Histone methylation and demethylation are an
important pair of histone epigenetic modifications, which play key
roles in gene transcription regulation. Methylation occurs on the
lysine and arginine residues of histones, including K9, K27 and K36
of histone H3, K20 of H4, R2, R17 of H3, and R3 of H4(3). The transcriptional regulation of
histone lysine methylation is closely associated with lysine
residue sites and methylation degree. The methylation of H3K4,
H3K36 and H3K79 is often accompanied by the activation of gene
transcription, while the methylation of H3K9, H3K27 and H4K20
inhibits gene transcription (3,4).
Histone lysine demethylases are mainly composed of
lysine specific demethylase (LSD) and Jumonji C
(JmjC)-domain-containing demethylases (JMJD) family demethylases
(5). The LSD family consists of two
members, lysine demethylase 1A (KDM1A)/LSD1 and KDM1B/LSD2, which
remove the monomethyl and dimethyl (me1/me2) of histone lysine
residues via amine oxidation reaction. In 2004, researchers at
Harvard Medical School were the first to report that LSD1 removes
dimethyl and monomethyl modifications of histone H3K4 in
vitro, in the presence of the co-factor FAD and a proton
nitrogen (6). In vivo,
dimethylation and methylation of histone H3K9 can be removed to
inhibit gene expression (6). LSD1
is a member of the monoamine oxidase family, and its action
requires the participation of an extra proton on the ε-N atom
(7). Thus, its demethylation is
limited by the substrate and cannot be modified by the
demethylation of trimethyllysine (7).
JMJD uses an oxygenase mechanism to remove
monomethyl, dimethyl and trimethyl (me1/me2/me3) lysine residues.
In 2006, researchers from the University of North Carolina at
Chapel Hill reported that JmjC domain histone demethylase 1A
(JHDM1A), also known as KDM2A, demethylated H3-methyl-K36
generating formaldehyde and succinate in the presence of Fe (II)
and alpha-ketoglutarate (8,9).
According to sequence homology and structural
similarity, JmjC domain demethylases are divided into seven
subfamilies with different functions (KDM2-8) (9,10). As
an important epigenetic modification, histone lysine demethylation
regulates important physiological and pathological processes, such
as tissue development and tumorigenesis (11). Notably, each subfamily of JMJD
demethylase inhibits specific substrates of different histone
lysine residues (Table I) (11).
The KDM2A transcripts annotated on the NCBI website
mainly have two types, and the longer isoform encoding protein
consists of a JmjC domain, a CXXC-zinc finger (ZF-CXXC) domain, a
plant homologous zinc finger (PHD) domain, an F-box-like domain and
mitogenic exit network protein 1 (AMN1) (13). Conversely, the short-form KDM2A has
no JmjC region, which is the catalytic core of demethylation
(8). The ZF-CXXC domain
specifically recognizes unmethylated CpG islands (14), and the recognition requires the
participation of linker DNA. KDM2A binds to CpG islands and
demethylates the dimethylated H3K36 residue, and exerts weak
activity for monomethylated H3K36 residue (15,16).
In addition to the two standard transcripts annotated on the NCBI
websites, several KDM2A transcripts have been predicted and
reported, such as the isoforms missing the N-terminal JmjC domain
or the AMN1 domain. In addition, there are also significant
functional differences between the subtypes (17). For example, the alternative isoform
of KDM2A lacking the N-terminal demethylase domain can negatively
regulate canonical Wnt signaling (12,17-20).
KDM2A is located in the nucleus and binds to
unmethylated CpG DNA through the ZF-CxxC domain (14), which is essential for maintaining
heterochromosomal homeostasis (21). KDM2A is extensively expressed in
different tissues, with high expression levels in the brain,
testis, ovaries and lungs (22). In
addition, KDM2A is highly expressed in most tumors except prostate
cancer (21,23-25).
As an epigenetic regulator, the expression and biological function
of KDM2A are affected by multiple external factors (26,27).
In pathological processes, such as gastric cancer and glioblastoma,
LINC00460, microRNA (miRNA/miR)-29b, miR-134-5p and miR-3666
directly bind to the KDM2A promoters to regulate KDM2A expression
(24,28-31).
Inflammation, hypoxia or reactive oxygen species production promote
KDM2A expression (26,32), and upregulation of KDM2A induced by
human papilloma virus (HPV)16E7 promotes tumorigenesis and
progression of cervical cancer (33). Metformin activates the AMPK
signaling pathway and decreases intracellular succinic acid levels,
while activation of KDM2A decreases ribosomal RNA (rRNA)
transcription (27). p300 can
directly acetylate KDM2A at position K409, which in turn decreases
demethylation of H3K36me2 and enhances the transcription of p21 and
PUMA, thereby inhibiting the growth and metastasis of osteosarcoma
(34). Mild glucose starvation
induces KDM2A-mediated demethylation of H3K36me2 via the AMPK
signaling pathway to decrease rRNA transcription and the
proliferation of breast cancer cells (35). In non-small cell lung cancer, the
carcinogen TPA activates cyclooxygenase-2 (COX-2) expression via
KDM2A-mediated H3K36 dimethylation near the COX-2 promoter
(36).
JmjC domain-containing histone lysine demethylases
(KDM2-7) are important epigenetic regulators and potential targets
for cancer (11). Thus, there is
great interest to investigate and identify selective and
therapeutic KDMs inhibitors (37).
Understanding the structure of lysine demethylases and their
modular synthetic approach has helped design and develop a series
of highly selective KDM2/7 inhibitors (38,39).
Some inhibitors exhibit antiproliferative activity, and so may be
used as candidates for anticancer agents (38). Human immunodeficiency virus and HPV
induce epigenetic alterations in host cells by altering the levels
of H3K36 methylation within the promoter region of CTLA-4 and
FOXP3, resulting in several diseases and different types of cancer
(40,41). Histone demethylase inhibitors
combined with checkpoint blockade may be used as a novel cancer
treatment strategy (41-43).
As an inhibitor of KDM2A, plant growth regulator Daminozide
has been reported to significantly abrogate the effect of KDM2A on
histone demethylation, and exhibits promising results as an
anticancer therapeutic strategy (44,45).
KDM2A is abnormally expressed in different tumors,
and it plays a vital role in tumorigenesis and progression
(12). Wagner et al
(46) demonstrated that KDM2A binds
to the dual-specificity phosphatase 3 (DUSP3) gene promoter region
and inhibits its expression, which in turn increases
phosphorylation of ERK1/2 and promotes the occurrence and
metastasis of non-small cell lung cancer. Another study reported
similar findings for KDM2A in non-small cell lung cancer, with
HDAC3 as the target gene (25). In
addition, it has been reported that c-Fos recruits KDM2A to the
COX-2 promoter region to promote the transcription of COX-2, and
treatment with the carcinogen TPA promotes the recruitment process
(36). Huang et al (47) demonstrated that KDM2A is highly
expressed in gastric cancer tissues, which promotes the
proliferation and metastasis of tumor cells by inhibiting
programmed cell death protein 4 expression. Furthermore, KDM2A
reverses epithelial-to-mesenchymal transition by regulating the
PI3K signaling pathway, which promotes the progression of ovarian
cancer (48). In colon cancer,
LINC01278 upregulates KDM2A expression to promote cancer
progression (49), and KDM2A
expression is associated with cyclin D1 expression and cell
proliferation (50). In breast
cancer, studies have reported that KDM2A binds to the promoter
region of genes, such as E2F1, which inhibits its transcriptional
regulation and results in the invasion and metastasis of breast
cancer cells (23,51). Conversely, another study has
revealed a completely different transcriptional regulation
mechanism in breast cancer. The results of this study demonstrated
that KDM2A directly upregulates JAG1 expression, which affects
breast cancer stem cell-related characteristics and angiogenesis
(45). Subsequently, it has been
reported that the short isoform of KDM2A (without the JmjC region)
is crucial for promoting the tumorigenesis and progression of
breast cancer, while the role of the long isoform remains unknown
(17). High expression levels of
KDM2A enhance cancer-associated fibroblasts (52), and combined expression of KDM2A and
KDM2B may be associated with clinical prognosis in patients with
breast cancer (53). Researchers at
Harvard Medical School cloned four cDNA subtypes of KDM2A and
demonstrated that only the KDM2A-N782 isoform (the longer isoform
N-terminal 782 amino acids, including JmjC, CXXC and Ring domains,
but not F-box and AMN1 domains) has a significant effect on
promoting cell proliferation (19).
Our previous study reported that KDM2A expression is significantly
upregulated in hepatocellular carcinoma (HCC) tissues compared with
adjacent normal tissues. In addition, high KDM2A expression is
associated with poor prognosis and overall survival in patients
with HCC (54). KDM2A augments stem
cell-like characteristics via demethylation of histone H3K36 at
promoters of stemness-associated transcription factors, such as
OCT4, NANOG and SOX2(54).
In most cases, KDM2A promotes the progression of
tumors. However, KDM2A knockdown in zebrafish has been reported to
disrupt the transcriptome and result in high frequencies of
spontaneous melanoma (55). In
addition, interference with KDM2A expression in HT29 cells
increases colony formation in soft agar (18). KDM2A affects the NF-κB pathway by
demethylating the K218/K221 site of p65 in HT29 cells (56,57).
In different types of tumors, KDM2A has different pro-oncogenic or
anti-oncogenic effects, which may be associated with intertumor
heterogeneity or the extensiveness of KDM2A downstream target genes
and demethylation sites. In addition, multiple splicing forms of
KDM2A play different roles in tumorigenesis and progression of
different types of cancer (12).
KDM2A has been reported to play a vital role in the
differentiation of stem cells and the development of embryos
(19). Several studies have
investigated the role of KDM2A in the proliferation and
differentiation of stem cells from the apical papilla (SCAPs)
(58,59). The transcription factor, BCOR,
interacts with KDM2A to affect the expression of stem-related
genes, such as SOX2(59). In
addition, KDM2A abrogates the inhibition of p15INK4B and
p27kip1, and demethylation of SFRP2 to regulate the
differentiation and proliferation of SCAPs into adipocytes and
chondrocytes (58-60).
KDM2A exhibits synergistic effects with BCL6, inhibits cell
proliferation and regulates osteogenic differentiation of
mesenchymal stem cells via the epidermal growth factor epiregulin,
EREG (61). Researchers cloned four
cDNA isoforms of KDM2A and demonstrated that transfection with the
transcript variant of KDM2A-N782 containing N-terminal 782AA can
significantly promote keratinocytes proliferation (19). Notably, KDM2A promotes the vitamin
C-dependent reprogramming process of adult cells, accelerates the
cell cycle and inhibits resting senescence, and plays an important
role in the process of induced pluripotent stem cells (62).
Researchers analyzed KDM2A expression in the teeth
of mice during embryonic and postnatal stages via reverse
transcription-quantitative PCR and immunohistochemistry analyses.
The results demonstrated that KDM2A may play essential roles in
cell proliferation and tooth differentiation of mice (63). In addition, KDM2A knockdown
exhibited severe embryonic lethality, accompanied by severe growth
defects and weight loss (64).
KDM2A knockdown affects the expression of cell cycle regulatory
factors, such as p21Cip1, which in turn decreases cell
proliferation and increases apoptosis (64). KDM2A plays a vital role in
maintaining embryonic stem cells by affecting the methylation
levels of H3K36me2 and H3K4me3, and regulating the expression of
germ cell-related genes (65). In
the xenopus model, the stability of nuclear β-catenin depends on
its methylation/demethylation, and the lysine demethylase Kdm2a/b
specifically demethylates and degrades non-phosphorylated β-catenin
in the nucleus to regulate the canonical Wnt signaling pathway,
which plays a decisive role in the formation of the
anterior-posterior body axis in xenopus embryos (66).
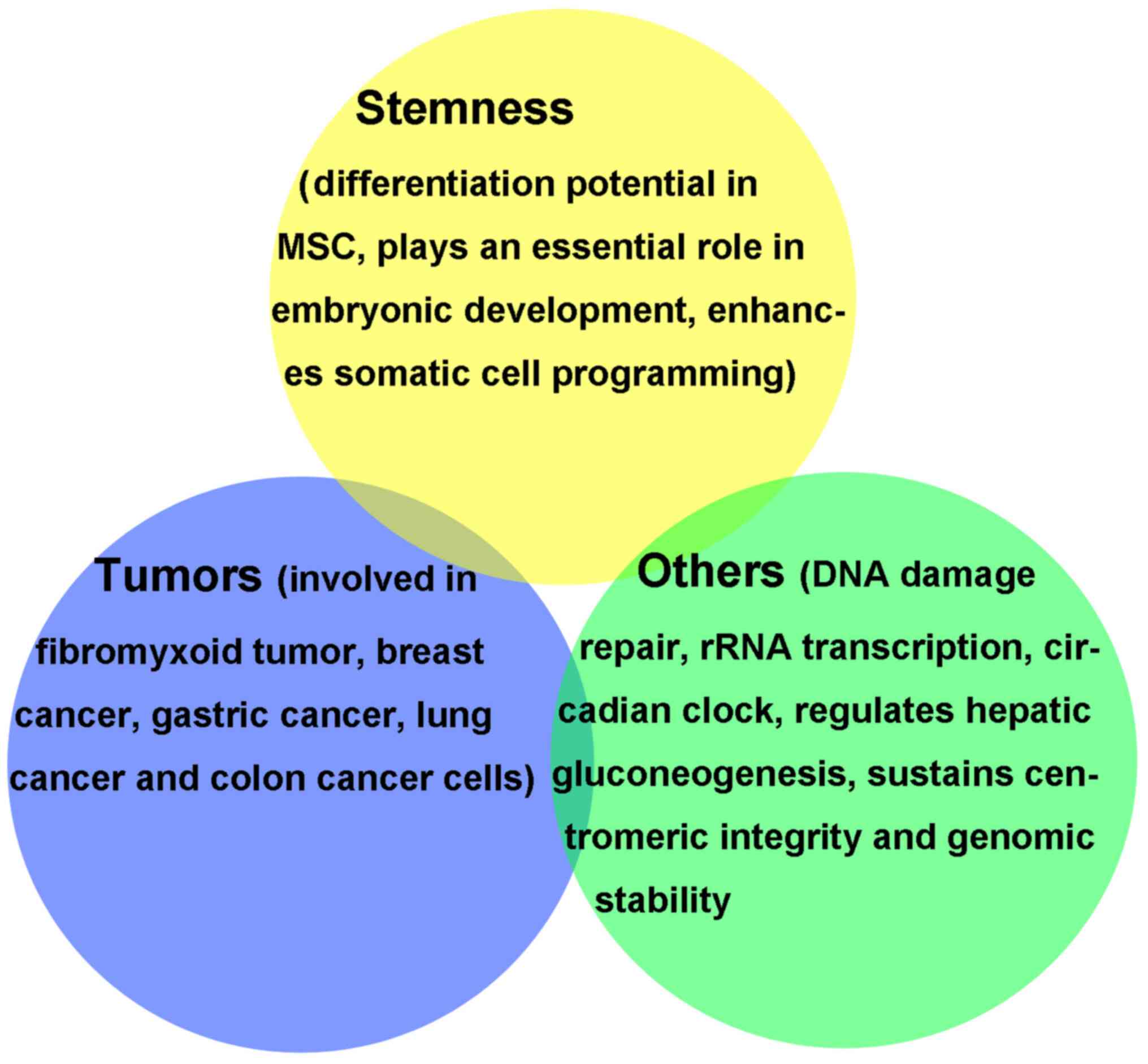
In addition to its involvement in embryonic
development and stem cell differentiation, KDM2A is also associated
with tumorigenesis and progression. KDM2A regulates several
physiological and pathological processes (Fig. 1) (12). Age-associated DNA methylation
changes in blood leukocytes during early childhood may reflect
epigenetic maturation, since histone modifiers and chromatin
remodeling factors, such as KDM2A, are locus susceptible and play
key roles in leukocyte biology (67). JmjC domain-containing histone lysine
demethylases (KDM2-7) are known to be altered with aging, which
could be associated with the regulation of the aging process and
age-related diseases (68). Pan
et al (69) demonstrated
that KDM2A negatively regulates gluconeogenesis-related genes in
vivo, which silences KDM2A expression and accelerates the
synthesis of liver glycogen. Overexpression of KDM2A decreases
blood sugar levels and this regulatory effect is achieved by the
demethylation of H3K36 in the C/EBPa promoter region by KDM2A
(69). Frescas et al
(21) reported that KDM2A is
involved in regulating the expression of small non-coding RNAs that
are encoded by the clusters of satellite repeats at the centromere
and are essential for maintaining heterochromatin homeostasis. The
transcription of rRNA genes is a rate-limiting step in ribosomal
synthesis, which changes in response to environmental stimuli
(26,27,70).
It has been demonstrated that the demethylase KDM2A containing the
JmjC domain remarkably decreases rDNA transcription in response to
starvation, which is accompanied by the demethylation of H3K36me2
in the rDNA promoter (71). In
addition, KDM2A binds to the unmethylated CpG sequence of rDNA
promoter under starvation stress via the CXXC-ZF domain, and
demethylates H3K36me2 in the rDNA promoter to decrease the
transcription of rDNA (70-72).
Reischl and Kramer (73)
demonstrated that KDM2A is an important circadian clock regulator.
KDM2A directly binds to the promoter regions of
CLOCK/BMAL1-regulated genes via a CXXC-ZF motif and regulates the
expression of clock genes, such as Nr1d1, which plays a key role in
maintaining the mammalian circadian rhythms (73). ATM protein (ataxia-telangiectasia
mutated) is an important signal molecule in DNA repair (74); however, its specific molecular
mechanism remains unclear. It has been reported that following DNA
damage, ATM specifically phosphorylates the KDM2A serine 632 site,
which decreases the chromosome binding capacity of KDM2A and
increases the degree of dimethylation of H3K36 at the DNA damage
site (75). This in turn recruits
the MRE11 complex to the injury site, which interacts with BRCT2 to
induce repair and cell survival (75). KDM2A is also recruited to DNA
double-strand breaks and interacts with 53BP1 to ensure the
stability of the genome (76).
Downregulated KDM2A expression increases H3K36me2 at DNA damage
sites to inhibit transcription and promote repair (77).
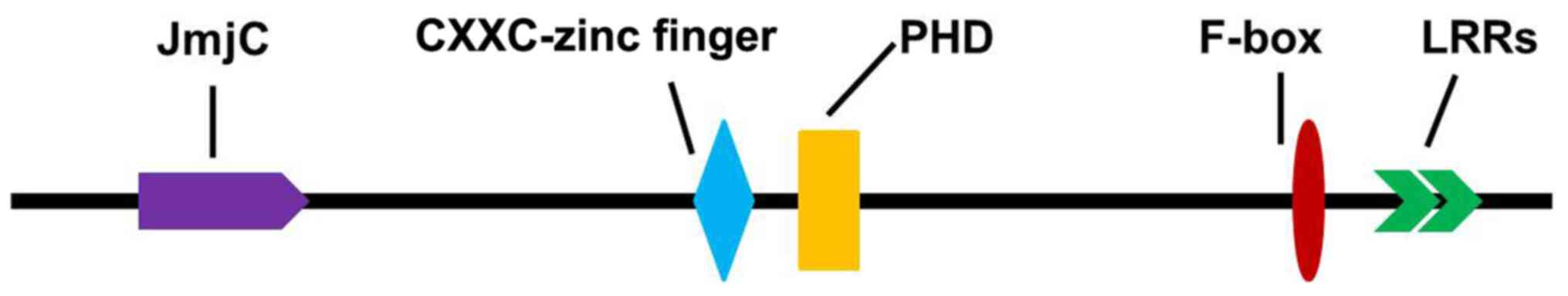
KDM2A belongs to the JMJD family, which consists of
a JmjC domain, a PHD zinc finger structure, LRR-AMN1 and F-box,
CXXC-zinc finger structure domain and other components (Fig. 2) (8). As a histone demethylase, KDM2A
specifically demethylates both monomethylated and dimethylated
lysine-36 of histone H3, and is an important epigenetic
modification (13). KDM2A also
contains SKP1-cullin-F-box, a subunit of the ubiquitin protein
ligase complex, which recognizes and binds to some phosphorylated
proteins and promotes their ubiquitination and degradation
(76).
The methylation and demethylation of histones has
always been relatively balanced, affecting gene regulation and
several biological functions. In addition to affecting chromosome
structure, the level of histone methylation also acts as a
scaffolding molecule to recognize and bind certain transcription
factors and affect transcription regulation (3). Abnormal KDM2A expression and an
imbalance of target gene methylation result in tumorigenesis and
the progression of different types of cancer (12).
In conclusion, KDM2A is closely associated with
physiological processes, such as stem cell differentiation, cell
rhythm and metabolism, as well as heterosomal stability and DNA
damage repair.
Not applicable.
Funding: The present review was supported by the Medicine and
Health Science Technology Development Program of Shandong province
(grant no. 2019WS198) and the Jinan Clinical Medical Science
Technology Innovation Program (grant no. 201704080).
Not applicable.
LL and QL conceived the present review and performed
the literature review. LL drafted the initial manuscript. JL
prepared the table and revised the manuscript for important
intellectual content. All authors have read and approved the final
manuscript.
Not applicable.
Not applicable.
The authors declare that they have no competing
interests.
|
1
|
Luger K, Mader AW, Richmond RK, Sargent DF
and Richmond TJ: Crystal structure of the nucleosome core particle
at 2.8 a resolution. Nature. 389:251–260. 1997.PubMed/NCBI View
Article : Google Scholar
|
|
2
|
Richmond TJ, Finch JT, Rushton B, Rhodes D
and Klug A: Structure of the nucleosome core particle at 7 a
resolution. Nature. 311:532–537. 1984.PubMed/NCBI View
Article : Google Scholar
|
|
3
|
Hyun K, Jeon J, Park K and Kim J: Writing,
erasing and reading histone lysine methylations. Exp Mol Med.
49(e324)2017.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Black JC, Van Rechem C and Whetstine JR:
Histone lysine methylation dynamics: Establishment, regulation, and
biological impact. Mol Cell. 48:491–507. 2012.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Hojfeldt JW, Agger K and Helin K: Histone
lysine demethylases as targets for anticancer therapy. Nat Rev Drug
Discov. 12:917–930. 2013.PubMed/NCBI View
Article : Google Scholar
|
|
6
|
Shi Y, Lan F, Matson C, Mulligan P,
Whetstine JR, Cole PA, Casero RA and Shi Y: Histone demethylation
mediated by the nuclear amine oxidase homolog LSD1. Cell.
119:941–953. 2004.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Fang Y, Liao G and Yu B: Targeting histone
lysine demethylase LSD1/KDM1A as a new avenue for cancer therapy.
Curr Top Med Chem. 19:889–891. 2019.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Tsukada Y, Fang J, Erdjument-Bromage H,
Warren ME, Borchers CH, Tempst P and Zhang Y: Histone demethylation
by a family of JmjC domain-containing proteins. Nature.
439:811–816. 2006.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Xiang Y, Zhu Z, Han G, Ye X, Xu B, Peng Z,
Ma Y, Yu Y, Lin H, Chen AP and Chen CD: JARID1B is a histone H3
lysine 4 demethylase up-regulated in prostate cancer. Proc Natl
Acad Sci USA. 104:19226–19231. 2007.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Klose RJ, Kallin EM and Zhang Y:
JmjC-domain-containing proteins and histone demethylation. Nat Rev
Genet. 7:715–727. 2006.PubMed/NCBI View
Article : Google Scholar
|
|
11
|
Cloos PA, Christensen J, Agger K and Helin
K: Erasing the methyl mark: Histone demethylases at the center of
cellular differentiation and disease. Genes Dev. 22:1115–1140.
2008.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Vacik T, Ladinovic D and Raska I: KDM2A/B
lysine demethylases and their alternative isoforms in development
and disease. Nucleus. 9:431–441. 2018.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Zlotorynski E: Protein folding:
Structure-function analysis of KDM2A. Nat Rev Mol Cell Biol.
15(630)2014.PubMed/NCBI View
Article : Google Scholar
|
|
14
|
Blackledge NP, Zhou JC, Tolstorukov MY,
Farcas AM, Park PJ and Klose RJ: CpG islands recruit a histone H3
lysine 36 demethylase. Mol Cell. 38:179–190. 2010.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Zhou JC, Blackledge NP, Farcas AM and
Klose RJ: Recognition of CpG island chromatin by KDM2A requires
direct and specific interaction with linker DNA. Mol Cell Biol.
32:479–489. 2012.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Iuchi S and Paulo JA: Lysine-specific
demethylase 2A enhances binding of various nuclear factors to
CpG-rich genomic DNAs by action of its CXXC-PHD domain. Sci Rep.
9(5496)2019.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Liu H, Liu L, Holowatyj A, Jiang Y and
Yang ZQ: Integrated genomic and functional analyses of histone
demethylases identify oncogenic KDM2A isoform in breast cancer. Mol
Carcinog. 55:977–990. 2016.PubMed/NCBI View
Article : Google Scholar
|
|
18
|
Cheng Z, Cheung P, Kuo AJ, Yukl ET, Wilmot
CM, Gozani O and Patel DJ: A molecular threading mechanism
underlies Jumonji lysine demethylase KDM2A regulation of methylated
H3K36. Genes Dev. 28:1758–1771. 2014.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Iuchi S and Green H: Lysine-specific
demethylase 2A (KDM2A) normalizes human embryonic stem cell derived
keratinocytes. Proc Natl Acad Sci USA. 109:9442–9447.
2012.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Ladinovic D, Pinkas D, Šopin T, Raška O,
Liška F, Raška I and Vacík T: Alternative isoforms of KDM2A and
KDM2B lysine demethylases negatively regulate canonical Wnt
signaling. PLoS One. 15(e236612)2020.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Frescas D, Guardavaccaro D, Kuchay SM,
Kato H, Poleshko A, Basrur V, Elenitoba-Johnson KS, Katz RA and
Pagano M: KDM2A represses transcription of centromeric satellite
repeats and maintains the heterochromatic state. Cell Cycle.
7:3539–3547. 2008.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Nagase T, Ishikawa K, Suyama M, Kikuno R,
Hirosawa M, Miyajima N, Tanaka A, Kotani H, Nomura N and Ohara O:
Prediction of the coding sequences of unidentified human genes.
XIII. The complete sequences of 100 new cDNA clones from brain
which code for large proteins in vitro. DNA Res. 6:63–70.
1999.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Chen JY, Luo CW, Lai YS, Wu CC and Hung
WC: Lysine demethylase KDM2A inhibits TET2 to promote DNA
methylation and silencing of tumor suppressor genes in breast
cancer. Oncogenesis. 6(e369)2017.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Kong Y, Zou S, Yang F, Xu X, Bu W, Jia J
and Liu Z: RUNX3-mediated up-regulation of miR-29b suppresses the
proliferation and migration of gastric cancer cells by targeting
KDM2A. Cancer Lett. 381:138–148. 2016.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Dhar SS, Alam H, Li N, Wagner KW, Chung J,
Ahn YW and Lee MG: Transcriptional repression of histone
deacetylase 3 by the histone demethylase KDM2A is coupled to
tumorigenicity of lung cancer cells. J Biol Chem. 289:7483–7496.
2014.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Tanaka Y, Obinata H, Konishi A, Yamagiwa N
and Tsuneoka M: Production of ROS by Gallic acid activates KDM2A to
reduce rRNA transcription. Cells. 9(2266)2020.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Tanaka Y, Konishi A, Obinata H and
Tsuneoka M: Metformin activates KDM2A to reduce rRNA transcription
and cell proliferation by dual regulation of AMPK activity and
intracellular succinate level. Sci Rep. 9(18694)2019.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Wang F, Liang S, Liu X, Han L, Wang J and
Du Q: LINC00460 modulates KDM2A to promote cell proliferation and
migration by targeting miR-342-3p in gastric cancer. Onco Targets
Ther. 11:6383–6394. 2018.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Shou T, Yang H, Lv J, Liu D and Sun X:
MicroRNA3666 suppresses the growth and migration of glioblastoma
cells by targeting KDM2A. Mol Med Rep. 19:1049–1055.
2019.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Li X, Wei C, Zhang Z, Jin Q and Xiao X:
MiR-134-5p regulates myocardial apoptosis and angiogenesis by
directly targeting KDM2A After myocardial infarction. Int Heart J.
61:815–821. 2020.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Zhao Y, Chen X, Jiang J, Wan X, Wang Y and
Xu P: Epigallocatechin gallate reverses gastric cancer by
regulating the long noncoding RNA LINC00511/miR-29b/KDM2A axis.
Biochim Biophys Acta Mol Basis Dis. 1866(165856)2020.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Yang H, Li G, Han N, Zhang X, Cao Y, Cao Y
and Fan Z: Secreted frizzled-related protein 2 promotes the
osteo/odontogenic differentiation and paracrine potentials of stem
cells from apical papilla under inflammation and hypoxia
conditions. Cell Prolif. 53(e12694)2020.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Ou R, Zhu L, Zhao L, Li W, Tao F, Lu Y, He
Q, Li J, Ren Y and Xu Y: HPV16 E7-induced upregulation of KDM2A
promotes cervical cancer progression by regulating miR-132-radixin
pathway. J Cell Physiol. 234:2659–2671. 2019.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Wang Y, Sun B, Zhang Q, Dong H and Zhang
J: p300 Acetylates JHDM1A to inhibit osteosarcoma carcinogenesis.
Artif Cells Nanomed Biotechnol. 47:2891–2899. 2019.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Tanaka Y, Yano H, Ogasawara S, Yoshioka S,
Imamura H, Okamoto K and Tsuneoka M: Mild glucose starvation
induces KDM2A-mediated H3K36me2 demethylation through AMPK To
reduce rRNA transcription and cell proliferation. Mol Cell Biol.
35:4170–4184. 2015.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Lu S, Yang Y, Du Y, Cao LL, Li M, Shen C,
Hou T, Zhao Y, Wang H, Deng D, et al: The transcription factor
c-Fos coordinates with histone lysine-specific demethylase 2A to
activate the expression of cyclooxygenase-2. Oncotarget.
6:34704–34717. 2015.PubMed/NCBI View Article : Google Scholar
|
|
37
|
McAllister TE, England KS, Hopkinson RJ,
Brennan PE, Kawamura A and Schofield CJ: Recent progress in histone
demethylase inhibitors. J Med Chem. 59:1308–1329. 2016.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Suzuki T, Ozasa H, Itoh Y, Zhan P, Sawada
H, Mino K, Walport L, Ohkubo R, Kawamura A, Yonezawa M, et al:
Identification of the KDM2/7 histone lysine demethylase subfamily
inhibitor and its antiproliferative activity. J Med Chem.
56:7222–7231. 2013.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Gerken PA, Wolstenhulme JR, Tumber A,
Hatch SB, Zhang Y, Müller S, Chandler SA, Mair B, Li F, Nijman SMB,
et al: Discovery of a highly selective cell-active inhibitor of the
histone lysine demethylases KDM2/7. Angew Chem Int Ed Engl.
56:15555–15559. 2017.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Zhou Q, Chen L, Song Y, Ma L, Xiao P, Chen
L, Zhen H, Han R, Chen X, Sun S, et al: Induction of co-inhibitory
molecule CTLA-4 by human papillomavirus E7 protein through
downregulation of histone methyltransferase JHDM1B expression.
Virology. 538:111–118. 2019.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Abdel-Hameed EA, Ji H and Shata MT:
HIV-induced epigenetic alterations in host cells. Adv Exp Med Biol.
879:27–38. 2016.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Sheng W, LaFleur MW, Nguyen TH, Chen S,
Chakravarthy A, Conway JR, Li Y, Chen H, Yang H, Hsu PH, et al:
LSD1 ablation stimulates anti-tumor immunity and enables checkpoint
blockade. Cell. 174:549–563.e19. 2018.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Boehm D and Ott M: Host methyltransferases
and demethylases: Potential new epigenetic targets for HIV cure
strategies and beyond. AIDS Res Hum Retroviruses. 33
(Suppl):S8–S22. 2017.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Rose NR, Woon EC, Tumber A, Walport LJ,
Chowdhury R, Li XS, King ON, Lejeune C, Ng SS, Krojer T, et al:
Plant growth regulator daminozide is a selective inhibitor of human
KDM2/7 histone demethylases. J Med Chem. 55:6639–6643.
2012.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Chen JY, Li CF, Chu PY, Lai YS, Chen CH,
Jiang SS, Hou MF and Hung WC: Lysine demethylase 2A promotes
stemness and angiogenesis of breast cancer by upregulating Jagged1.
Oncotarget. 7:27689–27710. 2016.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Wagner KW, Alam H, Dhar SS, Giri U, Li N,
Wei Y, Giri D, Cascone T, Kim JH, Ye Y, et al: KDM2A promotes lung
tumorigenesis by epigenetically enhancing ERK1/2 signaling. J Clin
Invest. 123:5231–5246. 2013.PubMed/NCBI View Article : Google Scholar
|
|
47
|
Huang Y, Liu Y, Yu L, Chen J, Hou J, Cui
L, Ma D and Lu W: Histone demethylase KDM2A promotes tumor cell
growth and migration in gastric cancer. Tumour Biol. 36:271–278.
2015.PubMed/NCBI View Article : Google Scholar
|
|
48
|
Lu DH, Yang J, Gao LK, Min J, Tang JM, Hu
M, Li Y, Li ST, Chen J and Hong L: Lysine demethylase 2A promotes
the progression of ovarian cancer by regulating the PI3K pathway
and reversing epithelialmesenchymal transition. Oncol Rep.
41:917–927. 2019.PubMed/NCBI View Article : Google Scholar
|
|
49
|
Xi C, Ye NY and Wang YB: LncRNA LINC01278
accelerates colorectal cancer progression via miR-134-5p/KDM2A
axis. Eur Rev Med Pharmacol Sci. 24:10526–10534. 2020.PubMed/NCBI View Article : Google Scholar
|
|
50
|
Cao LL, Du C, Liu H, Pei L, Qin L, Jia M
and Wang H: Lysine-specific demethylase 2A expression is associated
with cell growth and cyclin D1 expression in colorectal
adenocarcinoma. Int J Biol Markers: Apr 1, 2018 (Epub ahead of
print). doi: 10.1177/1724600818764069.
|
|
51
|
Rizwani W, Schaal C, Kunigal S, Coppola D
and Chellappan S: Mammalian lysine histone demethylase KDM2A
regulates E2F1-mediated gene transcription in breast cancer cells.
PLoS One. 9(e100888)2014.PubMed/NCBI View Article : Google Scholar
|
|
52
|
Chen JY, Li CF, Lai YS and Hung WC: Lysine
demethylase 2A expression in cancer-associated fibroblasts promotes
breast tumour growth. Br J Cancer. 124:484–493. 2020.PubMed/NCBI View Article : Google Scholar
|
|
53
|
De Nicola I, Guerrieri AN, Penzo M,
Ceccarelli C, De Leo A, Trerè D and Montanaro L: Combined
expression levels of KDM2A and KDM2B correlate with nucleolar size
and prognosis in primary breast carcinomas. Histol Histopathol.
35:1181–1187. 2020.PubMed/NCBI View Article : Google Scholar
|
|
54
|
Lin Q, Wu Z, Yue X, Yu X, Wang Z, Song X,
Xu L, He Y, Ge Y, Tan S, et al: ZHX2 restricts hepatocellular
carcinoma by suppressing stem cell-like traits through
KDM2A-mediated H3K36 demethylation. Ebiomedicine.
53(102676)2020.PubMed/NCBI View Article : Google Scholar
|
|
55
|
Scahill CM, Digby Z, Sealy IM,
Wojciechowska S, White RJ, Collins JE, Stemple DL, Bartke T,
Mathers ME, Patton EE and Busch-Nentwich EM: Loss of the chromatin
modifier Kdm2aa causes BrafV600E-independent spontaneous melanoma
in zebrafish. PLoS Genet. 13(e1006959)2017.PubMed/NCBI View Article : Google Scholar
|
|
56
|
Lu T, Jackson MW, Singhi AD, Kandel ES,
Yang M, Zhang Y, Gudkov AV and Stark GR: Validation-based
insertional mutagenesis identifies lysine demethylase FBXL11 as a
negative regulator of NFkappaB. Proc Natl Acad Sci USA.
106:16339–16344. 2009.PubMed/NCBI View Article : Google Scholar
|
|
57
|
Lu T, Jackson MW, Wang B, Yang M, Chance
MR, Miyagi M, Gudkov AV and Stark GR: Regulation of NF-kappaB by
NSD1/FBXL11-dependent reversible lysine methylation of p65. Proc
Natl Acad Sci USA. 107:46–51. 2010.PubMed/NCBI View Article : Google Scholar
|
|
58
|
Gao R, Dong R, Du J, Ma P, Wang S and Fan
Z: Depletion of histone demethylase KDM2A inhibited cell
proliferation of stem cells from apical papilla by de-repression of
p15INK4B and p27Kip1. Mol Cell Biochem. 379:115–122.
2013.PubMed/NCBI View Article : Google Scholar
|
|
59
|
Dong R, Yao R, Du J, Wang S and Fan Z:
Depletion of histone demethylase KDM2A enhanced the adipogenic and
chondrogenic differentiation potentials of stem cells from apical
papilla. Exp Cell Res. 319:2874–2882. 2013.PubMed/NCBI View Article : Google Scholar
|
|
60
|
Yu G, Wang J, Lin X, Diao S, Cao Y, Dong
R, Wang L, Wang S and Fan Z: Demethylation of SFRP2 by histone
demethylase KDM2A regulated osteo-/dentinogenic differentiation of
stem cells of the apical papilla. Cell Prolif. 49:330–340.
2016.PubMed/NCBI View Article : Google Scholar
|
|
61
|
Du J, Ma Y, Ma P, Wang S and Fan Z:
Demethylation of epiregulin gene by histone demethylase FBXL11 and
BCL6 corepressor inhibits osteo/dentinogenic differentiation. Stem
Cells. 31:126–136. 2013.PubMed/NCBI View Article : Google Scholar
|
|
62
|
Wang T, Chen K, Zeng X, Yang J, Wu Y, Shi
X, Qin B, Zeng L, Esteban MA, Pan G and Pei D: The histone
demethylases Jhdm1a/1b enhance somatic cell reprogramming in a
vitamin-C-dependent manner. Cell Stem Cell. 9:575–587.
2011.PubMed/NCBI View Article : Google Scholar
|
|
63
|
Yi Q, Cao Y, Liu OS, Lu YQ, Wang JS, Wang
SL, Yao R and Fan ZP: Spatial and temporal expression of histone
demethylase, Kdm2a, during murine molar development. Biotech
Histochem. 91:137–144. 2016.PubMed/NCBI View Article : Google Scholar
|
|
64
|
Kawakami E, Tokunaga A, Ozawa M, Sakamoto
R and Yoshida N: The histone demethylase Fbxl11/Kdm2a plays an
essential role in embryonic development by repressing cell-cycle
regulators. Mech Dev. 135:31–42. 2015.PubMed/NCBI View Article : Google Scholar
|
|
65
|
Fu E, Shen J, Dong Z, Zhang W, Zhang Y,
Chen F, Cheng Z, Zhao X, Shuai L and Lu X: Histone demethylase
Kdm2a regulates germ cell genes and endogenous retroviruses in
embryonic stem cells. Epigenomics. 11:751–766. 2019.PubMed/NCBI View Article : Google Scholar
|
|
66
|
Lu L, Gao Y, Zhang Z, Cao Q, Zhang X, Zou
J and Cao Y: Kdm2a/b lysine demethylases regulate canonical Wnt
signaling by modulating the stability of nuclear β-catenin. Dev
Cell. 33:660–674. 2015.PubMed/NCBI View Article : Google Scholar
|
|
67
|
Acevedo N, Reinius LE, Vitezic M, Fortino
V, Söderhäll C, Honkanen H, Veijola R, Simell O, Toppari J, Ilonen
J, et al: Age-associated DNA methylation changes in immune genes,
histone modifiers and chromatin remodeling factors within 5 years
after birth in human blood leukocytes. Clin Epigenetics.
7(34)2015.PubMed/NCBI View Article : Google Scholar
|
|
68
|
Salminen A, Kauppinen A and Kaarniranta K:
2-Oxoglutarate-dependent dioxygenases are sensors of energy
metabolism, oxygen availability, and iron homeostasis: Potential
role in the regulation of aging process. Cell Mol Life Sci.
72:3897–3914. 2015.PubMed/NCBI View Article : Google Scholar
|
|
69
|
Pan D, Mao C, Zou T, Yao AY, Cooper MP,
Boyartchuk V and Wang YX: The histone demethylase Jhdm1a regulates
hepatic gluconeogenesis. PLoS Genet. 8(e1002761)2012.PubMed/NCBI View Article : Google Scholar
|
|
70
|
Tanaka Y, Okamoto K, Teye K, Umata T,
Yamagiwa N, Suto Y, Zhang Y and Tsuneoka M: JmjC enzyme KDM2A is a
regulator of rRNA transcription in response to starvation. EMBO J.
29:1510–1522. 2010.PubMed/NCBI View Article : Google Scholar
|
|
71
|
Tanaka Y, Umata T, Okamoto K, Obuse C and
Tsuneoka M: CxxC-ZF domain is needed for KDM2A to demethylate
histone in rDNA promoter in response to starvation. Cell Struct
Funct. 39:79–92. 2014.PubMed/NCBI View Article : Google Scholar
|
|
72
|
Tsuneoka M, Tanaka Y and Okamoto K: A CxxC
domain that binds to unmethylated CpG is required for KDM2A to
control rDNA transcription. Yakugaku Zasshi. 135:11–21.
2015.PubMed/NCBI View Article : Google Scholar : (In Japanese).
|
|
73
|
Reischl S and Kramer A: Fbxl11 is a novel
negative element of the mammalian circadian clock. J Biol Rhythms.
30:291–301. 2015.PubMed/NCBI View Article : Google Scholar
|
|
74
|
Jin MH and Oh DY: ATM in DNA repair in
cancer. Pharmacol Ther. 203(107391)2019.PubMed/NCBI View Article : Google Scholar
|
|
75
|
Cao LL, Wei F, Du Y, Song B, Wang D, Shen
C, Lu X, Cao Z, Yang Q, Gao Y, et al: ATM-mediated KDM2A
phosphorylation is required for the DNA damage repair. Oncogene.
35(402)2016.PubMed/NCBI View Article : Google Scholar
|
|
76
|
Bueno M, Baldascini M, Richard S and
Lowndes NF: Recruitment of lysine demethylase 2A to DNA double
strand breaks and its interaction with 53BP1 ensures genome
stability. Oncotarget. 9:15915–15930. 2018.PubMed/NCBI View Article : Google Scholar
|
|
77
|
Rezazadeh S, Yang D, Biashad SA, Firsanov
D, Takasugi M, Gilbert M, Tombline G, Bhanu NV, Garcia BA, Seluanov
A and Gorbunova V: SIRT6 mono-ADP ribosylates KDM2A to locally
increase H3K36me2 at DNA damage sites to inhibit transcription and
promote repair. Aging (Albany NY). 12:11165–11184. 2020.PubMed/NCBI View Article : Google Scholar
|