Introduction
Cancer is one of the most serious threats to human
health worldwide and affects a variety of organs and tissues
(1). Colorectal cancer (CRC) is a
global public health concern (2).
The most common treatment strategies for CRC include surgery,
chemotherapy and radiotherapy; however, these strategies often do
not completely eliminate the disease (3,4). The
targeted treatment of tumors has attracted increasing attention and
several targeted drugs are extensively used in the clinic (4) with demonstrated effectiveness
(5). An increased understanding of
the molecular mechanism underlying the pathogenesis of CRC may aid
with the identification of effective therapeutic targets and novel
biomarkers for the risk assessment and early diagnosis of CRC
(6-8).
MicroRNAs (miRNAs/miRs) are small non-coding RNAs
that are ~22 nucleotides in length (9,10).
Previous studies have revealed that miRNAs may negatively regulate
gene expression by binding to the 3'-untranslated region (UTR) of
target mRNAs to silence gene expression (9-11).
It has been widely established that miRNAs serve critical roles in
the regulation of several biological process, including
proliferation, differentiation, apoptosis, tumorigenesis and
metastasis (6,12,13).
Furthermore, miRNAs may predict overall prognosis in certain types
of cancer, including lung cancer (14), gastric cancer (15) and CRC (16). Previous studies suggested that
miR-590 is abnormally expressed in CRC (5,17);
however, the mechanism underlying the action of miR-590-5p in the
pathogenesis of CRC, as well as its potential gene target have not
been identified.
The present study investigated the role of
miR-590-5p, as well as its underlying mechanism, in CRC. The
expression of miR-590-5p in CRC tissues were compared with healthy
adjacent tissues. Furthermore, in vitro studies were
designed to examine the effect of miR-590-5p knockdown on CRC cell
viability and invasion.
Materials and methods
Patients and clinical tissue
samples
A total of 30 CRC and healthy adjacent tissue
samples (distance from tumour margin, 3 cm) were collected from
patients with CRC (age, 48-65 years; mean age, 52.32±7.41; 17 male
patients and 13 female patients) at Taizhou People's Hospital
between May 2017 and February 2019. The tissue samples were
obtained from patients who had undergone resection but had not
received radiotherapy and/or chemotherapy. All tissues were
pathologically confirmed. The present study was approved by the
Ethical Review Board of Taizhou People's Hospital and written
informed consent was obtained from each patient.
Cell culture and transfection
Human CRC cell lines (HCT116, SW480, SW620 and LoVo)
and human normal colon epithelial cells (FHC) were purchased from
The Cell Bank of Type Culture Collection of the Chinese Academy of
Sciences. Cells were cultured in RPMI-1640 medium (Invitrogen;
Thermo Fisher Scientific, Inc.) supplemented with 10% fetal bovine
serum (Invitrogen; Thermo Fisher Scientific, Inc.) at 37˚C in a
humidified incubator with 5% CO2. Cells were plated in
24-well plates (1x105 cells/well) and transfected with
miR-590-5p mimic, miR-590-5p inhibitor, miR-590-5p mimic negative
control (NC), miR-590-5p inhibitor NC, PDCD4 small interfering
(si)RNA NC or PDCD4 siRNA (all, 100 pmol) using
Lipofectamine® 3000 (Invitrogen; Thermo Fisher
Scientific, Inc.) according to the manufacturer's protocol. The
sequences of the oligonucleotides were: miR-590-5p mimic,
5'-GAGCUUAUUCAUAAAAUGCAG-3'; miR-590-5p mimic NC,
5'-CGCCAAUAUCAUUAUACCUC-3'; miR-590-5p inhibitor,
5'-AAAUAUGCUGUAUGUCAUGUGUU-3'; miR-590-5p inhibitor NC,
5'-GUCCAGUGAAUUCCCAG-3'; PDCD4 siRNA, 5'-GGAGGTGGATGTGAAAGAT-3';
and PDCD4 siRNA NC, 5'-GGCCTTGCGATAGCGTAGC-3'. At 48 h
post-transfections, cells were used for subsequent experiments.
Reverse transcription-quantitative PCR
(RT-qPCR)
Total RNA was extracted from cells or clinical
tissue samples using TRIzol® reagent (cat. no. 15596026;
Invitrogen; Thermo Fisher Scientific, Inc.). Total RNA was reverse
transcribed into cDNA using PrimeScript RT Master mix (Takara
Biotechnology, Co., Ltd.) at 37˚C for 15 min followed by 85˚C for 5
sec. Subsequently, qPCR was performed using SYBR premix Ex Taq
(Takara Biotechnology, Co., Ltd.) and a thermocycler (Applied
Biosystems; Thermo Fisher Scientific, Inc.). The thermocycling
conditions used for qPCR were as follows: Initial denaturation 95˚C
for 30 sec; followed by 40 cycles at 95˚C for 5 sec and 60˚C for 30
sec. The sequences of the primers used for qPCR were as follows:
miR-590-5p forward, 5'-GAGCTTATTCATAAAAGT-3' and reverse,
5'-TCCACGACACGCACTGGATACGAC-3'; PDCD4 forward,
5'-GCAAAAAGGCGACTAAGGAAAAA-3' and reverse,
5'-TAAGGGCGTCACTCCCACT-3'; U6 forward, 5'-GTGCTCGCTTCGGCAGCACAT-3'
and reverse, 5'-TACCTTGCGAAGTGCTTAAAC-3'; and GAPDH forward,
5'-TGTGGGCATCAATGGATTTGG-3' and reverse,
5'-ACACCATGTATTCCGGGTCAAT-3'. miRNA and mRNA levels were quantified
using the 2-ΔΔCq method (18) and normalized to the internal
reference genes U6 and GAPDH, respectively.
Cell viability analysis
At 48 h post-transfection, the effect of miR-590-5p
on cell viability was measured using an MTT assay. Briefly, cells
were washed with PBS (pH 7.4), harvested by trypsinization and
seeded (5x105 cells/well) into a 96-well plate. The
plate was incubated in humidified incubator at 37˚C with 5%
CO2. Subsequently, 10 µl MTT solution was added to each
well and incubated for 1 to 4 h at 37˚C. DMSO was added to each
well to dissolve the formazan crystals. The absorbance of each well
was measured at a wavelength of 490 nm using a microplate
reader.
Wound healing assay
HCT116 and SW480 cells were seeded (8x105
cells/well) into 6-well plates. At 100% confluence, 8 µg/ml
mitomycin C (Beyotime Institute of Biotechnology) was added to each
well for 3 h at 37˚C to inhibit cell proliferation. Subsequently,
the cell monolayer was scratched with a 10 µl pipette tip and
incubated in RPMI-1640 medium supplemented with 1% FBS at 37˚C for
24 h. Images of the wounds were captured using an inverted light
microscope at 0 and 24 h (magnification, x200).
Transwell assay
The transwell plates were pre-coated with
Matrigel® at room temperature for 30 min. Cells were
seeded (5x104 cells/well) into the upper chamber of the
Transwell plate (24-well plates; Corning, Inc.) with serum-free
RPMI-1640 medium. The lower chamber was treated with RPMI-1640
medium containing 10% FBS Cells were incubated at 37˚C for 24 h.
Invading cells were fixed with 100% methanol for 20 min at room
temperature, stained with crystal violet for 10 min at room
temperature and visualized under a light microscope (magnification,
x200).
Western blotting
Total protein was isolated from cells using a
protease inhibitor cocktail (RIPA; Beyotime Institute of
Biotechnology). Total protein was quantified using a bicinchoninic
acid protein assay kit (Pierce; Thermo Fisher Scientific, Inc.).
Proteins (30 µg) were separated via 10% SDS-PAGE, transferred onto
PVDF membranes and blocked with 5% non-fat dry milk in TBST (pH
7.4; 0.05% Tween 20) at room temperature for 2 h. The membranes
were incubated at 4˚C overnight with the following primary
antibodies: Anti-PCDC4 (cat. no. ab80590; 1:1,000), anti-p-Smad2/3
(cat. no. ab63399; 1:1,000), anti-TGF-β (cat. no. ab92486; 1:1,000)
and anti-GAPDH (cat. no. ab9485; 1:2,000). Following primary
antibody incubation, the membranes were incubated with a
horseradish peroxidase-conjugated secondary antibody (cat. no.
ab6721; 1:5,000) at room temperature for 45 min. All antibodies
were purchased from Abcam. Proteins were visualized using the
Pierce™ ECL Western Blotting Substrate (Pierce; Thermo Fisher
Scientific, Inc.) and chemiluminescence signals were detected using
a Tanon-5200 Imaging system (Tanon Science and Technology, Co.,
Ltd.). Protein expression levels were quantified using ImageJ
software (version 1.47; National Institutes of Health) with GAPDH
as the loading control.
Dual-luciferase reporter assay
Wild-type PDCD4 3'-UTR (PDCD4-3'UTR), containing the
miR-590-5p binding site, and mutant PDCD4 3'UTR (PDCD4-MUT) were
cloned into the p-MIR-reporter plasmid (Thermo Fisher Scientific,
Inc.). 293T cells (Cell Bank of Type Culture Collection of the
Chinese Academy of Sciences) were seeded onto 6 well plates
(8x105 cells/well), co-transfected with PDCD4-3'UTR or
PDCD4-MUT (100 pmol) and miR-590-5p mimic or miR-590-5p mimic NC
(each, 50 pmol) using Lipofectamine® RNAi Max
(Invitrogen; Thermo Fisher Scientific, Inc.). At 48 h
post-transfection, luciferase activities were detected using the
Dual Luciferase Reporter Assay kit (Beyotime Institute of
Biotechnology). Firefly luciferase activity was normalized to
Renilla luciferase activity.
Statistical analysis
All experiments were repeated at least three times.
Statistical analyses were performed using SPSS software (version
17.0; SPSS, Inc.). Data are presented as the mean ± standard
deviation. Comparisons between two groups were analyzed using the
paired Student's t-test. Comparisons among >2 groups were
analyzed using one-way ANOVA followed by Tukey's post hoc test.
P<0.05 was considered to indicate a statistically significant
difference.
Results
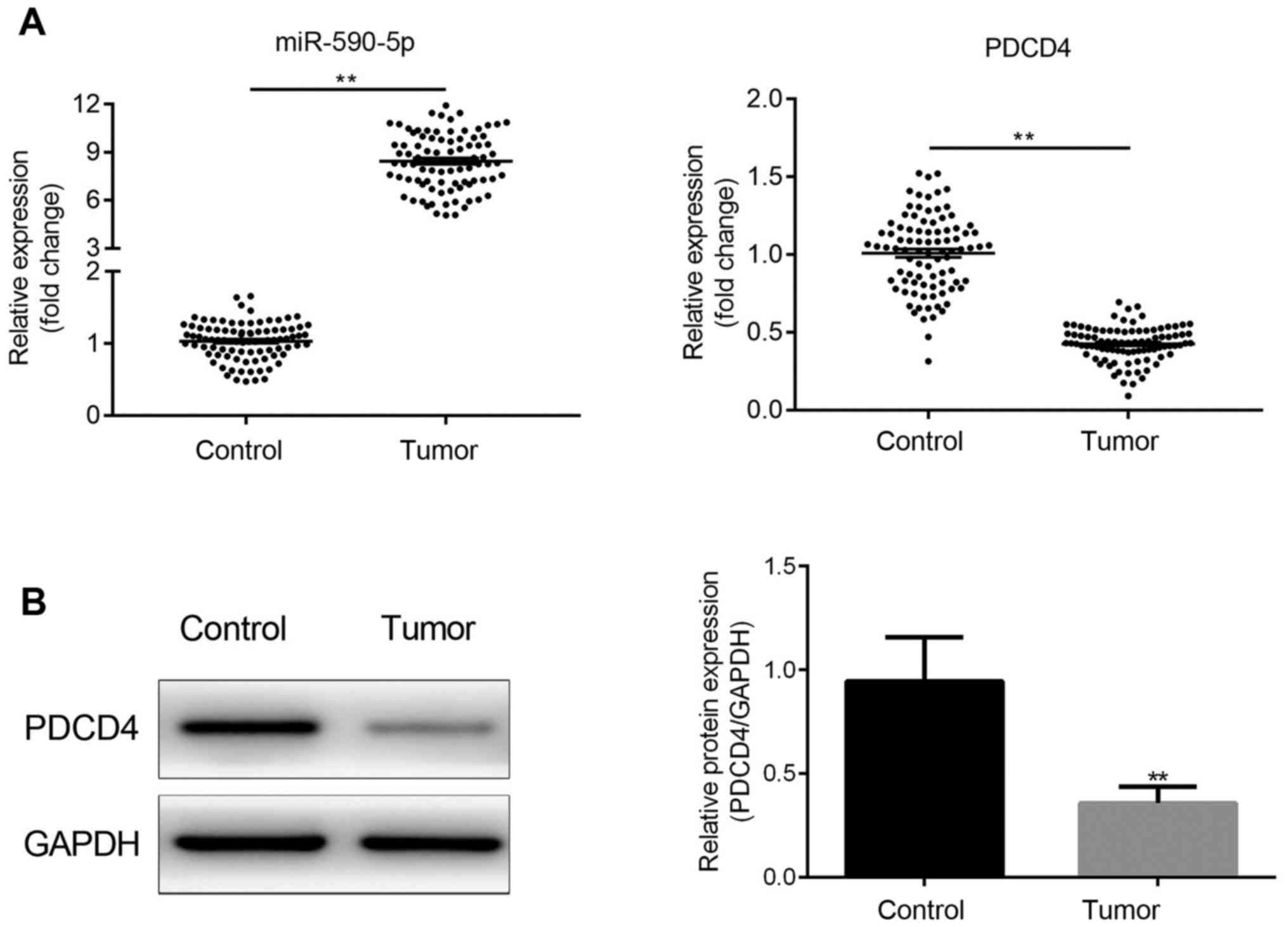
miR-590-5p expression is increased and
is negatively associated with PDCD4 in human CRC tissues
miR-590-5p and PDCD4 expression levels were detected
in all 30 paired samples of human CRC and adjacent healthy tissues.
The results indicated that miR-590-5p expression levels were
significantly increased in CRC tissues compared with adjacent
healthy tissues (P<0.01; Fig.
1A). By contrast, PDCD4 expression levels were significantly
decreased in CRC tissues compared with adjacent healthy tissues
(P<0.01; Fig. 1A and B).
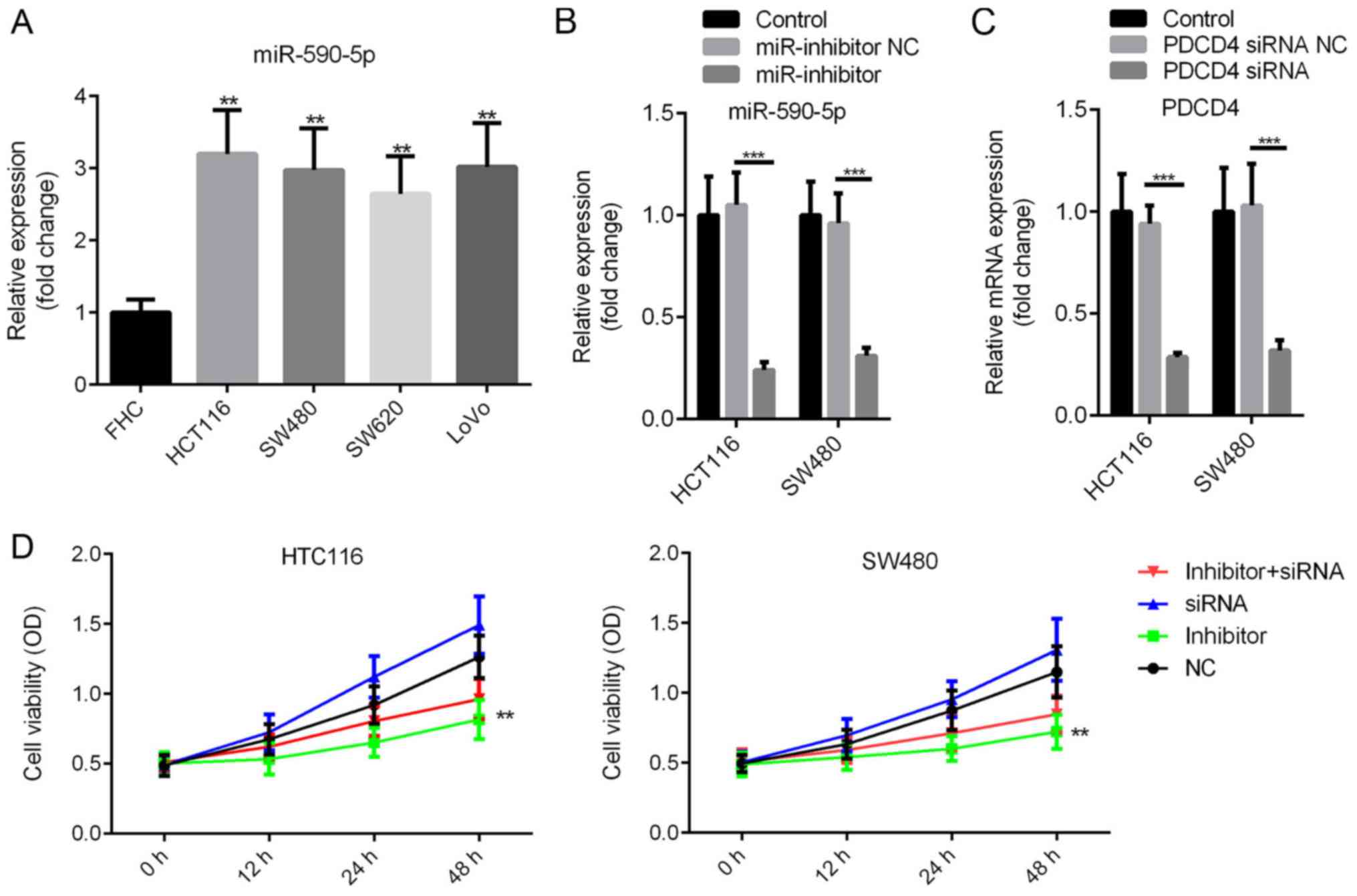
miR-590-5p inhibitor decreases human
CRC HCT116 and SW480 cell viability
miR-590-5p expression levels in CRC cells and human
normal colon epithelial cells (FHC) were compared. miR-590-5p
expression levels were significantly increased in CRC cells
compared with FHC cells (P<0.01; Fig. 2A). miR-590-5p inhibitor or
miR-590-5p inhibitor NC were transfected into HCT116 and SW480
cells. The MTT assay was conducted to assess the effect of
miR-590-5p on cell viability. miR-590-5p inhibitor and PDCD4 siRNA
significantly decreased the expression levels of miR-590-5p and
PDCD4, respectively, in HCT116 and SW480 cells (P<0.001;
Fig. 2B and C). Moreover, compared with the NC group,
miR-590-5p inhibitor decreased cell viability at 12 and 24 h
post-transfection, and significantly decreased cell viability at 48
h post-transfection (Fig. 2D;
P<0.01). Additionally, co-transfection of miR-590-5p inhibitor
and PDCD4 siRNA partially reversed the effects of miR-590-5p
inhibitor on cell viability (Fig.
2D).
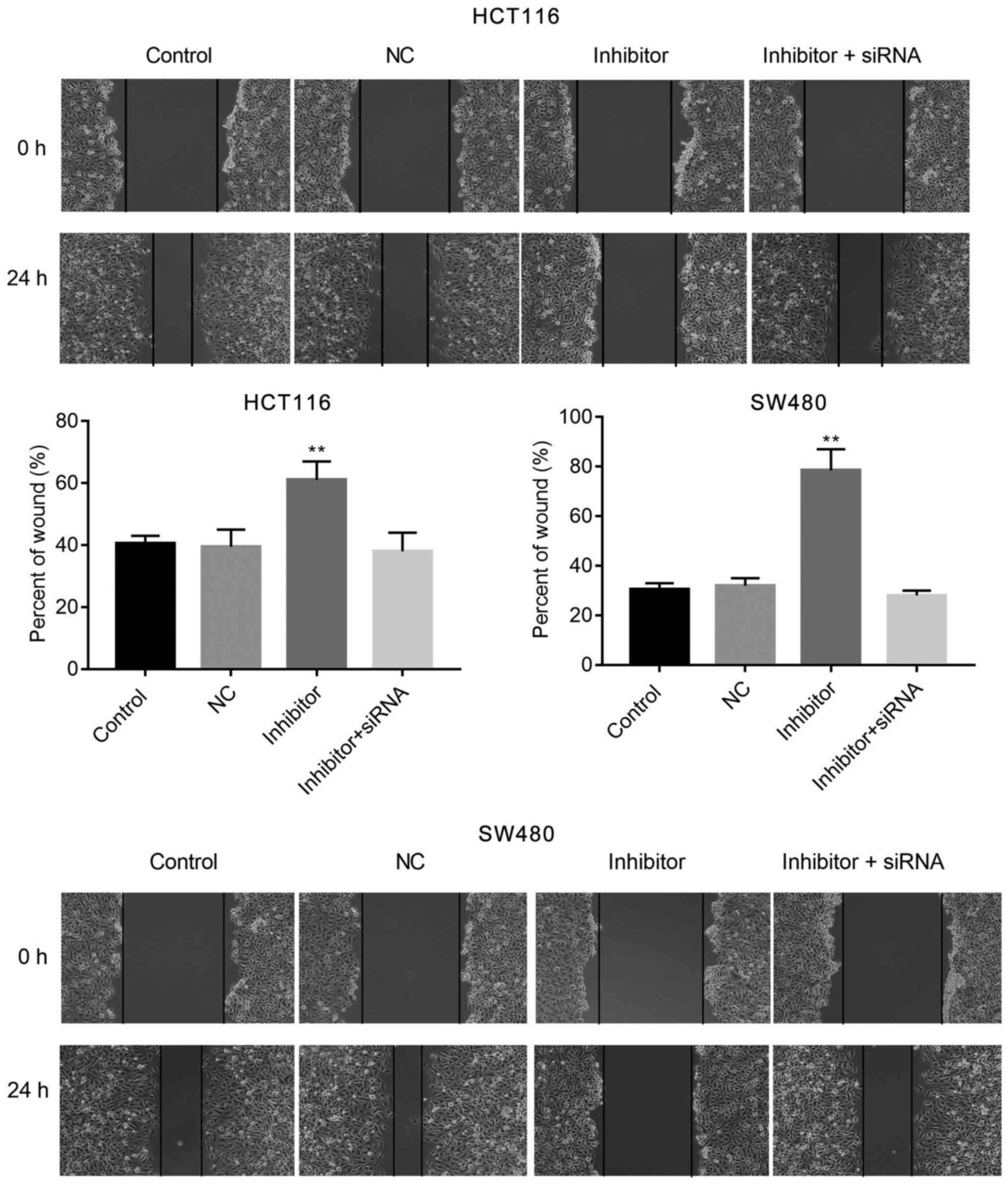
miR-590-5p inhibitor suppresses human
CRC HCT116 and SW480 cell migration and invasion
The effects of miR-590-5p inhibitor on cell
migration and invasion were examined by conducting wound healing
and Transwell assays, respectively. miR-590-5p inhibitor
significantly increased cell migration at 24 h post-transfection
(P<0.01; Fig. 3) when compared
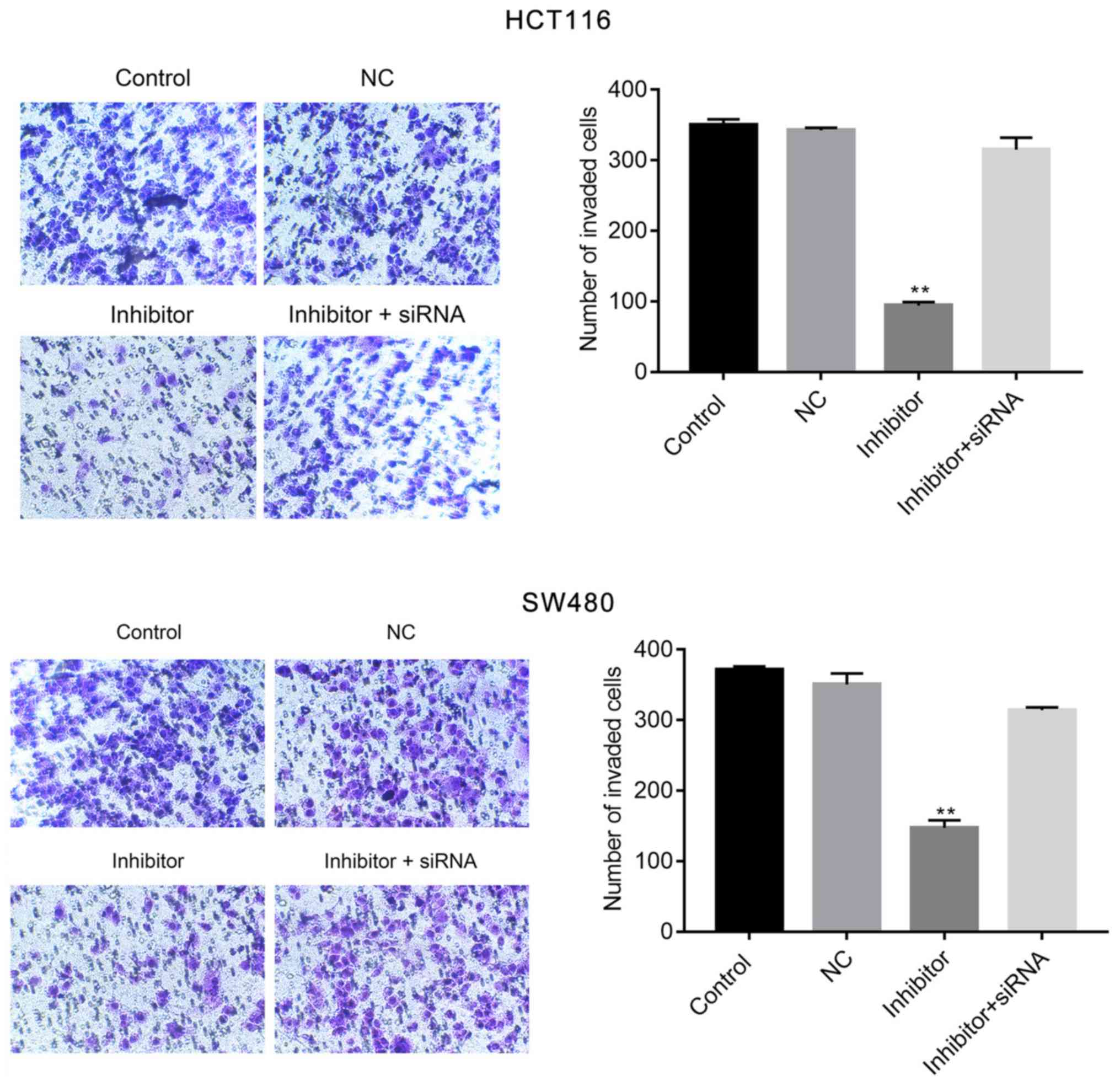
with the control. Moreover, the Transwell assay indicated that
miR-590-5p inhibitor significantly reduced HCT116 and SW480 cell
invasion compared with the control (P<0.01; Fig. 4). Additionally, co-transfection of
miR-590-5p inhibitor and PDCD4 siRNA reversed the effects of
miR-590-5p inhibitor on HCT116 and SW480 cell migration and
invasion (Figs. 3 and 4).
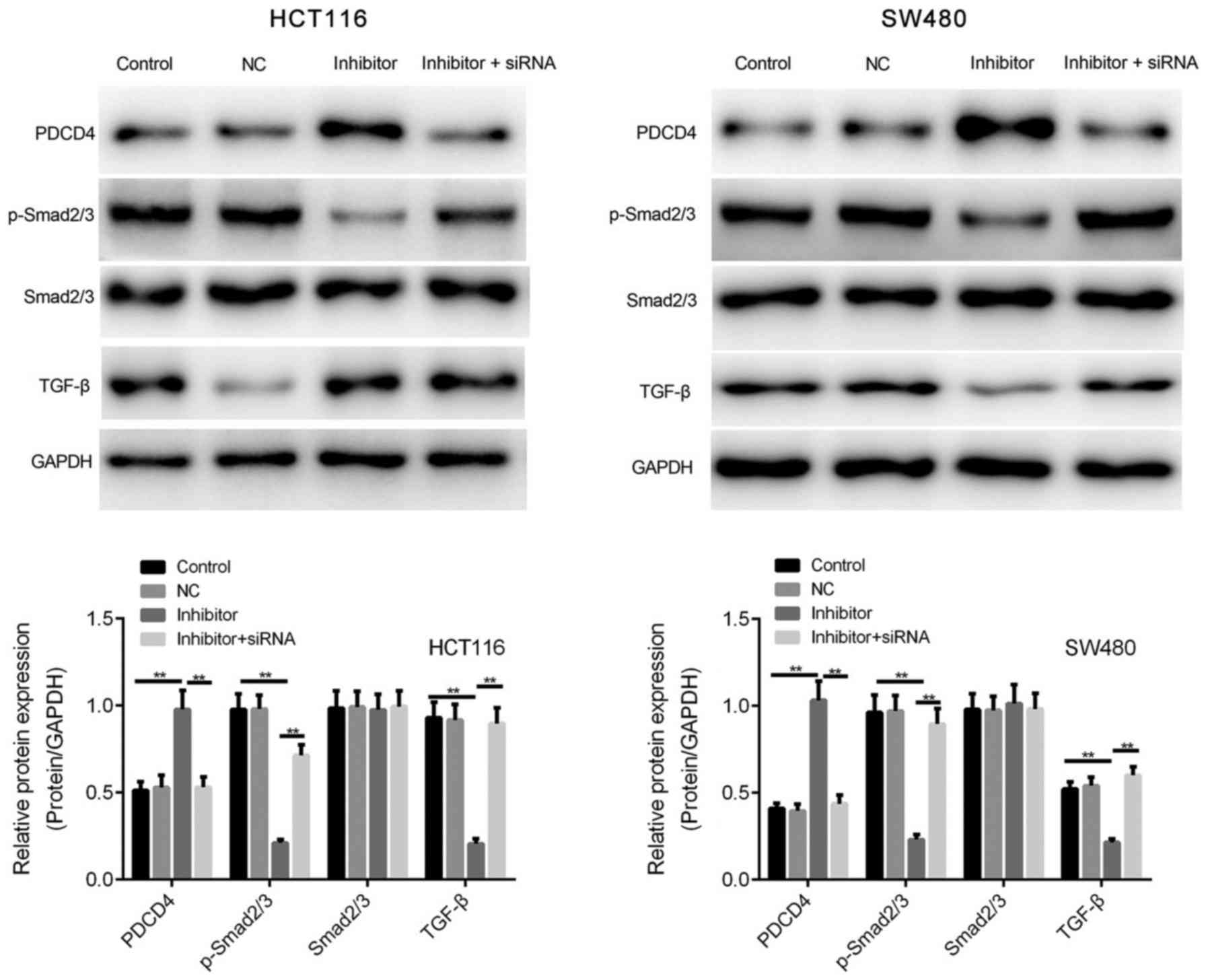
miR-590-5p inhibitor enhances the
expression of PDCD4 and TGF-β/Smad2/3 signaling
Based on the association between miR-590-5p and
PDCD4 expression observed in human CRC tissues, HCT116 and SW480
cells were transfected with miR-590-5p inhibitor to investigate
whether PDCD4 expression was enhanced by miR-590-5p inhibitor.
miR-590-5p inhibitor significantly increased PDCD4 protein
expression levels compared with the control group in HCT116 and
SW480 cells (P<0.01; Fig.
5).
The effect of miR-590-5p on the phosphorylation of
Smad2/3 was investigated. Compared with the control group,
miR-590-5p inhibitor significantly decreased the expression levels
of TGF-β and p-Smad2/3 (P<0.01; Fig.
5). However, co-transfection of miR-590-5p inhibitor and PDCD4
siRNA significantly reversed the effects of miR-590-5p inhibitor on
p-Smad2/3 expression levels (P<0.01; Fig. 5).
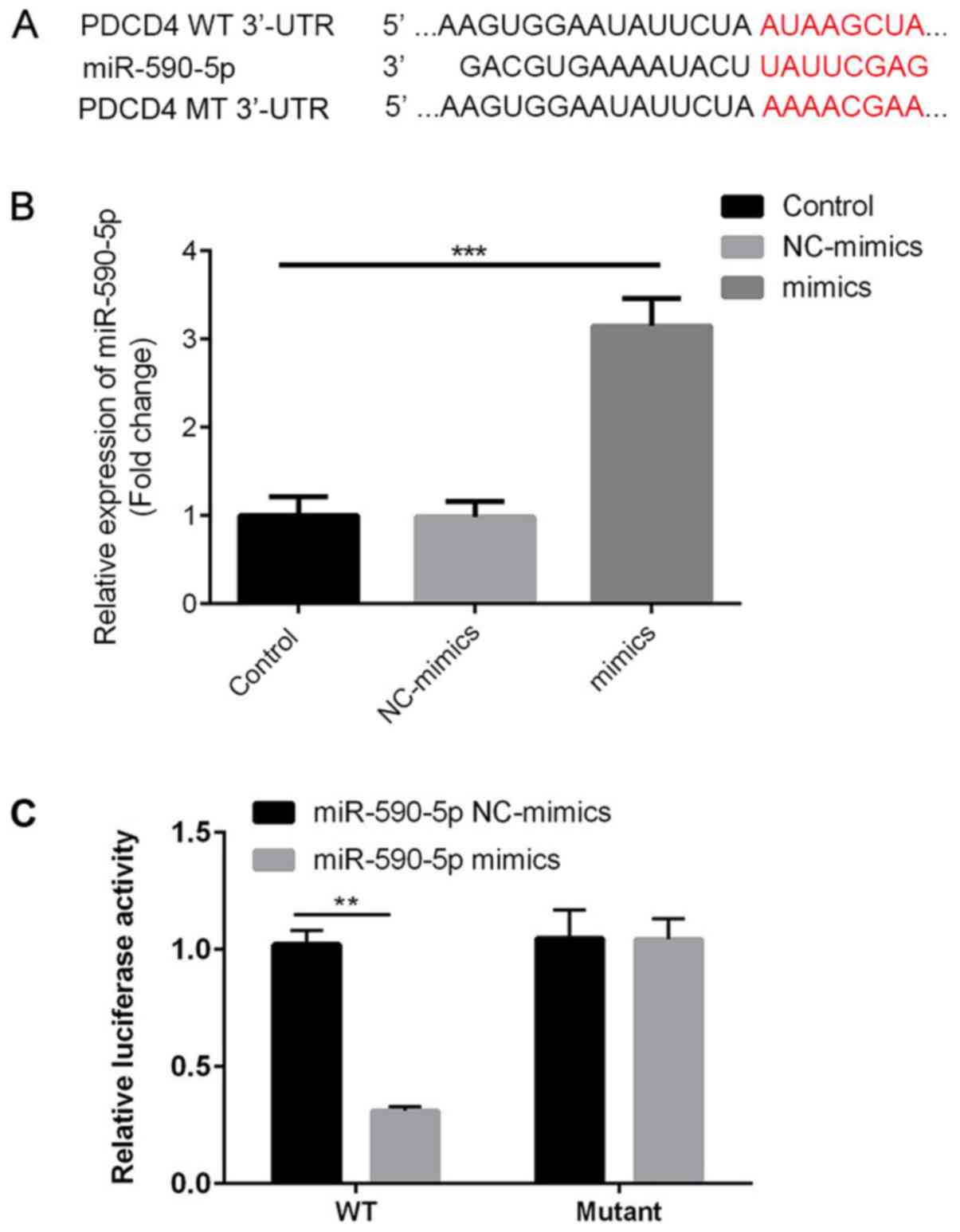
PDCD4 is a direct target of
miR-590-5p
The present study investigated whether the observed
reduction in PDCD4 was caused by direct binding of miR-590-5p to
the 3'UTR of PDCD4. Therefore, PDCD4-3'UTR containing the putative
binding site for miR-590-5p and the mutated segment were cloned in
a firefly luciferase reporter vector (Fig. 6A). miR-590-5p mimic significantly
increased the expression levels of miR-590-5p in 293T cells
compared with the control group (Fig.
6B). miR-590-5p mimic significantly decreased the luciferase
activity of PDCD4-3'UTR WT compared with miR-590-5p mimic NC
(P<0.01), but did not significantly alter the luciferase
activity of PDCD4-3'UTR MUT (Fig.
6C). The results suggested that miR-590-5p directly bound to
PDCD4-3'UTR, resulting in reduced PDCD4 protein expression
levels.
Discussion
miRNAs display great potential for the diagnosis and
treatment of CRC (19-21).
miR-590-5p has been reported to be strongly associated with CRC
progression (17,22). The present study indicated that
miR-590-5p expression levels were significantly increased in human
CRC tissues and cells compared with healthy adjacent tissues and
FHC cells, suggesting that miR-590-5p could function as an
oncogene.
The present study investigated the roles of
miR-590-5p in CRC pathogenesis by performing in vitro
studies in human CRC HCT116 and SW480 cells. Cell proliferation and
metastasis are important events associated with tumor progression
(23,24). The results of the present study
suggested that miR-590-5p inhibitor reduced HCT116 and SW480 cell
viability compared with the control. Furthermore, when compared
with the control group, miR-590-5p inhibitor decreased cell
migration and invasion in vitro, which was consistent with
the results obtained in a previous study (17). Of note, miR-590-5p inhibitor
suppressed the TGF-β/Smad2/3 signaling pathway, which regulates CRC
cell viability and migration (23,25,26).
Collectively, the results indicated that miR-590-5p knockdown
inhibited the TGF-β/Smad2/3 signaling pathway in HCT116 and SW480
cells.
Bioinformatics analysis identified PDCD4 as one of
the direct targets of miR-590-5p. PDCD4 expression levels were
significantly lower in human CRC tissues compared with healthy
adjacent tissues, which was consistent with previous studies
(27-30).
Therefore, it was hypothesized that miRNA-590-5p may directly
negatively regulate the expression of PDCD4. To investigate the
hypothesis, the present study investigated the effects of
miR-590-5p inhibitor in HCT116 and SW480 cells. The results
suggested that miR-590-5p inversely regulated the expression of
PDCD4. Subsequently, the direct binding of miR-590-5p to the
PDCD4-3'UTR was confirmed by conducting a dual-luciferase reporter
assay. The results indicated that co-transfection of miR-590-5p
inhibitor and PDCD4 siRNA partially reversed the antitumor effects
of miR-590-5p inhibitor. The results suggested that miR-590-5p may
exert its carcinogenic behaviors by downregulating the expression
of PDCD4.
Collectively, the results of the present study
suggested that miR-590-5p increased CRC cell viability and
migration, and negatively regulated the expression of PDCD4 by
directly binding to it. The results suggested that miR-590-5p may
serve an important role in CRC pathogenesis and serve as a novel
therapeutic strategy for CRC.
Acknowledgements
Not applicable.
Funding
The present work was funded by Taizhou High Level
Talents Research Project in 2015 (grant no. 201529).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
TG and JW performed the majority of experiments and
wrote the manuscript. GC performed the remaining experiments and
statistically analyzed the data. HH designed the present study,
provided funding and revised the manuscript. All authors read and
approved the final manuscript.
Ethics approval and consent to
participate
The present study was approved by the Ethical Review
Board of Taizhou People's Hospital and written informed consent was
obtained from each patient.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Jacobsen F, Kraft J, Schroeder C,
Hube-Magg C, Kluth M, Lang DS, Simon R, Sauter G, Izbicki JR,
Clauditz TS, et al: Up-regulation of biglycan is associated with
poor prognosis and PTEN deletion in patients with prostate cancer.
Neoplasia. 19:707–715. 2017.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Anderson JC and Levine JB: Age and crc
risk in the serrated pathway. J Clin Gastroenterol. 52:465–467.
2018.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Wang G, Fu Y, Hu F, Lan J, Xu F, Yang X,
Luo X, Wang J and Hu J: Loss of BRG1 induces CRC cell senescence by
regulating p53/p21 pathway. Cell Death Dis. 8(e2607)2017.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Orang AV and Barzegari A: MicroRNAs in
colorectal cancer: From diagnosis to targeted therapy. Asian Pac J
Cancer Prev. 15:6989–6999. 2014.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Sun ZQ, Shi K, Zhou QB, Zeng XY, Liu J,
Yang SX, Wang QS, Li Z, Wang GX, Song JM, et al: MiR-590-3p
promotes proliferation and metastasis of colorectal cancer via
Hippo pathway. Oncotarget. 8:58061–58071. 2017.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Chen M, Li D, Gong N, Wu H, Su C, Xie C,
Xiang H, Lin C and Li X: miR-133b down-regulates ABCC1 and enhances
the sensitivity of CRC to anti-tumor drugs. Oncotarget.
8:52983–52994. 2017.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Guglielmo A, Staropoli N, Giancotti M and
Mauro M: Personalized medicine in colorectal cancer diagnosis and
treatment: A systematic review of health economic evaluations. Cost
Eff Resour Alloc. 16(2)2018.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Dekker E and IJspeert JEG: Serrated
pathway: A paradigm shift in CRC prevention. Gut. 67:1751–1752.
2018.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Park YR, Lee ST, Kim SL, Zhu SM, Lee MR,
Kim SH, Kim IH, Lee SO, Seo SY and Kim SW: Down-regulation of miR-9
promotes epithelial mesenchymal transition via regulating
anoctamin-1 (ANO1) in CRC cells. Cancer Genet. 231-232:22–31.
2019.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Zhang Z, Li J, Huang Y, Peng W, Qian W, Gu
J, Wang Q, Hu T, Ji D, Ji B, et al: Upregulated miR-1258 regulates
cell cycle and inhibits cell proliferation by directly targeting
E2F8 in CRC. Cell Prolif. 51(e12505)2018.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Tong F, Ying Y, Pan H, Zhao W, Li H and
Zhan X: MicroRNA-466 (miR-466) functions as a tumor suppressor and
prognostic factor in colorectal cancer (CRC). Bosn J Basic Med Sci.
18:252–259. 2018.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Liu X and Cui M: MiRNA-98-5p inhibits the
progression of osteosarcoma by regulating cell cycle via targeting
CDC25A expression. Eur Rev Med Pharmacol Sci. 23:9793–9802.
2019.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Ling Z, Guan H, You Z, Wang C, Hu L, Zhang
L, Wang Y, Chen S, Xu B and Chen M: Aloperine executes antitumor
effects through the induction of apoptosis and cell cycle arrest in
prostate cancer in vitro and in vivo. Onco Targets Ther.
11:2735–2743. 2018.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Wang X, Shi J, Niu Z, Wang J and Zhang W:
MiR-216a-3p regulates the proliferation, apoptosis, migration, and
invasion of lung cancer cells via targeting COPB2. Biosci
Biotechnol Biochem: Jul 3, 2020 (Epub ahead of print).
|
|
15
|
Gao ZY, Liu H and Zhang Z: miR-144-3p
increases radiosensibility of gastric cancer cells by targeting
inhibition of ZEB1. Clin Transl Oncol: Jul 1, 2020 (Epub ahead of
print).
|
|
16
|
Chen S, Wang Y, Xu M, Zhang L, Su Y, Wang
B and Zhang X: miR-1184 regulates the proliferation and apoptosis
of colon cancer cells via targeting CSNK2A1. Mol Cell Probes.
101625:2020.(Epub ahead of print). PubMed/NCBI View Article : Google Scholar
|
|
17
|
Zhou Q, Zhu Y, Wei X, Zhou J, Chang L, Sui
H, Han Y, Piao D, Sha R and Bai Y: MiR-590-5p inhibits colorectal
cancer angiogenesis and metastasis by regulating nuclear factor
90/vascular endothelial growth factor A axis. Cell Death Dis.
7(e2413)2016.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Liu Q, Yang W, Luo Y, Hu S and Zhu L:
Correlation between miR-21 and miR-145 and the incidence and
prognosis of colorectal cancer. J BUON. 23:29–35. 2018.PubMed/NCBI
|
|
20
|
Hu JL, He GY, Lan XL, Zeng ZC, Guan J,
Ding Y, Qian XL, Liao WT, Ding YQ and Liang L: Inhibition of
ATG12-mediated autophagy by miR-214 enhances radiosensitivity in
colorectal cancer. Oncogenesis. 7(16)2018.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Liao D, Li T, Ye C, Zeng L, Li H, Pu X,
Ding C, He Z and Huang GL: miR-221 inhibits autophagy and targets
TP53INP1 in colorectal cancer cells. Exp Ther Med. 15:1712–1717.
2018.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Ou C, Sun Z, Li X, Li X, Ren W, Qin Z,
Zhang X, Yuan W, Wang J, Yu W, et al: MiR-590-5p, a
density-sensitive microRNA, inhibits tumorigenesis by targeting
YAP1 in colorectal cancer. Cancer Lett. 399:53–63. 2017.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Wang X, Lai Q, He J, Li Q, Ding J, Lan Z,
Gu C, Yan Q, Fang Y, Zhao X and Liu S: LncRNA SNHG6 promotes
proliferation, invasion and migration in colorectal cancer cells by
activating TGF-β/Smad signaling pathway via targeting UPF1 and
inducing EMT via regulation of ZEB1. Int J Med Sci. 16:51–59.
2019.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Peng H, Wang L, Su Q, Yi K, Du J and Wang
Z: MiR-31-5p promotes the cell growth, migration and invasion of
colorectal cancer cells by targeting NUMB. Biomed Pharmacother.
109:208–216. 2018.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Jiang Z, Cao Q, Dai G, Wang J, Liu C, Lv L
and Pan J: Celastrol inhibits colorectal cancer through
TGF-beta1/Smad signaling. Onco Targets Ther. 12:509–518.
2019.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Jin Y, Chen W, Yang H, Yan Z, Lai Z, Feng
J, Peng J and Lin J: Scutellaria barbata D. Don inhibits migration
and invasion of colorectal cancer cells via suppression of PI3K/AKT
and TGF-β/Smad signaling pathways. Exp Ther Med. 14:5527–5534.
2017.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Liu Y, Uzair-Ur-Rehman Guo Y, Liang H,
Cheng R, Yang F, Hong Y, Zhao C, Liu M, Yu M, et al: miR-181b
functions as an oncomiR in colorectal cancer by targeting PDCD4.
Protein Cell. 7:722–734. 2016.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Lee YR, Chen SH, Lin CY, Chao WY, Lim YP,
Yu HI and Lu CH: In vitro antitumor activity of aloperine on human
thyroid cancer cells through caspase-dependent apoptosis. Int J Mol
Sci. 19(312)2018.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Wang L, Zhao M, Guo C, Wang G, Zhu F, Wang
J, Wang X, Wang Q, Zhao W, Shi Y, et al: PDCD4 deficiency
aggravated colitis and Colitis-associated colorectal cancer via
promoting IL-6/STAT3 pathway in mice. Inflamm Bowel Dis.
22:1107–1118. 2016.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Lim SC and Hong R: Programmed cell death 4
(Pdcd4) expression in colorectal adenocarcinoma: Association with
clinical stage. Oncol Lett. 2:1053–1057. 2011.PubMed/NCBI View Article : Google Scholar
|