145 WT cells; p=0.001) in the ability to adhere to a
basement membrane (Fig. 3A, top
panel), indicating that the knockdown of MTSS1 expression in this
cell line resulted in a dramatic increase in the degree of
adhesion. However, a significant reduction (p=0.0001) in adhesive
ability was noted when PC-3 MTSS1Exp cells were compared to PC-3 WT
cells (Fig. 3A, bottom panel).
MTSS1 has differential effects on the
motile properties of the prostate cancer cell lines
The results revealed that the presence of MTSS1 had
a significant impact on the motile nature of the prostate cancer
cells (Fig. 3B). A significant
increase (p=0.0001) was observed in motility between the DU-145
MTSSIKD and DU-145 WT cells (Fig.
3B, top panel). However, forced expression of MTSS1 in the PC-3
cell line resulted in a dramatic reduction in the degree of
motility (PC-3 MTSS1Exp compared to PC-3 WT cells; p=0.002)
(Fig. 3B, bottom).
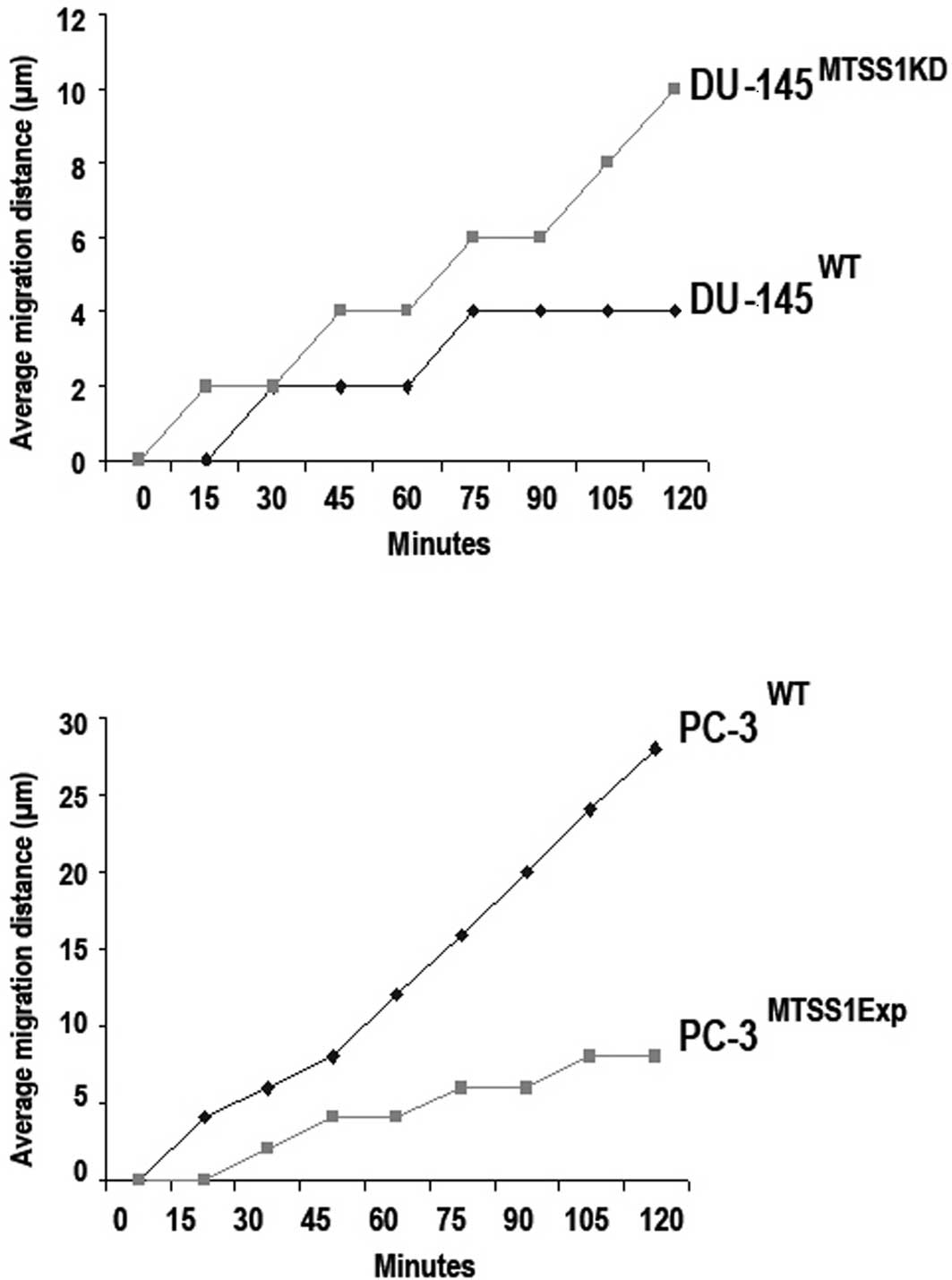
The DU-145 prostate cancer cells demonstrated motile
properties in the wound closure motility assay; however, the
absence of MTSS1 expression in these cells significantly increased
cell migration to close the wound as compared to the DU-145 WT
cells (p=0.001) (Fig. 4A). As
shown in Fig. 4B, the presence of
MTSS1 expression in the PC-3 MTSS1Exp cells significantly
suppressed cell migration when compared to the extent of cell
migration in the PC-3 WT cells (p=0.01).
Discussion
In the present study, we examined a possible
association between the expression of the metastasis suppressor
gene (MTSS1) and prostate tumour cell invasive behaviour. Supported
by a series of cellular function tests, the present study indicates
that MTSS1 acts as a powerful inhibitor to the aggressiveness of
prostate cancer cells. Our initial studies examined MTSS1
expression in a variety of human normal and cancer cell lines. We
reported that cancer cell lines, DU-145 and 3T3, expressed moderate
levels of MTTS1.
These cell lines are considered to be of a
low/non-invasive nature. The aggressive cell lines, MDA-MB-231 and
PC-3, are negative for MTSS1 expression. This is of note since it
may indicate that levels of MTSS1 expression are inversely
correlated with aggressiveness. This is reflected in our subsequent
expression modification studies, in which we created two cell
models, DU-145 MTSSIKD and PC-3 MTSS1Exp, with a differential
pattern of MTSS1 expression.
The invasive MTSS1-negative PC-3 cell line was
‘forced’ to overexpress MTSS1, while the non-invasive MTSS-positive
DU-145 cell line had its MTSS1 expression levels ‘knocked down’. We
used a range of biological function assays in vitro to
assess the affect of the modification of MTSS1 expression on the
metastatic nature of these prostate cancer cell lines. Our study
provides evidence that the forced expression of MTSS1 in the PC-3
prostate cancer cells greatly reduced the aggressive nature of
these cells by reducing their adhesive, growth and motile
properties.
In contrast, the elimination of MTSS1 expression in
the DU-145 cells exhibited an inverse effect. This low-invasive
cell line displayed significant increases in tumour cell migration,
growth and adhesion. Therefore, our findings suggest that MTSS1
plays a key role in determining the metastatic nature of prostate
cancer cells.
Notably, Parr et al (13) revealed that MTSS1 acts as an
indicator for survival for breast cancer patients. They
demonstrated that patients expressing high levels of MTSS1 had a
favourable prognosis in contrast to patients with low levels of
MTSS1 expression who were associated with a poor prognosis. Utikal
et al (12) suggested that
the mechanism for the down-regulation of MTSS1 may involve DNA
methylation. Overexpression of MTSS1 was found to enhance changes
in cell shape, such as an increase in the formation of membrane
ruffles and filopodia-like structures (7,10,14–16).
MTSS1 has been shown to be a metastatic suppressor
in bladder cancer (6). In
contrast, MTSS1 expression was found to be similar in metastatic
cell lines (14) and basal cell
carcinomas (9). This regulation
suggests that MTSS1 levels are likely to be controlled during cell
growth and development, and that MTSS1 expression is altered in
cancer cells, leading to changes in the signalling and architecture
of the cytoskeleton.
One study reported that the down-regulation of MTSS1
expression in bladder cancer may correlate with the transition of
tumour cells from a distinct epithelium-like morphology to less
differentiated carcinomas (17,18).
Another study found a dramatic increase in MTSS1 expression in
normal liver specimens compared to matched hepatocellular carcinoma
tumour tissue specimens (18).
Elevated MTSS1 expression was observed in early
stage disease, suggesting that MTSS1 plays a key role in promoting
the early development of hepatocellular carcinoma and may therefore
serve as a biomarker for the prediction of early tumour development
of hepatocellular carcinoma. Thus, it is likely that a metastasis
suppressor for MTSS1 will be highly dependent upon tumour type
(13).
In conclusion, our study indicates that MTSS1 is
capable of modulating the metastatic ability in prostate cancer
cells.
Acknowledgements
The authors wish to thank Cancer
Research Wales for supporting their work.
References
|
1.
|
Ferlay J, Parkin D and Pisani P: Estimates
of the worldwide incidence of 25 major cancers in 1990. Int J
Cancer. 80:158–162. 1999.
|
|
2.
|
Fidler IJ: The pathogenesis of cancer
metastasis: the ‘seed and soil’ hypothesis revisited. Nat Rev
Cancer. 3:453–458. 2003.
|
|
3.
|
Folkman J: Toward an understanding of
angiogenesis: search and discovery. Perspect Biol Med. 29:10–36.
1985. View Article : Google Scholar : PubMed/NCBI
|
|
4.
|
Boyer B, Valles AM and Thiery JP: Model
systems of carcinoma cell dispersion. Curr Top Microbiol Immunol.
213:179–194. 1996.PubMed/NCBI
|
|
5.
|
Stafford LJ, Vaidya KS and Welch DR:
Metastasis suppressor genes in cancer. Int J Biochem Cell Biol.
40:874–891. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
6.
|
Lee YG, Macoska JA, Korenchuk S and Pienta
KJ: MIM, a potential metastasis suppressor gene in bladder cancer.
Neoplasia. 4:291–294. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
7.
|
Loberg RD, Neeley CK and Adam-Day LL:
Differential expression analysis of MIM (MTSS1) splice variants and
a functional role of MIM in prostate cancer cell biology. Int J
Oncol. 26:1699–1705. 2005.PubMed/NCBI
|
|
8.
|
Callahan CA, Ofstad T, Horng T, Wang JK,
Zhen HH, Coulombe PA and Oro AE: MIM/BEG4, a Sonic hedgehog
responsive gene that potentiates Gli-dependent transcription. Genes
Dev. 18:2724–2729. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
9.
|
Mattila PK, Salminen M, Yamashiro T and
Lappalainen P: Mouse MIM, a tissue specific regulator of
cytoskeletal dynamics, interacts with ATP-actin monomers through
its C-terminal WH2 domain. J Biol Chem. 278:8452–8459. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
11.
|
Lin J, Liu J, Wang Y, Zhu J, Zhou K, Smith
N and Zhan X: Differential regulation of cortactin and
N-WASP-mediated actin polymerization by missing in metastasis (MIM)
protein. Oncogene. 24:2059–2066. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
12.
|
Utikal J, Gratchev A and Muller-Molinet I:
The expression of metastasis suppressor MIM/MTSS1 is regulated by
DNA methylation. Int J Cancer. 119:2287–2293. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
13.
|
Parr C and Jiang WG: Metastasis suppressor
1 (MTSS1) demonstrates prognostic value and anti-metastatic
properties in breast cancer. Int J Cancer. 45:1673–1683.
2008.PubMed/NCBI
|
|
14.
|
Liu K, Wang G, Ding H, Chen Y, Yu G and
Wang J: Downregulation of metastasis suppressor 1 (MTSS1) is
associated with nodal metastasis and poor outcome in Chinese
patients with gastric cancer. BMC Cancer. 10:4282010. View Article : Google Scholar : PubMed/NCBI
|
|
15.
|
Bompard G, Sharp SJ, Freiss G and Machesky
LM: Involvement of Rac in actin cytoskeleton rearrangements induced
by MIM-B. J Cell Sci. 118:5393–5403. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
16.
|
Gonzalez-Quevedo R, Shoffer M, Horng L and
Oro AE: Receptor tyrosine phosphatase-dependent cytoskeletal
remodeling by the hedgehog-responsive gene MIM/BEG4. J Cell Biol.
168:453–463. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
17.
|
Wang Y, Liu J and Smith E: Downregulation
of missing in metastasis gene (MIM) is associated with the
progression of bladder transitional carcinomas. Cancer Invest.
25:79–86. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
18.
|
Ma S, Guan XY, Lee TK and Chan KW:
Clinicopathological significance of missing in metastasis B
expression in hepatocellular carcinoma. Hum Pathol. 38:1201–1206.
2007. View Article : Google Scholar : PubMed/NCBI
|